Generation of Vertebra Micro-CT-like Image from MDCT: A Deep-Learning-Based Image Enhancement Approach
Abstract
:1. Introduction
2. Materials and Methods
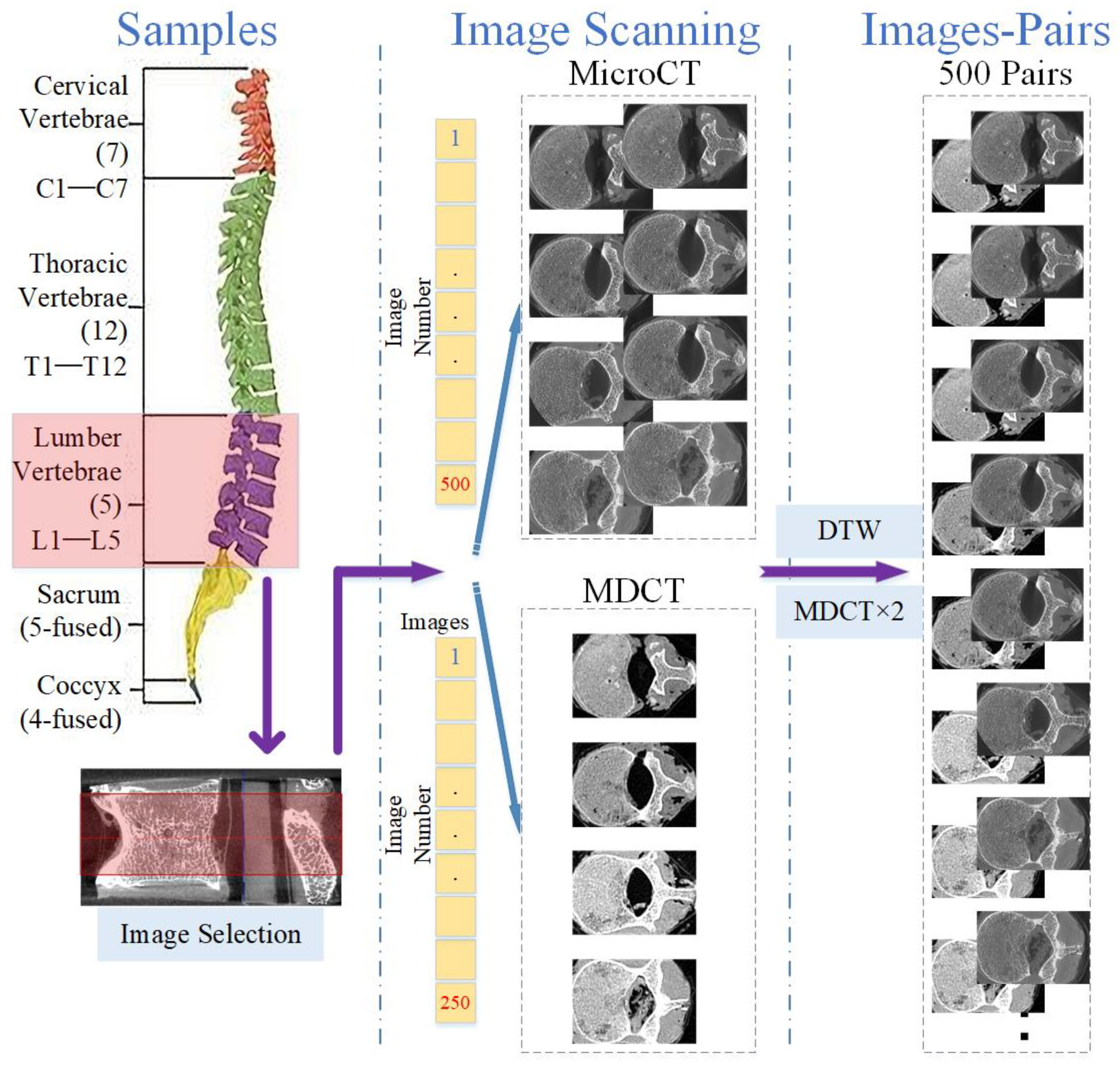
2.1. Specimens
2.2. Imaging Techniques
2.3. Image Co-Registration
2.4. Construction of Training Set and Testing Set
- (1)
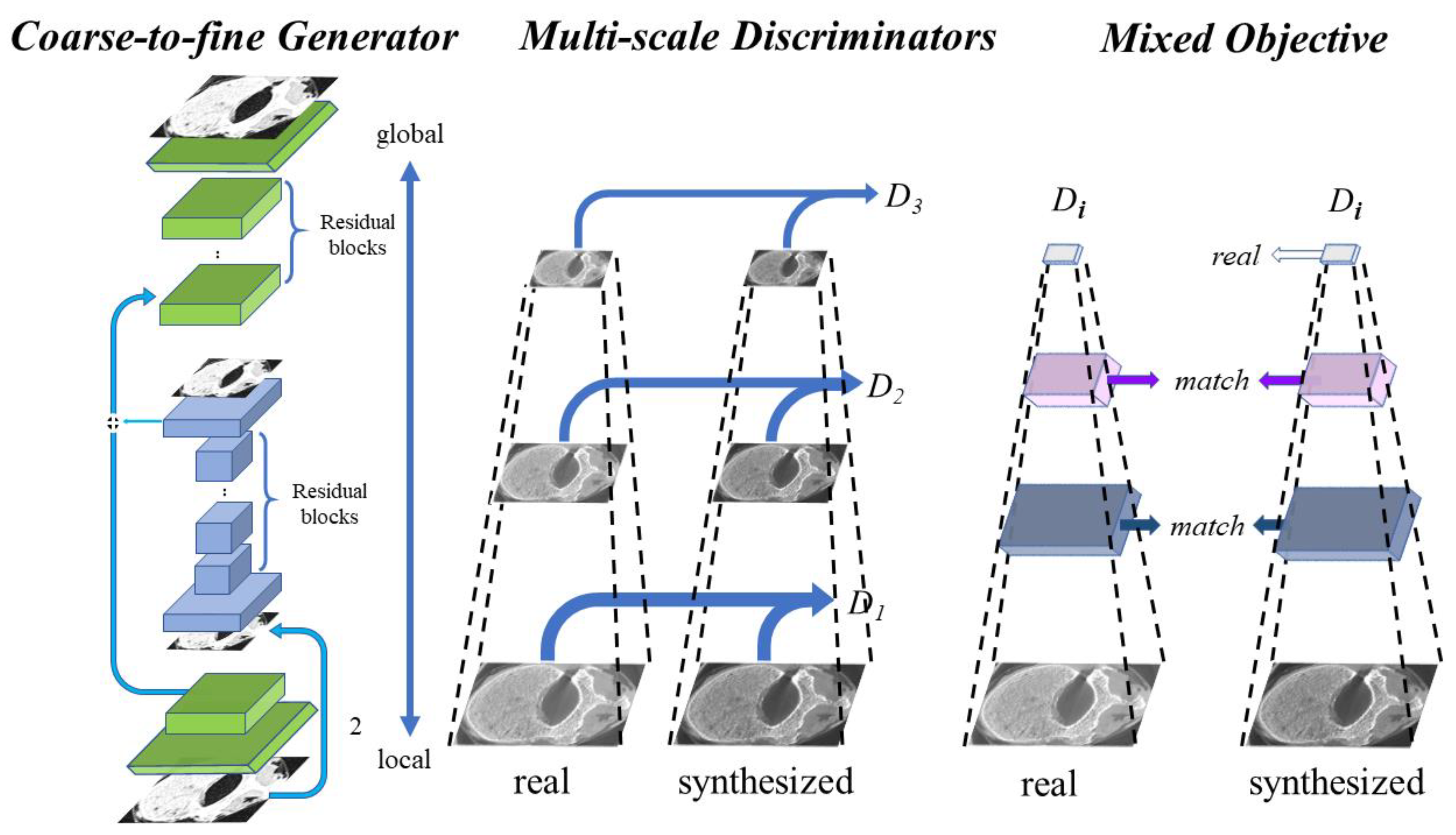
- Characteristics of the selected model. In this paper, we intended to map MDCT images to micro-CT-like images using an image-to-image method named pix2pixHD. This method is a supervised paired image learning method that maps images from the source MDCT domain to the target micro-CT domain and does not consider the continuity within the image domain. Image pairs are randomly selected for tuning the model during training, and no images of a particular vertebra are fed into the training as a set. In other words, in the framework of the selected technique, all image pairs are considered independent during training, and the correlation between different slices of images within a vertebra is ignored.
- (2)
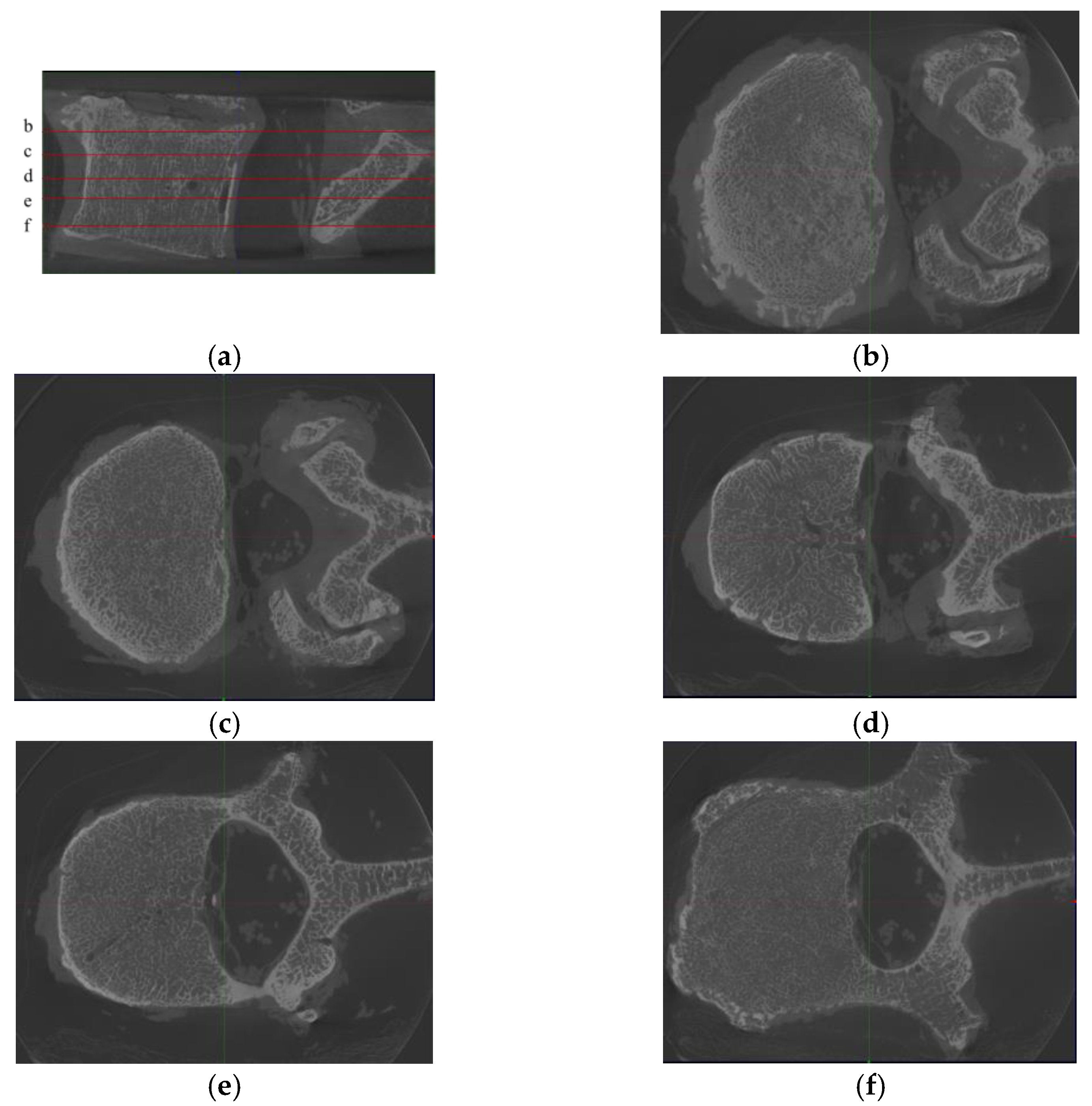
- Diversity within each vertebra. Due to the diversity of images at each slice inside vertebrae (see Figure 2), the images within a vertebra do not obey the same distribution. This diversity is even more pronounced in the presence of vertebral attachments. To better realize the training, we needed to use all pairs of images at all slices in vertebrae as the basic unit for model training.
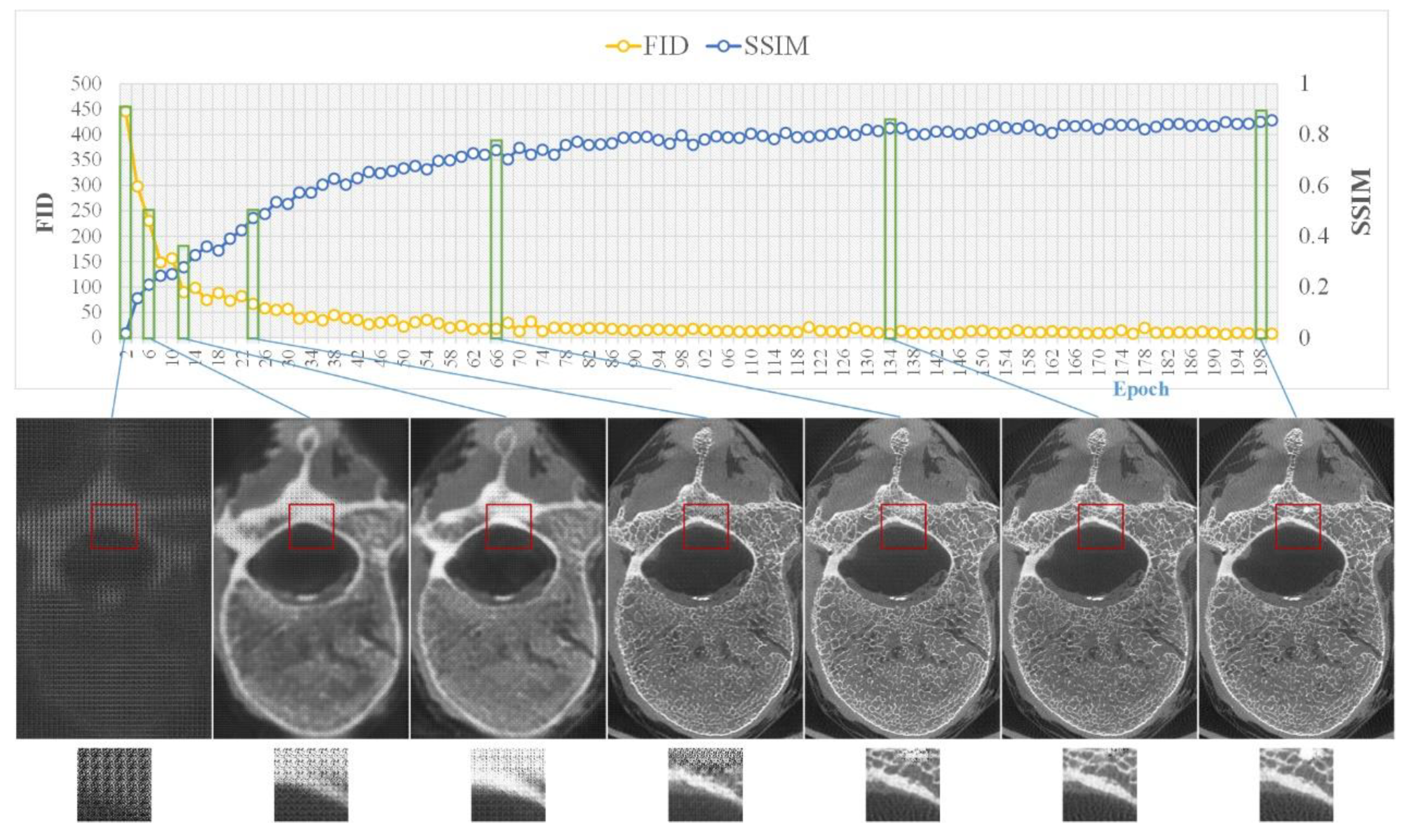
2.5. Model Training
2.6. Objective Assessment of Image Quality
2.7. Subjective Assessment of Image Quality
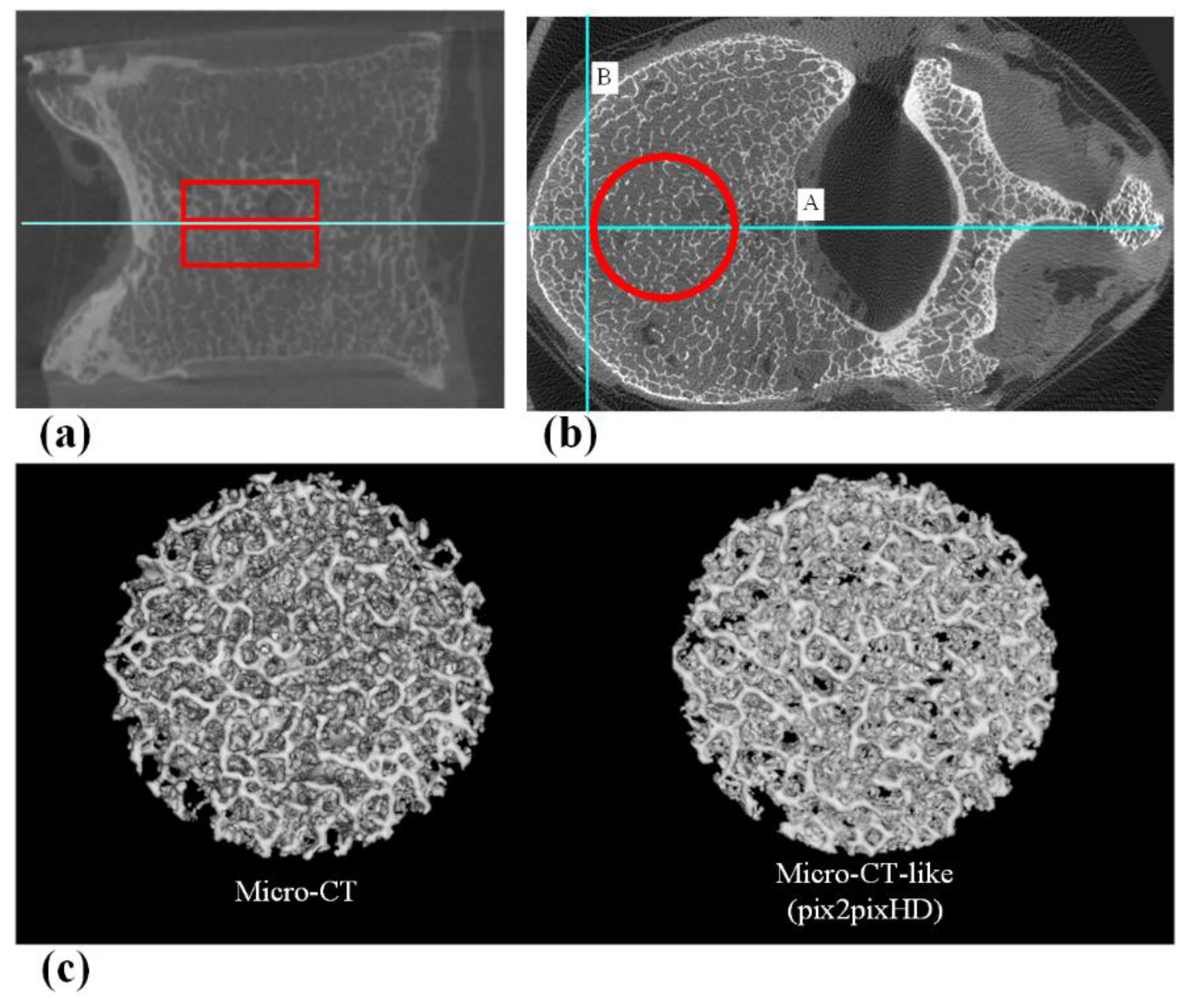
2.8. Assessment of the Trabecular Bone Microstructure
2.9. Statistics
3. Results
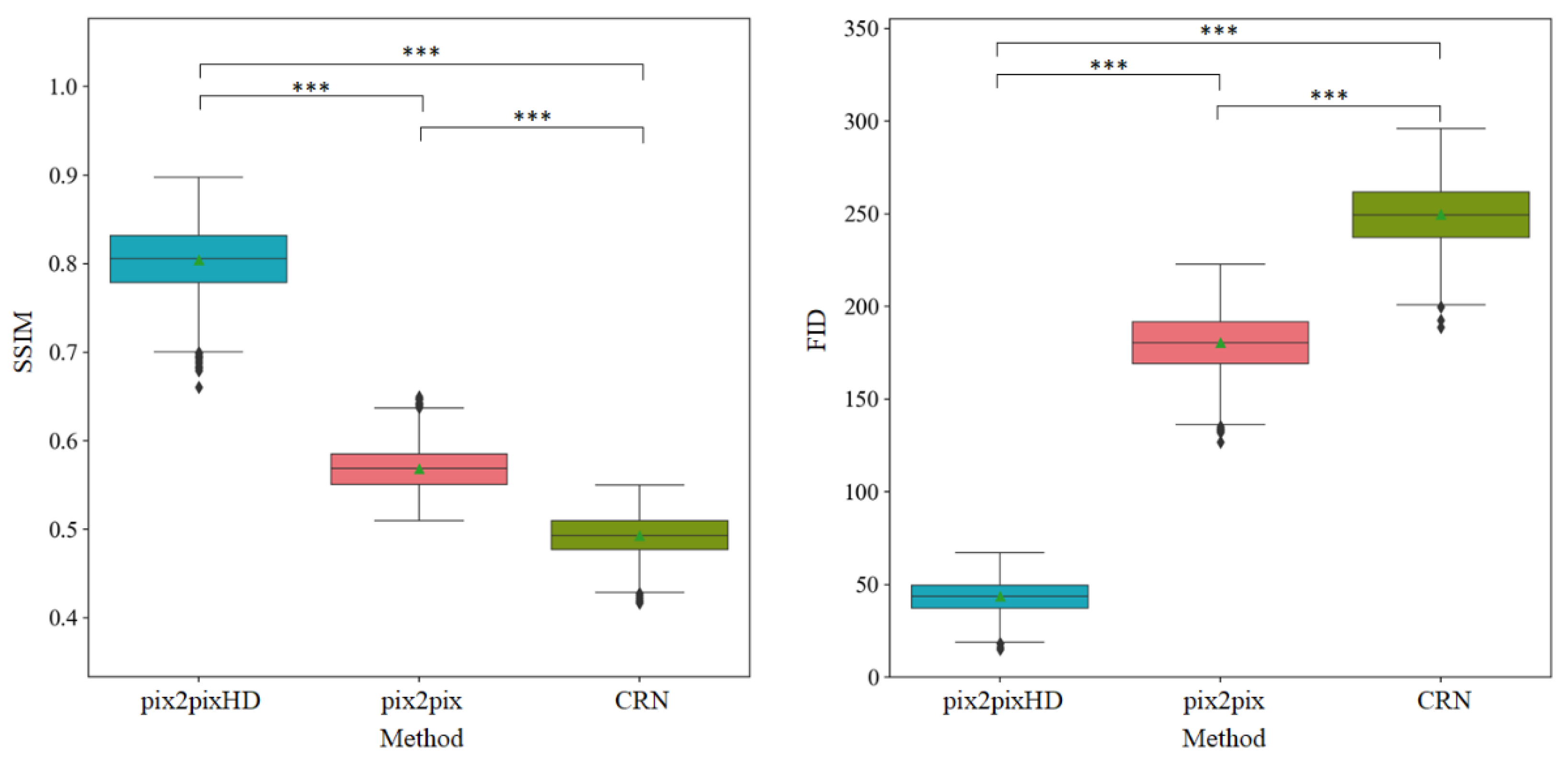
3.1. Objective Assessment of Micro-CT-like Image Quality of the Three Evaluated Methods
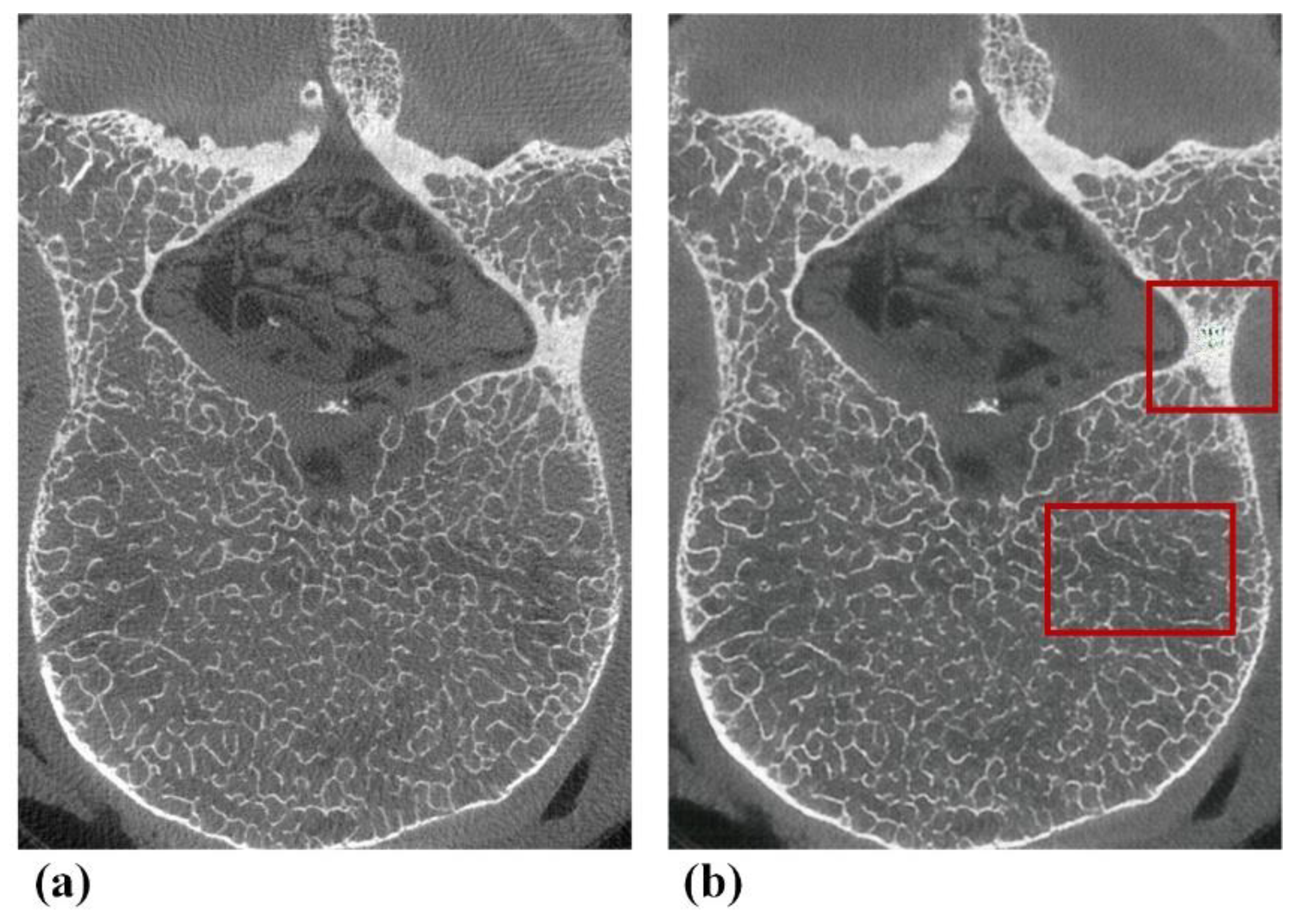
3.2. Subjective Assessment of pix2pixHD-Derived Micro-CT-like Image Quality
3.3. Assessment of Trabecular Bone Microstructure with pix2pixHD-Derived Micro-CT-like Images and Micro-CT Images
4. Discussion
5. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Klibanski, A.; Adams-Campbell, L.; Bassford, T.; Blair, S.N.; Boden, S.D.; Dickersin, K.; Gifford, D.R.; Glasse, L.; Goldring, S.R.; Hruska, K.; et al. Osteoporosis prevention, diagnosis, and therapy. J. Am. Med. Assoc. 2001, 285, 785–795. [Google Scholar]
- Wehrli, F.W.; Saha, P.K.; Gomberg, B.R.; Song, H.K.; Snyder, P.J.; Benito, M.; Wright, A.; Weening, R. Role of magnetic resonance for assessing structure and function of trabecular bone. Top. Magn. Reson. Imaging TMRI 2002, 13, 335–355. [Google Scholar] [CrossRef]
- Dalzell, N.; Kaptoge, S.; Morris, N.; Berthier, A.; Koller, B.; Braak, L.; Rietbergen, B.V.; Reeve, J. Bone micro-architecture and determinants of strength in the radius and tibia: Age-related changes in a population-based study of normal adults measured with high-resolution pqct. Osteoporos. Int. 2009, 20, 1683–1694. [Google Scholar] [CrossRef]
- Iwamoto, Y.; Sato, Y.; Uemura, K.; Takao, M.; Sugano, N.; Takeda, K.; Chen, Y.-W. Medical Imaging 2018: Physics of Medical Imaging. In Reconstruction of Micro CT-like Images from Clinical CT Images Using Machine Learning: A Preliminary Study; SPIE Medical Imaging: Houston, TX, USA, 2018; Volume 10573. [Google Scholar]
- Issever, A.S.; Link, T.M.; Kentenich, M.; Rogalla, P.; Schwieger, K.; Huber, M.B.; Burghardt, A.J.; Majumdar, S.; Diederichs, G. Trabecular bone structure analysis in the osteoporotic spine using a clinical in vivo setup for 64-slice mdct imaging: Comparison to microct imaging and microfe modeling. J. Bone Miner. Res. 2009, 24, 1628–1637. [Google Scholar] [CrossRef] [PubMed]
- Guha, I.; Klintström, B.; Klintström, E.; Zhang, X.; Smedby, Ö.; Moreno, R.; Saha, P.K. A comparative study of trabecular bone micro-structural measurements using different ct modalities. Phys. Med. Biol. 2020, 65, 235029. [Google Scholar] [CrossRef] [PubMed]
- Chen, C.; Zhang, X.; Guo, J.; Jin, D.; Letuchy, E.M.; Burns, T.L.; Levy, S.M.; Hoffman, E.A.; Saha, P.K. Quantitative imaging of peripheral trabecular bone micro-architecture using mdct. Med. Phys. 2018, 45, 236–249. [Google Scholar] [CrossRef]
- Wehrli, F.W. Structural and functional assessment of trabecular and cortical bone by micro magnetic resonance imaging. J. Magn. Reson. Imaging 2007, 25, 390–409. [Google Scholar] [CrossRef]
- Lespessailles, E.; Chappard, C.; Bonnet, N.; Benhamou, C.L. Imaging techniques for evaluating bone microarchitecture. Jt. Bone Spine 2006, 73, 254–261. [Google Scholar] [CrossRef]
- Greenspan, H. Super-resolution in medical imaging. Comput. J. 2009, 52, 43–63. [Google Scholar] [CrossRef]
- Park, S.C.; Park, M.K.; Kang, M.G. Super-resolution image reconstruction: A technical overview. IEEE Signal Process. Mag. 2003, 20, 21–36. [Google Scholar] [CrossRef] [Green Version]
- Soomro, T.A.; Afifi, A.J.; Ali Shah, A.; Soomro, S.; Baloch, G.A.; Zheng, L.; Yin, M.; Gao, J. Impact of image enhancement technique on cnn model for retinal blood vessels segmentation. IEEE Access 2019, 7, 158183–158197. [Google Scholar] [CrossRef]
- Munadi, K.; Muchtar, K.; Maulina, N.; Pradhan, B. Image enhancement for tuberculosis detection using deep learning. IEEE Access 2020, 8, 217897–217907. [Google Scholar] [CrossRef]
- Salvador, J. Chapter 2-a taxonomy of example-based super resolution. In Example-Based Super Resolution; Salvador, J., Ed.; Academic Press: New York, NY, USA, 2017; pp. 15–29. [Google Scholar]
- Fleet, D.; Pajdla, T.; Schiele, B.; Tuytelaars, T. Learning a deep convolutional network for image super-resolution. In European Conference on Computer Vision; Springer: Cham, Switzerland, 2014; Volume 8692, pp. 184–199. [Google Scholar]
- Shi, W.; Caballero, J.; Huszar, F.; Totz, J.; Aitken, A.P.; Bishop, R.; Rueckert, D.; Wang, Z. Real-time single image and video super-resolution using an efficient sub-pixel convolutional neural network. In Proceedings of the IEEE Computer Society Conference on Computer Vision and Pattern Recognition, Las Vegas, NV, USA, 27–30 June 2016; pp. 1874–1883. [Google Scholar]
- Kim, J.; Lee, J.K.; Lee, K.M. Deeply-recursive convolutional network for image super-resolution. In Proceedings of the 2016 IEEE Conference on Computer Vision and Pattern Recognition (CVPR), Las Vegas, NV, USA, 27–30 June 2016; pp. 1637–1645. [Google Scholar]
- Kim, J.; Lee, J.K.; Lee, K.M. Accurate image super-resolution using very deep convolutional networks. In Proceedings of the 2016 IEEE Conference on Computer Vision and Pattern Recognition (CVPR), Las Vegas, NV, USA, 27–30 June 2016; pp. 1646–1654. [Google Scholar]
- Pang, Y.; Lin, J.; Qin, T.; Chen, Z. Image-to-image translation: Methods and applications. arXiv 2021, arXiv:2101.08629. [Google Scholar] [CrossRef]
- Arjovsky, M.; Chintala, S.; Bottou, L. Wasserstein gan. arXiv 2017, arXiv:1701.07875. [Google Scholar]
- Isola, P.; Zhu, J.Y.; Zhou, T.; Efros, A.A. Image-to-image translation with conditional adversarial networks. In Proceedings of the 30th IEEE Conference on Computer Vision and Pattern Recognition, CVPR 2017, Honolulu, HI, USA, 21–26 July 2017; pp. 1125–1134. [Google Scholar]
- Zhu, J.Y.; Park, T.; Isola, P.; Efros, A.A. Unpaired image-to-image translation using cycle-consistent adversarial networks. In Proceedings of the 30th IEEE Conference on Computer Vision and Pattern Recognition, CVPR 2017, Honolulu, HI, USA, 21–26 July 2017; pp. 2223–2232. [Google Scholar]
- Radford, A.; Metz, L.; Chintala, S. Unsupervised representation learning with deep convolutional generative adversarial networks. arXiv 2015, arXiv:1511.06434. [Google Scholar]
- Wang, T.C.; Liu, M.Y.; Zhu, J.Y.; Tao, A.; Kautz, J.; Catanzaro, B. High-resolution image synthesis and semantic manipulation with conditional gans. In Proceedings of the IEEE Computer Society Conference on Computer Vision and Pattern Recognition 2018, Salt Lake City, UT, USA, 18–23 June 2018; pp. 8798–8807. [Google Scholar]
- Mattes, D.; Haynor, D.R.; Vesselle, H.; Lewellyn, T.K.; Eubank, W. Nonrigid multimodality image registration. Med. Imaging Image Process. 2001, 4322, 1609–1620. [Google Scholar]
- Liang, X.; Zhang, Z.; Gu, J.; Wang, Z.; Vandenberghe, B.; Jacobs, R.; Yang, J.; Ma, G.; Ling, H.; Ma, X. Comparison of micro-ct and cone beam ct on the feasibility of assessing trabecular structures in mandibular condyle. Dento Maxillo Facial Radiol. 2017, 46, 20160435. [Google Scholar] [CrossRef] [PubMed]
- Keogh, E.; Ratanamahatana, C.A. Exact indexing of dynamic time warping. Knowl. Inf. Syst. 2005, 7, 358–386. [Google Scholar] [CrossRef]
- Bradski, G. The opencv library. Dr. Dobb’s J. Softw. Tools 2000, 25, 120–123. [Google Scholar]
- Wang, S.; Li, J.; Sun, L.; Cai, J.; Wang, S.; Zeng, L.; Sun, S. Application of machine learning to predict the occurrence of arrhythmia after acute myocardial infarction. BMC Med. Inform. Decis. Mak. 2021, 21, 301. [Google Scholar] [CrossRef] [PubMed]
- Burt, P.J.; Adelson, E.H. The laplacian pyramid as a compact image code. In Readings in Computer Vision; Fischler, M.A., Firschein, O., Eds.; Morgan Kaufmann: San Francisco, CA, USA, 1987; pp. 671–679. [Google Scholar]
- Brown, M.; Lowe, D.G. Recognising panoramas. In Proceedings of the 9th IEEE International Conference on Computer Vision, Nice, France, 13–16 October 2003; Volume 1212, pp. 1218–1225. [Google Scholar]
- Huang, X.; Li, Y.; Poursaeed, O.; Hopcroft, J.; Belongie, S. Stacked Generative Adversarial Networks. 2016. Available online: https://arxiv.org/abs/1612.04357 (accessed on 1 August 2021).
- Chen, Q.; Koltun, V. Photographic image synthesis with cascaded refinement networks. In Proceedings of the 2017 IEEE International Conference on Computer Vision (ICCV), Honolulu, HI, USA, 21–26 July 2017. [Google Scholar]
- Wang, Z.; Bovik, A.C.; Sheikh, H.R.; Simoncelli, E.P. Image quality assessment: From error visibility to structural similarity. IEEE Trans. Image Process. 2004, 13, 600–612. [Google Scholar] [CrossRef] [Green Version]
- Heusel, M.; Ramsauer, H.; Unterthiner, T.; Nessler, B.; Hochreiter, S. Gans trained by a two time-scale update rule converge to a local nash equilibrium. Adv. Neural Inf. Process. Syst. 2017, 30, 6626–6637. [Google Scholar]
- Wang, Z.; Simoncelli, E.P.; Bovik, A.C. Multi-scale structural similarity for image quality assessment. In Proceedings of the Conference Record of the Asilomar Conference on Signals, Systems and Computers, Pacific Grove, CA, USA, 9–12 November 2003; pp. 1398–1402. [Google Scholar]
- Domander, R.; Felder, A.; Doube, M. Bonej2—refactoring established research software. Wellcome Open Res. 2021, 6, 37. [Google Scholar] [CrossRef]
- Schindelin, J.; Arganda-Carreras, I.; Frise, E.; Kaynig, V.; Longair, M.; Pietzsch, T.; Preibisch, S.; Rueden, C.; Saalfeld, S.; Schmid, B.; et al. Fiji: An open-source platform for biological-image analysis. Nat. Methods 2012, 9, 676–682. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Rueden, C.T.; Schindelin, J.; Hiner, M.C.; DeZonia, B.E.; Walter, A.E.; Arena, E.T.; Eliceiri, K.W. Imagej2: Imagej for the next generation of scientific image data. BMC Bioinform. 2017, 18, 1–26. [Google Scholar] [CrossRef] [PubMed]
- Schneider, C.A.; Rasband, W.S.; Eliceiri, K.W. Nih image to imagej: 25 years of image analysis. Nat. Methods 2012, 9, 671–675. [Google Scholar] [CrossRef]
- Ridler, T.W.; Calvard, S. Picture thresholding using an iterative selection method. IEEE Trans. Syst. Man Cybern. 1978, 8, 630–632. [Google Scholar]
- Dougherty, R.; Kunzelmann, K.H. Computing local thickness of 3d structures with imagej. Microsc. Microanal. 2007, 13, 1678–1679. [Google Scholar] [CrossRef]
- Simonyan, K.; Zisserman, A. Very deep convolutional networks for large-scale image recognition. arXiv 2014, arXiv:1409.1556; arXiv preprint. [Google Scholar]
- Goldhahn, J.; Suhm, N.; Goldhahn, S.; Blauth, M.; Hanson, B. Influence of osteoporosis on fracture fixation—A systematic literature review. Osteoporos. Int. 2008, 19, 761–772. [Google Scholar] [CrossRef]
- Hoppe, S.; Keel, M.J.B. Pedicle screw augmentation in osteoporotic spine: Indications, limitations and technical aspects. Eur. J. Trauma Emerg. Surg. 2017, 43, 3–8. [Google Scholar] [CrossRef] [PubMed]
- Halvorson, T.L.; Kelley, L.A.; Thomas, K.A.; Whitecloud, T.S.; Cook, S.D. Effects of bone mineral density on pedicle screw fixation. Spine 1994, 19, 2415–2420. [Google Scholar] [CrossRef] [PubMed]
- McCoy, S.; Tundo, F.; Chidambaram, S.; Baaj, A.A. Clinical considerations for spinal surgery in the osteoporotic patient: A comprehensive review. Clin. Neurol. Neurosurg. 2019, 180, 40–47. [Google Scholar] [CrossRef] [PubMed]
| Metrics | Scoring | |
|---|---|---|
| 1 | Contrast between the trabecular bone and bone marrow | 1. Too high or too low and unacceptable; 2. High or low but acceptable; 3. Optimal |
| 2 | Existence of noise | 1. Severe and unacceptable; 2. Marked but acceptable; 3. Moderate; 4. Mild; 5. None or minimal |
| 3 | Sharpness of the trabecular bone | 1. Severe blurring of the images and unacceptable; 2. Marked blurring of the images but acceptable; 3. Moderate blurring of the images; 4. Mild blurring of the images; 5. None or minimal blurring of the images |
| 4 | Obvious overlapping shadows | 1. Severe and unacceptable; 2. Marked but acceptable; 3. Moderate; 4. Mild; 5. None or minimal |
| 5 | Natural shape of the trabecular bone texture | 1. Poor and unacceptable; 2. Marked irregular and unnatural but acceptable; 3. Slightly irregular and unnatural; 4. almost defined and natural; 5. Completely defined and natural |
| Indexes | Methods | Observer | Score | Kendall’s W | p-Value † | ||||
|---|---|---|---|---|---|---|---|---|---|
| 1 | 2 | 3 | 4 | 5 | |||||
| Contrast | Micro-CT | Observer 1 | 0 | 7 | 23 | \ | \ | 0.912 | <0.001 |
| Observer 2 | 0 | 6 | 24 | \ | \ | ||||
| Observer 3 | 0 | 5 | 25 | \ | \ | ||||
| Micro-CT-like | Observer 1 | 0 | 7 | 23 | \ | \ | 0.959 | <0.001 | |
| Observer 2 | 0 | 7 | 23 | \ | \ | ||||
| Observer 3 | 0 | 6 | 24 | \ | \ | ||||
| Noise | Micro-CT | Observer 1 | 0 | 0 | 0 | 6 | 24 | 0.800 | <0.001 |
| Observer 2 | 0 | 0 | 0 | 9 | 21 | ||||
| Observer 3 | 0 | 0 | 0 | 4 | 26 | ||||
| Micro-CT-like | Observer 1 | 0 | 0 | 4 | 8 | 18 | 0.938 | <0.001 | |
| Observer 2 | 0 | 0 | 5 | 5 | 20 | ||||
| Observer 3 | 0 | 0 | 5 | 8 | 17 | ||||
| Sharpness | Micro-CT | Observer 1 | 0 | 0 | 0 | 4 | 26 | 0.817 | <0.001 |
| Observer 2 | 0 | 0 | 0 | 9 | 21 | ||||
| Observer 3 | 0 | 0 | 0 | 8 | 22 | ||||
| Micro-CT-like | Observer 1 | 0 | 0 | 4 | 8 | 18 | 0.888 | <0.001 | |
| Observer 2 | 0 | 0 | 6 | 3 | 21 | ||||
| Observer 3 | 0 | 0 | 5 | 10 | 15 | ||||
| Shadow | Micro-CT | Observer 1 | 0 | 0 | 0 | 0 | 30 | 0.000 | 1.000 |
| Observer 2 | 0 | 0 | 0 | 0 | 30 | ||||
| Observer 3 | 0 | 0 | 0 | 0 | 30 | ||||
| Micro-CT-like | Observer 1 | 0 | 0 | 0 | 0 | 30 | 0.000 | 1.000 | |
| Observer 2 | 0 | 0 | 0 | 0 | 30 | ||||
| Observer 3 | 0 | 0 | 0 | 0 | 30 | ||||
| Texture | Micro-CT | Observer 1 | 0 | 0 | 0 | 4 | 26 | 0.927 | <0.001 |
| Observer 2 | 0 | 0 | 0 | 3 | 27 | ||||
| Observer 3 | 0 | 0 | 0 | 3 | 27 | ||||
| Micro-CT-like | Observer 1 | 0 | 0 | 3 | 6 | 21 | 0.908 | <0.001 | |
| Observer 2 | 0 | 0 | 2 | 4 | 23 | ||||
| Observer 3 | 0 | 0 | 2 | 4 | 24 | ||||
| Micro-CT (n = 30) | Micro-CT-like (n = 30) | p-Value † | |
|---|---|---|---|
| Contrast | 2.8 ± 0.402 | 2.78 ± 0.418 | 0.716 |
| Noise | 4.79 ± 0.410 | 4.46 ± 0.752 | 0.002 |
| Sharpness | 4.77 ± 0.425 | 4.43 ± 0.765 | 0.004 |
| Shadow | 5.00 ± 0.00 | 5.00 ± 0.00 | 1.000 |
| Texture | 4.89 ± 0.316 | 4.68 ± 0.615 | 0.013 |
| n = 50 | Micro-CT | Micro-CT-like Images | p-Value † | Correlation Coefficient of Micro-CT-like and Micro-CT Images (R) | p-Value ‡ |
|---|---|---|---|---|---|
| BV/TV | 0.180 ± 0.016 | 0.175 ± 0.034 | 0.101 | 0.920 | <0.001 |
| Tb.Th (mm) | 0.220 ± 0.012 | 0.179 ± 0.027 | <0.001 | 0.905 | <0.001 |
| Tb.Sp (mm) | 0.934 ± 0.126 | 0.758 ± 0.479 | 0.002 | 0.885 | <0.001 |
| n = 50 | Micro-CT | MDCT | p-Value † | Correlation Coefficient of MDCT and Micro-CT Images (R) | p-Value ‡ |
|---|---|---|---|---|---|
| BV/TV | 0.180 ± 0.016 | 0.320 ± 0.067 | <0.001 | 0.514 | <0.001 |
| Tb.Th (mm) | 0.220 ± 0.012 | 0.680 ± 0.079 | <0.001 | 0.445 | <0.001 |
| Tb.Sp (mm) | 0.934 ± 0.126 | 0.870 ± 0.140 | <0.001 | 0.539 | <0.001 |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Jin, D.; Zheng, H.; Zhao, Q.; Wang, C.; Zhang, M.; Yuan, H. Generation of Vertebra Micro-CT-like Image from MDCT: A Deep-Learning-Based Image Enhancement Approach. Tomography 2021, 7, 767-782. https://doi.org/10.3390/tomography7040064
Jin D, Zheng H, Zhao Q, Wang C, Zhang M, Yuan H. Generation of Vertebra Micro-CT-like Image from MDCT: A Deep-Learning-Based Image Enhancement Approach. Tomography. 2021; 7(4):767-782. https://doi.org/10.3390/tomography7040064
Chicago/Turabian StyleJin, Dan, Han Zheng, Qingqing Zhao, Chunjie Wang, Mengze Zhang, and Huishu Yuan. 2021. "Generation of Vertebra Micro-CT-like Image from MDCT: A Deep-Learning-Based Image Enhancement Approach" Tomography 7, no. 4: 767-782. https://doi.org/10.3390/tomography7040064
APA StyleJin, D., Zheng, H., Zhao, Q., Wang, C., Zhang, M., & Yuan, H. (2021). Generation of Vertebra Micro-CT-like Image from MDCT: A Deep-Learning-Based Image Enhancement Approach. Tomography, 7(4), 767-782. https://doi.org/10.3390/tomography7040064





