Thoracic Aorta Calcium Detection and Quantification Using Convolutional Neural Networks in a Large Cohort of Intermediate-Risk Patients
Abstract
1. Introduction
2. Materials and Methods
2.1. Study Subjects and Image Acquisition
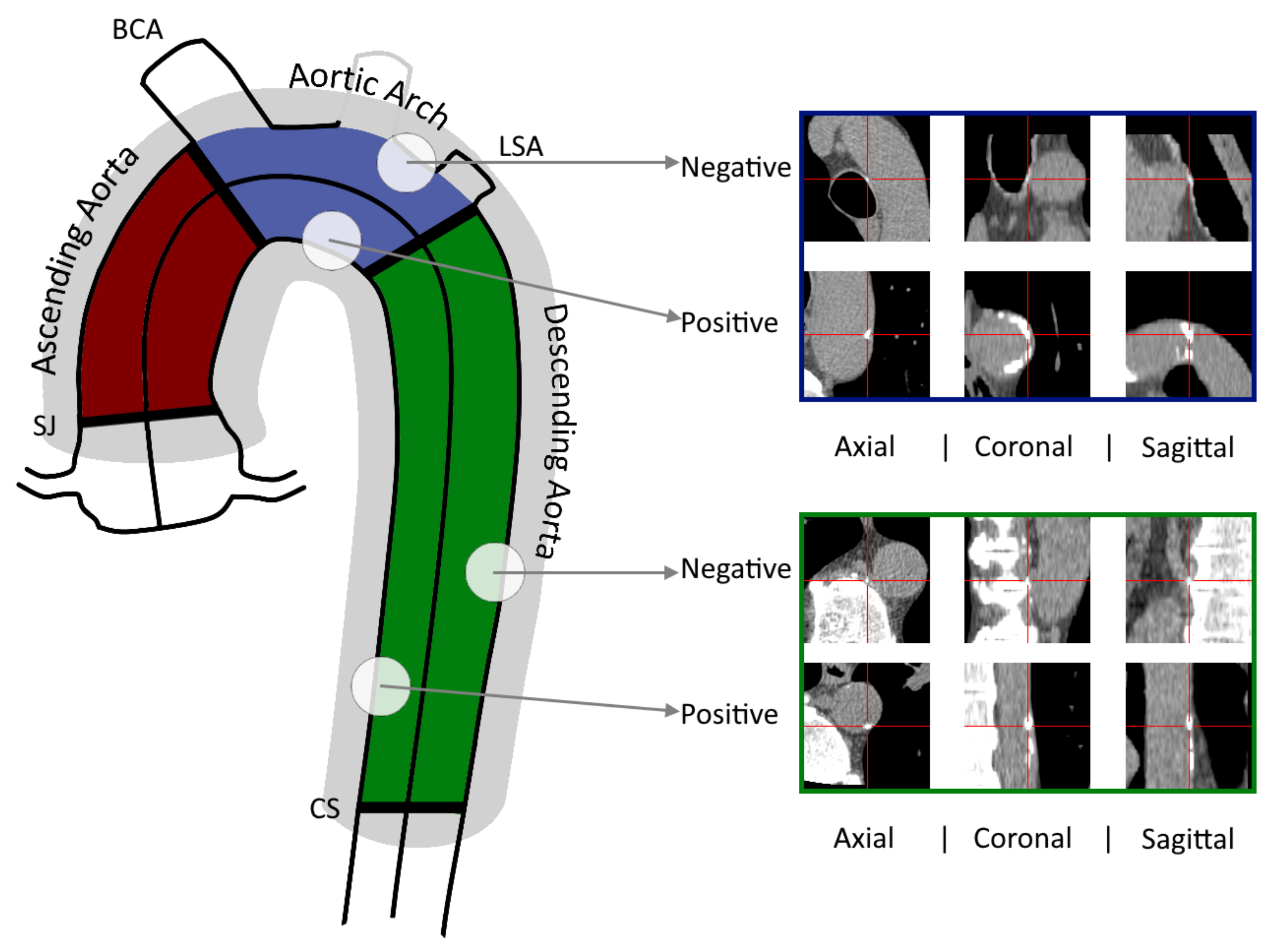
2.2. Detection of Aortic Calcifications
2.3. Datasets and Image processing
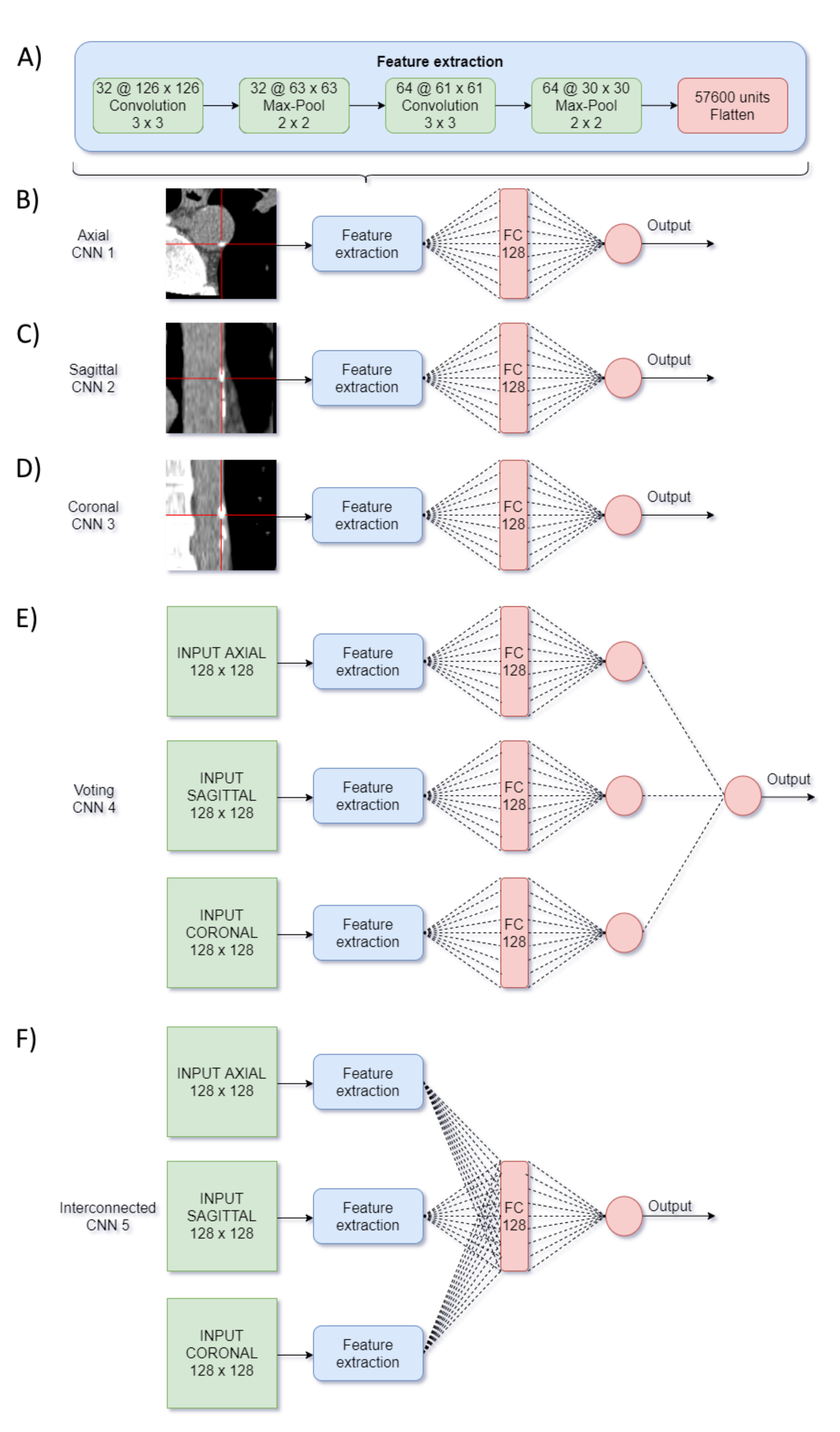
2.4. CNN Design
- Network #4 was based on [17] and was named “voting-network” because the processing of the three images remained independent until the last layer, where a neuron obtained as input a single value from each one, and hence voted among them.
- Network #5 was named "interconnected-network" and was inspired by in [18]. It combined the information of the three processed images by a concatenation and a FC layer.
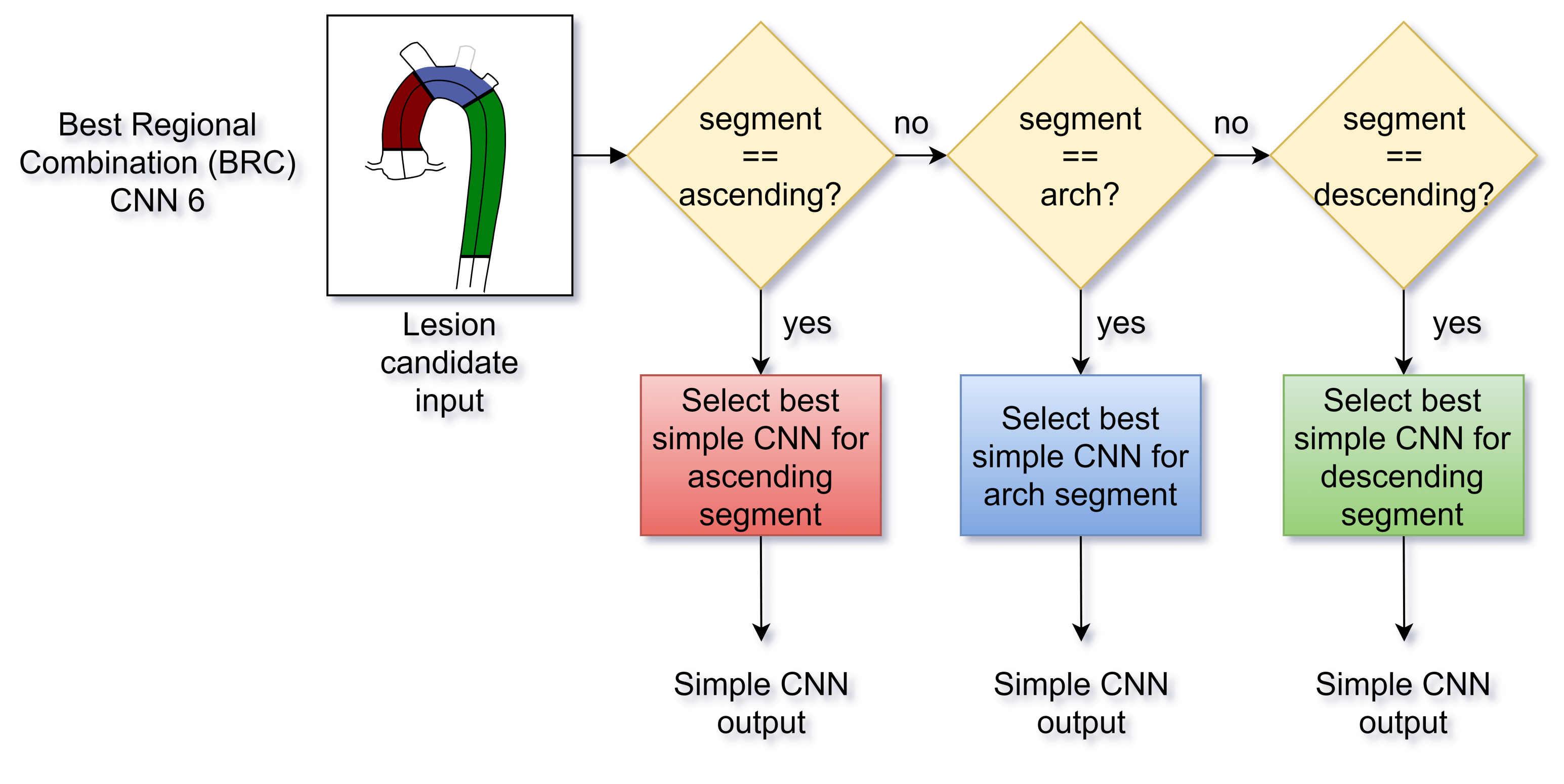
- Network #6 was named the Best Regional Combination (BRC) network and was conceived to improve the independent prediction of the single-input CNNs from the first group with a lesion location feature. Accordingly, using the 10 validation sets, the F1-Score of the 3 trained networks number 1-to-3 was compared with respect to the position of the candidate lesions in terms of the aortic region (ascending, arch and descending aorta). Then, the BRC network classified the lesions choosing either the CNN number #1, #2 or #3 depending on the candidate location (Figure 3).
2.5. Statistics and Evaluation Metrics
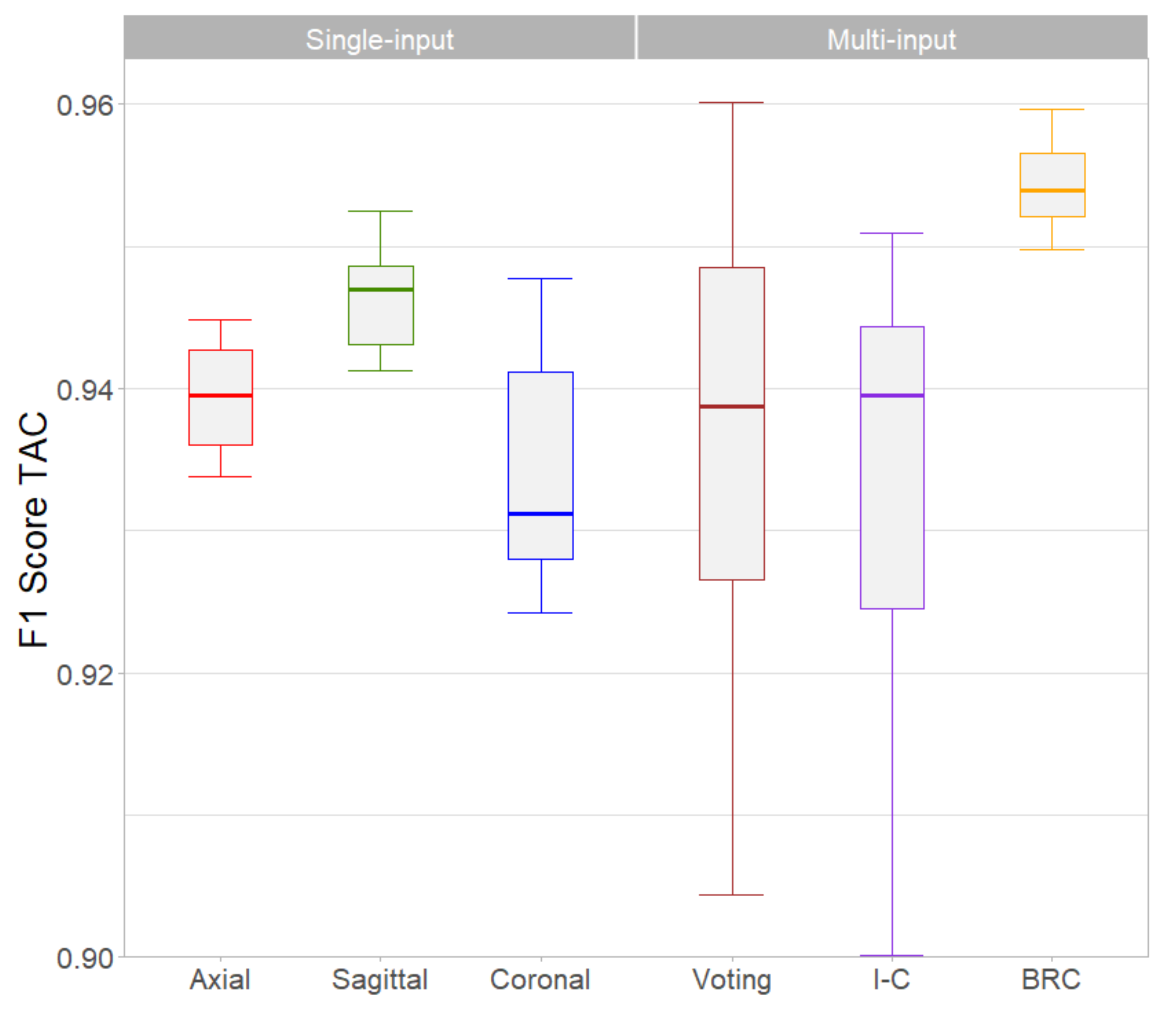
3. Results
4. Discussion
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Demer, L.L.; Tintut, Y. Vascular calcification: Pathobiology of a multifaceted disease. Circulation 2008, 117, 2938–2948. [Google Scholar] [CrossRef] [PubMed]
- Allison, M.A.; Hsi, S.; Wassel, C.L.; Morgan, C.; Ix, J.H.; Wright, C.M.; Criqui, M.H. Calcified atherosclerosis in different vascular beds and the risk of mortality. Arterioscler. Thromb. Vasc. Biol. 2012, 32, 140–146. [Google Scholar] [CrossRef] [PubMed]
- Agatston, A.S.; Janowitz, W.R.; Hildner, F.J.; Zusmer, N.R.; Viamonte, M.; Detrano, R. Quantification of coronary artery calcium using ultrafast computed tomography. J. Am. Coll. Cardiol. 1990, 15, 827–832. [Google Scholar] [CrossRef]
- Alluri, K.; Joshi, P.H.; Henry, T.S.; Blumenthal, R.S.; Nasir, K.; Blaha, M.J. Scoring of coronary artery calcium scans: History, assumptions, current limitations, and future directions. Atherosclerosis 2015, 239, 109–117. [Google Scholar] [CrossRef]
- Budoff, M.J.; Nasir, K.; Katz, R.; Takasu, J.; Carr, J.J.; Wong, N.D.; Allison, M.; Lima, J.A.; Detrano, R.; Blumenthal, R.S.; et al. Thoracic aortic calcification and coronary heart disease events: The multi-ethnic study of atherosclerosis (MESA). Atherosclerosis 2011, 215, 196–202. [Google Scholar] [CrossRef]
- Craiem, D.; Chironi, G.; Casciaro, M.E.; Graf, S.; Simon, A. Calcifications of the thoracic aorta on extended non-contrast-enhanced cardiac CT. PLoS ONE 2014, 9, e109584. [Google Scholar]
- Amarenco, P.; Cohen, A.; Tzourio, C.; Bertrand, B.; Hommel, M.; Besson, G.; Chauvel, C.; Touboul, P.J.; Bousser, M.G. Atherosclerotic disease of the aortic arch and the risk of ischemic stroke. N. Engl. J. Med. 1994, 331, 1474–1479. [Google Scholar] [CrossRef]
- Craiem, D.; Casciaro, M.; Pascaner, A.; Soulat, G.; Guilenea, F.; Sirieix, M.E.; Simon, A.; Mousseaux, E. Association of calcium density in the thoracic aorta with risk factors and clinical events. Eur. Radiol. 2020, 30, 3960–3967. [Google Scholar] [CrossRef]
- Craiem, D.; Chironi, G.; Casciaro, M.E.; Sirieix, M.E.; Mousseaux, E.; Simon, A. Association of thoracic aorta calcium and non cardiac vascular events in cardiac disease-free individuals. Atherosclerosis 2016, 245, 22–27. [Google Scholar] [CrossRef]
- Hecht, H.S.; Cronin, P.; Blaha, M.J.; Budoff, M.J.; Kazerooni, E.A.; Narula, J.; Yankelevitz, D.; Abbara, S. 2016 SCCT/STR guidelines for coronary artery calcium scoring of noncontrast noncardiac chest CT scans: A report of the Society of Cardiovascular Computed Tomography and Society of Thoracic Radiology. J. Cardiovasc. Comput. Tomogr. 2017, 11, 74–84. [Google Scholar] [CrossRef]
- Weininger, M.; Ritz, K.S.; Schoepf, U.J.; Flohr, T.G.; Vliegenthart, R.; Costello, P.; Hahn, D.; Beissert, M. Interplatform reproducibility of CT coronary calcium scoring software. Radiology 2012, 265, 70–77. [Google Scholar] [CrossRef]
- Abukhaled, N.F.; Alskaini, A.M.; Alziadey, H.; Suleiman, I.; Alsaileek, A.; Al-Mallah, M.H. The reproducibility of coronary calcium scoring on multiple software platforms. J. Saudi Heart Assoc. 2013, 25, 151–152. [Google Scholar] [CrossRef][Green Version]
- Arnett, D.K.; Blumenthal, R.S.; Albert, M.A.; Buroker, A.B.; Goldberger, Z.D.; Hahn, E.J.; Himmelfarb, C.D.; Khera, A.; Lloyd-Jones, D.; McEvoy, J.W.; et al. 2019 ACC/AHA guideline on the primary prevention of cardiovascular disease: A report of the American College of Cardiology/American Heart Association Task Force on Clinical Practice Guidelines. J. Am. Coll. Cardiol. 2019, 74, e177–e232. [Google Scholar] [CrossRef]
- Santini, G.; Della Latta, D.; Martini, N.; Valvano, G.; Gori, A.; Ripoli, A.; Susini, C.L.; Landini, L.; Chiappino, D. An automatic deep learning approach for coronary artery calcium segmentation. In EMBEC & NBC 2017; Springer: Berlin/Heidelberg, Germany, 2017; pp. 374–377. [Google Scholar]
- Shadmi, R.; Mazo, V.; Bregman-Amitai, O.; Elnekave, E. Fully-convolutional deep-learning based system for coronary calcium score prediction from non-contrast chest CT. In Proceedings of the 2018 IEEE 15th International Symposium on Biomedical Imaging (ISBI 2018), Washington, DC, USA, 4–7 April 2018; pp. 24–28. [Google Scholar]
- Išgum, I.; Rutten, A.; Prokop, M.; Staring, M.; Klein, S.; Pluim, J.P.; Viergever, M.A.; van Ginneken, B. Automated aortic calcium scoring on low-dose chest computed tomography. Med. Phys. 2010, 37, 714–723. [Google Scholar] [CrossRef]
- Lessmann, N.; Išgum, I.; Setio, A.A.; de Vos, B.D.; Ciompi, F.; de Jong, P.A.; Oudkerk, M.; Willem, P.T.M.; Viergever, M.A.; van Ginneken, B. Deep convolutional neural networks for automatic coronary calcium scoring in a screening study with low-dose chest CT. In Proceedings of the Medical Imaging 2016: Computer-Aided Diagnosis, San Diego, CA, USA, 27 February–3 March 2016; Volume 9785, p. 978511. [Google Scholar]
- Lessmann, N.; van Ginneken, B.; Zreik, M.; de Jong, P.A.; de Vos, B.D.; Viergever, M.A.; Išgum, I. Automatic calcium scoring in low-dose chest CT using deep neural networks with dilated convolutions. IEEE Trans. Med. Imaging 2017, 37, 615–625. [Google Scholar] [CrossRef]
- Cano-Espinosa, C.; González, G.; Washko, G.R.; Cazorla, M.; Estépar, R.S.J. Automated Agatston score computation in non-ECG gated CT scans using deep learning. In Proceedings of the Medical Imaging 2018: Image Processing, Houston, TX, USA, 10–15 February 2018; Volume 10574, p. 105742K. [Google Scholar]
- Chellamuthu, K.; Liu, J.; Yao, J.; Bagheri, M.; Lu, L.; Sandfort, V.; Summers, R.M. Atherosclerotic vascular calcification detection and segmentation on low dose computed tomography scans using convolutional neural networks. In Proceedings of the 2017 IEEE 14th International Symposium on Biomedical Imaging (ISBI 2017), Melbourne, VIC, Australia, 18–21 April 2017; pp. 388–391. [Google Scholar]
- van Velzen, S.G.; Lessmann, N.; Velthuis, B.K.; Bank, I.E.; van den Bongard, D.H.; Leiner, T.; de Jong, P.A.; Veldhuis, W.B.; Correa, A.; Terry, J.G.; et al. Deep learning for automatic calcium scoring in CT: Validation using multiple cardiac CT and chest CT protocols. Radiology 2020, 295, 66–79. [Google Scholar] [CrossRef]
- Graffy, P.M.; Liu, J.; O’Connor, S.; Summers, R.M.; Pickhardt, P.J. Automated segmentation and quantification of aortic calcification at abdominal CT: Application of a deep learning-based algorithm to a longitudinal screening cohort. Abdom. Radiol. 2019, 44, 2921–2928. [Google Scholar] [CrossRef]
- Liu, J.; Lu, L.; Yao, J.; Bagheri, M.; Summers, R.M. Pelvic artery calcification detection on CT scans using convolutional neural networks. In Proceedings of the Medical Imaging 2017: Computer-Aided Diagnosis, Orlando, FL, USA, 11–16 February 2017; Volume 10134, p. 101341A. [Google Scholar]
- Brains, J. PyCharm. 2018. Available online: https://www.jetbrains.com/pycharm (accessed on 1 October 2019).
- Wolterink, J.M.; Leiner, T.; Viergever, M.A.; Išgum, I. Automatic coronary calcium scoring in cardiac CT angiography using convolutional neural networks. In Proceedings of the International Conference on Medical Image Computing and Computer-Assisted Intervention, Munich, Germany, 5–9 October 2015; pp. 589–596. [Google Scholar]
- James, G.; Witten, D.; Hastie, T.; Tibshirani, R. An Introduction to Statistical Learning; Springer: Berlin/Heidelberg, Germany, 2013; Volume 112, pp. 177–178. [Google Scholar]
- Zou, H.; Hastie, T. Regularization and variable selection via the elastic net. J. R. Stat. Soc. Ser. B (Stat. Methodol.) 2005, 67, 301–320. [Google Scholar] [CrossRef]
- Srivastava, N.; Hinton, G.; Krizhevsky, A.; Sutskever, I.; Salakhutdinov, R. Dropout: A simple way to prevent neural networks from overfitting. J. Mach. Learn. Res. 2014, 15, 1929–1958. [Google Scholar]
- Cohen, J. Weighted kappa: Nominal scale agreement provision for scaled disagreement or partial credit. Psychol. Bull. 1968, 70, 213. [Google Scholar] [CrossRef]
- Bartko, J.J. The intraclass correlation coefficient as a measure of reliability. Psychol. Rep. 1966, 19, 3–11. [Google Scholar] [CrossRef] [PubMed]
- Krouwer, J.S.; Monti, K.L. A simple, graphical method to evaluate laboratory assays. Eur. J. Clin. Chem. Clin. Biochem. 1995, 33, 525–528. [Google Scholar] [PubMed]
- Gernaat, S.A.; Van Velzen, S.G.; Koh, V.; Emaus, M.J.; Išgum, I.; Lessmann, N.; Moes, S.; Jacobson, A.; Tan, P.W.; Grobbee, D.E.; et al. Automatic quantification of calcifications in the coronary arteries and thoracic aorta on radiotherapy planning CT scans of Western and Asian breast cancer patients. Radiother. Oncol. 2018, 127, 487–492. [Google Scholar] [CrossRef] [PubMed]
- Kurugol, S.; Come, C.E.; Diaz, A.A.; Ross, J.C.; Kinney, G.L.; Black-Shinn, J.L.; Hokanson, J.E.; Budoff, M.J.; Washko, G.R.; San Jose Estepar, R. Automated quantitative 3D analysis of aorta size, morphology, and mural calcification distributions. Med. Phys. 2015, 42, 5467–5478. [Google Scholar] [CrossRef] [PubMed]
- Noothout, J.M.; De Vos, B.D.; Wolterink, J.M.; Išgum, I. Automatic segmentation of thoracic aorta segments in low-dose chest CT. In Proceedings of the Medical Imaging 2018: Image Processing, Houston, TX, USA, 10–15 February 2018; Volume 10574, p. 105741S. [Google Scholar]

| Clinical Information | Mean, N | (SD), % |
|---|---|---|
| Age, y.o. | 57 | (9) |
| Male gender | 1066 | 75 |
| Hypertension | 661 | 47 |
| Hypercholesterolemia | 1161 | 82 |
| Current or past smoking | 760 | 54 |
| Framingham Risk Score at 10 years, % | 9.5 | 5.6 |
| CVD Risk I (TAC 10) | 592 | 42 |
| CVD Risk II (10 < TAC 100) | 258 | 18 |
| CVD Risk III (100 < TAC 400) | 257 | 18 |
| CVD Risk IV (TAC > 400) | 308 | 22 |
| Aortic | Candidates | Positives | Positives/ | ||
|---|---|---|---|---|---|
| Portion | Number, # | Area, cm | Number, # | Area, cm | Candidates |
| ascending | 1508 (8%) | 333.8 (13%) | 931 (8%) | 133.3 (9%) | 62% |
| arch | 6489 (33%) | 782.3 (29%) | 3168 (26%) | 488.6 (23%) | 49% |
| descending | 11793 (59%) | 1553.1 (58%) | 7942 (66%) | 1025.5 (68%) | 67% |
| Total | 19790 | 2669.2 | 12041 | 1647.4 | 61% |
| Axial | Sagittal | Coronal | Voting | Interconnected | BRC | |
|---|---|---|---|---|---|---|
| True positives, n (%) | 1095 | 1124 | 1136 | 1128 | 1129 | 1151 |
| (91.3%) | (93.7%) | (94.7%) | (94.0%) | (94.1%) | (95.9%) | |
| False positives, n (%) | 148 | 143 | 182 | 140 | 158 | 174 |
| (12.3%) | (11.9%) | (15.2%) | (11.7%) | (13.2%) | (14.5%) | |
| True positive Area | 96.8 | 99.4 | 100.4 | 100 | 100.3 | 101.0 |
| per patient, mm (%) | (93.9%) | (96.4%) | (97.5%) | (97.1%) | (97.3%) | (98.0%) |
| False positive Area | 7.94 | 9.4 | 13.4 | 9.8 | 10.0 | 9.1 |
| per patient, mm (%) | (7.7%) | (9.1%) | (13.0%) | (9.5%) | (9.7%) | (8.8%) |
| True positive TAC | 371 | 381 | 385 | 384 | 385 | 387 |
| per patient, AS (%) | (94.4%) | (97.0%) | (98.0%) | (97.8%) | (97.9%) | (98.4%) |
| False positive TAC | 26 | 32 | 48 | 34 | 34 | 30 |
| per patient, AS (%) | (6.7%) | (8.0%) | (12.1%) | (8.7%) | (8.7%) | (7.8%) |
| TAC Sensitivity | 0.944 | 0.970 | 0.979 | 0.978 | 0.979 | 0.984 |
| TAC F1-Score | 0.938 | 0.946 | 0.933 | 0.947 | 0.947 | 0.954 |
| TAC Risk Kappa, | 0.879 | 0.863 | 0.846 | 0.868 | 0.850 | 0.878 |
| value (95% CI) | (0.822–0.935) | (0.802–0.924) | (0.766–0.915) | (0.808–0.928) | (0.786–0.915) | (0.821–0.936) |
| TAC Score ICC, | 0.991 | 0.996 | 0.983 | 0.995 | 0.995 | 0.998 |
| value (95% CI) | (0.987–0.993) | (0.994–0.997) | (0.976–0.988) | (0.993–0.997) | (0.992–0.996) | (0.997–0.999) |
| CNN Prediction | ||||||
|---|---|---|---|---|---|---|
| I | II | III | IV | Total | ||
| Reference | I | 42 | 10 | 3 | 0 | 55 |
| II | 1 | 34 | 3 | 0 | 38 | |
| III | 0 | 0 | 21 | 1 | 22 | |
| IV | 0 | 0 | 0 | 26 | 26 | |
| Total | 43 | 44 | 27 | 27 | 141 | |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Guilenea, F.N.; Casciaro, M.E.; Pascaner, A.F.; Soulat, G.; Mousseaux, E.; Craiem, D. Thoracic Aorta Calcium Detection and Quantification Using Convolutional Neural Networks in a Large Cohort of Intermediate-Risk Patients. Tomography 2021, 7, 636-649. https://doi.org/10.3390/tomography7040054
Guilenea FN, Casciaro ME, Pascaner AF, Soulat G, Mousseaux E, Craiem D. Thoracic Aorta Calcium Detection and Quantification Using Convolutional Neural Networks in a Large Cohort of Intermediate-Risk Patients. Tomography. 2021; 7(4):636-649. https://doi.org/10.3390/tomography7040054
Chicago/Turabian StyleGuilenea, Federico N., Mariano E. Casciaro, Ariel F. Pascaner, Gilles Soulat, Elie Mousseaux, and Damian Craiem. 2021. "Thoracic Aorta Calcium Detection and Quantification Using Convolutional Neural Networks in a Large Cohort of Intermediate-Risk Patients" Tomography 7, no. 4: 636-649. https://doi.org/10.3390/tomography7040054
APA StyleGuilenea, F. N., Casciaro, M. E., Pascaner, A. F., Soulat, G., Mousseaux, E., & Craiem, D. (2021). Thoracic Aorta Calcium Detection and Quantification Using Convolutional Neural Networks in a Large Cohort of Intermediate-Risk Patients. Tomography, 7(4), 636-649. https://doi.org/10.3390/tomography7040054






