3.1. Proposed Solution Encoding Scheme
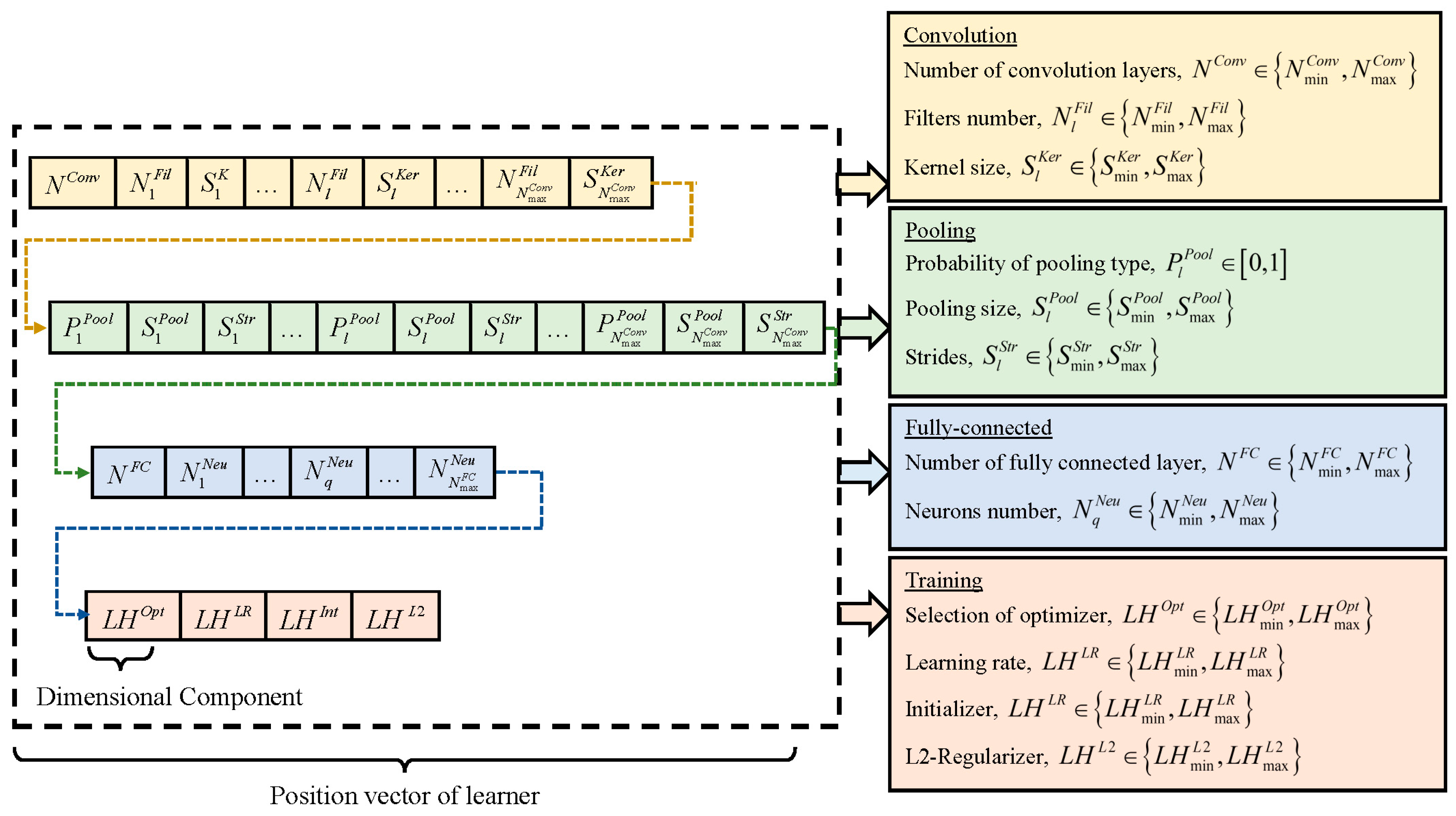
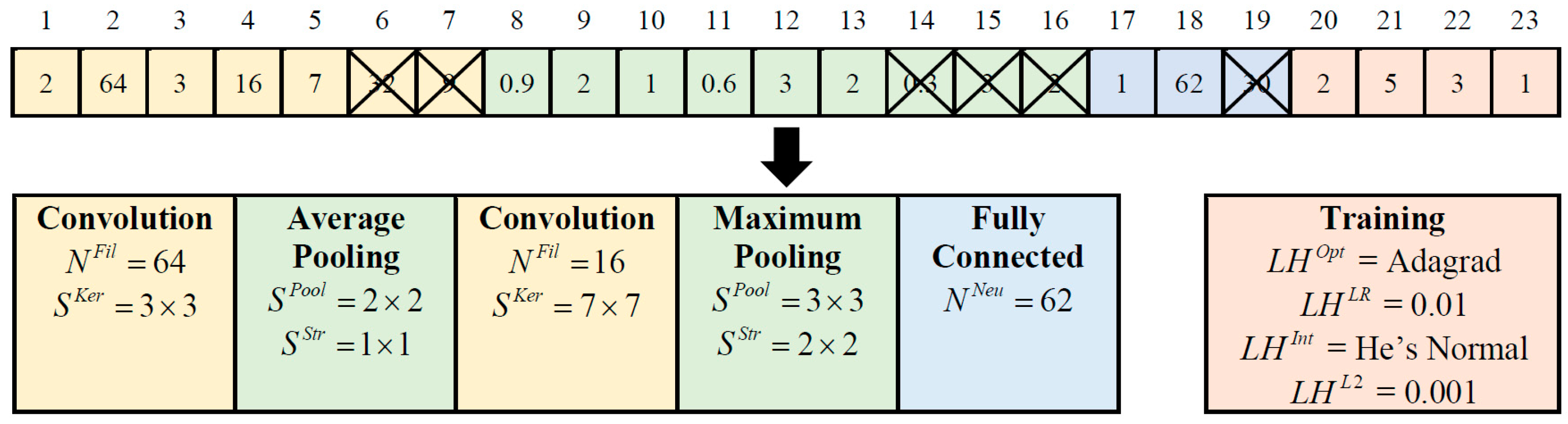
Constructing optimal CNN models involves determining network hyperparameters, including network depth, layer types, kernel size, filter numbers, pooling size, pooling stride, and neuron number. Furthermore, selecting appropriate combinations of learning hyperparameters, such as optimizer type, learning rate, initializer type, and L2 regularization, is vital for optimizing network training and achieving competitive classification performance. In light of these considerations, ETLBOCBL-CNN introduces an efficient solution encoding scheme, which enables each learner to effectively search for optimal network and learning hyperparameters. This scheme ensures that only valid architectures are created without limiting ETLBOCBL-CNN’s ability to discover novel and effective CNN architectures for image classification. As depicted in
Figure 4, each ETLBOCBL-CNN learner, denoted as the
n-th learner, is represented by a
D-dimensional position vector,
. Each
d-th dimension,
, corresponds to specific network or learning hyperparameters required for the construction of a unique CNN architecture. These hyperparameters are divided into four main sections: convolution, pooling, fully connected, and network training.
The CNN’s convolution section is characterized by three key hyperparameters, as illustrated in
Figure 4. The first hyperparameter,
, represents the number of convolution layers and is encoded in
with
, where
and
define the minimum and maximum allowable convolution layer counts. Each convolution layer is identified by an index number,
. For each
l-th convolution layer, there are two additional hyperparameters:
, which signifies the number of filters, and
, which denotes the kernel size of each filter. These hyperparameters are encoded in
, with
for
and
for
, where
. It is worth noting that while all ETLBOCBL-CNN learners possess position vectors
of the same dimension size
D, they can generate CNNs with varying numbers of convolution layers by referring to the value of
encoded in
with
. If
, only the first
values of
and
, encoded into
with
and
for
, are utilized to construct the CNN’s convolution section. Redundant network information stored in
, with
and
for
, is omitted from the network construction process.
Three hyperparameters are introduced to define the pooling section of the CNN. The first hyperparameter, , is encoded into , with for . It signifies the type of pooling layer connected to each l-th convolution layer according to the following guidelines: (a) no pooling layer is inserted when , (b) maximum pooling is applied when , and (c) average pooling is employed when . The minimum and maximum sizes of the pooling layers linked to each l-th convolutional layer are denoted as and , while and represent the minimum and maximum stride sizes of the pooling layer. Two more hyperparameters, and , represent the kernel size and stride size of the pooling layer associated with the l-th convolution layer. These hyperparameters are encoded into , with for and for , where . Similarly, only relevant network information of , , and contributes to the construction of the CNN’s pooling section. In cases where , only the first values of , , and , encoded into with , , and , respectively, for , are used for generating the pooling section of the CNN. Redundant network information of , , and stored in with , , and , respectively, for , is disregarded in network construction. It is essential to note that the network information of and is excluded if the corresponding falls within the range of since no pooling layer is introduced with the l-th convolution layer in this scenario.
The fully connected section of a CNN is constructed using two hyperparameters. The first hyperparameter, , represents the number of fully connected layers in the CNN. It is encoded into , with , where and define the minimum and maximum allowable numbers of fully connected layers, respectively. Each fully connected layer is identified by an index number, . The second hyperparameter, , indicates the number of neurons in the q-th fully connected layer. It is encoded into with and . Here, and represent the minimum and maximum numbers of neurons in a fully connected layer. Similarly to the convolution and pooling sections, only the first values of encoded into with and are used in generating the fully connected section of the CNN. Any redundant information of , stored in with and , is disregarded in network construction.
In addition to the network hyperparameters used in constructing the convolution, pooling, and fully connected sections of the CNN, four learning hyperparameters are integrated into the position vector of each ETLBOCBL-CNN learner. This integration enhances the optimization process, allowing ETLBOCBL-CNN to produce more accurate and effective CNN models. Notably, the optimization of learning hyperparameters in ETLBOCBL-CNN involves selecting the optimizer type, learning rate, initializer type, and L2-regularizer. These learning hyperparameters are represented by integer indices within the ranges of , , , and , respectively. To optimize the CNN training process for each ETLBOCBL-CNN learner, the selection of these learning hyperparameters is represented by the integer decision variables: , , , and . These variables are encoded into with the dimension indices of , , , and , respectively.
The process of constructing a CNN architecture from the network and learning hyperparameters, decoded from the position vector
of the
n-th ETLBOCBL-CNN learner, is visually represented in
Figure 5.
Table 1 summarizes the feasible search ranges for all network and learning hyperparameters based on [
62]. Meanwhile, the selection of the optimizer type, learning rate, initializer type, and L2-regularizer is based on the integer indices presented in
Table 2. In this demonstration, the maximum allowable numbers of convolution and fully connected layers are defined as
and
, respectively. Consequently, the total dimension size of the position vector
for each
n-th ETLBOCBL-CNN learner is calculated as
. To illustrate, the values of
and
encoded into
, with
and
, respectively, indicate that the constructed CNN has two convolution layers, up to two pooling layers, and one fully connected layer. Specifically, the first convolution layer is generated with values
and
, encoded into
with
and
, respectively. The second convolutional layer is established using values
and
encoded into
with
and
. In the pooling section of the CNN, a maximum pooling layer is inserted into the first convolution layer. This is achieved via the values of
,
, and
encoded into
with
to
. An averaging pooling layer is inserted into the second convolution layer based on values
,
, and
encoded into
with
to
. The fully connected layer of the CNN comprises 18 neurons, as indicated by the value of
encoded into
with
. The trainable parameters of the constructed CNN are initialized using He Normal and optimized using Adagrad with a learning rate of 0.01 and an L2-regularization value of 0.001. This information is revealed through the integer indices of
,
,
, and
encoded into
with
to
. It is important to note that certain network information, specifically related to the third convolutional layer, the third pooling layer, and the second fully connected layer, is excluded from the network construction.
3.3. Fitness Evaluation of ETLBOCBL-CNN
To assess each ETLBOCBL-CNN learner’s fitness, a two-step process is employed as detailed in Algorithm 2. In the first step, a potential CNN architecture is constructed and trained using the training set. The second step evaluates this trained CNN architecture using the validation set. The fitness of each learner is quantified by measuring the classification error of its respective CNN architecture, where lower error values indicate superior fitness. In this paper, ETLBOCBL-CNN’s objective is to discover the CNN model capable of achieving the minimum classification errors when solving given datasets.
The CNN model’s configuration is determined by the network and learning hyperparameters decoded from its corresponding position vector,. These hyperparameters encompass , , , , , , , , , , and , with and . Additionally, the CNN architecture is inserted with a fully connected layer containing an output neuron number matching the number of classes, denoted as , for classification purposes.
Referring to the learning hyperparameter
, which is decoded from
with
, a weight initializer is selected. This initializer is responsible for initializing the trainable parameters of all the convolutional and fully connected layers within the CNN. These weight parameters are denoted as
. Define
as the training dataset, which contains
samples and is used to train the potential CNN architecture constructed by every learner. To train each CNN architecture, multiple training steps of
are executed with a predefined batch size of
. During these training steps, the training data are input into the network in batches, i.e.,
Next, an optimizer is chosen based on the learning hyperparameter
decoded from
with
. This optimizer is employed to train the compiled CNN across a predetermined epoch number
, performed on
batches of data obtained from
. At each
i-th training step, where
, the cross-entropy loss function of CNN is obtained as
based on the current weight parameters
and the
i-th batch of data,
. Let
represents the learning rate determined based on the learning hyperparameter
decoded from
with
. Additionally,
refers to the gradient of cross-entropy loss. The new weight parameters
for the CNN model are then updated as follows:
The performance of a CNN is assessed with a validation dataset
of size
after the training. This evaluation process is carried out in
steps, i.e.,
In every
j-th step of evaluation, various batches from the validation dataset,
, are utilized to assess the trained CNN models. This results in distinct classification errors, denoted as
, where
. The mean classification error of the trained CNN model, considering all
batches of data in
, is calculated to derive the fitness value of each
n-th learner, i.e.,
, as follows:
Finding the best CNN architecture for a given dataset with ETLBOCBL-CNN poses a considerable challenge given the time-consuming nature of exhaustive training and evaluation for each potential solution. While exploring numerous alternatives is crucial for enhancing solutions in MSA-based methods like ETLBOCBL-CNN, the exhaustive training of each learner on
with a large
is often impractical due to the substantial computational load involved. To address this challenge, a fitness approximation method is employed. It involves training the potential CNN architecture represented by each learner using a reduced training epoch (e.g.,
) during fitness evaluation. This approach, while potentially leading to less precise evaluations, significantly alleviates the computational burden. The primary aim of the selection operator is to identify the next generation of the population through fair comparisons among the learners, rather than achieving precise fitness evaluations for each learner. Additionally, a potential CNN architecture demonstrating superior performance in the initial epochs is more likely to exhibit a competitive classification error in the final stage. Upon completing the search process with ETLBOCBL-CNN, the optimal CNN architecture, constructed based on network and learning hyperparameters decoded from the teacher solution, can be thoroughly trained with a higher
to obtain its final classification error.
| Algorithm 2: Fitness Evaluation of ETLBOCBL-CNN |
| Inputs: , , , , , , |
| 01: | Construct a candidate CNN architecture based on the network and learning hyperparameters decoded from and insert a fully connected layer with output neurons; |
| 02: | Compute and using Equations (4) and (6), respectively; |
| 03: | Generate the initial weights of the CNN model as using the selected weight initializer; |
| 04: | for to do |
| 05: | for to do |
| 06: | Calculate of CNN model; |
| 07: | Update the weights based on Equation (5); |
| 08: |
end for |
| 09: | end for |
| 10: | for to do |
| 11: | Classify the dataset using the trained CNN model; |
| 12: | Record the classification errors for solving the dataset as ; |
| 13: | end for |
| 14: | Calculate of the candidate CNN architecture built from with Equation (7); |
| Output:
|
3.5. Modified Learner Phase of ETLBOCBL-CNN
To encourage exploration and prevent convergence toward local optima, the original TLBO employs a repelling mechanism within its single peer interaction, as seen in Equation (3). However, the effectiveness of this mechanism diminishes over iterations, particularly as the population converges. This renders it inadequate for complex problems like automatic network architecture design. Furthermore, the single peer interaction neglects the dynamics of interactions among multiple peers in a classroom, interactions that foster more efficient knowledge enhancement and the inclination of learners to preserve their original useful knowledge. To rectify these shortcomings, the modified learning phase of ETLBOCBL-CNN introduces a stochastic peer interaction scheme, aiming to enhance its performance in the discovery of optimal CNN architectures.
In the modified learner phase of ETLBOCBL-CNN, a stochastic peer learning scheme is introduced. This scheme enables each learner to interact with different peers randomly, fostering the creation of new CNN architectures. The stochastic nature of these interactions allows ETLBOCBL-CNN to escape local optima and discover more diverse solutions. Moreover, by promoting interactions among multiple peers, this phase mimics the intricate learning dynamics found in classrooms, facilitating more effective knowledge exchange and retention.
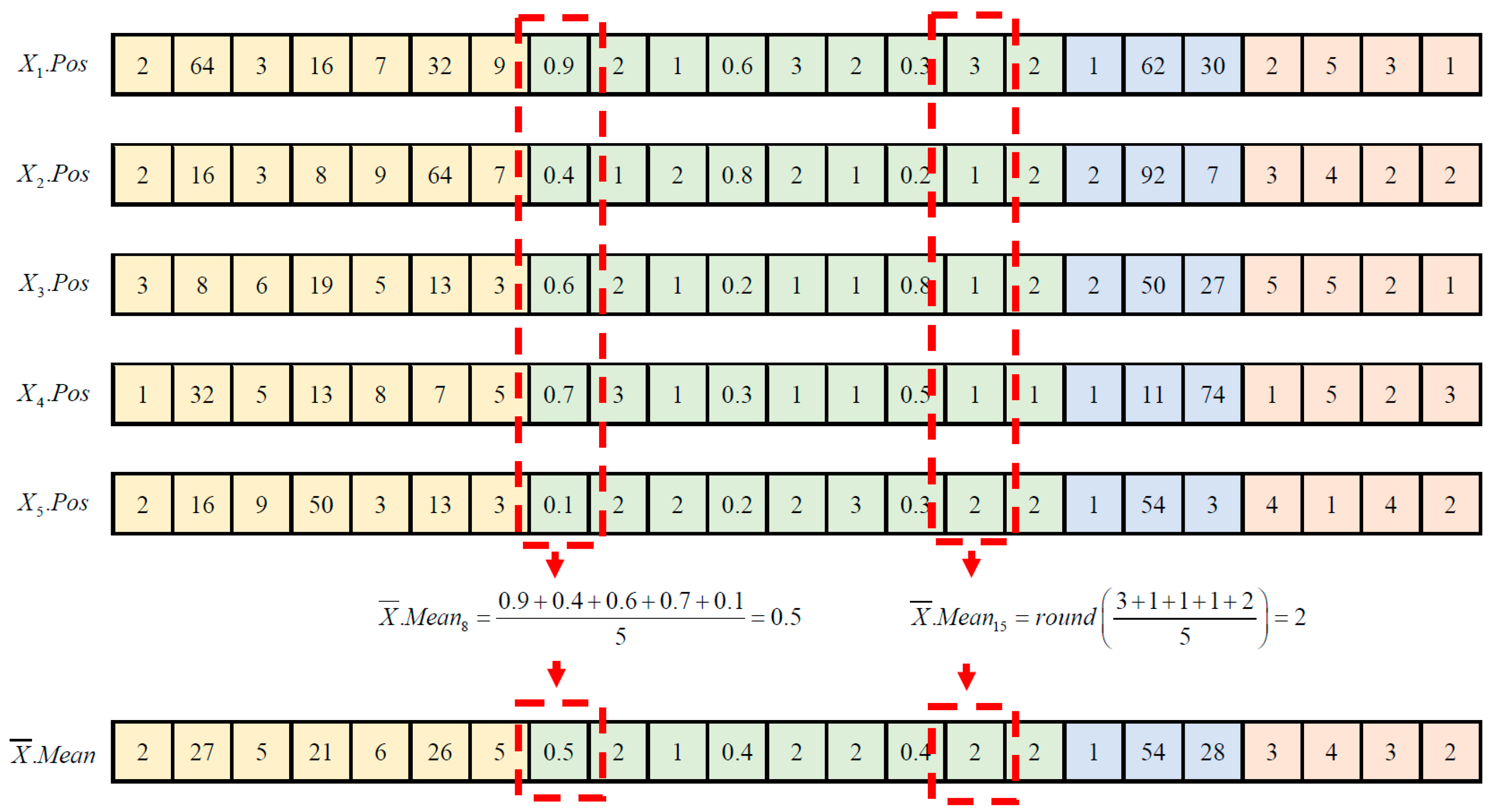
After completing the modified teacher phase, a clone population, denoted as , is formed by duplicating the offspring population and sorting it in ascending order based on its fitness values, denoted as for . From , two subsets of the population, and , are created to store the top 20% and top 50% of learners from the offspring population, respectively. In the context of the stochastic peer interaction scheme, three distinct strategies are employed to update the d-th dimension of the position vector for each n-th offspring learner, . The strategy applied is determined by a random variable . Specifically, (a) if , a multiple peer interaction is triggered to update ; (b) if , a modified single peer interaction is employed to update ; and (c) if , the original value of is retained.
Suppose two top-performing offspring learners, denoted as
and
, are randomly chosen from
, where
. If the random variable
rand falls within the range of 0 to 1/3, the multiple peer interaction condition is triggered to update for the
d-th component of the
n-th learner,
, as follows:
where
are the random numbers obtained from the uniform distribution.
Let
be a top-performing learner randomly chosen from
, and it is utilized to update the
d-th dimension of the
n-th offspring learner, i.e.,
, where
. This update occurs through a modified single peer interaction scheme when the random variable
rand falls in the range of 1/3 to 2/3. In this modified scheme,
is attracted towards
if
. In contrary,
is repelled from
if
. The formulation of this modified single peer interaction scheme for updating
in each
d-th dimension of the
n-th learner is as follows:
where
is a random number obtained from the uniform distribution.
The stochastic peer interaction scheme, introduced in the modified learner phase of ETLBOCBL-CNN and detailed in Algorithm 5, allows for unique updates in each dimension of the learners. These updates can involve multiple peer interactions, modified single peer interactions, or the retention of the original values, facilitating the generation of diverse candidate solutions and enhancing search capabilities. All dimensions of the updated are subject to a rounding operation using , except for those corresponding to for because they represent the selection probabilities of pooling layers associated with the l-th convolutional layer. Subsequently, Algorithm 2 is used to assess the fitness of each updated offspring learner, resulting in the computation of their classification error, . If the CNN architecture represented by the updated yields lower classification error than , the teacher solution is replaced by the n-th updated offspring learner .
3.6. Tri-Criterion Selection Scheme
In any optimization process employing MSAs, the choice of the selection scheme for constructing the next-generation population is pivotal. Conventional selection methods, like greedy selection and tournament selection, rely solely on the fitness values of solutions to determine their survival. For example, the original TLBO uses a greedy selection scheme to compare the fitness values of existing learners with those of new learners generated through teacher and learner phases. While these fitness-based selection schemes are straightforward to implement, they have the drawback of rejecting potentially valuable solutions with temporarily inferior fitness values that could substantially enhance the overall population quality over time. To address this limitation, the ETLBOCBL-CNN introduces a tri-criterion selection scheme. This scheme not only takes into account the fitness of learners but also factors in their diversity and improvement rate.
| Algorithm 5: Stochastic Peer Interaction in ETLBOCBL-CNN’s Modified Teacher Phase |
| Inputs: , , , , , , , , , |
| 01: | Initialize clone population set as ; |
| 02: | Construct by duplicating and sorting the offspring learners ascendingly by referring to their fitness values of for ; |
| 03: | Construct and by extracting the top 20% and 50% of offspring learners stored in ; |
| 04: | for to do |
| 05: | for to D do |
| 06: | Randomly generate from uniform distribution; |
| 07: | if then |
| 08: | Randomly select and from , where ; |
| 09: | Update using Equation (14); |
| 10: | else if then |
| 11: | Randomly select from , where ; |
| 12: | Update using Equation (15); |
| 13: | else if then |
| 14: | Retain the original value of ; |
| 15: |
end if |
| 16: | if with then |
| 17: | ; |
| 18: |
end if |
| 19: |
end for |
| 20: | Perform fitness evaluation on the updated to obtain new using Algorithm 2; |
| 21: | if then |
| 22: | , ; |
| 23: |
end if |
| 24: | end for |
| Output: Updated and |
After completing the modified learner phase, each
n-th offspring learner’s fitness in the updated population
is compared with that of its corresponding
n-th original learner from
. The improvement rate for each
n-th offspring learner is then determined as follows:
Here, in the numerator represents the change in fitness between the original and offspring learners, while in the denominator quantifies the Euclidean distance between these two learners. A positive indicates that the n-th offspring learner can yield a CNN architecture with a lower classification error compared to its original counterpart. The magnitude of plays a crucial role in assessing the effectiveness of each offspring learner in enhancing population quality. Higher values of suggest that the n-th offspring learner has achieved substantial improvement in classification error with a relatively small traversal in the solution space. This signifies that the offspring learner possesses valuable information for constructing a robust CNN architecture that is worth inheriting in the next generation. Notably, the improvement rate of each n-th original learner in is set to for , as these original learners serve as the baseline for comparison with their respective offspring learners.
After calculating the improvement rates for all offspring learners, the subsequent action involves creating a merged population
by combining the original
with the updated
. The total population size of
is 2
N and is represented as follows:
Each
n-th solution member in
, designated as
, can originate from either an original learner in
or an offspring learner in
. These solution members in
are subsequently arranged in ascending order based on the classification error of their corresponding CNN architecture, represented by
. Additionally,
indicates the Euclidean distance between the CNN architecture represented by the
n-th solution member in
(i.e.,
) and the current best CNN architecture, which is represented by the first solution member (i.e.,
), where
A tri-criterion selection scheme is designed to determine the next population of ETLBOCBL-CNN. This selection is based on the fitness, diversity, and improvement rate of each n-th solution within , denoted as , , and values, respectively, for . For the construction of the population in the next generation, a randomly generated integer is used. It serves to select the first solution members from , focusing on the fitness criterion. These solution members are directly selected from the subset of with the best values.
The diversity criterion is then applied to select the next
solution members for
, with
being a randomly generated integer. The solution members in
with population indices
, that were not initially chosen for
, are flagged with
, indicating their non-selection. Subsequently, the weighted fitness value
is computed for the remaining
) solution members in
with population indices
, taking into account their classification error (
) and diversity (
) values, i.e.,
The weight factor
is stochastically generated from a normal distribution with a mean of 0.9 and a standard deviation of 0.05, and it is constrained to fall within the range of 0.8 to 1.0 to maintain a balance between diversity and other selection factors. Let
and
represent the largest and smallest Euclidean distances measured from the best solution member
, respectively, while
and
denote the worst and best fitness values observed within
. Once
is computed for each solution member, a binary tournament strategy is employed to randomly select two solution members,
and
, from
, with
,
, and
. The solution member with the smaller weighted fitness value is designated as the new member of
, represented as
for
, where
The selection process based on the diversity criterion in Equation (20) continues until all solution members are chosen for . Once a solution member of is selected in based on the diversity criterion, it is flagged with to avoid its selection in subsequent binary tournaments, ensuring the population diversity in the next generation is not compromised.
The final
solution members of
are chosen from the remaining
) solution members of
based on the improvement rate criterion, considering their
values for
, with
. The same binary tournament strategy is applied to randomly select two solution members,
and
, from
, where
,
, and
. The solution member with the greater improvement rate is designated as the new solution member of
, i.e.,
, for
, where
The selection process based on the improvement rate criterion in Equation (21) continues until all solution members are included in . Similarly, any solution member of that has been chosen for based on the improvement rate criterion is flagged with to prevent it from being selected again in the next binary tournament, maintaining population diversity.
Algorithm 6 presents the pseudocode of the proposed tri-criterion selection scheme. Unlike traditional fitness-based selection methods like greedy selection and tournament selection, the proposed selection scheme not only retains the
elite solution members for
in the next iteration but also prioritizes the preservation of population diversity by simultaneously considering the diversity and improvement rate of solutions when selecting the remaining
and
solution members for
. This approach enriches population diversity by preserving promising individuals with various solutions, thus enhancing the search process. Furthermore, it encourages the selection of solutions with a higher improvement rate, leading to quicker convergence and improved overall optimization performance. By incorporating these three criteria, the tri-criterion selection scheme empowers ETLBOCBL-CNN with a more comprehensive selection process, resulting in the selection of higher-quality solutions in the next generation.
| Algorithm 6: Tri-Criterion Selection Scheme |
| Inputs: , , |
| 01: | Initialize ; |
| 02: | for to N do |
| 03: | Assign for each n-th original learner stored in ; |
| 04: | Calculate of every n-th offspring learner stored in with Equation (16); |
| 05: | end for |
| 06: | Construct the merged population using Equation (17); |
| 07: | Sort the solution members in ascendingly based on fitness values; |
| 08: | for to 2N do |
| 09: | Calculate of every n-th solution stored in with Equation (18); |
| 10: | end for |
| 11: | Randomly generate the integers of , and ; |
| 12: | for to do /*Fitness criterion*/ |
| 13: | ; |
| 14: | ; |
| 15: | end for |
| 12: | for to 2N do |
| 13: | Randomly generate based on a normal distribution of ; |
| 14: | Restrict the value of in between 0.8 and 1. |
| 15: | Compute the of each n-th solution stored in with Equation (19); |
| 16: | Initialize the flag variable of each n-th solution stored in as ; |
| 17: | end for |
| 18: | for to do /*Diversity criterion*/ |
| 19: | Randomly select and from , where , , and . |
| 20: | Determine with Equation (20); |
| 21: | ; |
| 22: | if is selected as then /*Prevent the selection of same solution members*/ |
| 23: | ; |
| 24: | else if is selected as then |
| 25: | ; |
| 26: |
end if |
| 27: | end for |
| 28: | for to do /*Improvement rate criterion*/ |
| 29: | Randomly select and from , where , e , and . |
| 30: | Determine using Equation (21); |
| 31: | ; |
| 32: | if is selected as then /*Prevent the selection of same solution members*/ |
| 33: | ; |
| 34: | else if is selected as then |
| 35: | ; |
| 36: |
end if |
| 37: | end for |
| Output:
|