Quantitative Comparison of Deep Learning-Based Image Reconstruction Methods for Low-Dose and Sparse-Angle CT Applications
Abstract
1. Introduction
1.1. Goal of This Study
1.1.1. Reconstruction of Low-Dose Medical CT Images
1.1.2. Reconstruction of Sparse-Angle CT Images
2. Dataset Description
2.1. LoDoPaB-CT Dataset
2.2. Apple CT Datasets
3. Algorithms
3.1. Learned Reconstruction Methods
3.1.1. Post-Processing
3.1.2. Fully Learned
3.1.3. Learned Iterative Schemes
3.1.4. Generative Approach
3.1.5. Unsupervised Methods
3.2. Classical Reconstruction Methods
4. Evaluation Methodology
4.1. Evaluation Metrics
4.1.1. Peak Signal-to-Noise Ratio
- PSNR: In this case , that is, the difference between the highest and lowest entry in . This allows for a PSNR value that is adapted to the range of the current ground truth image. The disadvantage is that the PSNR is image-dependent in this case.
- PSNR-FR: The same fixed L is chosen for all images. It is determined as the maximum entry computed over all training ground truth images, that is, for LoDoPaB-CT and for the Apple CT datasets. This can be seen as an (empirical) upper limit of the intensity range in the ground truth. In general, a fixed L is preferable because the scaling of the metric is image-independent in this case. This allows for a direct comparison of PSNR values calculated on different images. The downside for most CT applications is, that high values ( dense material) are not present in every scan. Therefore, the results can be too optimistic for these scans. However, based on Equation (7), all mean PSNR-FR values can be directly converted for another fixed choice of L.
4.1.2. Structural Similarity
4.1.3. Data Discrepancy
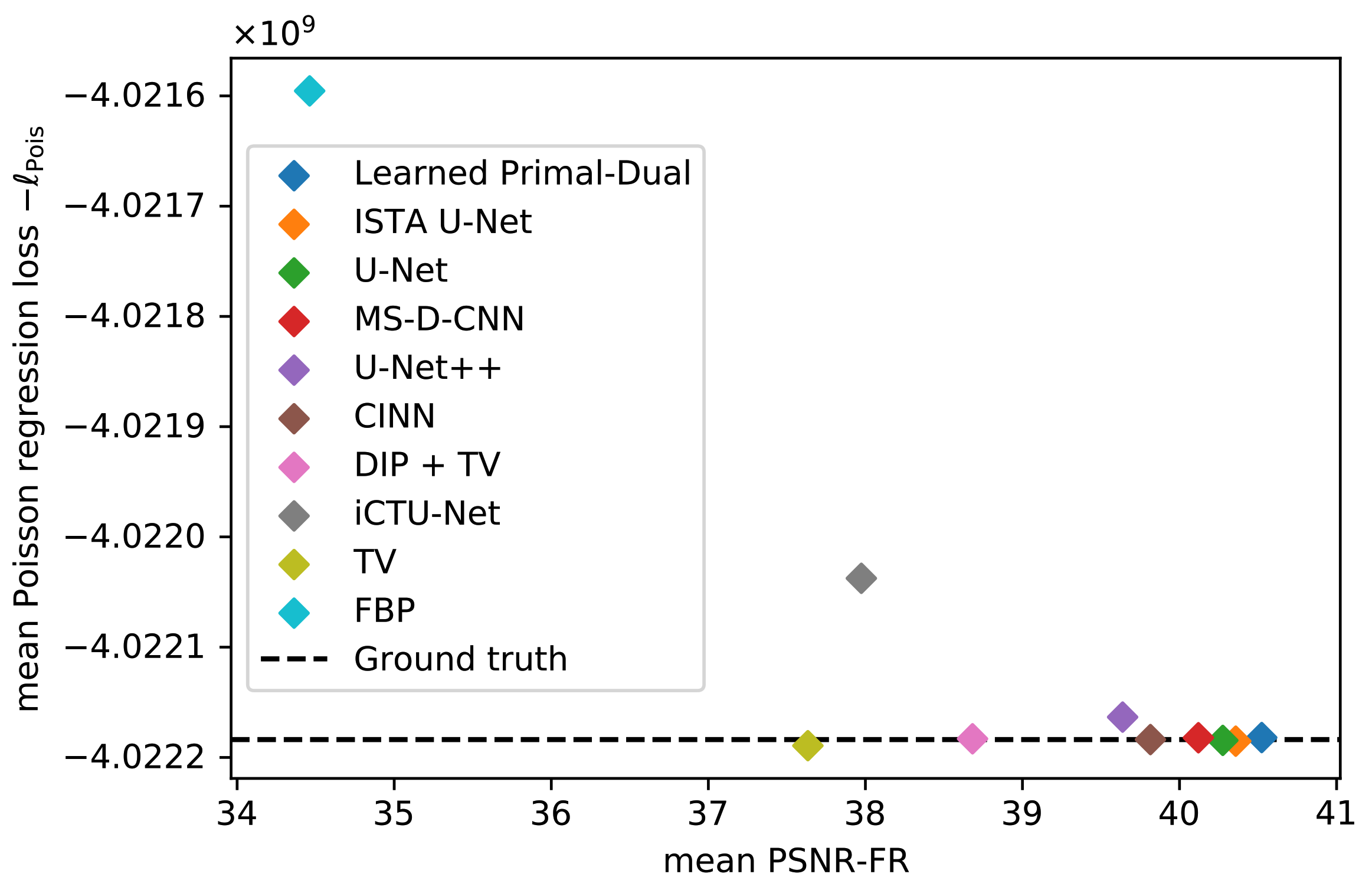
Poisson Regression Loss on LoDoPaB-CT Dataset
Mean Squared Error on Apple CT Data
4.2. Training Procedure
5. Results
5.1. LoDoPaB-CT Dataset
5.1.1. Reconstruction Performance
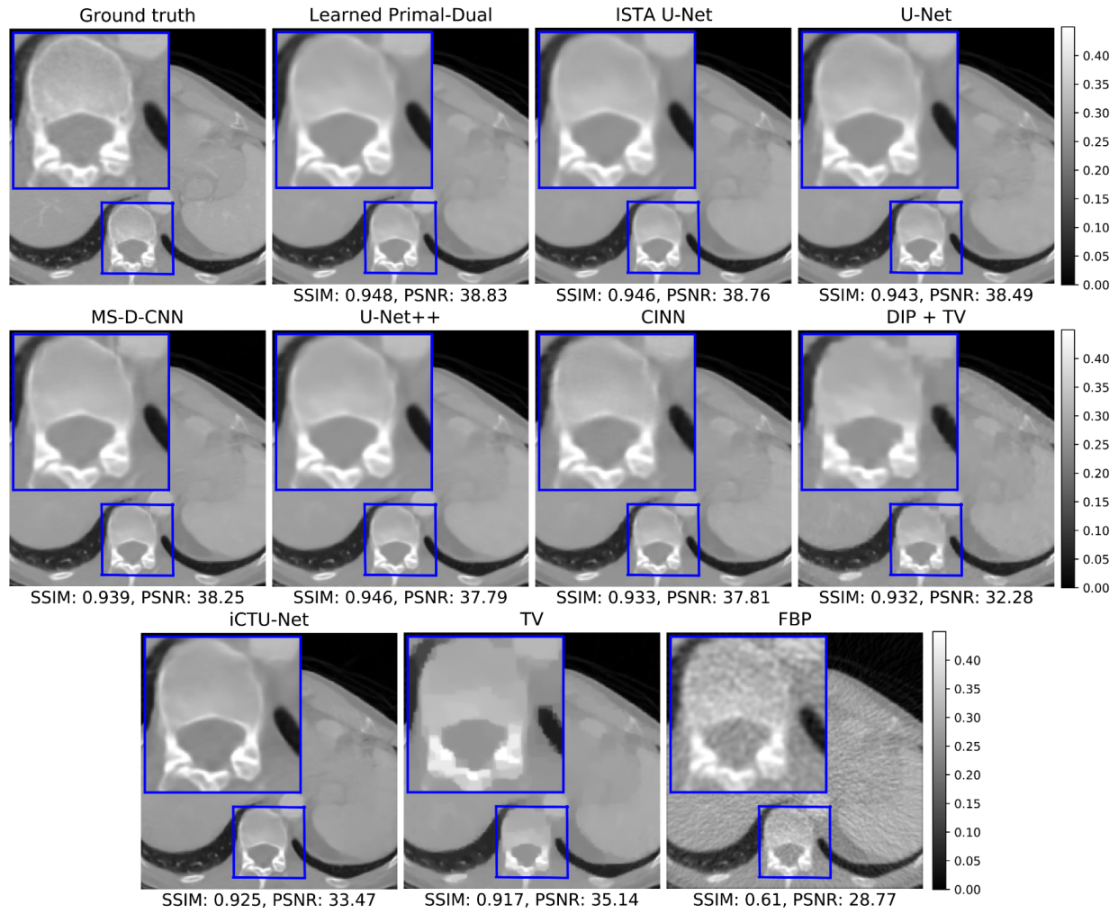
5.1.2. Visual Comparison
5.1.3. Data Consistency
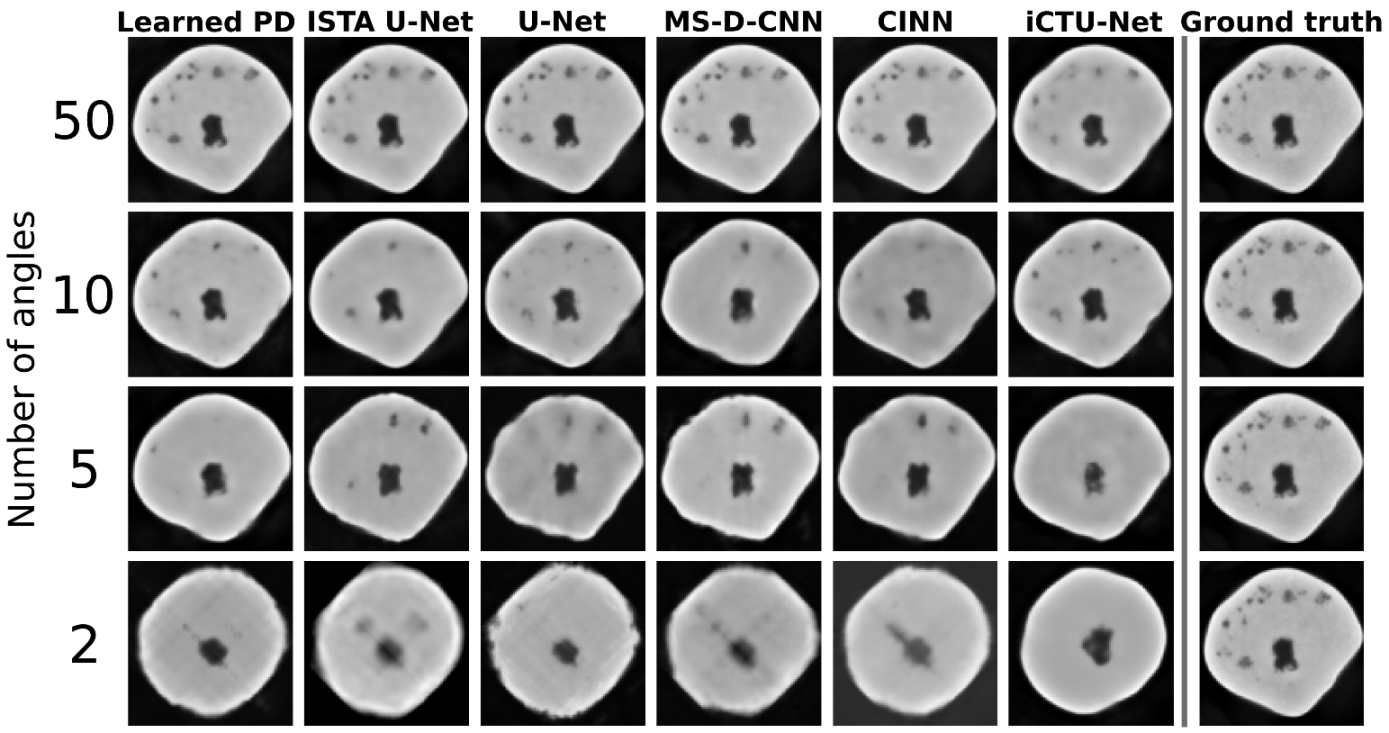
5.2. Apple CT Datasets
5.2.1. Reconstruction Performance
5.2.2. Visual Comparison
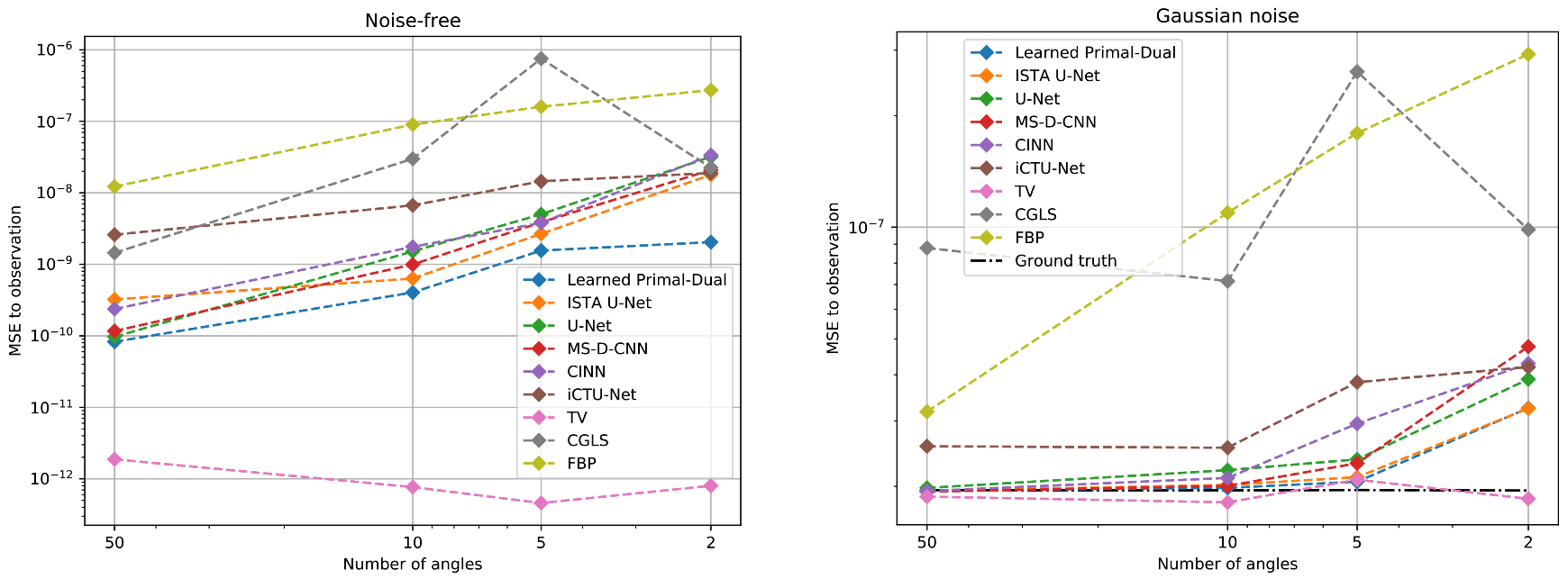
5.2.3. Data Consistency
6. Discussion
6.1. Computational Requirements and Reconstruction Speed
Transfer to 3D Reconstruction
6.2. Impact of the Datasets
6.2.1. Number of Training Samples
6.2.2. Observations on LoDoPaB-CT and Apple CT
6.2.3. Robustness to Changes in the Scanning Setup
6.2.4. Generalization to Other CT Setups
6.3. Conformance of Image Quality Scores and Requirements in Real Applications
6.4. Impact of Data Consistency
6.5. Recommendations and Future Work
7. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
Appendix A. Learned Reconstruction Methods
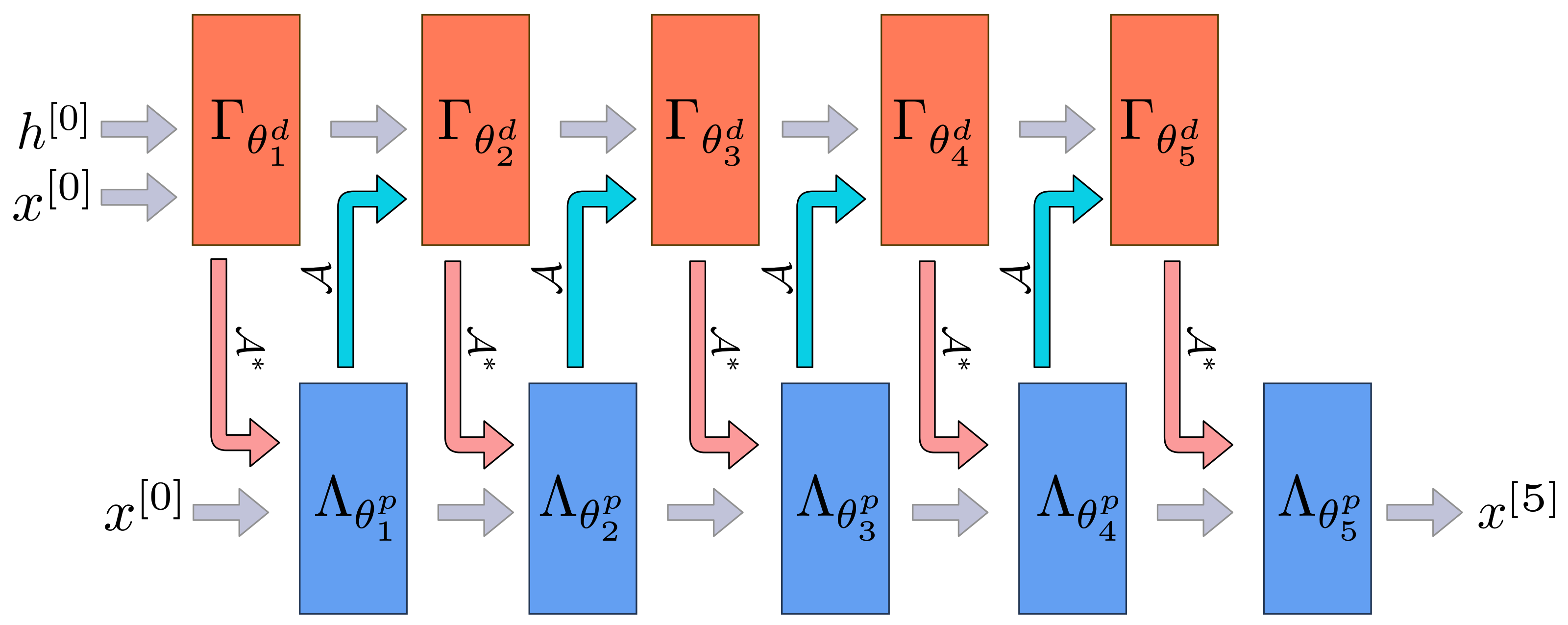
Appendix A.1. Learned Primal-Dual
| Algorithm A1 Learned Primal-Dual. |
Given learned proximal dual and primal operators for the reconstruction from noisy measurements is calculated as follows.
|
Appendix A.2. U-Net

Appendix A.3. U-Net++
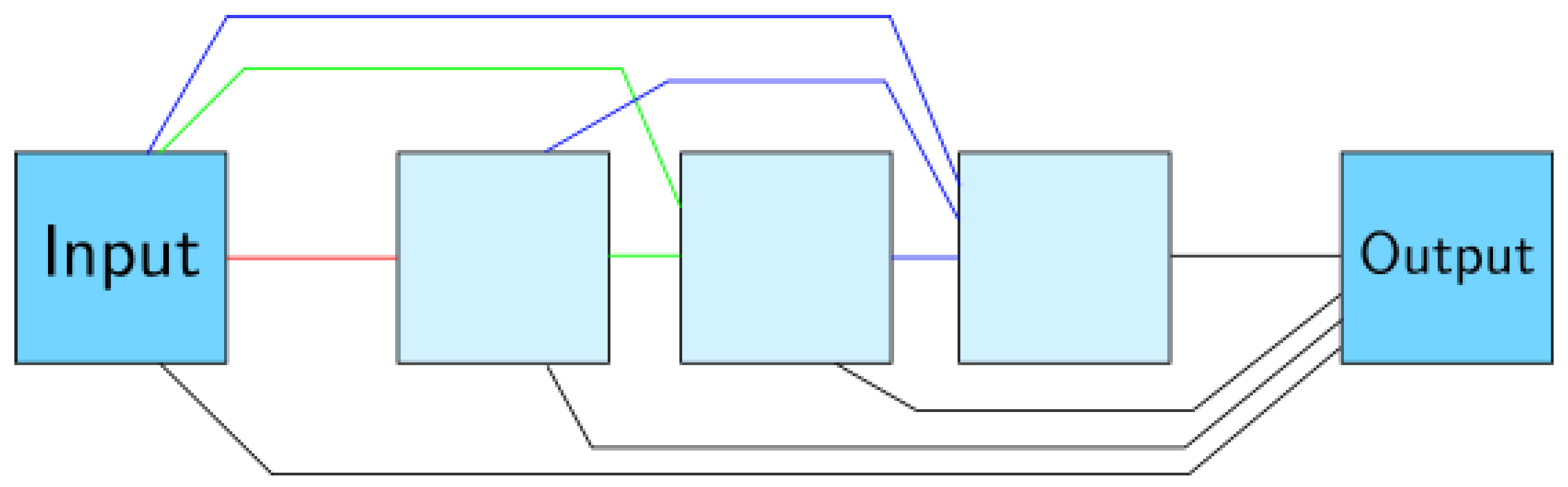
Appendix A.4. Mixed-Scale Dense Convolutional Neural Network
Appendix A.5. Conditional Invertible Neural Networks
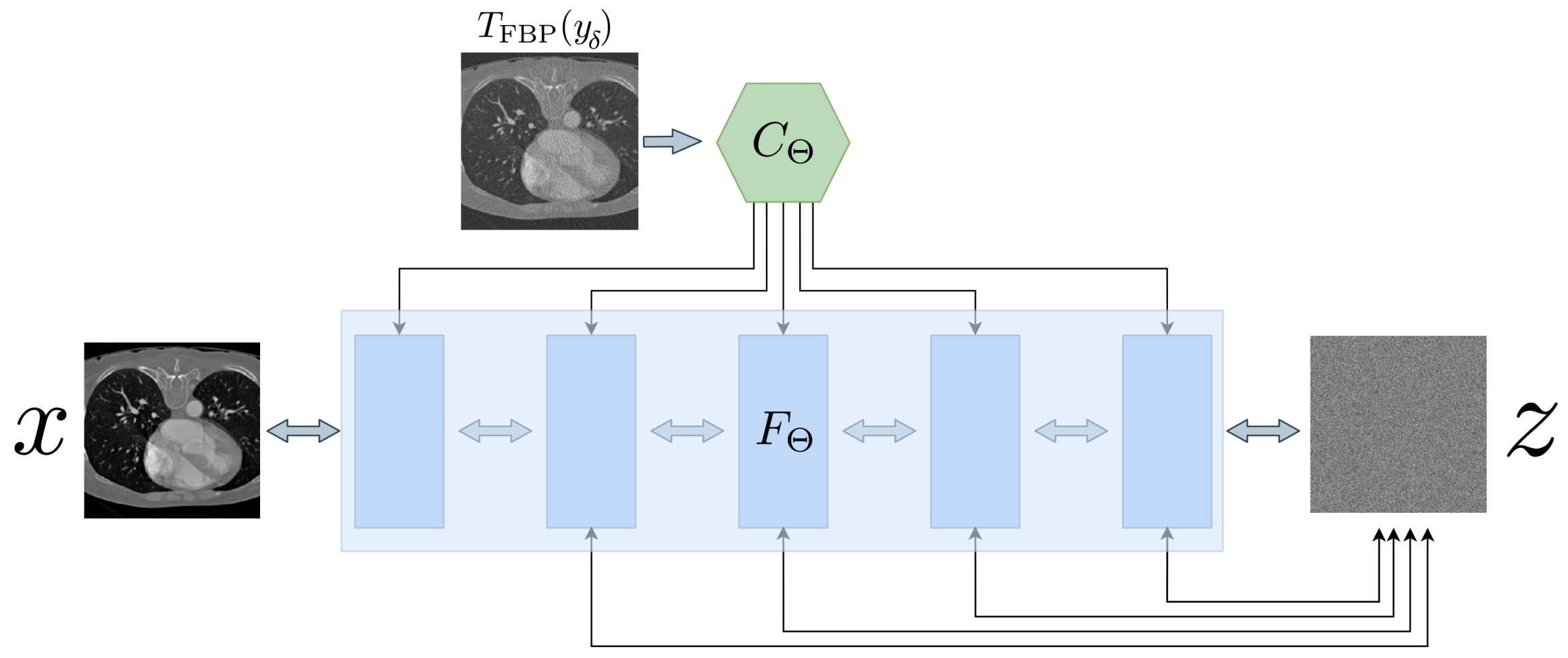
| Algorithm A2 Conditional Invertible Neural Network (CINN). |
Given a noisy measurement, , an invertible neural network F and a conditioning network C. Let be the number of random samples that should be drawn from a normal distribution . The algorithm calculates the mean and variance of the conditioned reconstructions.
|
Appendix A.6. ISTA U-Net
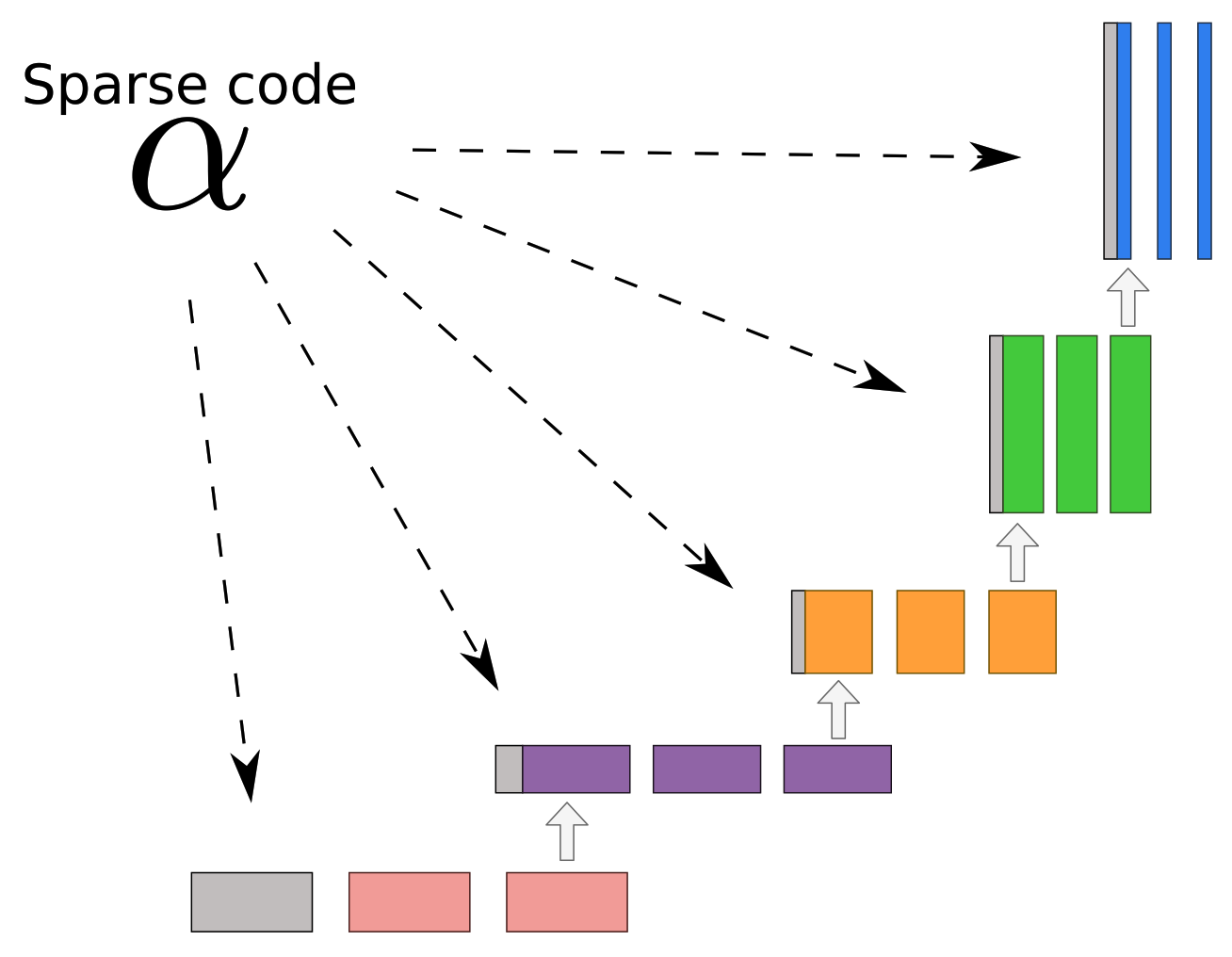
| Algorithm A3 ISTA U-Net. |
Given a noisy input , learned dictionaries and learned step sizes η and λ the reconstruction using the ISTA U-Net can be computed as follows.
|
Appendix A.7. Deep Image Prior with TV Denoising
| Algorithm A4 Deep Image Prior + Total Variation (DIP + TV). |
Given a noisy measurement , a neural network with initial parameterization , forward operator and a fixed random input z. The reconstruction is calculated iteratively over a number of iterations:
|
Appendix A.8. iCTU-Net
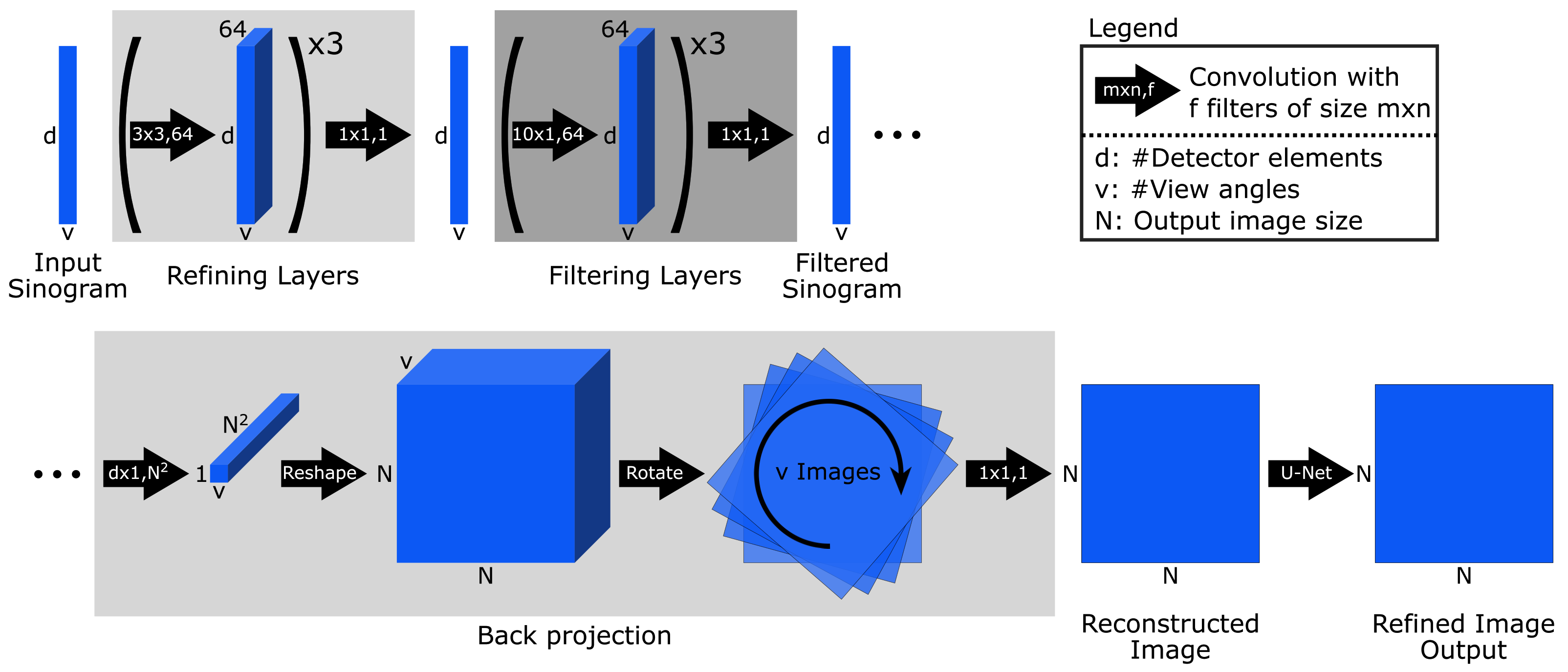
Appendix B. Classical Reconstruction Methods
Appendix B.1. Filtered Back-Projection (FBP)
| Window | Frequency Scaling | ||
|---|---|---|---|
| LoDoPaB-CT Dataset | Hann | 0.641 | |
| Apple CT Dataset A (Noise-free) | 50 angles | Cosine | 0.11 |
| 10 angles | Cosine | 0.013 | |
| 5 angles | Hann | 0.011 | |
| 2 angles | Hann | 0.011 | |
| Apple CT Dataset B (Gaussian noise) | 50 angles | Cosine | 0.08 |
| 10 angles | Cosine | 0.013 | |
| 5 angles | Hann | 0.011 | |
| 2 angles | Hann | 0.011 | |
| Apple CT Dataset C (Scattering) | 50 angles | Cosine | 0.09 |
| 10 angles | Hann | 0.018 | |
| 5 angles | Hann | 0.011 | |
| 2 angles | Hann | 0.009 |
Appendix B.2. Conjugate Gradient Least Squares
| Algorithm A5 Conjugate Gradient Least Squares (CGLS). |
Given a geometry matrix, A, a data vector and a zero solution vector (a black image) as the starting point, the algorithm below gives the solution at kth iteration.
|
Appendix B.3. Total Variation Regularization
| Discrepancy | Iterations | Step Size | |||
|---|---|---|---|---|---|
| LoDoPaB-CT Dataset | 5000 | 0.001 | 20.56 | ||
| Apple CT Dataset A (Noise-free) | 50 angles | MSE | 600 | ||
| 10 angles | MSE | 75,000 | |||
| 5 angles | MSE | 146,000 | |||
| 2 angles | MSE | 150,000 | |||
| Apple CT Dataset B (Gaussian noise) | 50 angles | MSE | 900 | ||
| 10 angles | MSE | 66,000 | |||
| 5 angles | MSE | 100,000 | |||
| 2 angles | MSE | 149,000 | |||
| Apple CT Dataset C (Scattering) | 50 angles | MSE | 400 | ||
| 10 angles | MSE | 13,000 | |||
| 5 angles | MSE | 149,000 | |||
| 2 angles | MSE | 150,000 |
| Algorithm A6 Total Variation Regularization (TV). |
Given a noisy measurement , an initial reconstruction , a weight and a maximum number of iterations K.
|
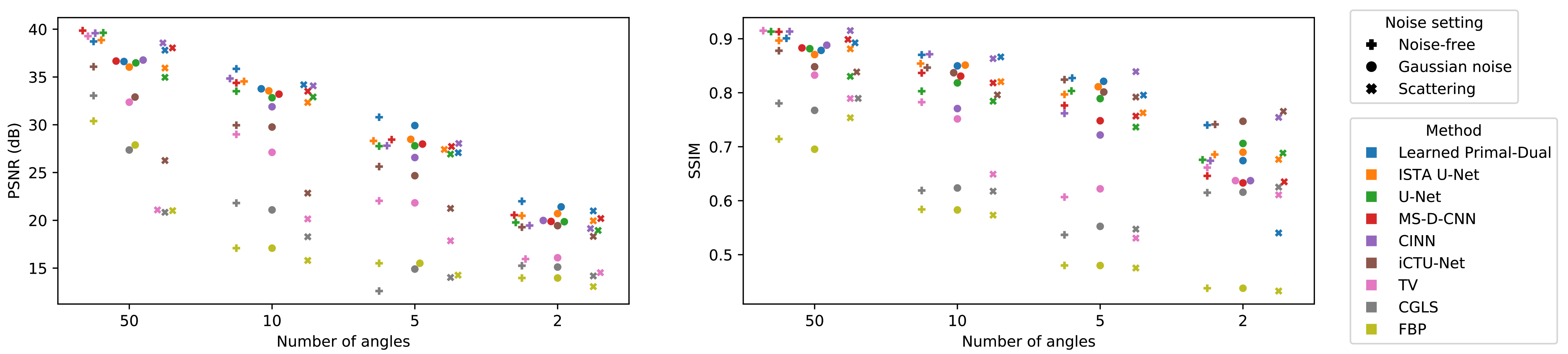
Appendix C. Further Results
| Noise-Free | Standard Deviation of PSNR | Standard Deviation of SSIM | ||||||
|---|---|---|---|---|---|---|---|---|
| Number of Angles | 50 | 10 | 5 | 2 | 50 | 10 | 5 | 2 |
| Learned Primal-Dual | 1.51 | 1.63 | 1.97 | 2.58 | 0.022 | 0.016 | 0.014 | 0.022 |
| ISTA U-Net | 1.40 | 1.77 | 2.12 | 2.13 | 0.018 | 0.018 | 0.022 | 0.037 |
| U-Net | 1.56 | 1.61 | 2.28 | 1.63 | 0.021 | 0.019 | 0.025 | 0.031 |
| MS-D-CNN | 1.51 | 1.65 | 1.81 | 2.09 | 0.021 | 0.020 | 0.024 | 0.022 |
| CINN | 1.40 | 1.64 | 1.99 | 2.17 | 0.016 | 0.019 | 0.023 | 0.027 |
| iCTU-Net | 1.68 | 2.45 | 1.92 | 1.93 | 0.024 | 0.027 | 0.030 | 0.028 |
| TV | 1.60 | 1.29 | 1.21 | 1.49 | 0.022 | 0.041 | 0.029 | 0.023 |
| CGLS | 0.69 | 0.48 | 2.94 | 0.70 | 0.014 | 0.027 | 0.029 | 0.039 |
| FBP | 0.80 | 0.58 | 0.54 | 0.50 | 0.021 | 0.023 | 0.028 | 0.067 |
| Gaussian Noise | Standard Deviation of PSNR | Standard Deviation of SSIM | ||||||
| Number of Angles | 50 | 10 | 5 | 2 | 50 | 10 | 5 | 2 |
| Learned Primal-Dual | 1.56 | 1.63 | 2.00 | 2.79 | 0.021 | 0.018 | 0.021 | 0.022 |
| ISTA U-Net | 1.70 | 1.76 | 2.27 | 2.12 | 0.025 | 0.021 | 0.022 | 0.038 |
| U-Net | 1.66 | 1.59 | 1.99 | 2.22 | 0.023 | 0.020 | 0.025 | 0.026 |
| MS-D-CNN | 1.66 | 1.75 | 1.79 | 1.79 | 0.025 | 0.024 | 0.019 | 0.022 |
| CINN | 1.53 | 1.51 | 1.62 | 2.06 | 0.023 | 0.017 | 0.017 | 0.020 |
| iCTU-Net | 1.98 | 2.06 | 1.89 | 1.91 | 0.031 | 0.032 | 0.039 | 0.027 |
| TV | 1.38 | 1.26 | 1.09 | 1.62 | 0.036 | 0.047 | 0.039 | 0.030 |
| CGLS | 0.78 | 0.49 | 1.76 | 0.68 | 0.014 | 0.026 | 0.029 | 0.037 |
| FBP | 0.91 | 0.58 | 0.54 | 0.50 | 0.028 | 0.023 | 0.028 | 0.067 |
| Scattering Noise | Standard Deviation of PSNR | Standard Deviation of SSIM | ||||||
| Number of Angles | 50 | 10 | 5 | 2 | 50 | 10 | 5 | 2 |
| Learned Primal-Dual | 1.91 | 1.80 | 1.71 | 2.47 | 0.017 | 0.016 | 0.016 | 0.060 |
| ISTA U-Net | 1.48 | 1.59 | 2.05 | 1.81 | 0.023 | 0.019 | 0.019 | 0.038 |
| U-Net | 1.76 | 1.56 | 1.81 | 1.47 | 0.015 | 0.021 | 0.027 | 0.024 |
| MS-D-CNN | 2.04 | 1.78 | 1.85 | 2.03 | 0.023 | 0.022 | 0.015 | 0.020 |
| CINN | 1.82 | 1.92 | 2.32 | 2.25 | 0.019 | 0.024 | 0.029 | 0.030 |
| iCTU-Net | 1.91 | 2.09 | 1.78 | 2.29 | 0.030 | 0.031 | 0.033 | 0.040 |
| TV | 2.53 | 2.44 | 1.86 | 1.59 | 0.067 | 0.076 | 0.035 | 0.062 |
| CGLS | 2.38 | 1.32 | 1.71 | 0.95 | 0.020 | 0.020 | 0.026 | 0.032 |
| FBP | 2.23 | 0.97 | 0.80 | 0.68 | 0.044 | 0.025 | 0.023 | 0.058 |
| Noise-Free | PSNR-FR | SSIM-FR | ||||||
|---|---|---|---|---|---|---|---|---|
| Number of Angles | 50 | 10 | 5 | 2 | 50 | 10 | 5 | 2 |
| Learned Primal-Dual | 45.33 | 42.47 | 37.41 | 28.61 | 0.971 | 0.957 | 0.935 | 0.872 |
| ISTA U-Net | 45.48 | 41.15 | 34.93 | 27.10 | 0.967 | 0.944 | 0.907 | 0.823 |
| U-Net | 46.24 | 40.13 | 34.38 | 26.39 | 0.975 | 0.917 | 0.911 | 0.830 |
| MS-D-CNN | 46.47 | 41.00 | 35.06 | 27.17 | 0.975 | 0.936 | 0.898 | 0.808 |
| CINN | 46.20 | 41.46 | 34.43 | 26.07 | 0.975 | 0.958 | 0.896 | 0.838 |
| iCTU-Net | 42.69 | 36.57 | 32.24 | 25.90 | 0.957 | 0.938 | 0.920 | 0.861 |
| TV | 45.89 | 35.61 | 28.66 | 22.57 | 0.976 | 0.904 | 0.746 | 0.786 |
| CGLS | 39.66 | 28.43 | 19.22 | 21.87 | 0.901 | 0.744 | 0.654 | 0.733 |
| FBP | 37.01 | 23.71 | 22.12 | 20.58 | 0.856 | 0.711 | 0.596 | 0.538 |
| Gaussian Noise | PSNR-FR | SSIM-FR | ||||||
| Number of Angles | 50 | 10 | 5 | 2 | 50 | 10 | 5 | 2 |
| Learned Primal-Dual | 43.24 | 40.38 | 36.54 | 28.03 | 0.961 | 0.944 | 0.927 | 0.823 |
| ISTA U-Net | 42.65 | 40.17 | 35.09 | 27.32 | 0.956 | 0.942 | 0.916 | 0.826 |
| U-Net | 43.09 | 39.45 | 34.42 | 26.47 | 0.961 | 0.924 | 0.904 | 0.843 |
| MS-D-CNN | 43.28 | 39.82 | 34.60 | 26.50 | 0.962 | 0.932 | 0.886 | 0.797 |
| CINN | 43.39 | 38.50 | 33.19 | 26.60 | 0.966 | 0.904 | 0.878 | 0.816 |
| iCTU-Net | 39.51 | 36.38 | 31.29 | 26.06 | 0.939 | 0.932 | 0.905 | 0.867 |
| TV | 38.98 | 33.73 | 28.45 | 22.70 | 0.939 | 0.883 | 0.770 | 0.772 |
| CGLS | 33.98 | 27.71 | 21.52 | 21.73 | 0.884 | 0.748 | 0.668 | 0.734 |
| FBP | 34.50 | 23.70 | 22.12 | 20.58 | 0.839 | 0.711 | 0.596 | 0.538 |
| Scattering Noise | PSNR-FR | SSIM-FR | ||||||
| Number of Angles | 50 | 10 | 5 | 2 | 50 | 10 | 5 | 2 |
| Learned Primal-Dual | 44.42 | 40.80 | 33.69 | 27.60 | 0.967 | 0.954 | 0.912 | 0.760 |
| ISTA U-Net | 42.55 | 38.95 | 34.03 | 26.57 | 0.959 | 0.922 | 0.887 | 0.816 |
| U-Net | 41.58 | 39.52 | 33.55 | 25.56 | 0.932 | 0.910 | 0.877 | 0.828 |
| MS-D-CNN | 44.66 | 40.13 | 34.34 | 26.81 | 0.969 | 0.927 | 0.889 | 0.796 |
| CINN | 45.18 | 40.69 | 34.66 | 25.76 | 0.976 | 0.952 | 0.936 | 0.878 |
| iCTU-Net | 32.88 | 29.46 | 27.86 | 24.93 | 0.931 | 0.901 | 0.896 | 0.873 |
| TV | 27.71 | 26.76 | 24.48 | 21.15 | 0.903 | 0.799 | 0.674 | 0.743 |
| CGLS | 27.46 | 24.89 | 20.64 | 20.80 | 0.896 | 0.738 | 0.659 | 0.736 |
| FBP | 27.63 | 22.42 | 20.88 | 19.68 | 0.878 | 0.701 | 0.589 | 0.529 |
| Noise-Free | Standard Deviation of PSNR-FR | Standard Deviation of SSIM-FR | ||||||
|---|---|---|---|---|---|---|---|---|
| Number of Angles | 50 | 10 | 5 | 2 | 50 | 10 | 5 | 2 |
| Learned Primal-Dual | 1.49 | 1.67 | 2.03 | 2.54 | 0.007 | 0.006 | 0.010 | 0.019 |
| ISTA U-Net | 1.37 | 1.82 | 2.21 | 2.21 | 0.005 | 0.010 | 0.020 | 0.034 |
| U-Net | 1.53 | 1.66 | 2.33 | 1.68 | 0.006 | 0.012 | 0.019 | 0.026 |
| MS-D-CNN | 1.46 | 1.71 | 1.90 | 2.15 | 0.006 | 0.011 | 0.021 | 0.015 |
| CINN | 1.35 | 1.65 | 2.09 | 2.21 | 0.004 | 0.007 | 0.023 | 0.025 |
| iCTU-Net | 1.82 | 2.54 | 2.03 | 1.91 | 0.014 | 0.017 | 0.020 | 0.023 |
| TV | 1.54 | 1.32 | 1.28 | 1.36 | 0.006 | 0.023 | 0.026 | 0.018 |
| CGLS | 0.71 | 0.51 | 2.96 | 0.56 | 0.009 | 0.029 | 0.033 | 0.045 |
| FBP | 0.77 | 0.46 | 0.38 | 0.41 | 0.011 | 0.015 | 0.029 | 0.088 |
| Gaussian Noise | Standard Deviation of PSNR-FR | Standard Deviation of SSIM-FR | ||||||
| Number of Angles | 50 | 10 | 5 | 2 | 50 | 10 | 5 | 2 |
| Learned Primal-Dual | 1.52 | 1.68 | 2.04 | 2.83 | 0.006 | 0.008 | 0.013 | 0.016 |
| ISTA U-Net | 1.65 | 1.78 | 2.36 | 2.17 | 0.008 | 0.010 | 0.018 | 0.034 |
| U-Net | 1.61 | 1.62 | 2.05 | 2.24 | 0.007 | 0.012 | 0.019 | 0.024 |
| MS-D-CNN | 1.62 | 1.80 | 1.84 | 1.84 | 0.008 | 0.011 | 0.015 | 0.014 |
| CINN | 1.50 | 1.59 | 1.65 | 2.09 | 0.007 | 0.016 | 0.017 | 0.019 |
| iCTU-Net | 2.07 | 2.12 | 1.93 | 1.90 | 0.020 | 0.021 | 0.026 | 0.024 |
| TV | 1.30 | 1.26 | 1.15 | 1.50 | 0.014 | 0.027 | 0.030 | 0.019 |
| CGLS | 0.63 | 0.45 | 1.76 | 0.53 | 0.012 | 0.028 | 0.034 | 0.043 |
| FBP | 0.83 | 0.46 | 0.38 | 0.41 | 0.014 | 0.015 | 0.029 | 0.088 |
| Scattering Noise | Standard Deviation of PSNR-FR | Standard Deviation of SSIM-FR | ||||||
| Number of Angles | 50 | 10 | 5 | 2 | 50 | 10 | 5 | 2 |
| Learned Primal-Dual | 1.92 | 1.85 | 1.81 | 2.51 | 0.005 | 0.007 | 0.014 | 0.038 |
| ISTA U-Net | 1.56 | 1.68 | 2.17 | 1.89 | 0.010 | 0.014 | 0.014 | 0.035 |
| U-Net | 1.72 | 1.63 | 1.91 | 1.59 | 0.010 | 0.012 | 0.024 | 0.024 |
| MS-D-CNN | 2.02 | 1.84 | 1.96 | 2.08 | 0.008 | 0.012 | 0.016 | 0.019 |
| CINN | 1.74 | 1.97 | 2.41 | 2.21 | 0.005 | 0.011 | 0.016 | 0.022 |
| iCTU-Net | 1.96 | 2.14 | 1.79 | 2.32 | 0.016 | 0.023 | 0.022 | 0.030 |
| TV | 2.43 | 2.35 | 1.80 | 1.49 | 0.048 | 0.074 | 0.040 | 0.051 |
| CGLS | 2.28 | 1.24 | 1.67 | 0.83 | 0.016 | 0.021 | 0.030 | 0.035 |
| FBP | 2.14 | 0.87 | 0.66 | 0.55 | 0.028 | 0.016 | 0.020 | 0.078 |

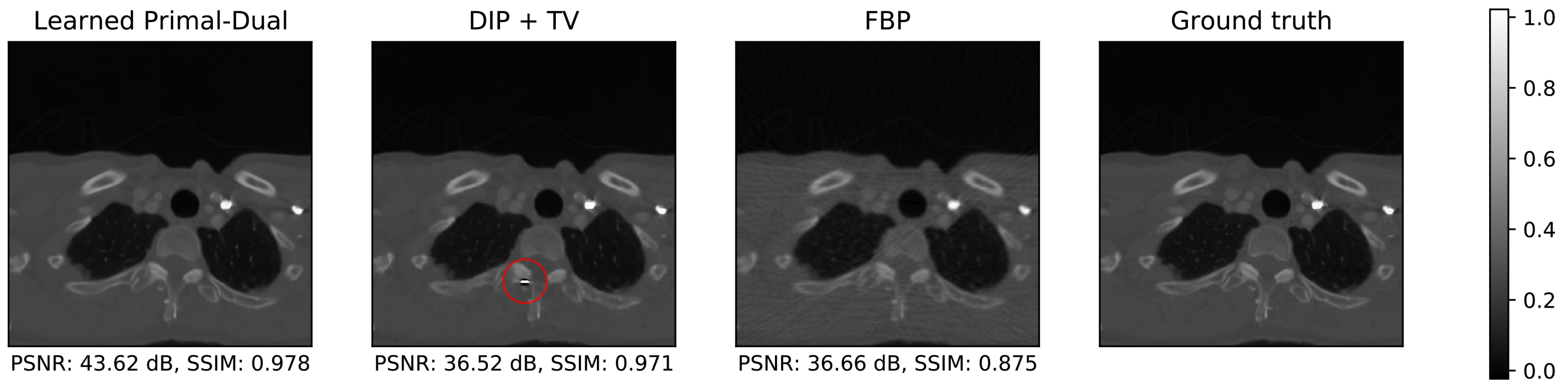
| Noise Free | MSE | |||
|---|---|---|---|---|
| Number of Angles | 50 | 10 | 5 | 2 |
| Learned Primal-Dual | ||||
| ISTA U-Net | ||||
| U-Net | ||||
| MS-D-CNN | ||||
| CINN | ||||
| iCTU-Net | ||||
| TV | ||||
| CGLS | ||||
| FBP | ||||
| Ground truth | ||||
| Gaussian Noise | MSE | |||
| Number of Angles | 50 | 10 | 5 | 2 |
| Learned Primal-Dual | ||||
| ISTA U-Net | ||||
| U-Net | ||||
| MS-D-CNN | ||||
| CINN | ||||
| iCTU-Net | ||||
| TV | ||||
| CGLS | ||||
| FBP | ||||
| Ground truth | ||||
| Scattering Noise | MSE | |||
| Number of Angles | 50 | 10 | 5 | 2 |
| Learned Primal-Dual | ||||
| ISTA U-Net | ||||
| U-Net | ||||
| MS-D-CNN | ||||
| CINN | ||||
| iCTU-Net | ||||
| TV | ||||
| CGLS | ||||
| FBP | ||||
| Ground truth | ||||
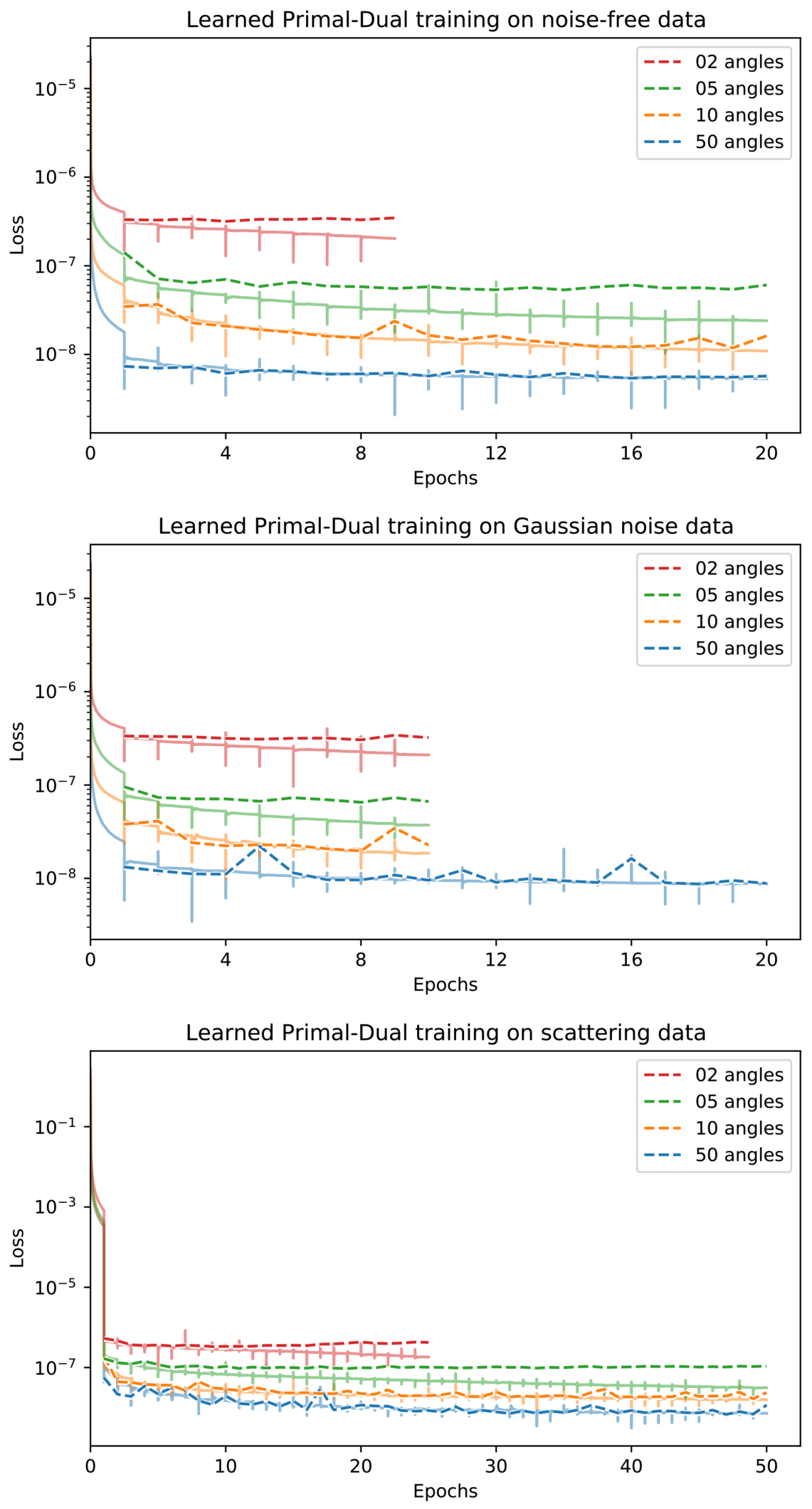
Appendix D. Training Curves
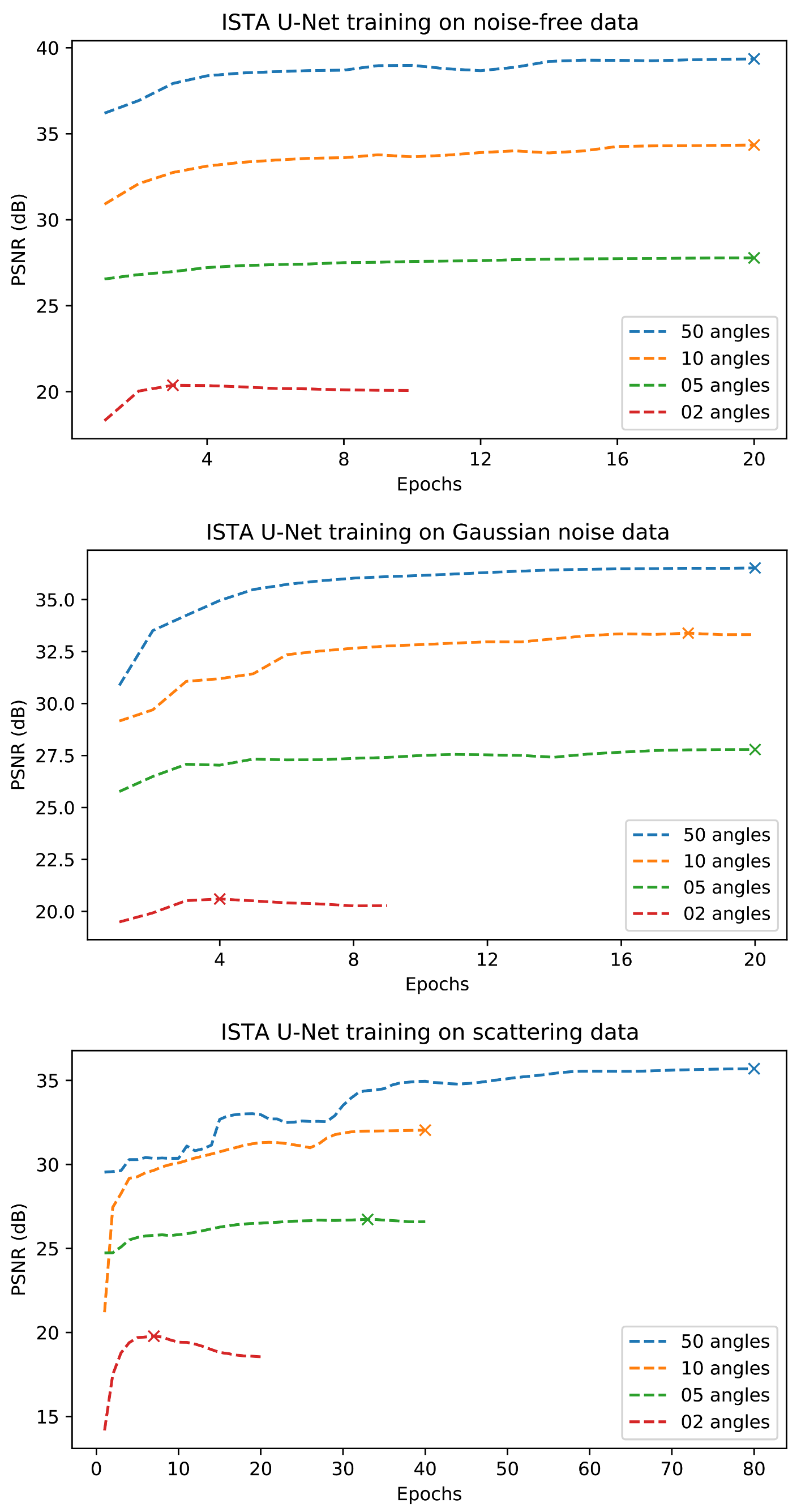

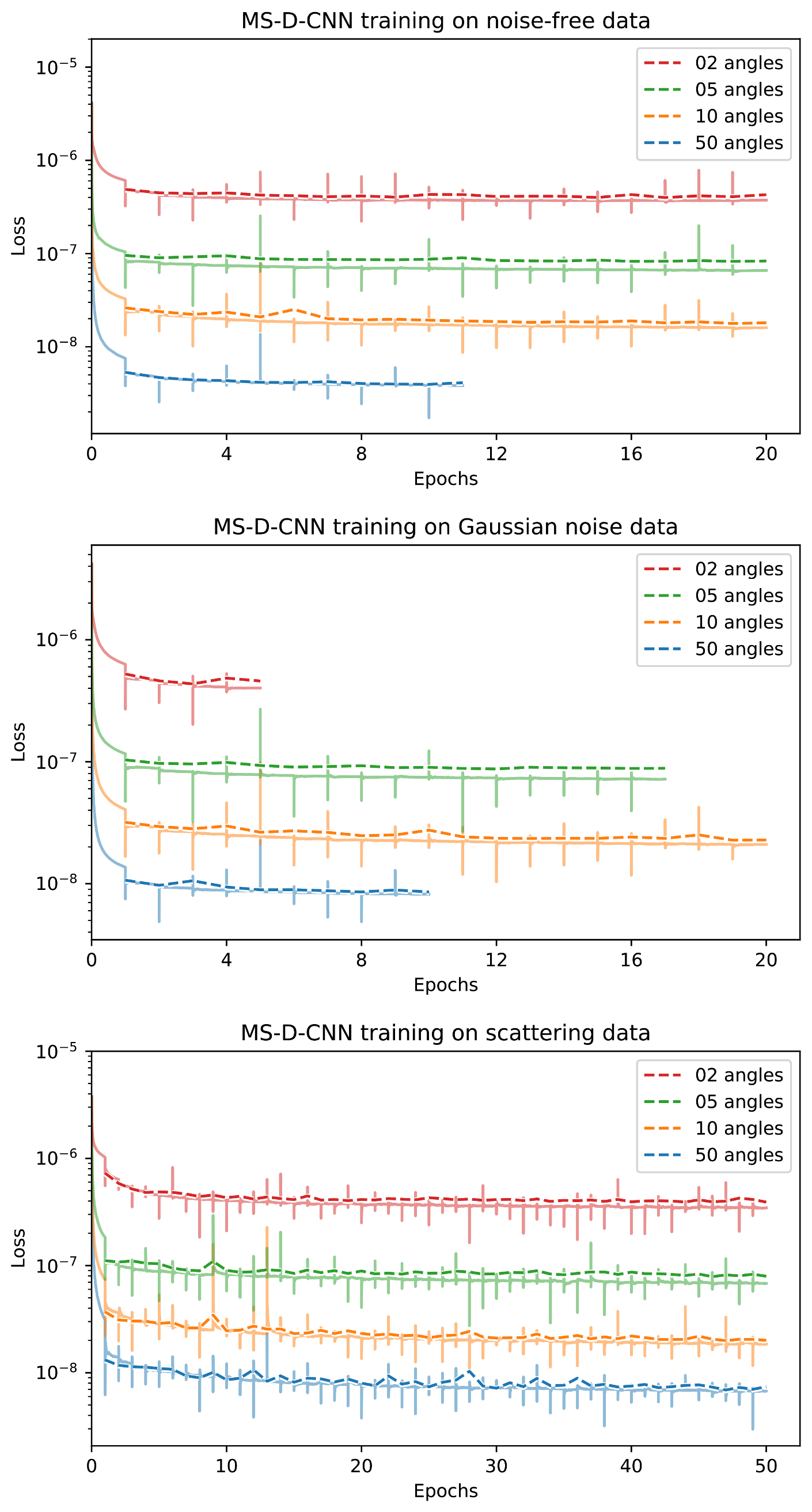
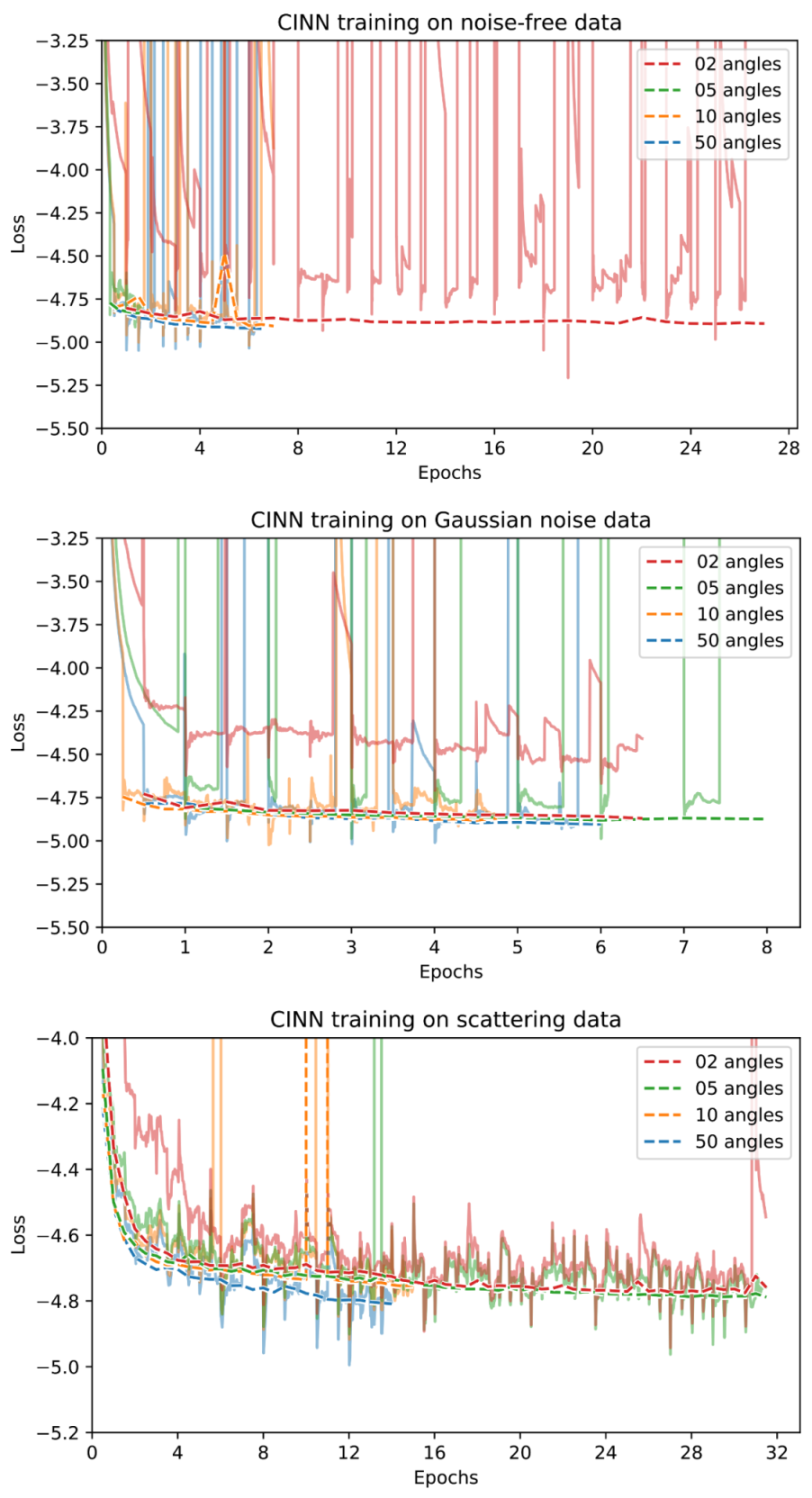
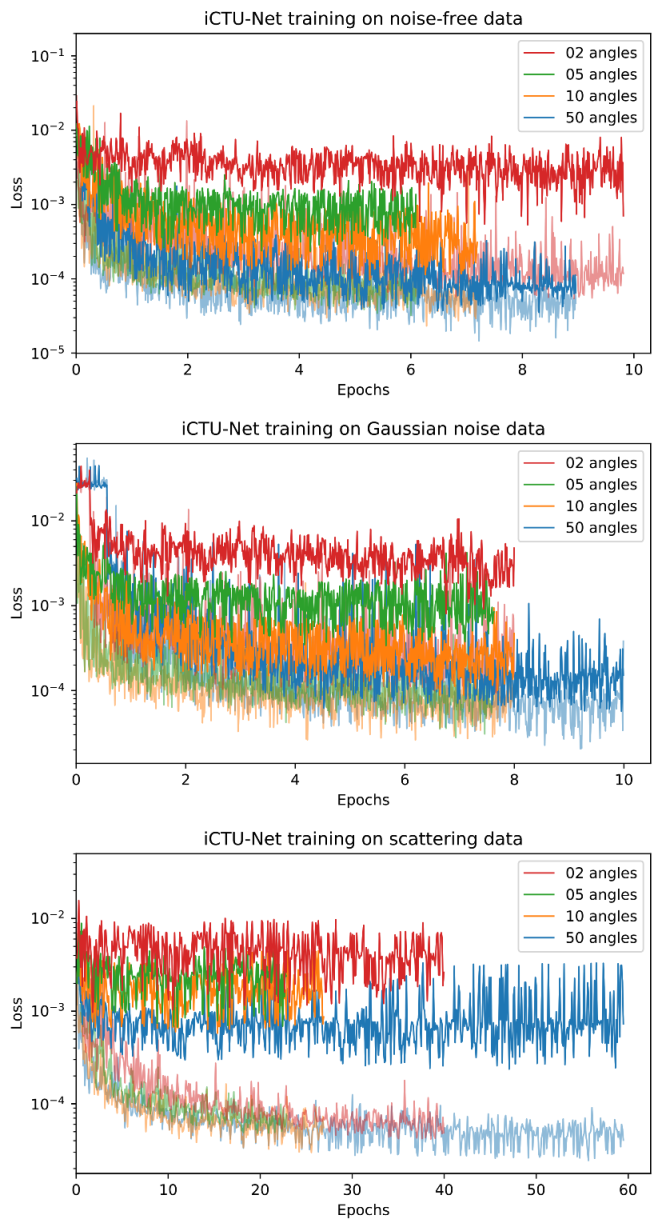
References
- Liguori, C.; Frauenfelder, G.; Massaroni, C.; Saccomandi, P.; Giurazza, F.; Pitocco, F.; Marano, R.; Schena, E. Emerging clinical applications of computed tomography. Med. Devices 2015, 8, 265. [Google Scholar]
- National Lung Screening Trial Research Team. Reduced lung-cancer mortality with low-dose computed tomographic screening. N. Engl. J. Med. 2011, 365, 395–409. [Google Scholar] [CrossRef]
- Yoo, S.; Yin, F.F. Dosimetric feasibility of cone-beam CT-based treatment planning compared to CT-based treatment planning. Int. J. Radiat. Oncol. Biol. Phys. 2006, 66, 1553–1561. [Google Scholar] [CrossRef]
- Swennen, G.R.; Mollemans, W.; Schutyser, F. Three-dimensional treatment planning of orthognathic surgery in the era of virtual imaging. J. Oral Maxillofac. Surg. 2009, 67, 2080–2092. [Google Scholar] [CrossRef]
- De Chiffre, L.; Carmignato, S.; Kruth, J.P.; Schmitt, R.; Weckenmann, A. Industrial applications of computed tomography. CIRP Ann. 2014, 63, 655–677. [Google Scholar] [CrossRef]
- Mees, F.; Swennen, R.; Van Geet, M.; Jacobs, P. Applications of X-ray Computed Tomography in the Geosciences; Special Publications; Geological Society: London, UK, 2003; Volume 215, pp. 1–6. [Google Scholar]
- Morigi, M.; Casali, F.; Bettuzzi, M.; Brancaccio, R.; d’Errico, V. Application of X-ray computed tomography to cultural heritage diagnostics. Appl. Phys. A 2010, 100, 653–661. [Google Scholar] [CrossRef]
- Coban, S.B.; Lucka, F.; Palenstijn, W.J.; Van Loo, D.; Batenburg, K.J. Explorative Imaging and Its Implementation at the FleX-ray Laboratory. J. Imaging 2020, 6, 18. [Google Scholar] [CrossRef]
- McCollough, C.H.; Bartley, A.C.; Carter, R.E.; Chen, B.; Drees, T.A.; Edwards, P.; Holmes, D.R., III.; Huang, A.E.; Khan, F.; Leng, S.; et al. Low-dose CT for the detection and classification of metastatic liver lesions: Results of the 2016 Low Dose CT Grand Challenge. Med Phys. 2017, 44, e339–e352. [Google Scholar] [CrossRef] [PubMed]
- Radon, J. On the determination of functions from their integral values along certain manifolds. IEEE Trans. Med Imaging 1986, 5, 170–176. [Google Scholar] [CrossRef]
- Natterer, F. The mathematics of computerized tomography (classics in applied mathematics, vol. 32). Inverse Probl. 2001, 18, 283–284. [Google Scholar]
- Boas, F.E.; Fleischmann, D. CT artifacts: Causes and reduction techniques. Imaging Med. 2012, 4, 229–240. [Google Scholar] [CrossRef]
- Wang, G.; Ye, J.C.; Mueller, K.; Fessler, J.A. Image Reconstruction is a New Frontier of Machine Learning. IEEE Trans. Med. Imaging 2018, 37, 1289–1296. [Google Scholar] [CrossRef] [PubMed]
- Sidky, E.Y.; Pan, X. Image reconstruction in circular cone-beam computed tomography by constrained, total-variation minimization. Phys. Med. Biol. 2008, 53, 4777. [Google Scholar] [CrossRef]
- Niu, S.; Gao, Y.; Bian, Z.; Huang, J.; Chen, W.; Yu, G.; Liang, Z.; Ma, J. Sparse-view X-ray CT reconstruction via total generalized variation regularization. Phys. Med. Biol. 2014, 59, 2997. [Google Scholar] [CrossRef] [PubMed]
- Hestenes, M.R.; Stiefel, E. Methods of conjugate gradients for solving linear systems. J. Res. Natl. Bur. Stand. 1952, 49, 409–436. [Google Scholar] [CrossRef]
- Arridge, S.; Maass, P.; Öktem, O.; Schönlieb, C.B. Solving inverse problems using data-driven models. Acta Numer. 2019, 28, 1–174. [Google Scholar] [CrossRef]
- Lunz, S.; Öktem, O.; Schönlieb, C.B. Adversarial Regularizers in Inverse Problems. In Advances in Neural Information Processing Systems; Bengio, S., Wallach, H., Larochelle, H., Grauman, K., Cesa-Bianchi, N., Garnett, R., Eds.; Curran Associates, Inc.: Red Hook, NY, USA, 2018; Volume 31, pp. 8507–8516. [Google Scholar]
- Adler, J.; Öktem, O. Learned Primal-Dual Reconstruction. IEEE Trans. Med. Imaging 2018, 37, 1322–1332. [Google Scholar] [CrossRef]
- Jin, K.H.; McCann, M.T.; Froustey, E.; Unser, M. Deep Convolutional Neural Network for Inverse Problems in Imaging. IEEE Trans. Image Process. 2017, 26, 4509–4522. [Google Scholar] [CrossRef]
- Pelt, D.M.; Batenburg, K.J.; Sethian, J.A. Improving Tomographic Reconstruction from Limited Data Using Mixed-Scale Dense Convolutional Neural Networks. J. Imaging 2018, 4, 128. [Google Scholar] [CrossRef]
- Chen, H.; Zhang, Y.; Zhang, W.; Liao, P.; Li, K.; Zhou, J.; Wang, G. Low-dose CT via convolutional neural network. Biomed. Opt. Express 2017, 8, 679–694. [Google Scholar] [CrossRef]
- Chen, H.; Zhang, Y.; Kalra, M.K.; Lin, F.; Chen, Y.; Liao, P.; Zhou, J.; Wang, G. Low-dose CT with a residual encoder-decoder convolutional neural network. IEEE Trans. Med. Imaging 2017, 36, 2524–2535. [Google Scholar] [CrossRef]
- Yang, Q.; Yan, P.; Kalra, M.K.; Wang, G. CT image denoising with perceptive deep neural networks. arXiv 2017, arXiv:1702.07019. [Google Scholar]
- Yang, Q.; Yan, P.; Zhang, Y.; Yu, H.; Shi, Y.; Mou, X.; Kalra, M.K.; Zhang, Y.; Sun, L.; Wang, G. Low-dose CT image denoising using a generative adversarial network with Wasserstein distance and perceptual loss. IEEE Trans. Med. Imaging 2018, 37, 1348–1357. [Google Scholar] [CrossRef]
- Feng, R.; Rundle, D.; Wang, G. Neural-networks-based Photon-Counting Data Correction: Pulse Pileup Effect. arXiv 2018, arXiv:1804.10980. [Google Scholar]
- Zhu, B.; Liu, J.Z.; Cauley, S.F.; Rosen, B.R.; Rosen, M.S. Image reconstruction by domain-transform manifold learning. Nature 2018, 555, 487–492. [Google Scholar] [CrossRef]
- He, J.; Ma, J. Radon inversion via deep learning. IEEE Trans. Med. Imaging 2020, 39, 2076–2087. [Google Scholar] [CrossRef] [PubMed]
- Li, Y.; Li, K.; Zhang, C.; Montoya, J.; Chen, G.H. Learning to reconstruct computed tomography images directly from sinogram data under a variety of data acquisition conditions. IEEE Trans. Med Imaging 2019, 38, 2469–2481. [Google Scholar] [CrossRef]
- European Society of Radiology (ESR). The new EU General Data Protection Regulation: What the radiologist should know. Insights Imaging 2017, 8, 295–299. [Google Scholar] [CrossRef] [PubMed]
- Kaissis, G.A.; Makowski, M.R.; Rückert, D.; Braren, R.F. Secure, privacy-preserving and federated machine learning in medical imaging. Nat. Mach. Intell. 2020, 2, 305–311. [Google Scholar] [CrossRef]
- Leuschner, J.; Schmidt, M.; Baguer, D.O.; Maass, P. The LoDoPaB-CT Dataset: A Benchmark Dataset for Low-Dose CT Reconstruction Methods. arXiv 2020, arXiv:1910.01113. [Google Scholar]
- Armato, S.G., III; McLennan, G.; Bidaut, L.; McNitt-Gray, M.F.; Meyer, C.R.; Reeves, A.P.; Zhao, B.; Aberle, D.R.; Henschke, C.I.; Hoffman, E.A.; et al. The Lung Image Database Consortium (LIDC) and Image Database Resource Initiative (IDRI): A Completed Reference Database of Lung Nodules on CT Scans. Med. Phys. 2011, 38, 915–931. [Google Scholar] [CrossRef]
- Baguer, D.O.; Leuschner, J.; Schmidt, M. Computed tomography reconstruction using deep image prior and learned reconstruction methods. Inverse Probl. 2020, 36, 094004. [Google Scholar] [CrossRef]
- Armato, S.G., III; McLennan, G.; Bidaut, L.; McNitt-Gray, M.F.; Meyer, C.R.; Reeves, A.P.; Zhao, B.; Aberle, D.R.; Henschke, C.I.; Hoffman, E.A.; et al. Data From LIDC-IDRI; The Cancer Imaging Archive: Frederick, MD, USA, 2015. [Google Scholar] [CrossRef]
- Buzug, T. Computed Tomography: From Photon Statistics to Modern Cone-Beam CT; Springer: Berlin/Heidelberg, Germany, 2008. [Google Scholar] [CrossRef]
- Coban, S.B.; Andriiashen, V.; Ganguly, P.S. Apple CT Data: Simulated Parallel-Beam Tomographic Datasets; Zenodo: Geneva, Switzerland, 2020. [Google Scholar] [CrossRef]
- Coban, S.B.; Andriiashen, V.; Ganguly, P.S.; van Eijnatten, M.; Batenburg, K.J. Parallel-beam X-ray CT datasets of apples with internal defects and label balancing for machine learning. arXiv 2020, arXiv:2012.13346. [Google Scholar]
- Leuschner, J.; Schmidt, M.; Ganguly, P.S.; Andriiashen, V.; Coban, S.B.; Denker, A.; van Eijnatten, M. Source Code and Supplementary Material for “Quantitative comparison of deep learning-based image reconstruction methods for low-dose and sparse-angle CT applications”. Zenodo 2021. [Google Scholar] [CrossRef]
- Ronneberger, O.; Fischer, P.; Brox, T. U-net: Convolutional networks for biomedical image segmentation. In International Conference on Medical Image Computing and Computer-Assisted Intervention; Springer: Heidelberg, Germany, 2015; pp. 234–241. [Google Scholar]
- Zhou, Z.; Siddiquee, M.M.R.; Tajbakhsh, N.; Liang, J. Unet++: A nested u-net architecture for medical image segmentation. In Deep Learning in Medical Image Analysis and Multimodal Learning for Clinical Decision Support; Springer: Heidelberg, Germany, 2018; pp. 3–11. [Google Scholar]
- Liu, T.; Chaman, A.; Belius, D.; Dokmanić, I. Interpreting U-Nets via Task-Driven Multiscale Dictionary Learning. arXiv 2020, arXiv:2011.12815. [Google Scholar]
- Comelli, A.; Dahiya, N.; Stefano, A.; Benfante, V.; Gentile, G.; Agnese, V.; Raffa, G.M.; Pilato, M.; Yezzi, A.; Petrucci, G.; et al. Deep learning approach for the segmentation of aneurysmal ascending aorta. Biomed. Eng. Lett. 2020, 1–10. [Google Scholar]
- Dashti, M.; Stuart, A.M. The Bayesian Approach to Inverse Problems. In Handbook of Uncertainty Quantification; Springer International Publishing: Cham, Switzerland, 2017; pp. 311–428. [Google Scholar] [CrossRef]
- Adler, J.; Öktem, O. Deep Bayesian Inversion. arXiv 2018, arXiv:1811.05910. [Google Scholar]
- Goodfellow, I.; Pouget-Abadie, J.; Mirza, M.; Xu, B.; Warde-Farley, D.; Ozair, S.; Courville, A.; Bengio, Y. Generative adversarial nets. arXiv 2014, arXiv:1406.2661. [Google Scholar]
- Ardizzone, L.; Lüth, C.; Kruse, J.; Rother, C.; Köthe, U. Guided image generation with conditional invertible neural networks. arXiv 2019, arXiv:1907.02392. [Google Scholar]
- Denker, A.; Schmidt, M.; Leuschner, J.; Maass, P.; Behrmann, J. Conditional Normalizing Flows for Low-Dose Computed Tomography Image Reconstruction. arXiv 2020, arXiv:2006.06270. [Google Scholar]
- Hadamard, J. Lectures on Cauchy’s Problem in Linear Partial Differential Equations; Dover: New York, NY, USA, 1952. [Google Scholar]
- Nashed, M. A new approach to classification and regularization of ill-posed operator equations. In Inverse and Ill-Posed Problems; Engl, H.W., Groetsch, C., Eds.; Academic Press: Cambridge, MA, USA, 1987; pp. 53–75. [Google Scholar] [CrossRef]
- Natterer, F.; Wübbeling, F. Mathematical Methods in Image Reconstruction; Society for Industrial and Applied Mathematics: Philadelphia, PA, USA, 2001. [Google Scholar]
- Saad, Y. Iterative Methods for Sparse Linear Systems, 2nd ed.; Society for Industrial and Applied Mathematics: Philadelphia, PA, USA, 2003. [Google Scholar]
- Björck, Å.; Elfving, T.; Strakos, Z. Stability of conjugate gradient and Lanczos methods for linear least squares problems. SIAM J. Matrix Anal. Appl. 1998, 19, 720–736. [Google Scholar] [CrossRef]
- Chen, H.; Wang, C.; Song, Y.; Li, Z. Split Bregmanized anisotropic total variation model for image deblurring. J. Vis. Commun. Image Represent. 2015, 31, 282–293. [Google Scholar] [CrossRef]
- Wang, Z.; Bovik, A.C.; Sheikh, H.R.; Simoncelli, E.P. Image quality assessment: From error visibility to structural similarity. IEEE Trans. Image Process. 2004, 13, 600–612. [Google Scholar] [CrossRef] [PubMed]
- Goodfellow, I.; Bengio, Y.; Courville, A. Deep Learning; MIT Press: Cambridge, MA, USA, 2016; Available online: http://www.deeplearningbook.org (accessed on 1 March 2021).
- Leuschner, J.; Schmidt, M.; Ganguly, P.S.; Andriiashen, V.; Coban, S.B.; Denker, A.; van Eijnatten, M. Supplementary Material for Experiments in “Quantitative comparison of deep learning-based image reconstruction methods for low-dose and sparse-angle CT applications”. Zenodo 2021. [Google Scholar] [CrossRef]
- Leuschner, J.; Schmidt, M.; Baguer, D.O.; Bauer, D.; Denker, A.; Hadjifaradji, A.; Liu, T. LoDoPaB-CT Challenge Reconstructions compared in “Quantitative comparison of deep learning-based image reconstruction methods for low-dose and sparse-angle CT applications”. Zenodo 2021. [Google Scholar] [CrossRef]
- Leuschner, J.; Schmidt, M.; Ganguly, P.S.; Andriiashen, V.; Coban, S.B.; Denker, A.; van Eijnatten, M. Apple CT Test Reconstructions compared in “Quantitative comparison of deep learning-based image reconstruction methods for low-dose and sparse-angle CT applications”. Zenodo 2021. [Google Scholar] [CrossRef]
- Leuschner, J.; Schmidt, M.; Otero Baguer, D.; Erzmann, D.; Baltazar, M. DIVal Library. Zenodo 2021. [Google Scholar] [CrossRef]
- Knoll, F.; Murrell, T.; Sriram, A.; Yakubova, N.; Zbontar, J.; Rabbat, M.; Defazio, A.; Muckley, M.J.; Sodickson, D.K.; Zitnick, C.L.; et al. Advancing machine learning for MR image reconstruction with an open competition: Overview of the 2019 fastMRI challenge. Magn. Reson. Med. 2020, 84, 3054–3070. [Google Scholar] [CrossRef]
- Putzky, P.; Welling, M. Invert to Learn to Invert. In Advances in Neural Information Processing Systems; Wallach, H., Larochelle, H., Beygelzimer, A., dAlch’e-Buc, F., Fox, E., Garnett, R., Eds.; Curran Associates, Inc.: Red Hook, NY, USA, 2019; Volume 32, pp. 446–456. [Google Scholar]
- Etmann, C.; Ke, R.; Schönlieb, C. iUNets: Learnable Invertible Up- and Downsampling for Large-Scale Inverse Problems. In Proceedings of the 30th IEEE International Workshop on Machine Learning for Signal Processing (MLSP 2020), Espoo, Finland, 21–24 September 2020; pp. 1–6. [Google Scholar] [CrossRef]
- Ziabari, A.; Ye, D.H.; Srivastava, S.; Sauer, K.D.; Thibault, J.; Bouman, C.A. 2.5D Deep Learning For CT Image Reconstruction Using A Multi-GPU Implementation. In Proceedings of the 2018 52nd Asilomar Conference on Signals, Systems, and Computers, Pacific Grove, CA, USA, 28–31 October 2018; pp. 2044–2049. [Google Scholar] [CrossRef]
- Scherzer, O.; Weickert, J. Relations Between Regularization and Diffusion Filtering. J. Math. Imaging Vis. 2000, 12, 43–63. [Google Scholar] [CrossRef]
- Perona, P.; Malik, J. Scale-space and edge detection using anisotropic diffusion. IEEE Trans. Pattern Anal. Mach. Intell. 1990, 12, 629–639. [Google Scholar] [CrossRef]
- Mendrik, A.M.; Vonken, E.; Rutten, A.; Viergever, M.A.; van Ginneken, B. Noise Reduction in Computed Tomography Scans Using 3-D Anisotropic Hybrid Diffusion With Continuous Switch. IEEE Trans. Med Imaging 2009, 28, 1585–1594. [Google Scholar] [CrossRef] [PubMed]
- Adler, J.; Lunz, S.; Verdier, O.; Schönlieb, C.B.; Öktem, O. Task adapted reconstruction for inverse problems. arXiv 2018, arXiv:1809.00948. [Google Scholar]
- Boink, Y.E.; Manohar, S.; Brune, C. A partially-learned algorithm for joint photo-acoustic reconstruction and segmentation. IEEE Trans. Med. Imaging 2019, 39, 129–139. [Google Scholar] [CrossRef]
- Handels, H.; Deserno, T.M.; Maier, A.; Maier-Hein, K.H.; Palm, C.; Tolxdorff, T. Bildverarbeitung für die Medizin 2019; Springer Fachmedien Wiesbaden: Wiesbaden, Germany, 2019. [Google Scholar] [CrossRef]
- Mason, A.; Rioux, J.; Clarke, S.E.; Costa, A.; Schmidt, M.; Keough, V.; Huynh, T.; Beyea, S. Comparison of objective image quality metrics to expert radiologists’ scoring of diagnostic quality of MR images. IEEE Trans. Med. Imaging 2019, 39, 1064–1072. [Google Scholar] [CrossRef] [PubMed]
- Coban, S.B.; Lionheart, W.R.B.; Withers, P.J. Assessing the efficacy of tomographic reconstruction methods through physical quantification techniques. Meas. Sci. Technol. 2021. [Google Scholar] [CrossRef]
- Goodfellow, I.J.; Shlens, J.; Szegedy, C. Explaining and Harnessing Adversarial Examples. In Proceedings of the 3rd International Conference on Learning Representations (ICLR 2015), San Diego, CA, USA, 7–9 May 2015. [Google Scholar]
- Antun, V.; Renna, F.; Poon, C.; Adcock, B.; Hansen, A.C. On instabilities of deep learning in image reconstruction and the potential costs of AI. Proc. Natl. Acad. Sci. USA 2020. [Google Scholar] [CrossRef]
- Gottschling, N.M.; Antun, V.; Adcock, B.; Hansen, A.C. The troublesome kernel: Why deep learning for inverse problems is typically unstable. arXiv 2020, arXiv:2001.01258. [Google Scholar]
- Schwab, J.; Antholzer, S.; Haltmeier, M. Deep null space learning for inverse problems: Convergence analysis and rates. Inverse Probl. 2019, 35, 025008. [Google Scholar] [CrossRef]
- Chambolle, A.; Pock, T. A first-order primal-dual algorithm for convex problems with applications to imaging. J. Math. Imaging Vis. 2011, 40, 120–145. [Google Scholar] [CrossRef]
- Kingma, D.P.; Ba, J. Adam: A Method for Stochastic Optimization. In Proceedings of the 3rd International Conference on Learning Representations (ICLR 2015), San Diego, CA, USA, 7–9 May 2015. [Google Scholar]
- Hinton, G.; Srivastava, N.; Swersky, K. Neural networks for machine learning lecture 6a overview of mini-batch gradient descent. Lect. Notes 2012, 14, 1–31. [Google Scholar]
- Winkler, C.; Worrall, D.; Hoogeboom, E.; Welling, M. Learning likelihoods with conditional normalizing flows. arXiv 2019, arXiv:1912.00042. [Google Scholar]
- Dinh, L.; Sohl-Dickstein, J.; Bengio, S. Density estimation using Real NVP. In Proceedings of the 5th International Conference on Learning Representations (ICLR 2017), Toulon, France, 24–26 April 2017. [Google Scholar]
- Dinh, L.; Krueger, D.; Bengio, Y. NICE: Non-linear Independent Components Estimation. In Proceedings of the 3rd International Conference on Learning Representations (ICLR 2015), San Diego, CA, USA, 7–9 May 2015. [Google Scholar]
- Kingma, D.P.; Dhariwal, P. Glow: Generative Flow with Invertible 1x1 Convolutions. In Proceedings of the Advances in Neural Information Processing Systems 31: Annual Conference on Neural Information Processing Systems 2018 (NeurIPS 2018), Montréal, QC, Canada, 3–8 December 2018; Bengio, S., Wallach, H.M., Larochelle, H., Grauman, K., Cesa-Bianchi, N., Garnett, R., Eds.; pp. 10236–10245. [Google Scholar]
- Daubechies, I.; Defrise, M.; De Mol, C. An iterative thresholding algorithm for linear inverse problems with a sparsity constraint. Commun. Pure Appl. Math. 2004, 57, 1413–1457. [Google Scholar] [CrossRef]
- Gregor, K.; LeCun, Y. Learning fast approximations of sparse coding. In Proceedings of the 27th International Conference on International Conference on Machine Learning, Haifa, Israel, 21–24 June 2010; pp. 399–406. [Google Scholar]
- Lempitsky, V.; Vedaldi, A.; Ulyanov, D. Deep Image Prior. In Proceedings of the 2018 IEEE/CVF Conference on Computer Vision and Pattern Recognition, Salt Lake City, UT, USA, 18–23 June 2018; pp. 9446–9454. [Google Scholar] [CrossRef]
- Dittmer, S.; Kluth, T.; Maass, P.; Otero Baguer, D. Regularization by Architecture: A Deep Prior Approach for Inverse Problems. J. Math. Imaging Vis. 2019, 62, 456–470. [Google Scholar] [CrossRef]
- Chakrabarty, P.; Maji, S. The Spectral Bias of the Deep Image Prior. arXiv 2019, arXiv:1912.08905. [Google Scholar]
- Heckel, R.; Soltanolkotabi, M. Denoising and Regularization via Exploiting the Structural Bias of Convolutional Generators. Int. Conf. Learn. Represent. 2020. [Google Scholar]
- Adler, J.; Kohr, H.; Ringh, A.; Moosmann, J.; Banert, S.; Ehrhardt, M.J.; Lee, G.R.; Niinimäki, K.; Gris, B.; Verdier, O.; et al. Operator Discretization Library (ODL). Zenodo 2018. [Google Scholar] [CrossRef]
- Van Aarle, W.; Palenstijn, W.J.; De Beenhouwer, J.; Altantzis, T.; Bals, S.; Batenburg, K.J.; Sijbers, J. The ASTRA Toolbox: A platform for advanced algorithm development in electron tomography. Ultramicroscopy 2015, 157, 35–47. [Google Scholar] [CrossRef] [PubMed]
- Coban, S. SophiaBeads Dataset Project Codes. Zenodo. 2015. Available online: http://sophilyplum.github.io/sophiabeads-datasets/ (accessed on 10 June 2020).
- Wang, T.; Nakamoto, K.; Zhang, H.; Liu, H. Reweighted Anisotropic Total Variation Minimization for Limited-Angle CT Reconstruction. IEEE Trans. Nucl. Sci. 2017, 64, 2742–2760. [Google Scholar] [CrossRef]
- Paszke, A.; Gross, S.; Massa, F.; Lerer, A.; Bradbury, J.; Chanan, G.; Killeen, T.; Lin, Z.; Gimelshein, N.; Antiga, L.; et al. PyTorch: An Imperative Style, High-Performance Deep Learning Library. In Advances in Neural Information Processing Systems 32; Wallach, H., Larochelle, H., Beygelzimer, A., dAlch’e-Buc, F., Fox, E., Garnett, R., Eds.; Curran Associates, Inc.: Red Hook, NY, USA, 2019; pp. 8024–8035. [Google Scholar]
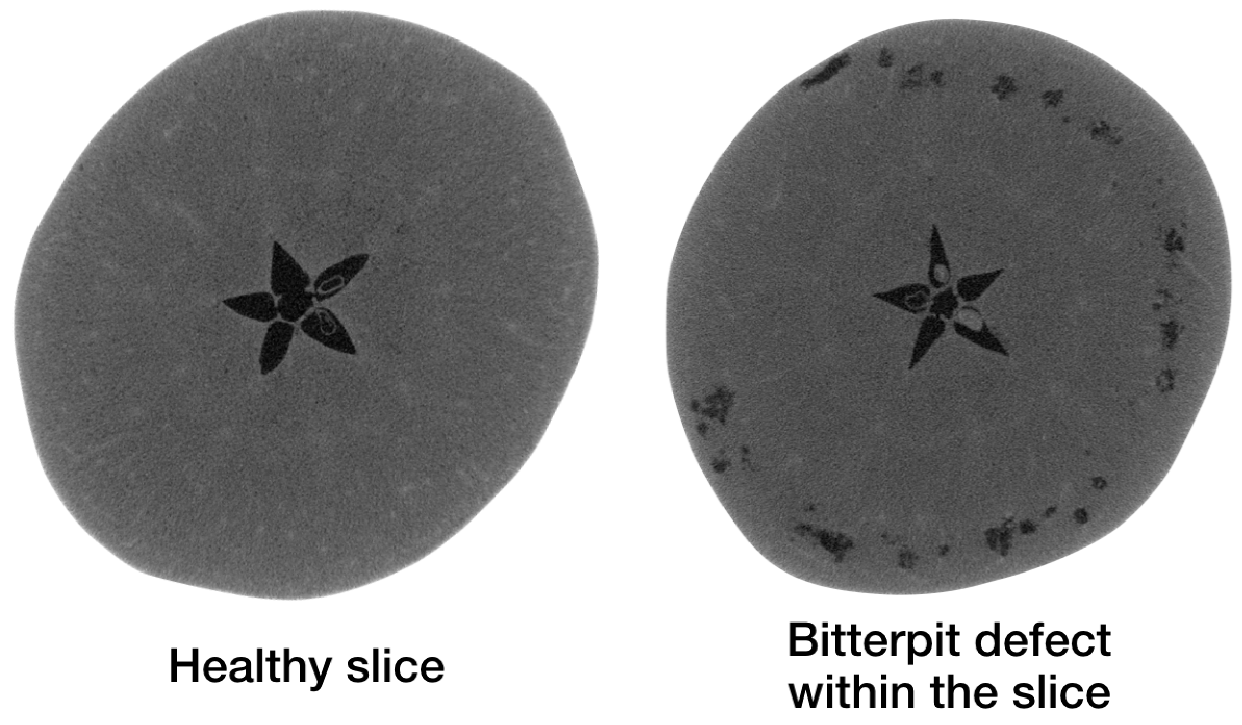

| Property | LoDoPaB-CT | Apple CT |
|---|---|---|
| Subject | Human thorax | Apples |
| Scenario | low photon count | sparse-angle |
| Challenge | 3678 reconstructions | 100 reconstructions |
| Image size | ||
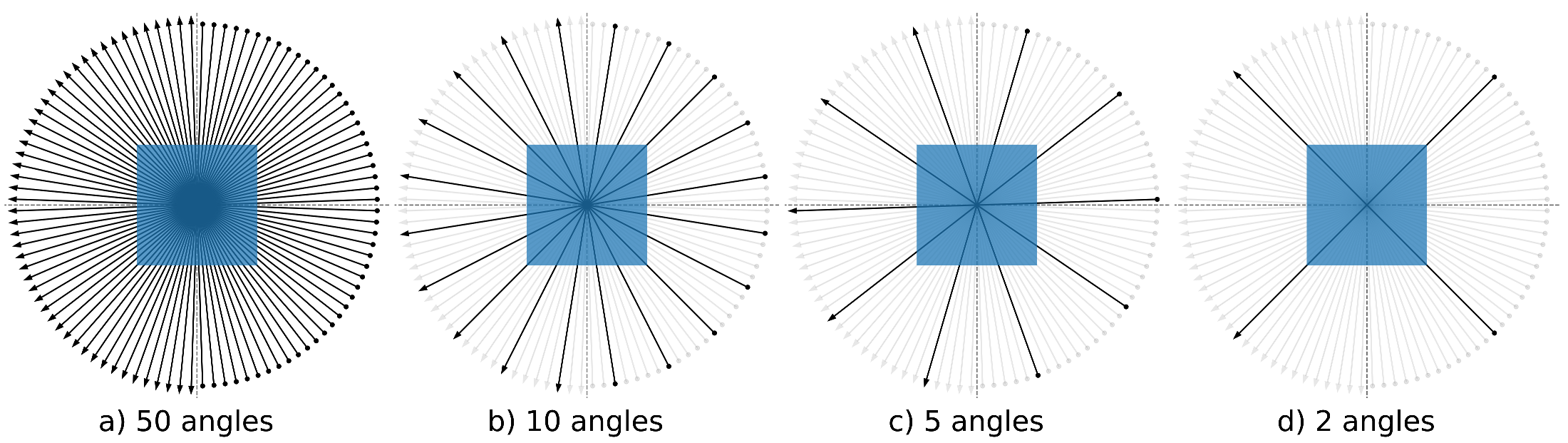
| Angles | 1000 | 50, 10, 5, 2 |
| Detector bins | 513 | 1377 |
| Sampling ratio | ≈3.9 | ≈0.07– |
| Model | PSNR | PSNR-FR | SSIM | SSIM-FR | Number of Parameters |
|---|---|---|---|---|---|
| Learned P.-D. | 36.25 ± 3.70 | 40.52 ± 3.64 | 0.866 ± 0.115 | 0.926 ± 0.076 | 874,980 |
| ISTA U-Net | 36.09 ± 3.69 | 40.36 ± 3.65 | 0.862 ± 0.120 | 0.924 ± 0.080 | 83,396,865 |
| U-Net | 36.00 ± 3.63 | 40.28 ± 3.59 | 0.862 ± 0.119 | 0.923 ± 0.079 | 613,322 |
| MS-D-CNN | 35.85 ± 3.60 | 40.12 ± 3.56 | 0.858 ± 0.122 | 0.921 ± 0.082 | 181,306 |
| U-Net++ | 35.37 ± 3.36 | 39.64 ± 3.40 | 0.861 ± 0.119 | 0.923 ± 0.080 | 9,170,079 |
| CINN | 35.54 ± 3.51 | 39.81 ± 3.48 | 0.854 ± 0.122 | 0.919 ± 0.081 | 6,438,332 |
| DIP + TV | 34.41 ± 3.29 | 38.68 ± 3.29 | 0.845 ± 0.121 | 0.913 ± 0.082 | hyperp. |
| iCTU-Net | 33.70 ± 2.82 | 37.97 ± 2.79 | 0.844 ± 0.120 | 0.911 ± 0.081 | 147,116,792 |
| TV | 33.36 ± 2.74 | 37.63 ± 2.70 | 0.830 ± 0.121 | 0.903 ± 0.082 | (hyperp.) |
| FBP | 30.19 ± 2.55 | 34.46 ± 2.18 | 0.727 ± 0.127 | 0.836 ± 0.085 | (hyperp.) |
| Method | |
|---|---|
| Learned Primal-Dual | |
| ISTA U-Net | |
| U-Net | |
| MS-D-CNN | |
| U-Net++ | |
| CINN | |
| DIP + TV | |
| iCTU-Net | |
| TV | |
| FBP | |
| Ground truth |
| Noise-Free | PSNR | SSIM | ||||||
|---|---|---|---|---|---|---|---|---|
| Number of Angles | 50 | 10 | 5 | 2 | 50 | 10 | 5 | 2 |
| Learned Primal-Dual | 38.72 | 35.85 | 30.79 | 22.00 | 0.901 | 0.870 | 0.827 | 0.740 |
| ISTA U-Net | 38.86 | 34.54 | 28.31 | 20.48 | 0.897 | 0.854 | 0.797 | 0.686 |
| U-Net | 39.62 | 33.51 | 27.77 | 19.78 | 0.913 | 0.803 | 0.803 | 0.676 |
| MS-D-CNN | 39.85 | 34.38 | 28.45 | 20.55 | 0.913 | 0.837 | 0.776 | 0.646 |
| CINN | 39.59 | 34.84 | 27.81 | 19.46 | 0.913 | 0.871 | 0.762 | 0.674 |
| iCTU-Net | 36.07 | 29.95 | 25.63 | 19.28 | 0.878 | 0.847 | 0.824 | 0.741 |
| TV | 39.27 | 29.00 | 22.04 | 15.95 | 0.915 | 0.783 | 0.607 | 0.661 |
| CGLS | 33.05 | 21.81 | 12.60 | 15.25 | 0.780 | 0.619 | 0.537 | 0.615 |
| FBP | 30.39 | 17.09 | 15.51 | 13.97 | 0.714 | 0.584 | 0.480 | 0.438 |
| Gaussian Noise | PSNR | SSIM | ||||||
| Number of Angles | 50 | 10 | 5 | 2 | 50 | 10 | 5 | 2 |
| Learned Primal-Dual | 36.62 | 33.76 | 29.92 | 21.41 | 0.878 | 0.850 | 0.821 | 0.674 |
| ISTA U-Net | 36.04 | 33.55 | 28.48 | 20.71 | 0.871 | 0.851 | 0.811 | 0.690 |
| U-Net | 36.48 | 32.83 | 27.80 | 19.86 | 0.882 | 0.818 | 0.789 | 0.706 |
| MS-D-CNN | 36.67 | 33.20 | 27.98 | 19.88 | 0.883 | 0.831 | 0.748 | 0.633 |
| CINN | 36.77 | 31.88 | 26.57 | 19.99 | 0.888 | 0.771 | 0.722 | 0.637 |
| iCTU-Net | 32.90 | 29.76 | 24.67 | 19.44 | 0.848 | 0.837 | 0.801 | 0.747 |
| TV | 32.36 | 27.12 | 21.83 | 16.08 | 0.833 | 0.752 | 0.622 | 0.637 |
| CGLS | 27.36 | 21.09 | 14.90 | 15.11 | 0.767 | 0.624 | 0.553 | 0.616 |
| FBP | 27.88 | 17.09 | 15.51 | 13.97 | 0.695 | 0.583 | 0.480 | 0.438 |
| Scattering Noise | PSNR | SSIM | ||||||
| Number of Angles | 50 | 10 | 5 | 2 | 50 | 10 | 5 | 2 |
| Learned Primal-Dual | 37.80 | 34.19 | 27.08 | 20.98 | 0.892 | 0.866 | 0.796 | 0.540 |
| ISTA U-Net | 35.94 | 32.33 | 27.41 | 19.95 | 0.881 | 0.820 | 0.763 | 0.676 |
| U-Net | 34.96 | 32.91 | 26.93 | 18.94 | 0.830 | 0.784 | 0.736 | 0.688 |
| MS-D-CNN | 38.04 | 33.51 | 27.73 | 20.19 | 0.899 | 0.818 | 0.757 | 0.635 |
| CINN | 38.56 | 34.08 | 28.04 | 19.14 | 0.915 | 0.863 | 0.839 | 0.754 |
| iCTU-Net | 26.26 | 22.85 | 21.25 | 18.32 | 0.838 | 0.796 | 0.792 | 0.765 |
| TV | 21.09 | 20.14 | 17.86 | 14.53 | 0.789 | 0.649 | 0.531 | 0.611 |
| CGLS | 20.84 | 18.28 | 14.02 | 14.18 | 0.789 | 0.618 | 0.547 | 0.625 |
| FBP | 21.01 | 15.80 | 14.26 | 13.06 | 0.754 | 0.573 | 0.475 | 0.433 |
| Evaluation | 50 Angles | 10 Angles | 5 Angles | 2 Angles | |||||
|---|---|---|---|---|---|---|---|---|---|
| Training | PSNR | SSIM | PSNR | SSIM | PSNR | SSIM | PSNR | SSIM | |
| 50 angles | 39.62 | 0.913 | 16.39 | 0.457 | 11.93 | 0.359 | 8.760 | 0.252 | |
| 10 angles | 27.59 | 0.689 | 33.51 | 0.803 | 18.44 | 0.607 | 9.220 | 0.394 | |
| 5 angles | 24.51 | 0.708 | 26.19 | 0.736 | 27.77 | 0.803 | 11.85 | 0.549 | |
| 2 angles | 15.57 | 0.487 | 14.59 | 0.440 | 15.94 | 0.514 | 19.78 | 0.676 | |
| Model | Reconstruction Error (Image Metrics) | Training Time | Recon- Struction Time | GPU Memory | Learned Para- Meters | Uses Discre- Pancy | Operator Required | |
|---|---|---|---|---|---|---|---|---|
| Learned P.-D. | no | |||||||
| ISTA U-Net | no | |||||||
| U-Net | no | |||||||
| MS-D-CNN | no | |||||||
| U-Net++ | - | no | ||||||
| CINN | no | |||||||
| DIP + TV | - | - | 3+ | yes | ||||
| iCTU-Net | no | |||||||
| TV | - | 3 | yes | |||||
| CGLS | - | - | 1 | yes | ||||
| FBP | - | 2 | no | |||||
| Legend | LoDoPaB | Apple CT | Rough values for Apple CT Dataset B | |||||
| Avg. improv. over FBP | (varying for different setups and datasets) | |||||||
| 0% | 0–15% | >2 weeks | >10 min | >10 GiB | > | Direct | ||
| 12–16% | 25–30% | >5 days | >30 s | >3 GiB | > | In network | ||
| 17–20% | 40–45% | >1 day | >0.1 s | >1.5 GiB | > | For input | ||
| 50–60% | ≤0.02 s | ≤1 GiB | ≤ | Only concept | ||||
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Leuschner, J.; Schmidt, M.; Ganguly, P.S.; Andriiashen, V.; Coban, S.B.; Denker, A.; Bauer, D.; Hadjifaradji, A.; Batenburg, K.J.; Maass, P.; et al. Quantitative Comparison of Deep Learning-Based Image Reconstruction Methods for Low-Dose and Sparse-Angle CT Applications. J. Imaging 2021, 7, 44. https://doi.org/10.3390/jimaging7030044
Leuschner J, Schmidt M, Ganguly PS, Andriiashen V, Coban SB, Denker A, Bauer D, Hadjifaradji A, Batenburg KJ, Maass P, et al. Quantitative Comparison of Deep Learning-Based Image Reconstruction Methods for Low-Dose and Sparse-Angle CT Applications. Journal of Imaging. 2021; 7(3):44. https://doi.org/10.3390/jimaging7030044
Chicago/Turabian StyleLeuschner, Johannes, Maximilian Schmidt, Poulami Somanya Ganguly, Vladyslav Andriiashen, Sophia Bethany Coban, Alexander Denker, Dominik Bauer, Amir Hadjifaradji, Kees Joost Batenburg, Peter Maass, and et al. 2021. "Quantitative Comparison of Deep Learning-Based Image Reconstruction Methods for Low-Dose and Sparse-Angle CT Applications" Journal of Imaging 7, no. 3: 44. https://doi.org/10.3390/jimaging7030044
APA StyleLeuschner, J., Schmidt, M., Ganguly, P. S., Andriiashen, V., Coban, S. B., Denker, A., Bauer, D., Hadjifaradji, A., Batenburg, K. J., Maass, P., & van Eijnatten, M. (2021). Quantitative Comparison of Deep Learning-Based Image Reconstruction Methods for Low-Dose and Sparse-Angle CT Applications. Journal of Imaging, 7(3), 44. https://doi.org/10.3390/jimaging7030044







