1. Introduction
The point spread function (PSF) is the image of a point source of light and the basic unit that makes up any image ([
1], Chapter 14.4.1, pp. 335–337). Its precise knowledge is fundamental for the characterization of any imaging system, as well as for obtaining reliable results in applications like deconvolution [
2], object localization [
3] or superresolution imaging [
4,
5], to mention some. In conventional fluorescence microscopy, PSF determination has become a common task, due to its strong dependence on acquisition conditions. From its acquaintance, it is possible to determine instrument capacities to resolve details, restore out of focus information, and so forth. However, it is always restricted to a bounded and specific set of acquisition conditions. In fact, in optical sectioning microscopy, the three-dimensional (3D) PSF presents a significant spatial variance along the optical axis as a consequence of the aberrations introduced by the technique. Therefore, a single 3D PSF determination is not enough for a complete description of the system.
The PSF can be determined in three ways, namely experimental, theoretical or analytical ([
1], Chapter 14.4.1, pp. 335–337) [
6]. In the experimental approach, fluorescent sub-resolution microspheres are used for this purpose. The microsphere images must be acquired under the same conditions in which the specimen is registered. This is methodologically difficult to do, and the experimental PSFs just represent a portion of the whole space. However, it has the advantage of generating a more realistic PSF, evidencing aberrations that are difficult to model. The theoretical determination, in turn, responds to a set of mathematical equations that describe the physical model of the optics. The model’s parameter values are filled with the instrument data-sheet and the capture conditions. The modeling of certain aberrations and the absence of noise are its main advantages. The third way to determine PSF was originated from parametric blind deconvolution algorithms; it has the advantage of estimating the model parameters from real data, but its main application has been image restoration.
Along the last decade, alternative ways of 3D PSF determination have appeared because of the emergence of new applications in optical microscopy (e.g., localization microscopy; see recent review articles [
3,
7,
8]). In a certain sense, these forms are similar to the PSF estimation by blind parametric deconvolution, since they fit a parametric model to an experimental 3D PSF. The contributions in this field are mainly represented by the works carried out by Hanser et al. [
9], Aguet et al. [
10], Mortensen et al. [
11] and Kirshner et al. [
12] and are aimed at connecting the physical formulation of the image formation and the experimental PSF. Hanser et al. [
9] developed a methodology to estimate the microscopy parameters using recovery algorithms of the phase pupil function from experimental 3D PSF. Meanwhile, Aguet et al. [
10] and Kirshner et al. [
12] contributed with algorithms to estimate the 3D PSF parameters of the Gibson and Lanni theoretical model [
13] from real data. Furthermore, Aguet et al. [
10] investigated the feasibility of particle localization, from blurred sections, analyzing the Cramer–Rao theoretical lower bound.
Regarding parameter estimation method comparison applied to optical microscopy, one of the first method comparison studies was made by Cheezum et al. [
14], who quantitatively compared the performance of four commonly-used methods for particle tracking problems: cross-correlation, sum-absolute difference, centroid and direct Gaussian fit. They found that the cross-correlation algorithm method was the most accurate for large particles, and for smaller point sources, direct Gaussian fit to the intensity distribution was superior in terms of both accuracy and precision, as well as robustness at a low signal-to-noise ratio (SNR). Abraham et al. [
15] studied comparatively the performance of maximum likelihood and non-linear least square estimation methods for fitting single molecule data under different scenarios. Their results showed that both estimators, on average, are able to recover the true planar location of the single molecule, in all the scenarios they examined. In particular, they found that under model misspecifications and low noise levels, maximum likelihood is more precise than the non-linear least square method. Abraham et al. [
15] used Gaussian and Airy profiles to model the PSF and to study the effect of pixelation on the results of the estimation methods.
All of these have been important advances in the research and development of tools for the precise determination of the PSF. However, according to the available literature, just a few ways of parameter estimation methods for theoretical PSF have been developed and, to our knowledge, there have been no comparative studies analyzing methods for parameter estimation of a more realistic PSF physical model. Thus, to contribute in this regard, in this article, a comparative computational study of three methods of 3D PSF parameter estimation of a wide-field fluorescence microscope is described. These were applied to improve the precision of the position on the optical axis of a point source using the model developed by Gibson and Lanni [
13]. This parameter was selected because of its importance in applications such as depth-variant deconvolution [
16,
17] and localization microscopy [
3,
7], which is also an unknown parameter in a variant and non-linear image formation physical model.
The method comparison described in this report is based on sampling theory statistics, although it had not been devised in this way. Thus, it is essential to mention that there are several analyses based on the Bayesian paradigm of image interpretation. Shaevitz [
18] developed an algorithm to localize a particle in the axial direction, which is also compared with other previously-developed methods that account for the PSF axial asymmetry. Rees et al. [
19] analyzed the resolution achieved in localization microscopy experiments, and they highlighted the importance of assessing the resolution directly from the image samples by analyzing their particular datasets, which can significantly differ from the resolution evaluated from a calibration sample. De Santis et al. [
5] obtained a precision expression for axial localization based on standard deviation measurements of the PSF intensity profile fitted to a Gaussian profile. Finally, a specific application can be found in the works of Cox et al. [
20] and Walde et al. [
21], who used the Bayesian approach for revealing the podosome biodynamics.
The remainder of this report is organized as follows. First, a description of the physical PSF model is presented; next, three forms of 3D PSF parameter estimation are shown, namely Csiszár
I-divergence minimization (our own preceding development), maximum likelihood [
10] and least square [
12]. Since all expressions to optimize are of a non-linear nature, they are approximated using the same series representation, obtaining three iterative algorithms that differ in the comparison criteria between the data and the model. Following, in the Results Section, the performance of the algorithms has been extensively analyzed using synthetic data generated with different sources and noise levels. This was carried out evaluating: success percentage, number of iterations, computation time, accuracy and precision. As an application, the position on the optical axis of an experimental 3D PSF is estimated by the three methods, taking the difference between the full width at half maximum (FWHM) between the estimated and experimental 3D PSF as the comparison criterion. Results are analyzed in the Discussion Section. Finally, the main conclusions of this research are presented.
2. PSF Model
The model developed by Gibson and Lanni [
13] provides an accurate way to determine the 3D PSF for a fluorescence microscope. This is based on the calculation of phase aberration (or aberration function)
W [
1,
22,
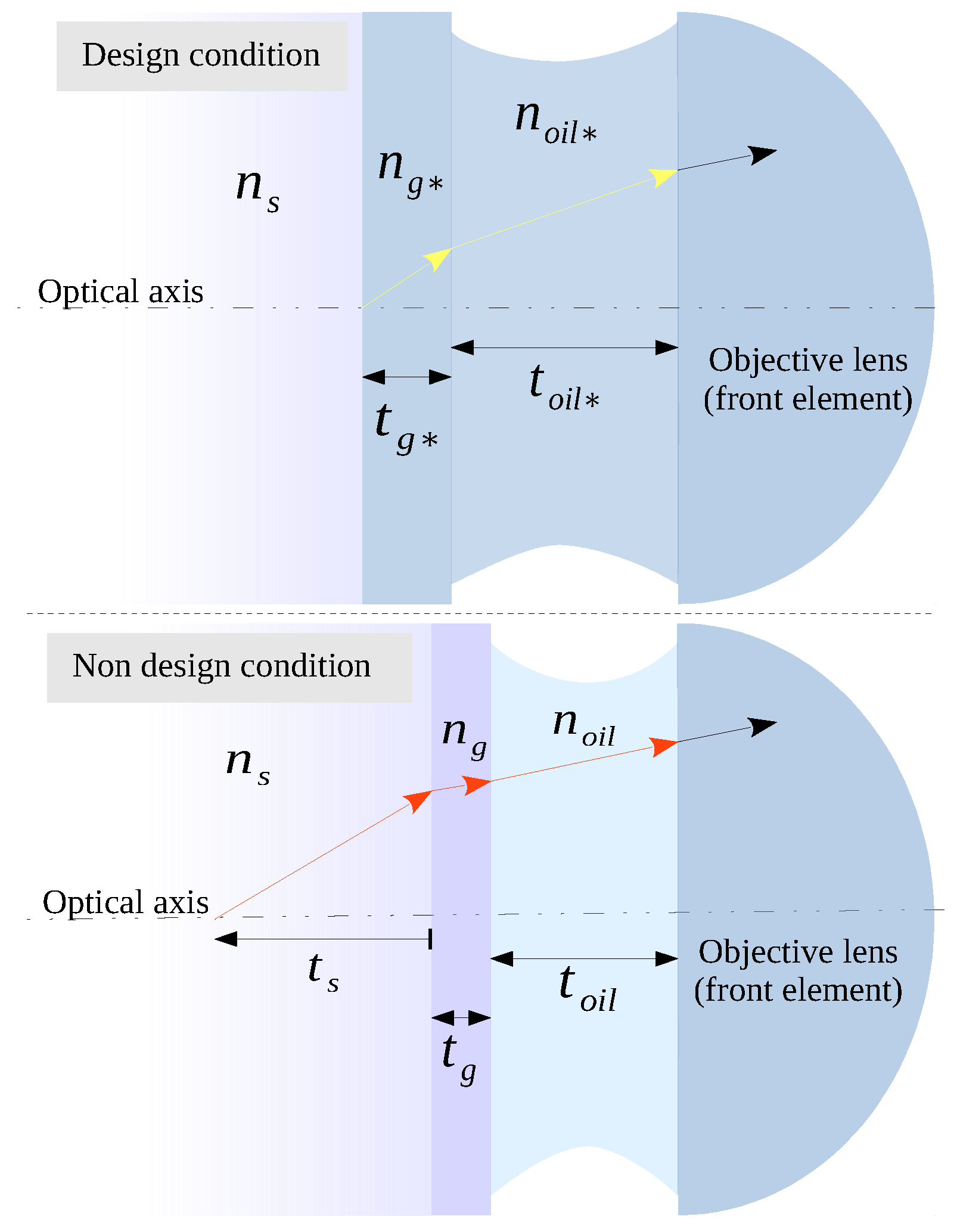
23], from the optical path difference (OPD) between two rays, originating from different conditions. One supposes the optical system used according to optimal settings given by the manufacturer, or design conditions, and the other under non-design conditions, a much more realistic situation. In design conditions (see
Figure 1 and
Table 1 for a reference), the object plane, immediately below the coverslip, is in focus in the design plane of the detector (origin of the yellow arrow). However, in non-design conditions, to observe a point source located at the depth
of the specimen, the stage must be moved along the optical axis towards the lens until the desired object is in focus in the plane of the detector (origin red arrow). This displacement produces a decrease in the thickness of the immersion oil layer that separates the front element of the objective lens from the coverslip. However, this shift is not always the same because, if the rays must also pass through no nominal thicknesses and refractive indices, both belonging to the coverslip and the immersion oil, the correct adjustment of the position will depend on the combination of all of these parameters. In any case, in non-design conditions, the diffraction pattern will be degraded from the ideal situation. Indeed, the more apart an acquisition setup is from design conditions, the more degraded the quality of the images will be.
In design conditions, the image of a point source of an ideal objective lens is the diffraction pattern produced by the spherical wave converging to the Gaussian image point in the design plane of the detector that is diffracted by the lens aperture. In this ideal case, the image is considered aberration-free and diffraction-limited. A remarkable characteristic of this model is that the OPD depends on a parameter set easily obtainable from the instrument data sheets. With this initial statement, Gibson and Lanni obtained an approximation to the OPD given by the following expression:
whose parameters are described in
Table 1. In particular,
is defined as the best geometrical focus of the system,
is the radius of the projection of the limiting aperture and
ρ is the normalized radius, these last two being defined in the back focal plane of the objective lens.
With the phase aberration, defined as
, where
is the wave number and
λ the wavelength of the source, the intensity field on the detector position
of the image space
due to a point source placed at the position
of the object space
of the system in non-design conditions can be computed by Kirchhoff’s diffraction integral:
being
C a constant complex amplitude,
the set of the
L fixed parameters listed in
Table 1 and
is the Bessel function of the first kind of first order.
It is of practical interest to relate the detector coordinates
in the image space
with the coordinates
in the object space. This can be done projecting the image plane
to the front focal plane. This projection counteracts the magnification and the 180 degree rotation with respect to the optical axis, placing the image on the specimen’s coordinate system [
23]. In order to reduce expressions, it will be assumed that
represents, generically, spatial position.
From a statistic point of view, the Gibson and Lanni model, appropriately normalized, represents the probability distribution of detecting photons in different spatial places. Therefore, without loss of generality, it can be assumed that [
24,
25]:
Alternatively, in the discrete formulation, which will be used in this work,
where
N is the total number of voxels and
n the voxel position in the object space. Conditional probability is represented by
instead of
, in order to reduce expressions.
2.1. OPD Function Implementation
In the OPD approximation (
1), there are terms that cancel if the corresponding nominal and actual parameters are equal. The implementation in GNU Octave [
26] coded for this work takes into account all parameters of the given formula. However, it is previously built with the parameters that will be used, evaluating the difference between the corresponding nominal and real parameters and binding those terms to the corresponding parameters whose difference is not zero. This reduces the computational time in those cases where the terms with corresponding parameters are equal.
2.2. Numerical Integration of the Model
The integral of Equation (
2) was tested with several numerical integration methods available in GNU Octave [
26]. All return similar qualitative results. However, in terms of speed of computation, the Gauss–Kronrod quadrature [
27] resolved the integral in a shorter time than the other methods, and, because of this, it was selected to compute the integral of the Equation (
2). Since this method could not converge (e.g., by stack overflow of the recursive calls), an alternative method that uses the adaptive Simpson rule is used. The implementation of the Gibson and Lanni model of this work adds a control logic that determines, as a function of the refraction index
and the numerical aperture (NA), an upper limit to the normalized radius of integration,
ρ. This avoids that expressions within square roots in Equation (
1) turn out to be negative.
2.3. Implementation of the 3D PSF Function
A GNU Octave function [
26] to generate 3D PSF was implemented. It takes as input the PSF model as an octave handle function, pixel size, plane separation, row number, column number, plane number, normalization type and shift of the PSF peak along the optical axis. The normalization parameter allows two forms of data normalization. The main form for this work corresponds to the total sum of intensities for each plane and relative to the plane with more accumulated intensity. This form simulates the intensity distributions in optical sectioning for a stable source and fixed exposure time. The peak shift parameter allows to center the PSF peak in a specific plane.
2.4. General Model for a Noisy PSF
The digital capture of optical images is a process that can be considered as an additional cause of aberration, due to the incidence of the noise sources involved in the acquisition process ([
1], Chapter 12, p. 251). As a general model, it can be considered that the number of photons
in each voxel is given by:
where
represents the intrinsic noise of the signal,
c is a constant that converts intensities of
to photon flux and
represents extrinsic noise sources of a given signal (e.g., read-out noise); assuming for each noise source a probability distribution model [
1,
10]. Simulated or acquired data will be called
.
3. Comparing Experimental and Theoretical 3D PSF
The building of a method for parameter estimation requires the prior definition of a comparison criterion, or objective function, between the parametric model and the data. Then, this function is optimized to find the optimal parameters that, according to the criterion, make the model more similar to the data.
In this report, parameter estimation by least square error, maximum likelihood and Csiszár
I-divergence minimization (our own development) are described. These estimation methods lead to non-linear function optimizations, and since the purpose of this study is to compare them, all were approximated to the first order term of their Taylor series representations, obtaining iterative estimators. In other words, assuming that the objective function is given by
, the problem is to find which
θ makes
optimum. This is,
The basic Taylor series expansion of the non-linear function
is given by,
where
is an arbitrary point near
around which
is evaluated. Therefore, if
is a root of
, then the left side of Equation (
7) is zero, and the right side of Equation (
7) can be truncated to the first order term of the series to get a linear approximation of the root. That is,
This approximation can be improved by successive substitutions, obtaining the iterative method known as Newton–Raphson ([
28], Chapter 5, pp. 127–129), which can be written as,
where the superscript
indicates the iteration number. The initial estimate (i.e.,
) has to be near the root to ensure the convergence of the method.
3.1. Least Square Error
Historically, in the signal processing field, the square error (SE) has been used as a quantitative metric of performance. From an optimization point of view, SE exhibits convexity, symmetry and differentiability properties, and its solutions usually have analytic closed forms; when not, there exist numerical solutions that are easy to formulate [
29]. Kirshner et al. [
12] developed and evaluated a fitting method based on the SE optimization between data and the Gibson and Lanni model [
13]. They used the Levenberg–Marquardt algorithm to solve the non-linear least square problem, obtaining low computation times, as well as accurate and precise approximation to the point source localization. In the present work, this algorithm was not considered. Instead, an iterative formulation for the estimator was obtained, considering the SE objective function given by:
Following the procedure given by Equations (
6)–(
9), the minimum of SE
is found making its partial derivatives equal to zero, that is:
The representation of Equation (
11) to the first order term of the Taylor series leads to the following iterative formulation of the estimator:
3.2. Csiszár I-Divergence Minimization
It has been argued that in cases where data do not belong to the set of positive real numbers, the SE does not reach a minimum [
30,
31], this being a fact in optical images, because the intensities captured are always nonnegative. An alternative is to use Csiszár
I-divergence, which is a generalization of the Kullback–Leibler information measure between two probability mass functions (or density, in the continuous case)
p and
r. It is defined as [
30]:
Compared with the Kullback–Leibler information measure, Csiszár adjusts the
p and
r functions for those cases in which their integrals are not equal [
31], adding the last summation term on the right side of the Equation (
13). Because of this,
has the following properties:
Consequently, for a fixed
r,
has a unique minimum, which is reached when
. The probability mass function
r operates as a reference from which the function
p follows peaks and valleys of
r. In this work, it is proposed that the reference function will be the data normalized so that their sum is equal to one. This normalized data form, which will be called
, represents the relative frequency of the photon count in each spatial position. This is,
Then, replacing
in Equation (
13) by
and
by
, the
I-divergence between the normalized data and the theoretical distribution
is given by:
To minimize
, its partial derivatives must be equal to zero, that is:
Representing Equation (
16) in Taylor series and approximating to the term of first order, the following iterative estimator is obtained:
3.3. Maximum Likelihood
The maximum likelihood principle establishes that data that occur must have, presumably, the maximum probability of occurrence [
25]. Aguet et al. [
10] used this idea to build an estimation method of the position of a point source along the optical axis. The likelihood function (or log-likelihood), represented by the expected number of photons in a point of the space, is given by:
which is built under the assumptions of a forward model given by Equation (
5), with
following a Poisson distribution and
representing read-out noise of the acquisition device,
, which follows a Gaussian distribution. With these assumptions, the authors build an iterative formulation of the estimator based on the maximization for the log-likelihood function between captured data and the Gibson and Lanni model. They also used a numerical approximation to the term of the first order of the Taylor series, obtaining:
3.4. Implementation of the Estimation Methods
The above described estimation methods were implemented as GNU Octave functions [
26], taking as input values the experimental PSF data properly normalized, a function handle to compute theoretical PSF and, as output conditions, an absolute tolerance value and a maximum number of iterations in case the methods do not converge. The derivatives of the Gibson and Lanni model in the iterative Equations (
12), (
17) and (
19) were evaluated numerically with a forward step. The outputs of these Octave functions return the estimation for each iteration.
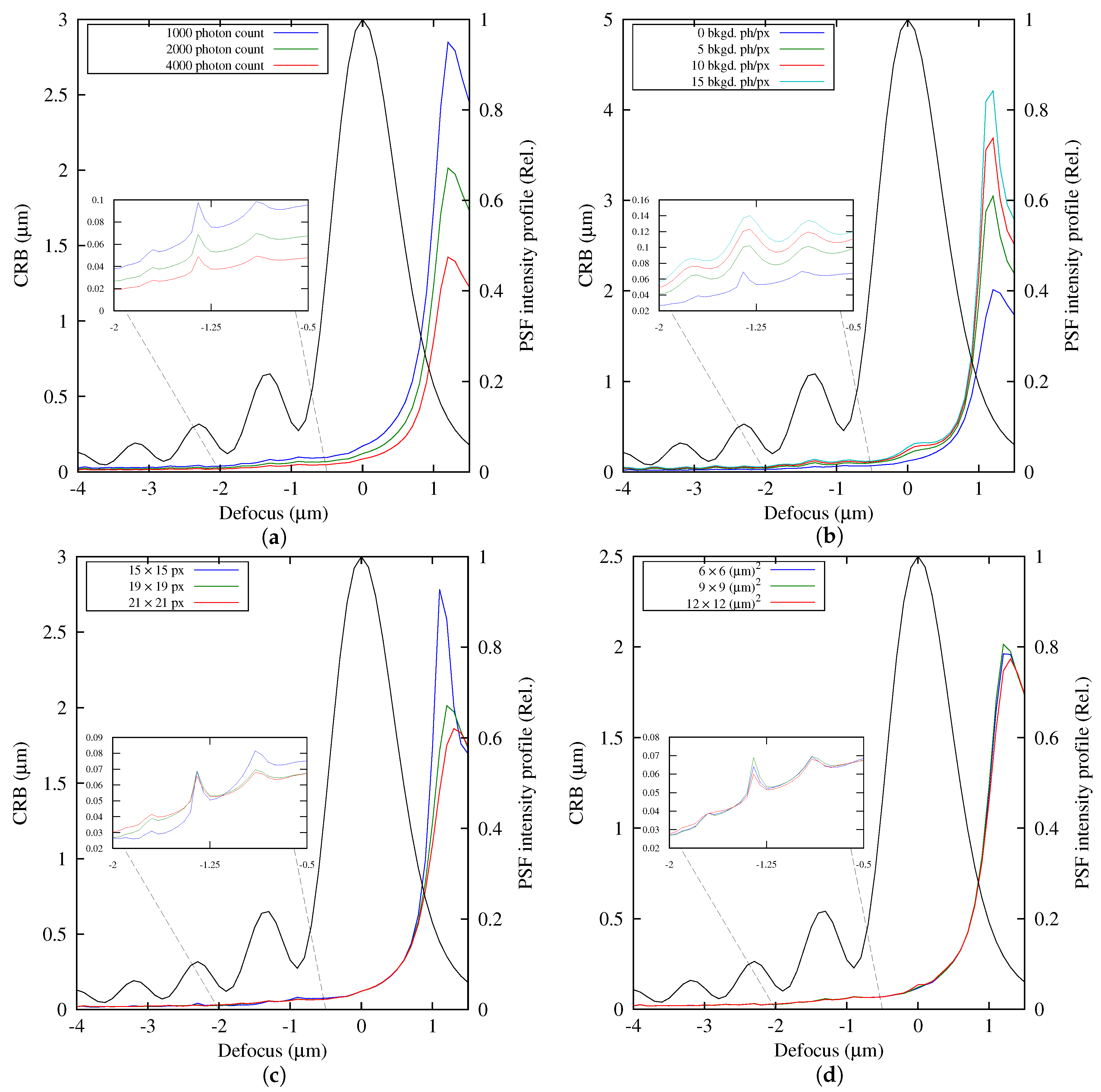
3.5. Lower Bound of the Estimation Error
Independently of the estimation method used to fit the parameters of a model, if the estimator is unbiased, its variance is bounded. This lower bound to the estimation method precision is commonly known as the Cramer–Rao bound. It is a general result that depends only on the image formation model ([
25], Chapter 17, pp. 391–392). Aguet et al. [
10] obtained a form for the Cramer–Rao bound calculation for axial localization, which is based on the image formation model given by Equation (
5) with intrinsic and extrinsic noise sources that are Poisson distributed. Abraham et al. [
15] used two forms of this bound for planar localization: a fundamental (or theoretical) one, which considers ideal detection conditions, and a practical bound, which ponders the limitations of the detector and noise sources that corrupt the signal. In the current work, the form obtained by Aguet et al. [
10] was used.
5. Discussion
To our knowledge, just a few ways of parameter estimation methods for theoretical PSF have been developed, and there are no comparative studies of parameter estimation methods applied to microscopy PSF theoretical models. The aim of this work was to contribute to these particular aspects, by carrying out a comparative computational study of three parameter estimation methods for the theoretical model of Gibson and Lanni PSF [
13]. This was applied to the position estimation of a point source along the optical axis. The parameter estimation methods analyzed were: Csiszár
I-divergence minimization (a previously unpublished own method), maximum likelihood [
10] and least square [
12]. Since all expressions to optimize are of a non-linear nature, they were approximated using the same numerical representation, obtaining three iterative algorithms that differ in the comparison criteria between the data and model. As an application, the position on the optical axis of an experimental 3D PSF is estimated by the three methods, taking the difference of the FWHM between the estimated and experimental 3D PSF as a comparison criterion.
5.1. Gibson and Lanni Model
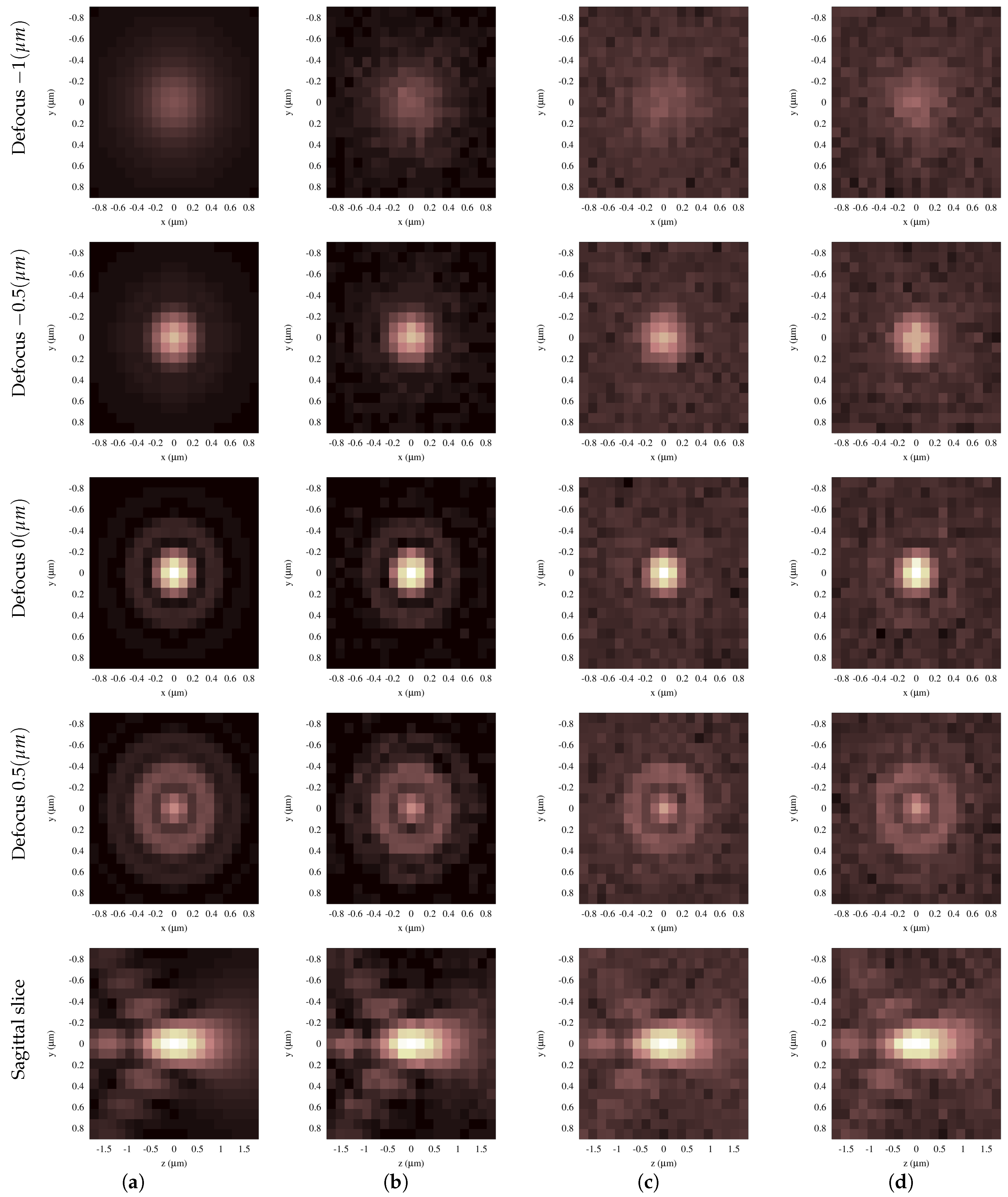
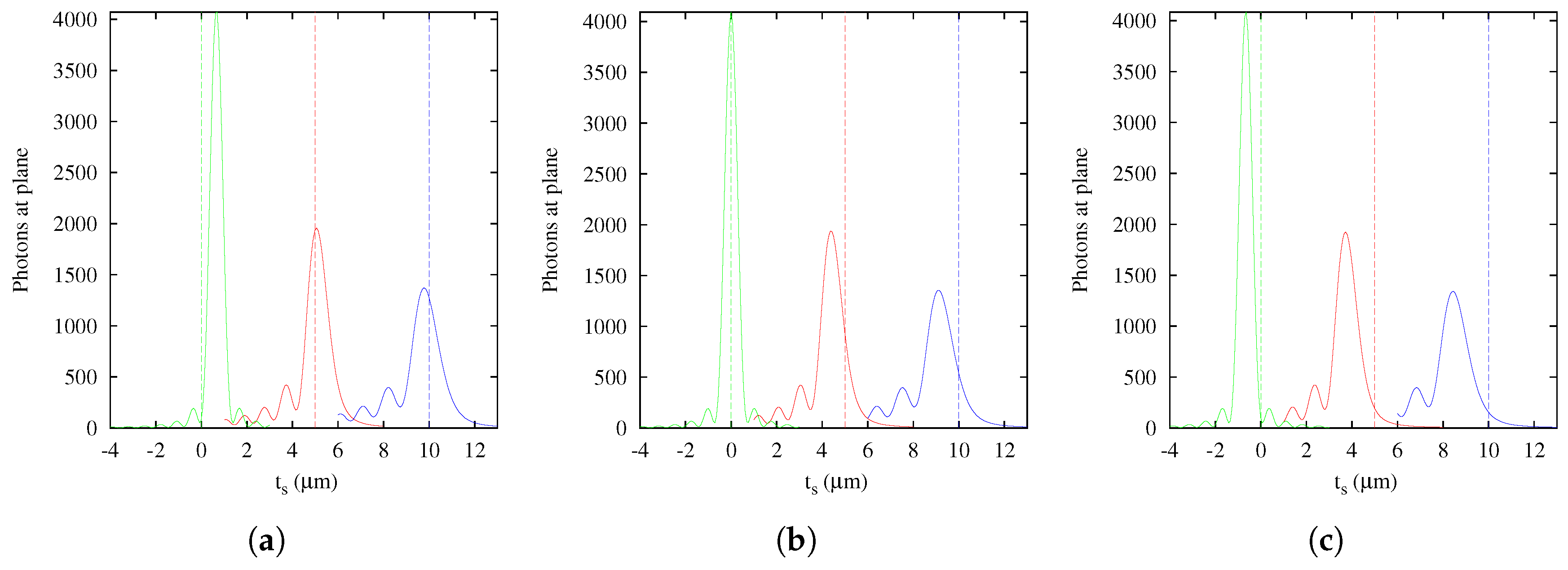
The first step in this study was the computational implementation of the Gibson and Lanni theoretical PSF model [
13]. This is a computationally-convenient scalar model that represents appropriately the spherical aberration produced by using a microscope system under non-design conditions. The optical sectioning technique is an example of this use in which, in the best scenario, the spherical aberration depends only on the depth of the fluorescent point source. In this case, the model appropriately describes the 3D PSF and evidences a strong spatial variance in the optical axis, showing at least three effects easily viewed in an intensity profile: morphology changes, including asymmetry relative to the peak, intensity changes and focal shift of the 3D PSF. This last effect reveals that the axial positions of the focused structures in images acquired by optical sectioning are not necessarily coincident with the positions directly measured from the relative distance between the cover slip and the objective lens. Therefore, in real situations where other factors are involved, it is practically impossible to know with certainty the spatial position from direct measurement between glasses and the objective lens. In fact, any small change in other parameters of the model (e.g., refraction index of immersion oil
) produces a different shift (
Figure 12). In this sense, Rees et al. [
19] have recently arrived at the same conclusion in their analysis, which included those modes of superresolution microscopy that use conventional fluorescence microscopes. In turn, this shows that estimators obtained via the sampling theory can be sufficient to provide useful information about precision and resolution. This highlights the importance of the estimation methods and the selection of an adequate image formation model for localization and deconvolution applications [
16,
17,
37].
Regarding deconvolution, a valid concern could come up for users. If the parameter estimation methods are performed over a limited axial support, what would its validity be for the larger PSF extension needed for deconvolution? It is still valid, because it is possible to have a good estimation of the PSF parameters with a smaller extension than the one used in restoration. In fact, according to the results obtained, using data from the optical axis would be enough for a good estimation of the axial position of the PSF, which, depending on the signal level, could even be in the nanometer order. Thus, parameter estimation methods could be applied to a reduced dataset to find the optimal PSF before restoration. This would need less computation time than using the full 3D PSF dataset.
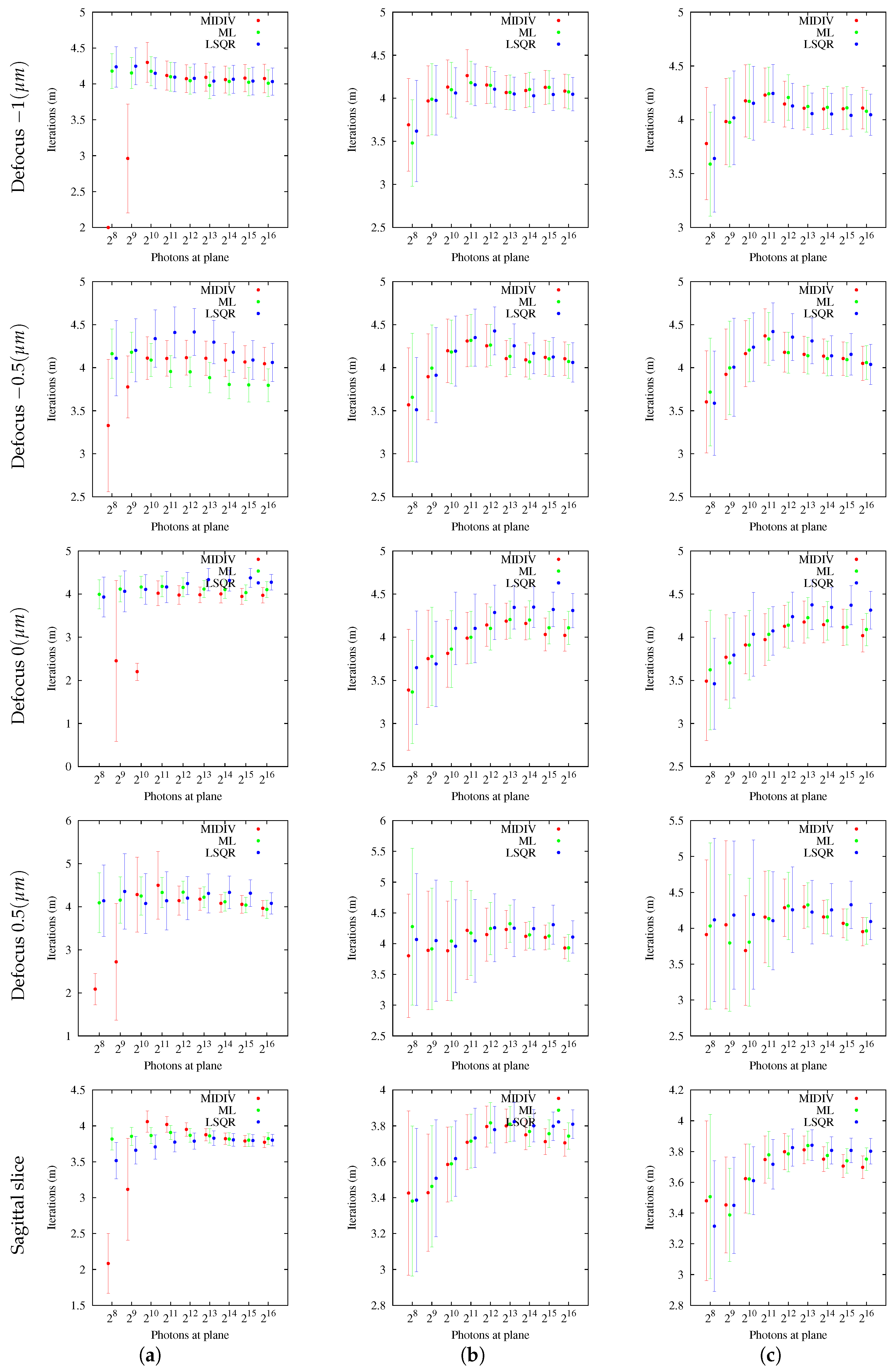
5.2. Computational Convergence
Regarding method convergence, it is necessary to point out that the numerical optimization method has a theoretical quadratic convergence. However, the tests carried out on simulated images, with and without noise, showed a logarithmic linear convergence (
Table 2) of the estimation methods considered here. A downside of the approximation method used for optimization, also discussed by Aguet et al. [
10], is that it requires a good initial estimate. The speed of the estimation methods could be improved taking into account more terms of the Taylor representation of the optimization method. However, this would require an additional performance evaluation because there might be more computations of the model due to the use of higher order derivatives.
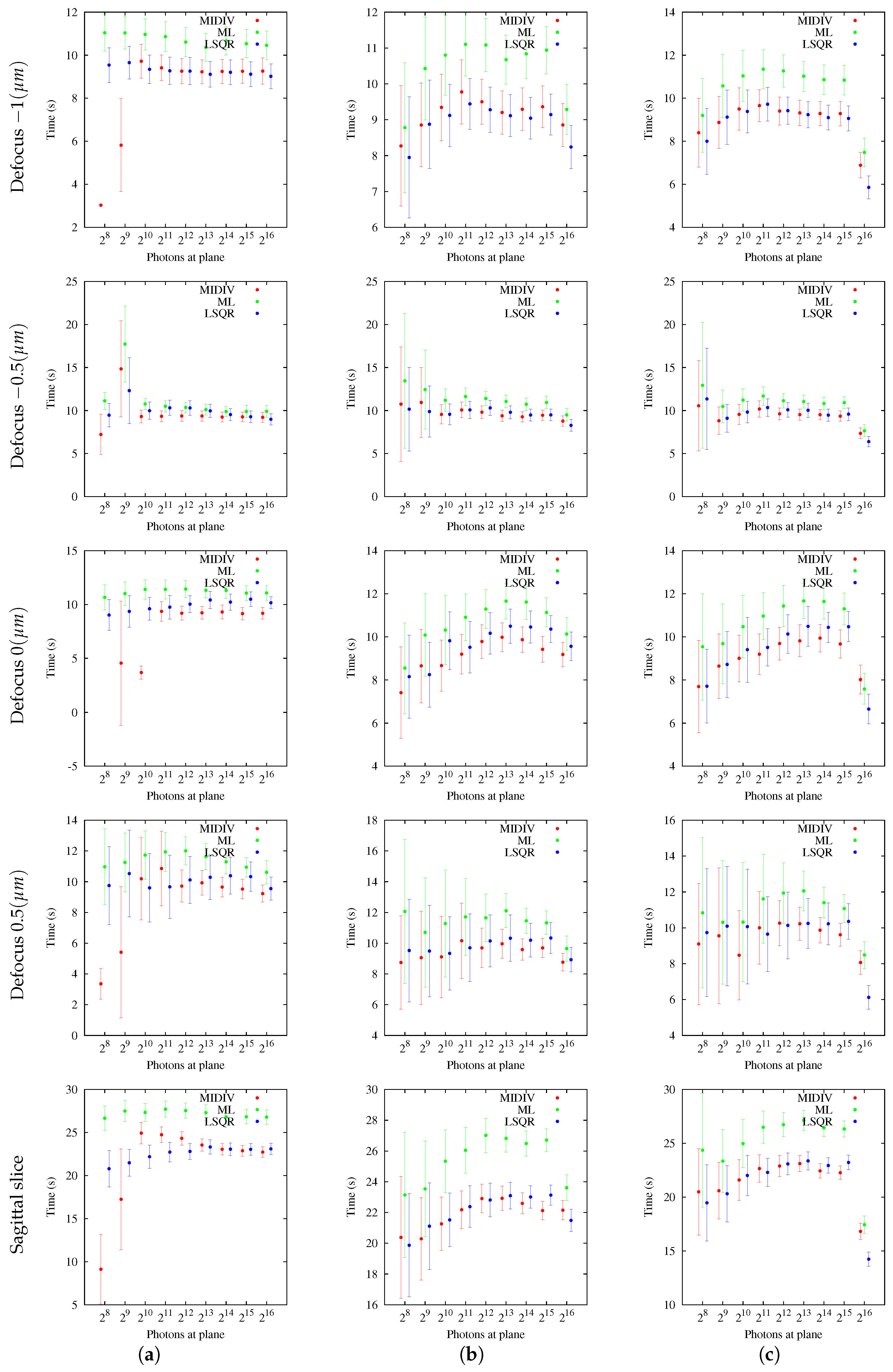
5.3. General Performance of Methods
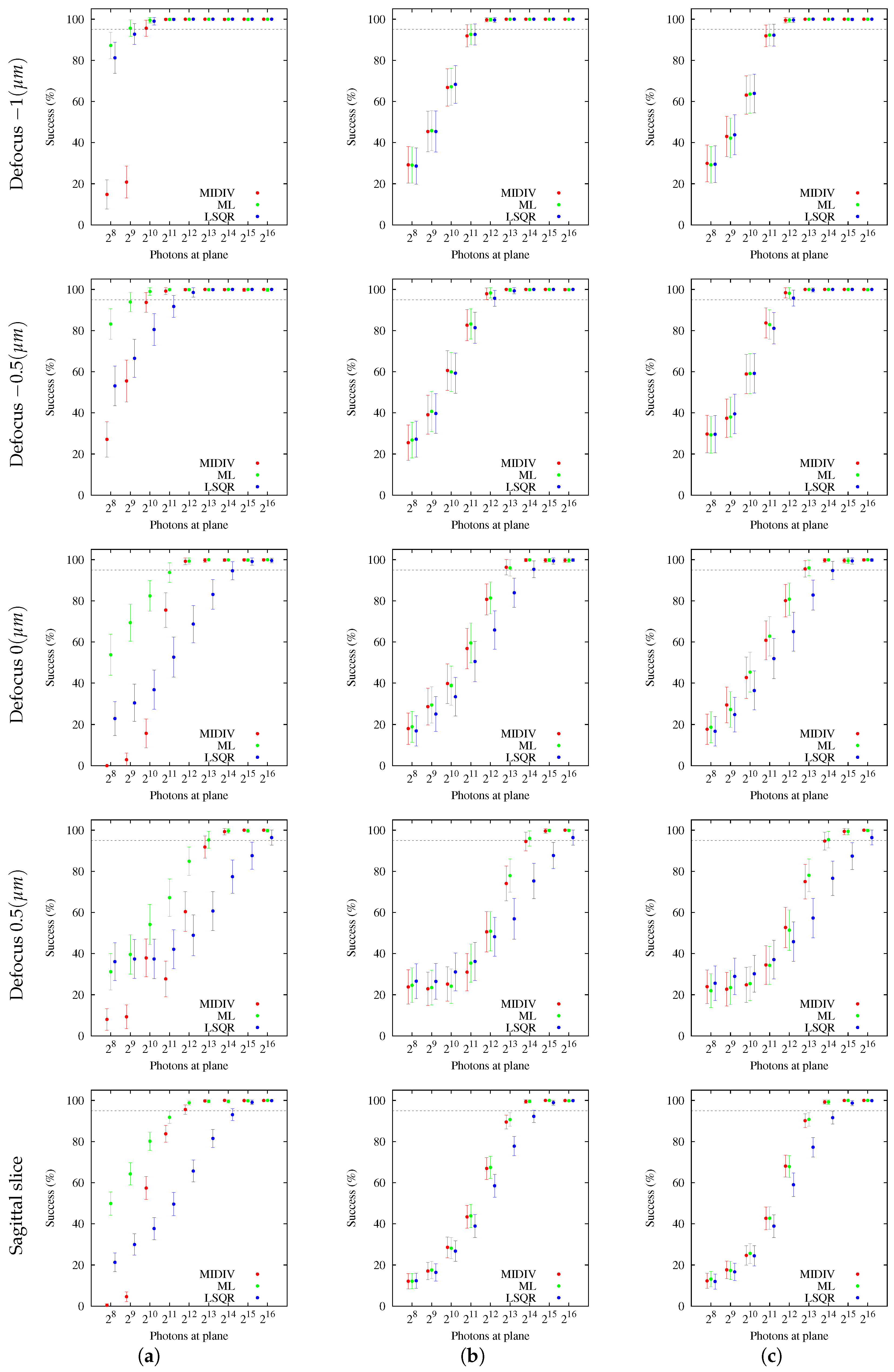
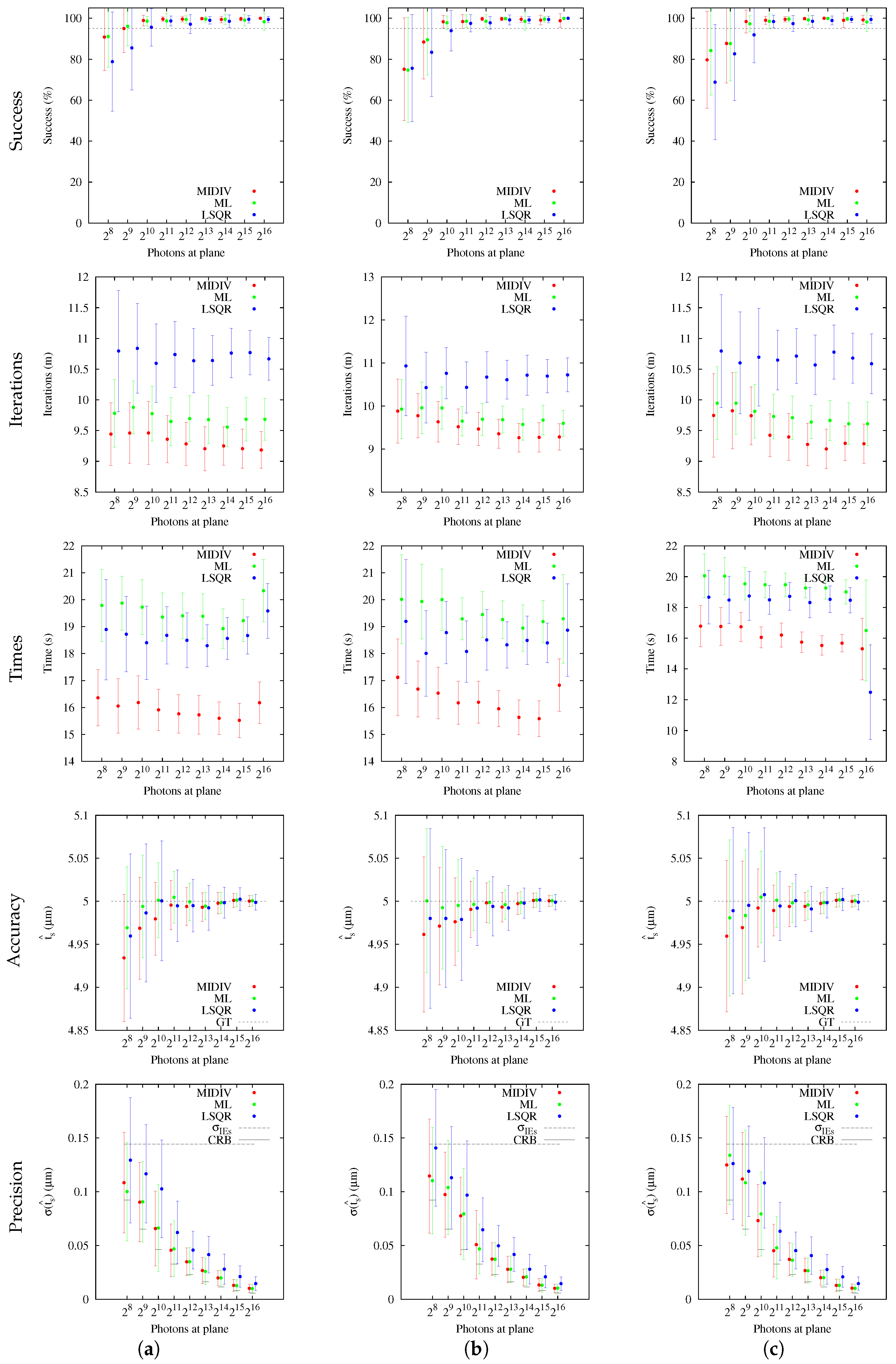
The analysis of success percentage was shown to be a useful tool for comparative analysis of the methods. Users could be confident that the methods are achieving good accuracy and precision, but this could not have any statistical value, since the chance of success could have low probability. This situation became evident with the results obtained using sagittal slices. In these cases, both CRBs as estimation methods were shown to be better than those obtained with optical sections for the same photon count. However, more photon counting is required for the sagittal slice to have a more reasonable success percentage in comparison with the optical section at −1
m, for example. In this sense, it should be noted that the CRB establishes a bound to the variance of the unbiased estimator, but nothing can be deduced from the method by which the estimator is computed. This assigns a truly methodological importance to the simulation. It could be argued that the range of initial estimates is wide and that the success percentage would increase making it narrower. The argument is valid in the theoretical sense, in the same way that the success percentage will improve when the photon count increases. However, in practical applications, the range of the initial estimates is limited by the real information of the parameter to be estimated. In fact, the range of initial estimates of the sagittal slice is less than the one used for optical sections, since using the same range, the results obtained were much less satisfying than the ones presented in the last rows of
Figure 5,
Figure 6,
Figure 7,
Figure 8 and
Figure 9.
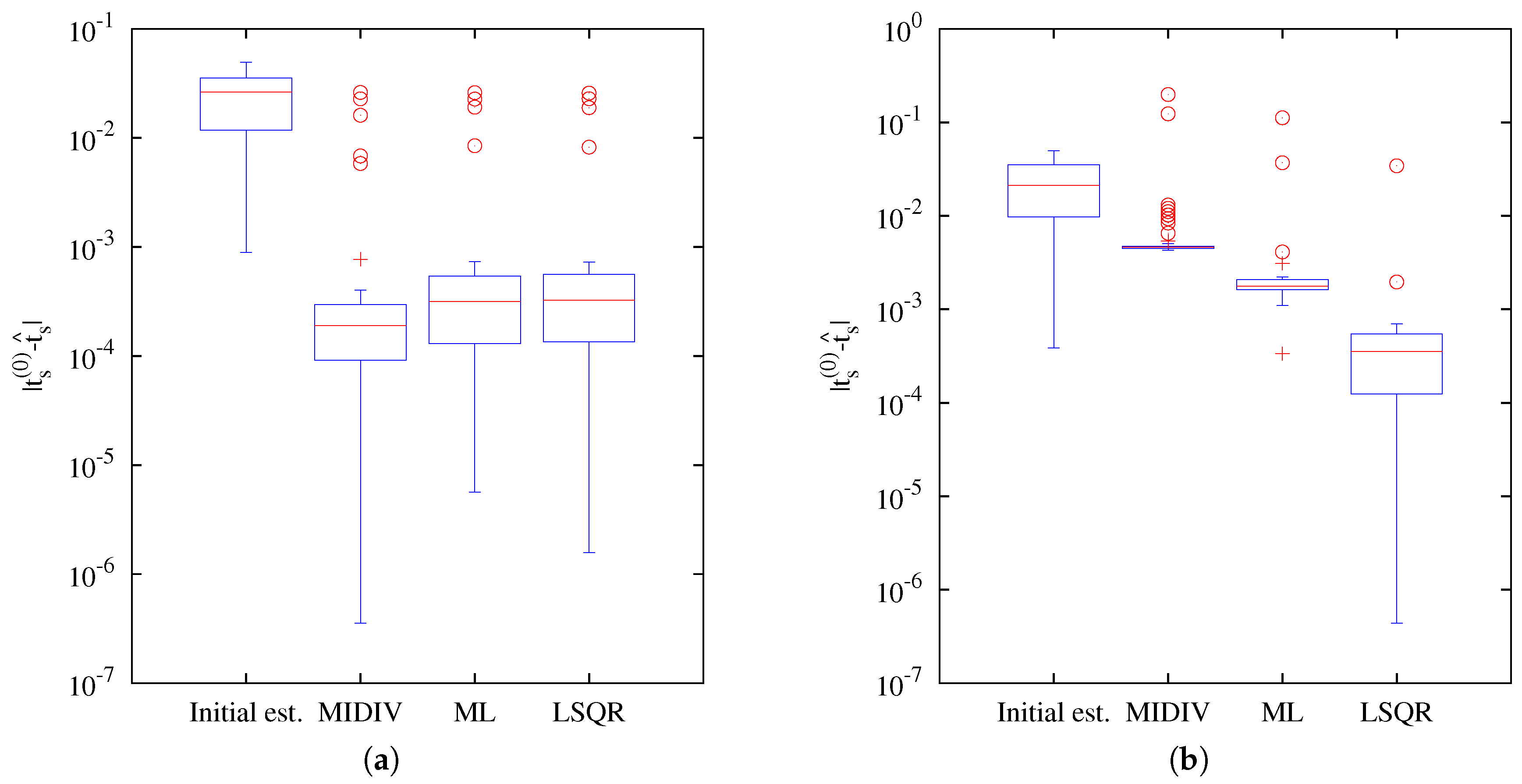
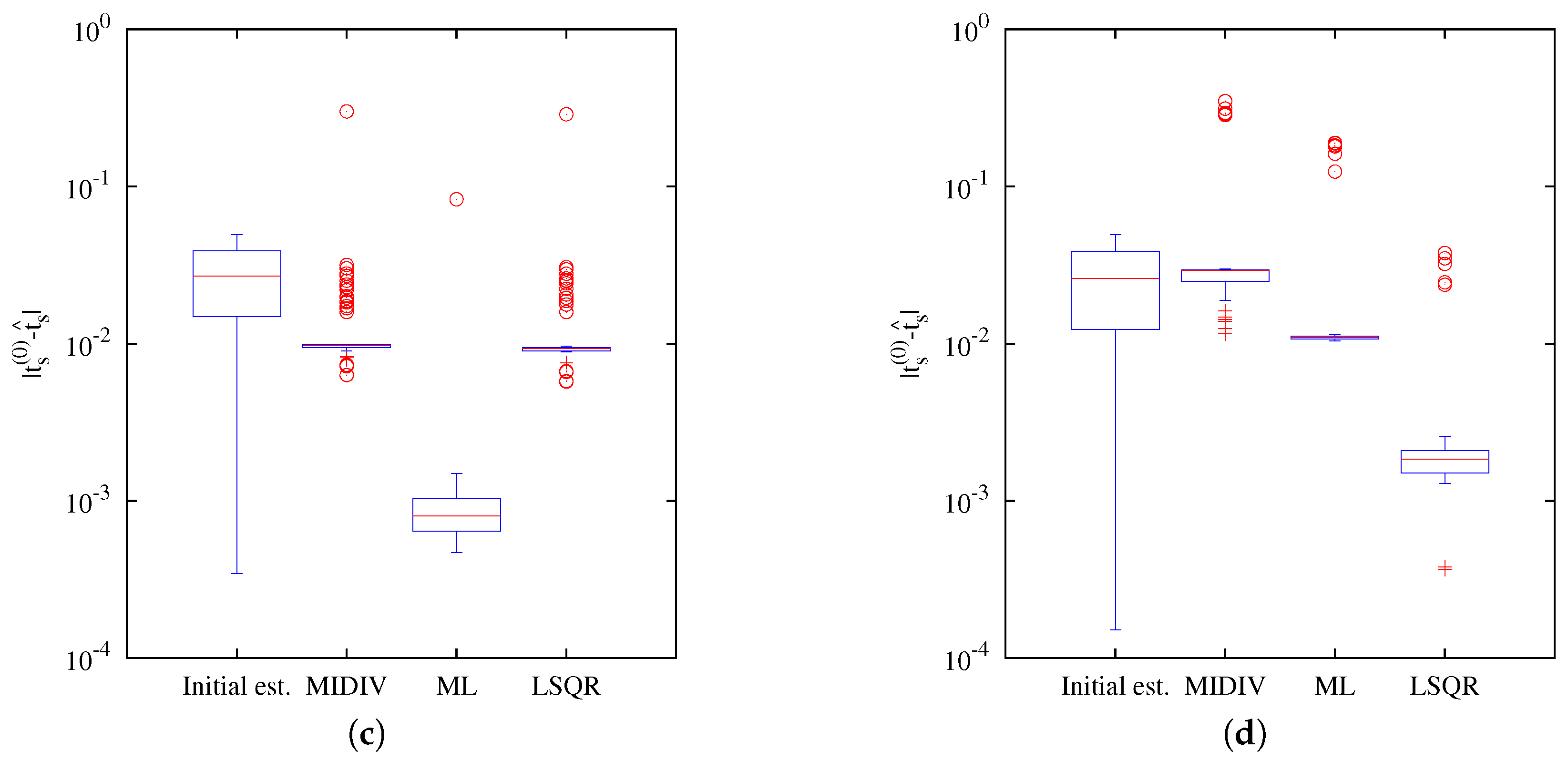
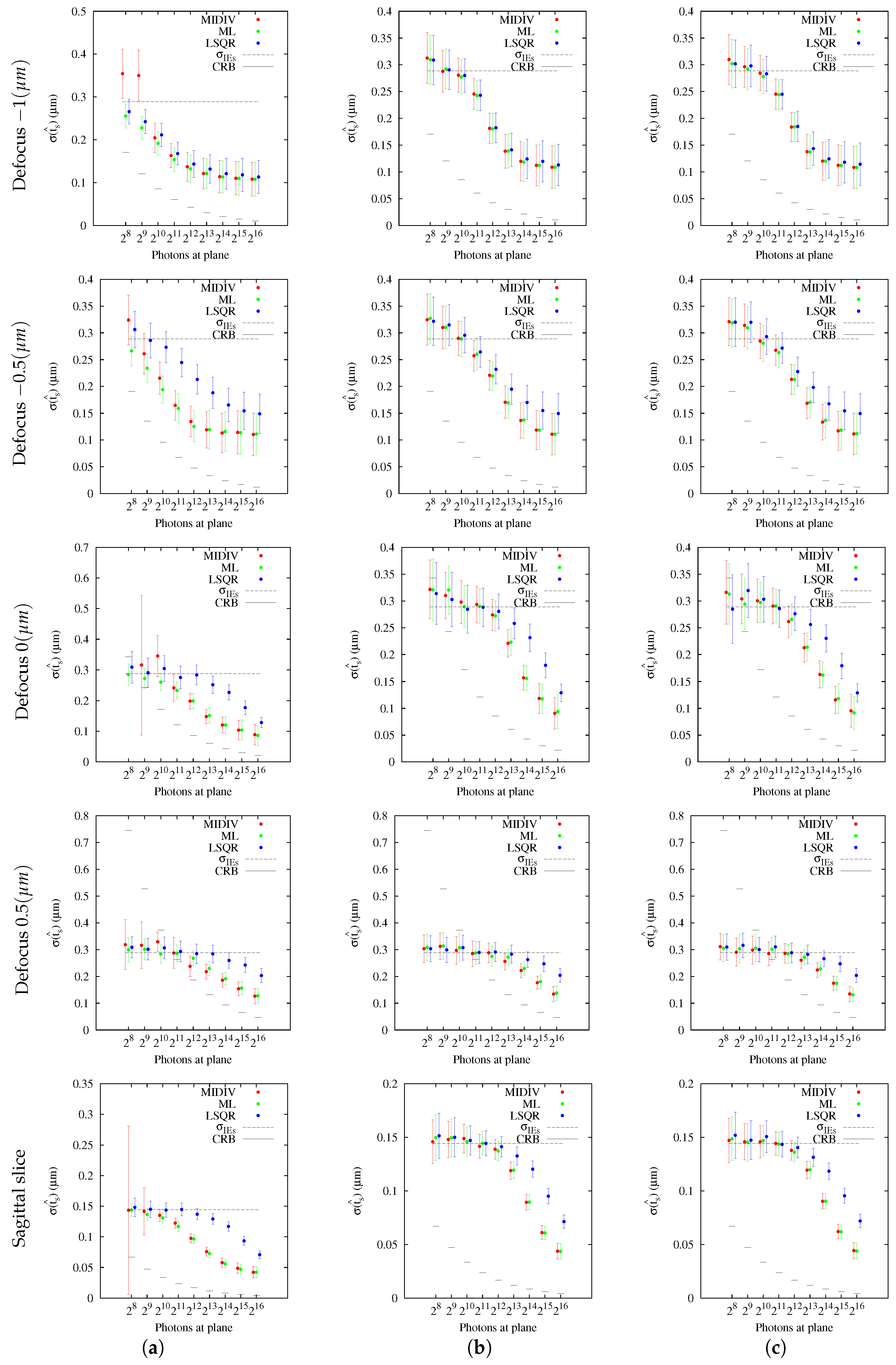
Our results indicate that the estimation of the axial position using the Gibson and Lanni model requires a high SNR to achieve
of success and higher still to be close to the CRB. In fact, only the ML and MIDIV methods (last row of
Figure 10) achieved the theoretical precision given by the CRB when data collected from the optical axis were used. A test done using more −DF optical sections did not improve the precision shown in the first row of
Figure 9. However, it must be noted that, in practice, exposition time is generally fixed during optical sectioning acquisition; thus, the more DF the optical section acquired is, the less the photon count will be.
The incidence of the extrinsic noise sources (NCII or NCIII) showed that success percentage estimation worsened, but no difference was found between noise sources. In fact, when columns NCII and NCIII of
Figure 5,
Figure 6,
Figure 7,
Figure 8 and
Figure 9 are compared, the qualitative result is very similar. In other words, estimation methods have the chance to successfully converge in cases where there are additive noise sources not considered in the forward model.
Finally, a reference to the MIDIV, our own method, is required. In this approach, there were no assumptions about noise, which is an advantage over ML and LSQR. It also should be noted that MIDIV has achieved the CRB in similar conditions to ML. However, a strong hypothesis was made about the experimental PSF representing a probability density in accordance with the law of large numbers. This assumption is true when the photon number tends to infinity, which does not hold in practice. In real circumstances, the number of photons is finite, and the variability of the mean level of intensity in each pixel is different. In pixels that capture the PSF peak, the variability of the mean level of intensity is less than in those that capture darker areas of the PSF. This may be influencing the method’s performance.
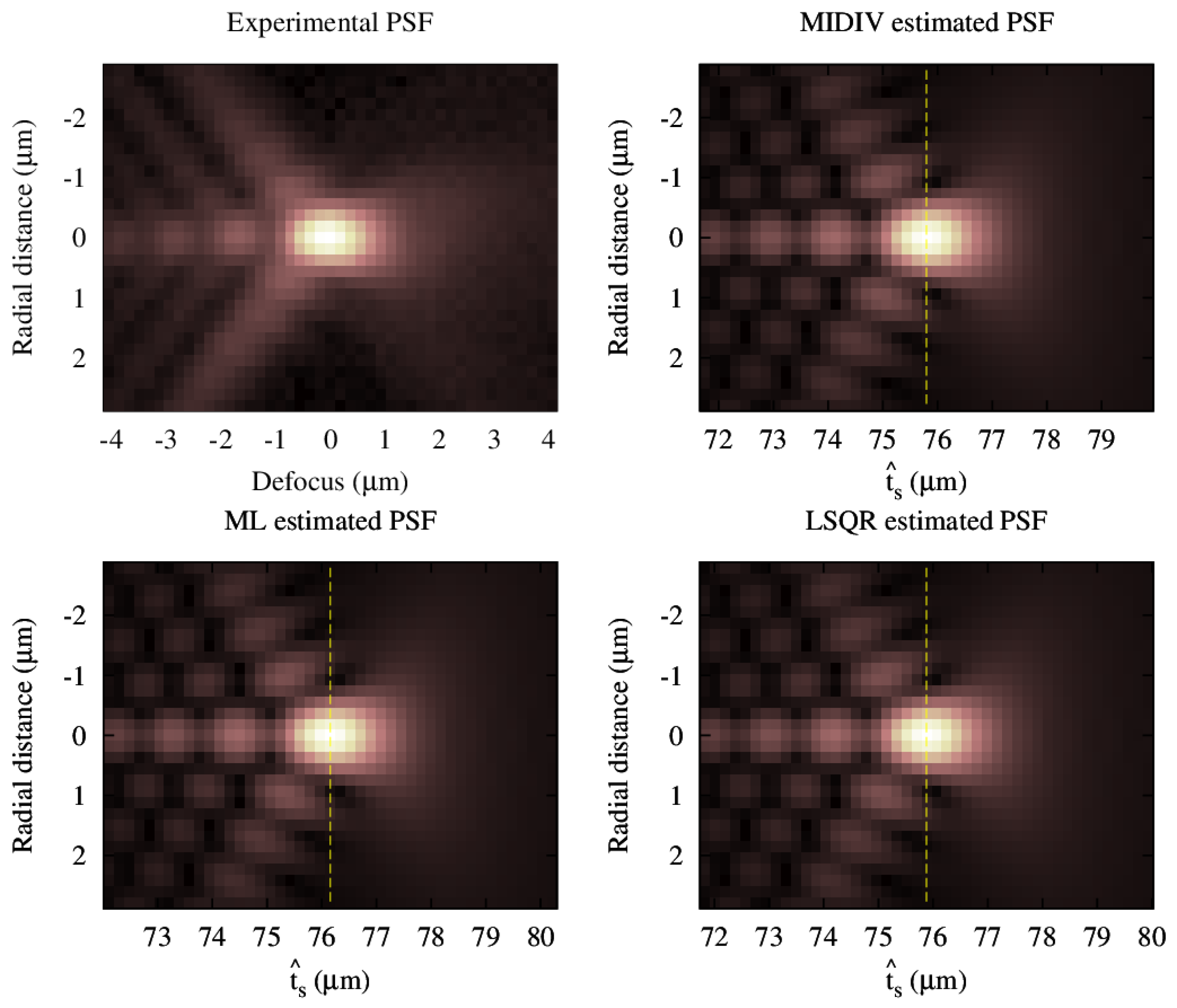
5.4. Test with an Experimental PSF
Estimations using an experimental PSF showed similar qualitative results. FWHM in both lateral and axial directions were evaluated, and all methods showed the same results up to the third decimal (nanometer). However, to obtain a sounder conclusion in this aspect, it is necessary to run a more complete and controlled experiment, which is beyond the purpose of this work.
6. Conclusions
Optical sectioning is, without corrected optics for this purpose, a shift-variant tomographic technique. In the best case, this is due to the spherical aberration that depends only on the light source position along the optical axis.
The Gibson and Lanni model appropriately predicts the 3D PSF for optical sectioning, evidencing changes in morphology and intensity, as well as focal shift.
Parameter estimation methods can be used together with the Gibson and Lanni model to efficiently estimate the position of a point light source along the optical axis from experimental data.
The iterative forms of the three parameter estimation methods tested in this work showed logarithmic linear convergence.
The estimation of the axial position using the Gibson and Lanni model requires a high SNR to achieve of success and higher still to be close to the CRB.
ML achieved a higher success percentage at lower signal levels compared with MIDIV and LSQR for the intrinsic noise source NCI.
Only the ML and MIDIV methods achieved the theoretical precision given by the CRB, but only with data belonging to the optical axis and high SNR.
Extrinsic noise sources NCII and NCII worsened the success percentage of all methods, but no difference was found between noise sources for the same method, for all methods studied.
The success percentage and CRB analysis are useful and necessary tools for comparative analysis of estimation methods. They are also useful for simulation before experimental applications.