The Genome-Wide Identification of Stable Internal Reference Genes Related to Delayed Spoilage for Accurate qRT-PCR Normalization in Ethephon-Treated Pueraria thomsonii Benth.
Abstract
:1. Introduction
2. Materials and Methods
2.1. Plant Material and Treatment
2.2. Assessment of PPD in P. thomsonii Tuberous Roots
2.3. Determination of MDA, H2O2 Content, and Antioxidant Enzyme Activities
2.4. Total RNA Isolation and First-Strand cDNA Synthesis
2.5. Transcriptomic Analysis
2.6. Selection of Candidate IRGs and Primer Design
2.7. qRT-PCR Analysis
2.8. Gene Expression Stability Analysis of Candidate IRGs
2.9. Membership Function Analysis of IRGs
2.10. Validation of the Selected IRGs for qRT-PCR Normalization
2.11. Statistical Analysis
3. Results
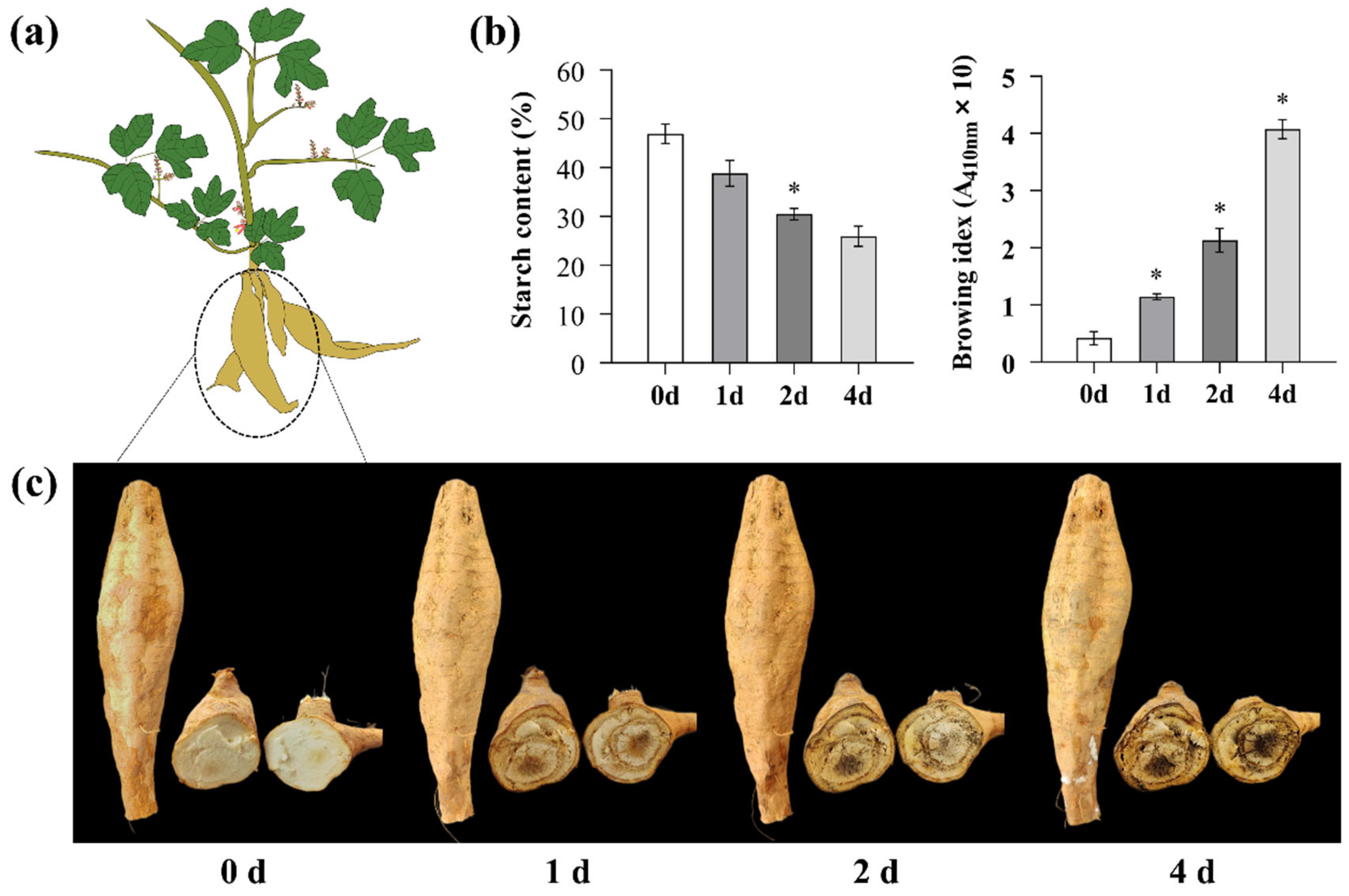
3.1. Exogenous Application of Ethephon Delayed PPD of P. thomsonii Tuberous Roots
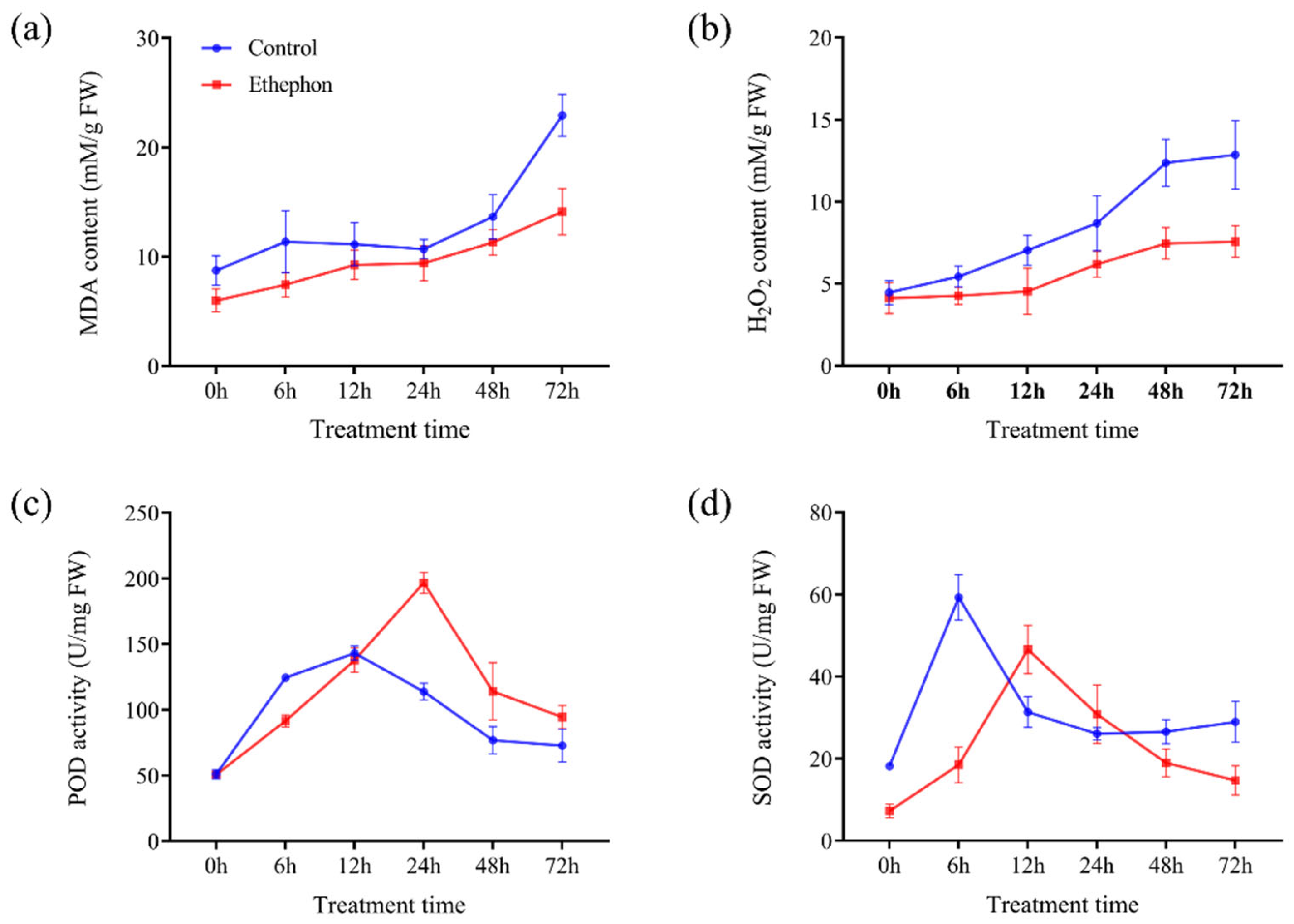
3.2. Exogenous Ethephon Application Decreased MDA and H2O2 Content and Enhanced POD and SOD Activities during PPD Process
3.3. Transcriptome Analysis of PPD delay in P. thomsonii during Ethephon Treatment Based on RNA-Seq
3.4. Selection of Candidate IRGs for Delaying PPD in P. thomsonii
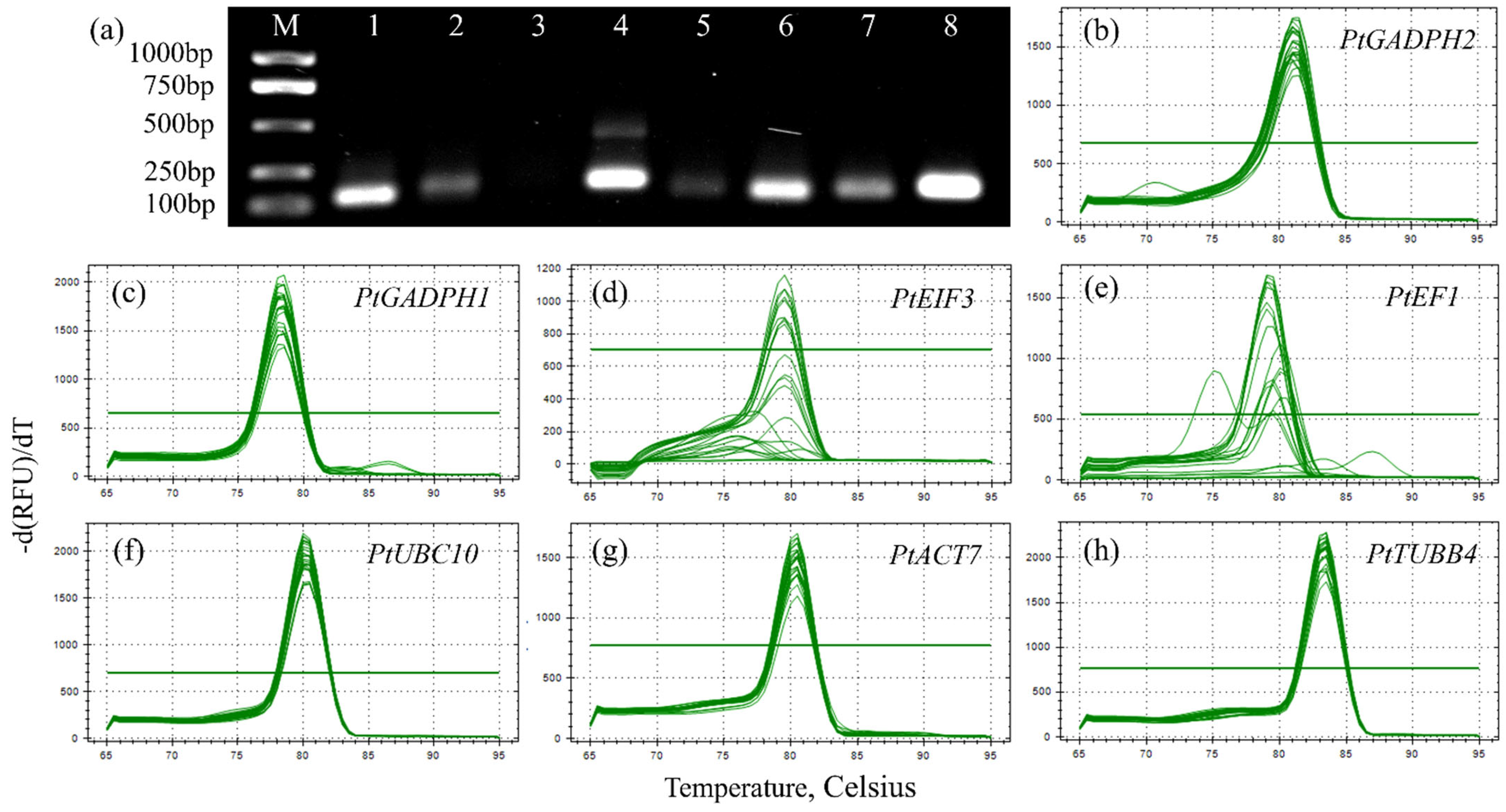
3.5. Assessment of Primer Specificity and PCR Amplification Efficiency
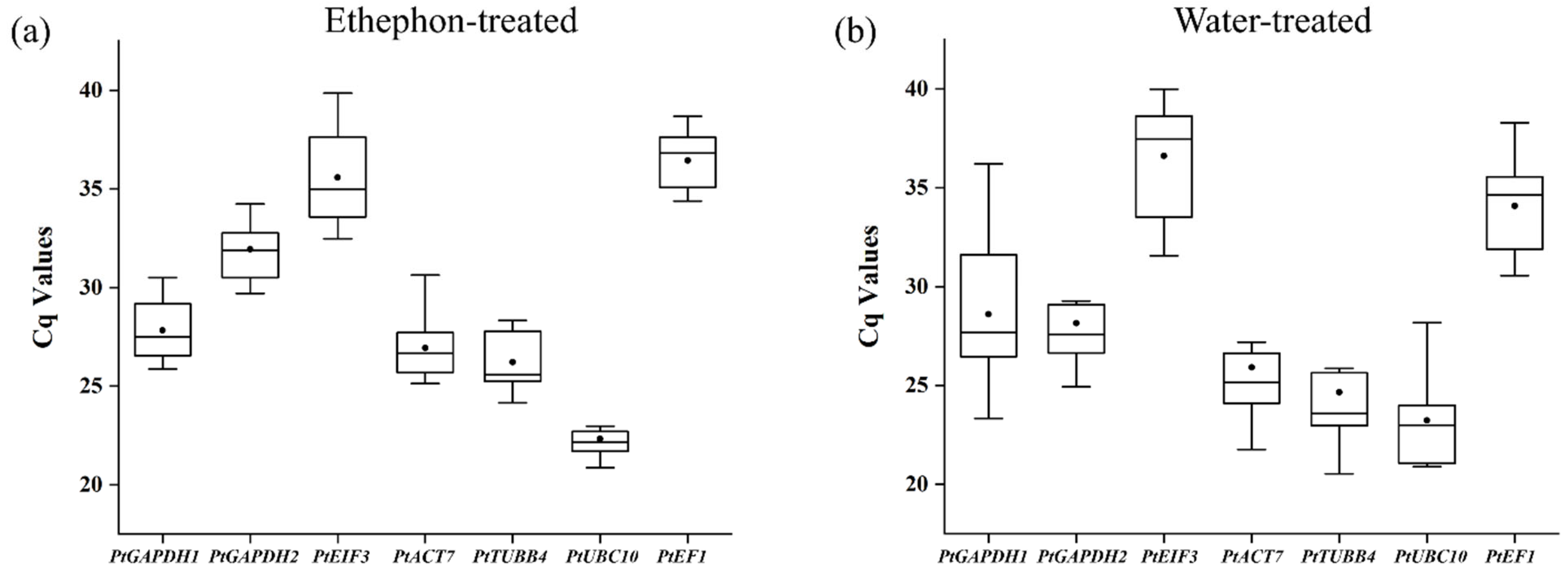
3.6. Cq Values of Candidate IRGs
3.7. Transcription Stability Analyses of Candidate IRGs
3.7.1. ΔCt Method
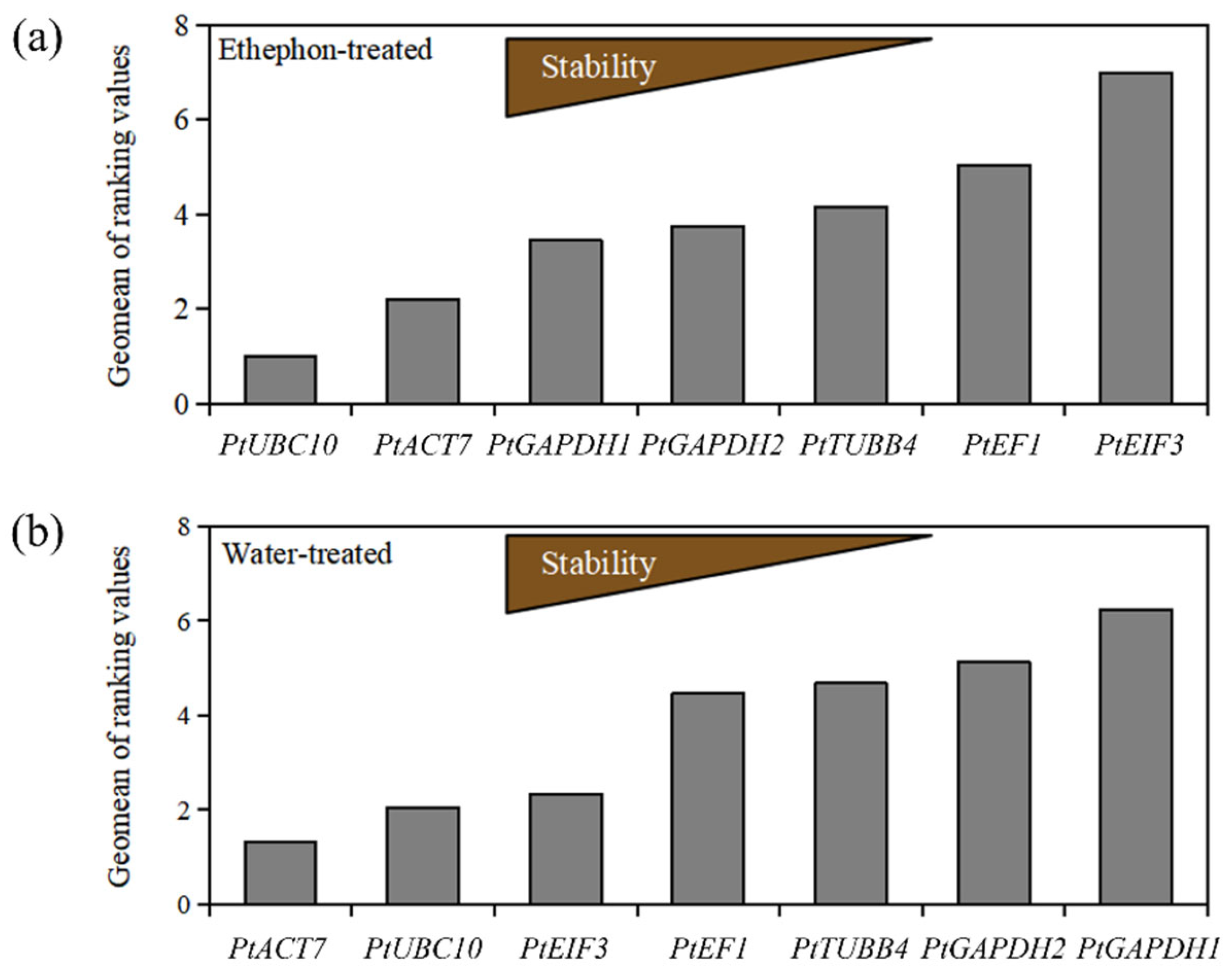
3.7.2. GeNorm Analysis
3.7.3. NormFinder Analysis
3.7.4. BestKeeper Analysis
3.7.5. RefFinder Analysis
3.7.6. Membership Functions Analysis
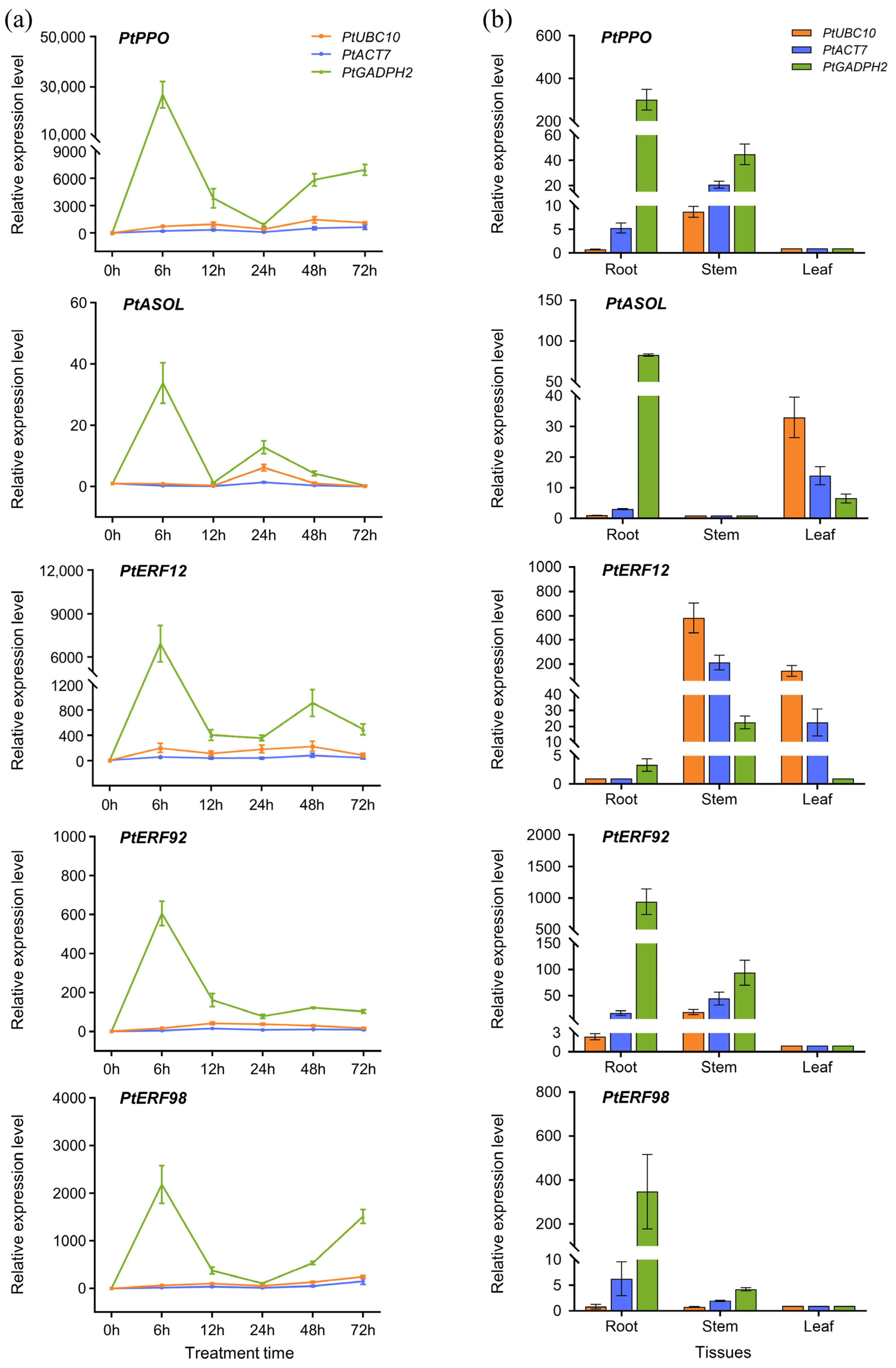
3.8. Validation of Recommended IRGs in P. thomsonii
4. Discussion
4.1. Exogenous Application of Ethephon Delayed PPD of P. thomsonii Tuberous Roots
4.2. Selection of Candidate IRGs Is Indispensable to Analyzing the Gene Expression Patterns after Ethephon Treatment
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Liu, X.; Wang, F.; Chen, Y.; An, Y.; Cheng, L.; Wang, L.; Kong, D.; Zhao, W.; Tian, J.; Niu, Y.; et al. Research Progress on Chemical Components and Pharmacological Action of Solanum Lyratum Thunb. J. Pharm. Pharmacol. 2023, 75, 328–362. [Google Scholar] [CrossRef] [PubMed]
- Zhou, S.; Li, Y.; Hong, Y.; Zhong, Z.; Zhao, M. Puerarin Protects against Sepsis-Associated Encephalopathy by Inhibiting NLRP3/Caspase-1/GSDMD Pyroptosis Pathway and Reducing Blood-Brain Barrier Damage. Eur. J. Pharmacol. 2023, 945, 175616. [Google Scholar] [CrossRef] [PubMed]
- Ding, Y.; Li, L.; Peng, S.; Zhou, G.; Chen, S.; Wei, Y.; Xu, J.; Gu, H.; Li, J.; Liu, S.; et al. Puerarin Protects against Myocardial Ischemia/Reperfusion Injury by Inhibiting Ferroptosis. Biol. Pharm. Bull. 2023, 46, 524–532. [Google Scholar] [CrossRef]
- Liang, T.; Xu, X.; Ye, D.; Chen, W.; Gao, B.; Huang, Y. Correction to: Caspase/AIF/Apoptosis Pathway: A New Target of Puerarin for Diabetes Mellitus Therapy. Mol. Biol. Rep. 2022, 49, 3369–3372. [Google Scholar] [CrossRef] [PubMed]
- Mo, Q.; Li, S.; You, S.; Wang, D.; Zhang, J.; Li, M.; Wang, C. Puerarin Reduces Oxidative Damage and Photoaging Caused by UVA Radiation in Human Fibroblasts by Regulating Nrf2 and MAPK Signaling Pathways. Nutrients 2022, 14, 4724. [Google Scholar] [CrossRef] [PubMed]
- Zhang, X.; Lang, Y.; Li, X.; Li, Z.; Xu, Y.; Chu, H. The Protective Effect of Puerarin-Loaded Mesoporous Silicon Nanoparticles on Alcoholic Hepatitis through MTOR-Mediated Autophagy Pathway. Biomed. Microdevices 2022, 24, 37. [Google Scholar] [CrossRef]
- Cai, Y.; Gao, Q.; Meng, J.; Chen, L. Puerarin Suppresses Glycolysis and Increases Cisplatin Chemosensitivity in Oral Squamous Cell Carcinoma via FBXW7/MTOR Signaling. Nutr. Cancer 2023, 75, 1028–1037. [Google Scholar] [CrossRef]
- Lang, J.; Guo, Z.; Xing, S.; Sun, J.; Qiu, B.; Shu, Y.; Wang, Z.; Liu, G. Inhibitory Role of Puerarin on the A549 Lung Cancer Cell Line. Transl. Cancer Res. 2022, 11, 4117–4125. [Google Scholar] [CrossRef]
- Liu, X.; Huang, R.; Wan, J. Puerarin: A Potential Natural Neuroprotective Agent for Neurological Disorders. Biomed. Pharmacother. 2023, 162, 114581. [Google Scholar] [CrossRef]
- Zhang, W.; Xu, M.; Wen, S.; Wang, L.; Zhang, K.; Zhang, C.; Zou, H.; Gu, J.; Liu, X.; Bian, J.; et al. Puerarin Alleviates Cadmium-Induced Rat Neurocyte Injury by Alleviating Nrf2-Mediated Oxidative Stress and Inhibiting Mitochondrial Unfolded Protein Response. Ecotoxicol. Environ. Saf. 2022, 247, 114239. [Google Scholar] [CrossRef]
- Qu, F.; Sun, Z.J.; Xiong, W.Z.; Wang, S.; Sun, Z. Thoughts on Protection of Geographical Indication Intellectual Property of Kudzu. Acta Agric. Jiangxi 2011, 23, 172–175. [Google Scholar] [CrossRef]
- Ma, C.; Zheng, S.Y.; Zhuo, H.B. Study on Micropropagation of Pueraria thomsonii Benth. In Vitro. Guangdong Agric. Sci. 2013, 40, 28–35. [Google Scholar] [CrossRef]
- Zhang, G.; Liu, J.; Gao, M.; Kong, W.; Zhao, Q.; Shi, L.; Wang, Q. Tracing the Edible and Medicinal Plant Pueraria Montana and Its Products in the Marketplace Yields Subspecies Level Distinction Using DNA Barcoding and DNA Metabarcoding. Front. Pharmacol. 2020, 11, 336. [Google Scholar] [CrossRef]
- Cui, X.; Cai, Y.; Chen, R.; Liu, Q. First Report of Bacterial Leaf Spot Disease on Pueraria Montana var. thomsonii Caused by Robbsia Andropogonis in China. Plant Dis. 2022, 106, 2985. [Google Scholar] [CrossRef] [PubMed]
- Vanderschuren, H.; Nyaboga, E.; Poon, J.S.; Baerenfaller, K.; Grossmann, J.; Hirsch-Hoffmann, M.; Kirchgessner, N.; Nanni, P.; Gruissem, W. Large-Scale Proteomics of the Cassava Storage Root and Identification of a Target Gene to Reduce Postharvest Deterioration. Plant Cell 2014, 26, 1913–1924. [Google Scholar] [CrossRef] [PubMed]
- Dong, X.; Mei, Y.; Wang, J.; Huang, N.; Cai, S. Research progress on a medicinal and edible plant Radix puerariae. J. Chang. Veg. 2020, 43–47. [Google Scholar]
- Zhang, L.; Song, K.; Hui, G.U.; Jia, Z.; Chang, J.; Hong, K.; Hou, X. Effect of Citric Acid Treatment on Browning of Fresh-Cut Pueraria. Chin. J. Trop. Crops 2017, 38, 1143–1148. [Google Scholar] [CrossRef]
- Liu, S.; Du, X. Enzymatic Browning and Its Reaction under Control during Radix Pueraria Processing. Acad. Period. Farm Prod. Process. 2011, 40–45. [Google Scholar]
- Ma, C.; Wu, Y.; Chen, S.; Wang, G. Effect of Storage Time and Manner on the Quality of Pueraria thomsonii Benth. North. Hortic. 2016, 367, 133–135. [Google Scholar] [CrossRef]
- Ren, M.Y.; Feng, R.J.; Shi, H.R.; Lu, L.F.; Yun, T.Y.; Peng, M.; Guan, X.; Zhang, H.; Wang, J.Y.; Zhang, X.Y.; et al. Expression Patterns of Members of the Ethylene Signaling-Related Gene Families in Response to Dehydration Stresses in Cassava. PLoS ONE 2017, 12, e0177621. [Google Scholar] [CrossRef]
- Moscoso-Ramírez, P.A.; Palou, L. Effect of Ethylene Degreening on the Development of Postharvest Penicillium Molds and Fruit Quality of Early Season Citrus Fruit. Postharvest Biol. Technol. 2014, 91, 1–8. [Google Scholar] [CrossRef]
- Mayuoni, L.; Sharabi-Schwager, M.; Feldmesser, E.; Porat, R. Effects of Ethylene Degreening on the Transcriptome of Mandarin Flesh. Postharvest Biol. Technol. 2011, 60, 75–82. [Google Scholar] [CrossRef]
- Dai, Z.J.; Wang, Y.; Wang, X.T.; Yang, Q.; Zhang, Y.Y.; Bao-Quan, L.I.; Wang, L.P.; Centre, C.D. Reference Genes Selection for Real-Time Quantitative PCR under Rough Dwarf Virus Induced Disease in Maize. Plant Physiol. J. 2019, 55, 1545–1553. [Google Scholar] [CrossRef]
- Shakeel, M.; Rodriguez, A.; Tahir, U.B.; Jin, F. Gene Expression Studies of Reference Genes for Quantitative Real-Time PCR: An Overview in Insects. Biotechnol. Lett. 2018, 40, 227–236. [Google Scholar] [CrossRef] [PubMed]
- Guénin, S.; Mauriat, M.; Pelloux, J.; Van Wuytswinkel, O.; Bellini, C.; Gutierrez, L. Normalization of QRT-PCR Data: The Necessity of Adopting a Systematic, Experimental Conditions-Specific, Validation of References. J. Exp. Bot. 2009, 60, 487–493. [Google Scholar] [CrossRef] [PubMed]
- Hu, A.; Yang, X.; Zhu, J.; Wang, X.; Liu, J.; Wang, J.; Wu, H.; Zhang, H.; Zhang, H. Selection and Validation of Appropriate Reference Genes for RT-QPCR Analysis of Nitraria Sibirica under Various Abiotic Stresses. BMC Plant Biol. 2022, 22, 592. [Google Scholar] [CrossRef]
- Yao, J.; Zhu, G.; Liang, D.; He, B.; Wang, Y.; Cai, Y.; Zhang, Q. Reference Gene Selection for QPCR Analysis in Schima Superba under Abiotic Stress. Genes 2022, 13, 1887. [Google Scholar] [CrossRef] [PubMed]
- Zhou, Q.; Xu, F.; Zeng, L.; Zhu, L.; Wang, Y.; Wang, B.; Ma, X. Selection and validation of appropriate reference genes for quantitative real-time PCR analysis in Panax vietnamensis var. fuscidiscus. Plant Physiol. J. 2020, 56, 329–335. [Google Scholar] [CrossRef]
- Hu, W.; Tie, W.; Ou, W.; Yan, Y.; Kong, H.; Zuo, J.; Ding, X.; Ding, Z.; Liu, Y.; Wu, C.; et al. Crosstalk between Calcium and Melatonin Affects Postharvest Physiological Deterioration and Quality Loss in Cassava. Postharvest Biol. Technol. 2018, 140, 42–49. [Google Scholar] [CrossRef]
- Trapnell, C.; Pachter, L.; Salzberg, S.L. TopHat: Discovering Splice Junctions with RNA-Seq. Bioinformatics 2009, 25, 1105–1111. [Google Scholar] [CrossRef]
- Trapnell, C.; Roberts, A.; Goff, L.; Pertea, G.; Kim, D.; Kelley, D.R.; Pimentel, H.; Salzberg, S.L.; Rinn, J.L.; Pachter, L. Differential Gene and Transcript Expression Analysis of RNA-Seq Experiments with TopHat and Cufflinks. Nat. Protoc. 2012, 7, 562–578. [Google Scholar] [CrossRef] [PubMed]
- Wang, L.; Feng, Z.; Wang, X.; Wang, X.; Zhang, X. DEGseq: An R Package for Identifying Differentially Expressed Genes from RNA-Seq Data. Bioinformatics 2010, 26, 136–138. [Google Scholar] [CrossRef]
- Ovesná, J.; Kučera, L.; Vaculová, K.; Štrymplová, K.; Svobodová, I.; Milella, L. Validation of the β-Amy1 Transcription Profiling Assay and Selection of Reference Genes Suited for a RT-QPCR Assay in Developing Barley Caryopsis. PLoS ONE 2012, 7, e41886. [Google Scholar] [CrossRef] [PubMed]
- Lin, S.; Xu, S.; Huang, L.; Qiu, F.; Wu, J. Selection and Validation of Reference Genes for Normalization of RT-QPCR Analysis in Developing and Abiotic-Stressed Tissues of Loquat (Eriobotrya Japonica). Phyton-Int. J. Exp. Bot. 2020, 92, 1181–1201. [Google Scholar] [CrossRef]
- Kaczmarczyk, A.; Bowra, S.; Elek, Z.; Vincze, E. Quantitative RT-PCR Based Platform for Rapid Quantification of the Transcripts of Highly Homologous Multigene Families and Their Members during Grain Development. BMC Plant Biol. 2012, 12, 184. [Google Scholar] [CrossRef] [PubMed]
- Guo, L.; Li, Y.; Wei, Z.; Wang, C.; Hou, X. Reference Genes Selection of Paeonia Ostii “Fengdan” under Osmotic Stresses and Hormone Treatments by RT-QPCR. Mol. Biol. Rep. 2023, 50, 133–143. [Google Scholar] [CrossRef] [PubMed]
- Yin, H.; Yin, D.; Zhang, M.; Gao, Z.; Tuluhong, M.; Li, X.; Li, J.; Li, B.; Cui, G. Validation of Sppropriate Reference Genes for QRT-PCR Normalization in Oat (Avena Sativa L.) under UV-B and High-Light Stresses. Int. J. Mol. Sci. 2022, 23, 11187. [Google Scholar] [CrossRef]
- Vinje, M.A.; Willis, D.K.; Duke, S.H.; Henson, C.A. Differential RNA Expression of Bmy1 during Barley Seed Development and the Association with β-Amylase Accumulation, Activity, and Total Protein. Plant Physiol. Biochem. 2011, 49, 39–45. [Google Scholar] [CrossRef]
- Ferdous, J.; Li, Y.; Reid, N.; Langridge, P.; Shi, B.; Tricker, P.J. Identification of Reference Genes for Quantitative Expression Analysis of MicroRNAs and MRNAs in Barley under Various Stress Conditions. PLoS ONE 2015, 10, e0118503. [Google Scholar] [CrossRef]
- Kundu, A.; Patel, A.; Pal, A. Defining Reference Genes for QPCR Normalization to Study Biotic and Abiotic Stress Responses in Vigna Mungo. Plant Cell Rep. 2013, 32, 1647–1658. [Google Scholar] [CrossRef]
- Vandesompele, J.; De Preter, K.; Pattyn, F.; Poppe, B.; Van Roy, N.; De Paepe, A.; Speleman, F. Accurate Normalization of Real-Time Quantitative RT-PCR Data by Geometric Averaging of Multiple Internal Control Genes. Genome Biol. 2002, 3, research0034. [Google Scholar] [CrossRef] [PubMed]
- Andersen, C.L.; Jensen, J.L.; Ørntoft, T.F. Normalization of Real-Time Quantitative Reverse Transcription-PCR Data: A Model-Based Variance Estimation Approach to Identify Genes Suited for Normalization, Applied to Bladder and Colon Cancer Data Sets. Cancer Res. 2004, 64, 5245–5250. [Google Scholar] [CrossRef] [PubMed]
- Pfaffl, M.W.; Tichopad, A.; Prgomet, C.; Neuvians, T.P. Determination of Stable Housekeeping Genes, Differentially Regulated Target Genes and Sample Integrity: BestKeeper-Excel-Based Tool Using Pair-Wise Correlations. Biotechnol. Lett. 2004, 26, 509–515. [Google Scholar] [CrossRef] [PubMed]
- Silver, N.; Best, S.; Jiang, J.; Thein, S.L. Selection of Housekeeping Genes for Gene Expression Studies in Human Reticulocytes Using Real-Time PCR. BMC Mol. Biol. 2006, 7, 33. [Google Scholar] [CrossRef] [PubMed]
- Long, X.; He, B.; Gao, X.; Qin, Y.; Yang, J.; Fang, Y.; Qi, J.; Tang, C. Validation of Reference Genes for Quantitative Real-Time PCR during Latex Regeneration in Rubber Tree. Gene 2015, 563, 190–195. [Google Scholar] [CrossRef] [PubMed]
- Xie, F.; Xiao, P.; Chen, D.; Xu, L.; Zhang, B. miRDeepFinder: A MiRNA Analysis Tool for Deep Sequencing of Plant Small RNAs. Plant Mol. Biol. 2012, 80, 75–84. [Google Scholar] [CrossRef]
- Xie, F.; Wang, J.; Zhang, B. RefFinder: A Web-based Tool for Comprehensively Analyzing and Identifying Reference Genes. Funct. Integr. Genomics. 2023, 23, 125. [Google Scholar] [CrossRef]
- Li, Q.; Zhu, C.; Li, X.; Wu, B.; Yi, J.; Fu, H.; Chen, D.; Zhang, H. Comprehensive evaluation and identification index of drought resistance at germination stage of rice. Acta Agric. Boreali-Occident. Sin. 2023, 32, 18–32. [Google Scholar] [CrossRef]
- Zhang, R.; Hussain, S.; Wang, Y.; Liu, Y.; Li, Q.; Chen, Y.; Wei, H.; Gao, P.; Dai, Q. Comprehensive Evaluation of Salt Tolerance in Rice (Oryza Sativa L.) Germplasm at the Germination Stage. Agronomy 2021, 11, 1569. [Google Scholar] [CrossRef]
- Taylor, S.; Wakem, M.; Dijkman, G.; Alsarraj, M.; Nguyen, M. A Practical Approach to RT-QPCR-Publishing Data That Conform to the MIQE Guidelines. Methods 2010, 50, S1–S5. [Google Scholar] [CrossRef]
- Li, F.; Sun, J.; Men, J.; Li, H.; Wang, J. Selection and Validation of Reference Genes for RT-QPCR Analysis in the Pericarp of Litchi Chinensis. Biol. Plant. 2020, 66, 103–111. [Google Scholar] [CrossRef]
- Shang, X.; Cao, S.; Yan, H.; Chen, Y.; Xiao, L.; Wang, Y.; Ou, K.; Qin, X.; Gan, X. Development status and countermeasures of Pueraria lobata (Willd.) Ohwi var. thomsoni (Benth.) van der Maesen industry in Guangxi. J. South. Agric. 2021, 52, 1510–1519. [Google Scholar] [CrossRef]
- Zidenga, T.; Leyva-Guerrero, E.; Moon, H.; Siritunga, D.; Sayre, R. Extending Cassava Root Shelf Life via Reduction of Reactive Oxygen Species Production. Plant Physiol. 2012, 159, 1396–1407. [Google Scholar] [CrossRef]
- Lv, J.; Zhou, Y.; Ge, Y.; Li, C.; Zhang, M. Effects of 1-Methylcyclopropene and Ethephon Treatment on Physiological Changes and Reactive Oxygen Metabolism of Postharvest “Golden Delicious” Apple Fruit during Storage at Ambient Temperature. Sci. Technol. Food Ind. 2017, 38, 297–302. [Google Scholar] [CrossRef]
- Song, K.H.; Zhang, L.B.; Hui, G.U.; Jia, Z.W.; Chang, J.M. Study on the Mechanism of NaCl Treatment Delaying Browning of Fresh-Cut Pueraria. Sci. Technol. Food Ind. 2018, 39, 285–288+295. [Google Scholar] [CrossRef]
- Liu, W.; Deng, Y.; Zhou, Y.; Chen, H.; Dong, Y.; Wang, N.; Li, X.; Jameel, A.; Yang, H.; Zhang, M.; et al. Normalization for Relative Quantification of MRNA and MicroRNA in Soybean Exposed to Various Abiotic Stresses. PLoS ONE 2016, 11, e0155606. [Google Scholar] [CrossRef] [PubMed]
- Li, J.; Han, J.; Hu, Y.; Yang, J. Selection of Reference Genes for Quantitative Real-Time PCR during Flower Development in Tree Peony (Paeonia Suffruticosa Andr.). Front. Plant Sci. 2016, 7, 516. [Google Scholar] [CrossRef] [PubMed]
- Lü, J.; Yang, C.; Zhang, Y.; Pan, H. Selection of Reference Genes for the Normalization of RT-QPCR Data in Gene Expression Studies in Insects: A Systematic Review. Front. Physiol. 2018, 9, 1560. [Google Scholar] [CrossRef] [PubMed]
- Zhao, Z.; Zhang, Z.; Ding, Z.; Meng, H.; Shen, R.; Tang, H.; Liu, Y.-G.; Chen, L. Public-Transcriptome-Database-Assisted Selection and Validation of Reliable Reference Genes for QRT-PCR in Rice. Sci. China Life Sci. 2020, 63, 92–101. [Google Scholar] [CrossRef]
- Mafra, V.; Kubo, K.S.; Alves-Ferreira, M.; Ribeiro-Alves, M.; Stuart, R.M.; Boava, L.P.; Rodrigues, C.M.; Machado, M.A. Reference Genes for Accurate Transcript Normalization in Citrus Genotypes under Different Experimental Conditions. PLoS ONE 2012, 7, e31263. [Google Scholar] [CrossRef]
- Xiao, C.; Yan, J.W.; Long, G.Y.; Dai, S.M.; Li, D.; Deng, Z.N. Stability Evaluation of Reference Genes in Citrus. J. Fruit Sci. 2012, 29, 978–984. [Google Scholar] [CrossRef]
- Ye, X.; Zhu, H.; Lin, H.; Liu, J.; Wang, B.; Chen, M.; Wen, Q. Screening and evaluation of reference genes for RT-qPCR in wax gourd (Benincasa hispida Cogn.). J. Nucl. Agric. Sci. 2019, 33, 473–481. [Google Scholar] [CrossRef]
- Ransbotyn, V.; Reusch, T.B.H. Housekeeping Gene Selection for Quantitative Real-Time PCR Assays in the Seagrass Zostera Marina Subjected to Heat Stress. Limnol. Oceanogr. Methods 2006, 4, 367–373. [Google Scholar] [CrossRef]
- Hu, R.; Qi, G.; Kong, Y.; Kong, D.; Gao, Q.; Zhou, G. Comprehensive Analysis of NAC Domain Transcription Factor Gene Family in Populus Trichocarpa. BMC Plant Biol. 2010, 10, 145. [Google Scholar] [CrossRef]


| No. | Gene | Gene ID | Primer Sequences (Forward/Reverse) | Amplicom Length (bp) | Tm (°C) | E (%) | R2 |
|---|---|---|---|---|---|---|---|
| 1 | PtGAPDH1 [33] | Pmon004G01500 | 5’-GAAAGTTCTTGGTGTCGGA-3’ 5’-TTAAAAGGTGGAATGGCTA-3’ | 103 bp | 56.0 | 90 | 0.994 |
| 2 | PtGAPDH2 | Pmon007G03494 | 5’-TCACCATCTTTATCCACACTC-3’ 5’-CACTTCTCTTCACCTTTTCCT-3’ | 127 bp | 57.0 | 106 | 0.995 |
| 3 | PtEIF3 [34] | Pmon010G01280 | 5’-TCGTATAAAAGAGAGAGAC-3’ 5’-TTGAGTAAAAACTGACATC-3’ | 149 bp | 55.2 | - | - |
| 4 | PtUBQ10 [35] | Pmon003G02895 | 5’-AAAGGCAAAGATCCAAGAC-3’ 5’-ATACCTCCCCTCAGACGAA-3’ | 153 bp | 55.4 | - | - |
| 5 | PtEF1 [36] | Novel05073 | 5’-AGGTTCTTCTGGATGGTTC-3’ 5’-TTGTCGGATTTTCTTGGTA-3’ | 115 bp | 56.3 | 304 | 0.969 |
| 6 | PtUBC10 [37] | Pmon010G02541 | 5’-AACTATTTCGAAGGTGTTG-3’ 5’-CGTACTTGTTCCTGTCTGT-3’ | 113 bp | 57.4 | 103 | 0.992 |
| 7 | PtACT7 [38] | Pmon003G05119 | 5’-TGTTCTTAGTGGTGGCTCA-3’ 5’-ATATTTTCTTTCTGGTGGA-3’ | 121 bp | 59.1 | 107 | 0.995 |
| 8 | PtTUBB4 [39] | Pmon002G02874 | 5’-TGGAAACTCGACCTCGATT-3’ 5’-ATGTTGCTCTCTGCCTCTG-3’ | 141 bp | 58.8 | 104 | 0.993 |
| Candidate Gene | Ethephon-Treated | Water-Treated | Membership Function Analysis | |||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Delta CT | GeNorm (M1) | Normfinder (M2) | Bestkeeper (SD ± CV) | Delta CT | GeNorm (M1) | Normfinder (M2) | Bestkeeper (SD ± CV) | Principal Components Value | Membership Function Value | D-Value | Ranking | |||||||||||||
| Stability | Ranking | Stability | Ranking | Stability | Ranking | Stability | Ranking | Stability | Ranking | Stability | Ranking | Stability | Ranking | Stability | Ranking | X1 | X2 | X3 | μ1 | μ2 | μ3 | |||
| PtACT7 | 1.67 | 2 | 0.0413 | 1 | 0.05 | 2 | 4.53 ± 1.22 | 4 | 2.17 | 1 | 0.0362 | 1 | 1.04 | 1 | 8.4 ± 2.18 | 6 | −1.3454 | 0.4620 | 0.3636 | 0.4168 | −0.2128 | −0.3268 | 0.1076 | 2 |
| PtTUBB4 | 1.82 | 4 | 0.0413 | 1 | 0.07 | 5 | 4.67 ± 1.22 | 5 | 2.38 | 2 | 0.0362 | 1 | 1.57 | 3 | 6.75 ± 2.47 | 5 | −0.8446 | −0.1316 | 0.7332 | 0.2080 | 0.0232 | −0.4676 | 0.0192 | 3 |
| PtUBC10 | 1.48 | 1 | 0.0431 | 3 | 0.04 | 1 | 3.43 ± 0.76 | 3 | 2.47 | 3 | 0.08 | 3 | 1.56 | 2 | 6.99 ± 1.62 | 1 | −0.9374 | −0.7342 | −0.7009 | 0.2467 | 0.2628 | 0.0788 | 0.2145 | 1 |
| PtGADPH1 | 1.71 | 3 | 0.0662 | 4 | 0.05 | 3 | 5.03 ± 1.4 | 6 | 3.01 | 6 | 0.1341 | 7 | 2.41 | 6 | 10.25 ± 2.93 | 2 | 0.5027 | 1.0172 | −1.4941 | −0.3535 | −0.4336 | 0.3810 | −0.2150 | 5 |
| PtGADPH2 | 1.87 | 5 | 0.0759 | 5 | 0.06 | 4 | 3.41 ± 1.09 | 2 | 3.12 | 7 | 0.1266 | 6 | 2.58 | 7 | 6.59 ± 1.86 | 3 | 0.7629 | −1.0732 | −0.8311 | −0.4619 | 0.3976 | 0.1284 | −0.1310 | 4 |
| PtEIF3 | 2.28 | 7 | 0.0834 | 6 | 0.09 | 6 | 5.97 ± 2.12 | 7 | 2.73 | 4 | 0.1183 | 5 | 1.91 | 4 | 9.36 ± 2.31 | 7 | 0.8078 | 1.4419 | 0.7989 | −0.4806 | −0.6024 | −0.4927 | −0.5122 | 7 |
| PtEF1 | 2.22 | 6 | 0.0945 | 7 | 0.11 | 7 | 3.19 ± 1.16 | 1 | 2.84 | 5 | 0.109 | 4 | 2.16 | 5 | 6.75 ± 2.3 | 4 | 1.0539 | −0.9822 | 1.1303 | −0.5832 | 0.3614 | −0.6190 | −0.3663 | 6 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2023 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Liu, Y.; Li, Y.; He, B.; Yu, B.; Liu, B.; Ma, C.; Chen, J.; Zhang, T.; Zhang, H.; Liu, Y.; et al. The Genome-Wide Identification of Stable Internal Reference Genes Related to Delayed Spoilage for Accurate qRT-PCR Normalization in Ethephon-Treated Pueraria thomsonii Benth. Horticulturae 2023, 9, 1014. https://doi.org/10.3390/horticulturae9091014
Liu Y, Li Y, He B, Yu B, Liu B, Ma C, Chen J, Zhang T, Zhang H, Liu Y, et al. The Genome-Wide Identification of Stable Internal Reference Genes Related to Delayed Spoilage for Accurate qRT-PCR Normalization in Ethephon-Treated Pueraria thomsonii Benth. Horticulturae. 2023; 9(9):1014. https://doi.org/10.3390/horticulturae9091014
Chicago/Turabian StyleLiu, Yujia, Ya Li, Binrong He, Baiyin Yu, Boting Liu, Chongjian Ma, Jie Chen, Tianhua Zhang, Hongrui Zhang, Yuanlong Liu, and et al. 2023. "The Genome-Wide Identification of Stable Internal Reference Genes Related to Delayed Spoilage for Accurate qRT-PCR Normalization in Ethephon-Treated Pueraria thomsonii Benth." Horticulturae 9, no. 9: 1014. https://doi.org/10.3390/horticulturae9091014
APA StyleLiu, Y., Li, Y., He, B., Yu, B., Liu, B., Ma, C., Chen, J., Zhang, T., Zhang, H., Liu, Y., & Li, X. (2023). The Genome-Wide Identification of Stable Internal Reference Genes Related to Delayed Spoilage for Accurate qRT-PCR Normalization in Ethephon-Treated Pueraria thomsonii Benth. Horticulturae, 9(9), 1014. https://doi.org/10.3390/horticulturae9091014






