Functional Verification of the Four Splice Variants from Ajania purpurea NST1 in Transgenic Tobacco
Abstract
1. Introduction
2. Materials and Methods
2.1. Ajania purpurea Growth and RNA Isolation
2.2. Cloning of ApNST1, ApNST1.1, ApNST1.2 and ApNST1.3 and Plasmid Construction
2.3. Tobacco Transformation
2.4. Quantitative Real-Time PCR Assay
2.5. Transgenic Tobacco Growing Conditions
2.6. Abiotic Stress Treatment
2.6.1. Salt, Abscisic Acid and Low-Temperature Treatment under Aseptic Conditions
2.6.2. Salt, Drought and Low-Temperature Treatment under Natural Conditions
2.7. Fresh Weight, Root Length and Stem Height Were Measured
2.8. Sequence Alignment
2.9. Paraffin Section
2.10. Analysis of Data
3. Results
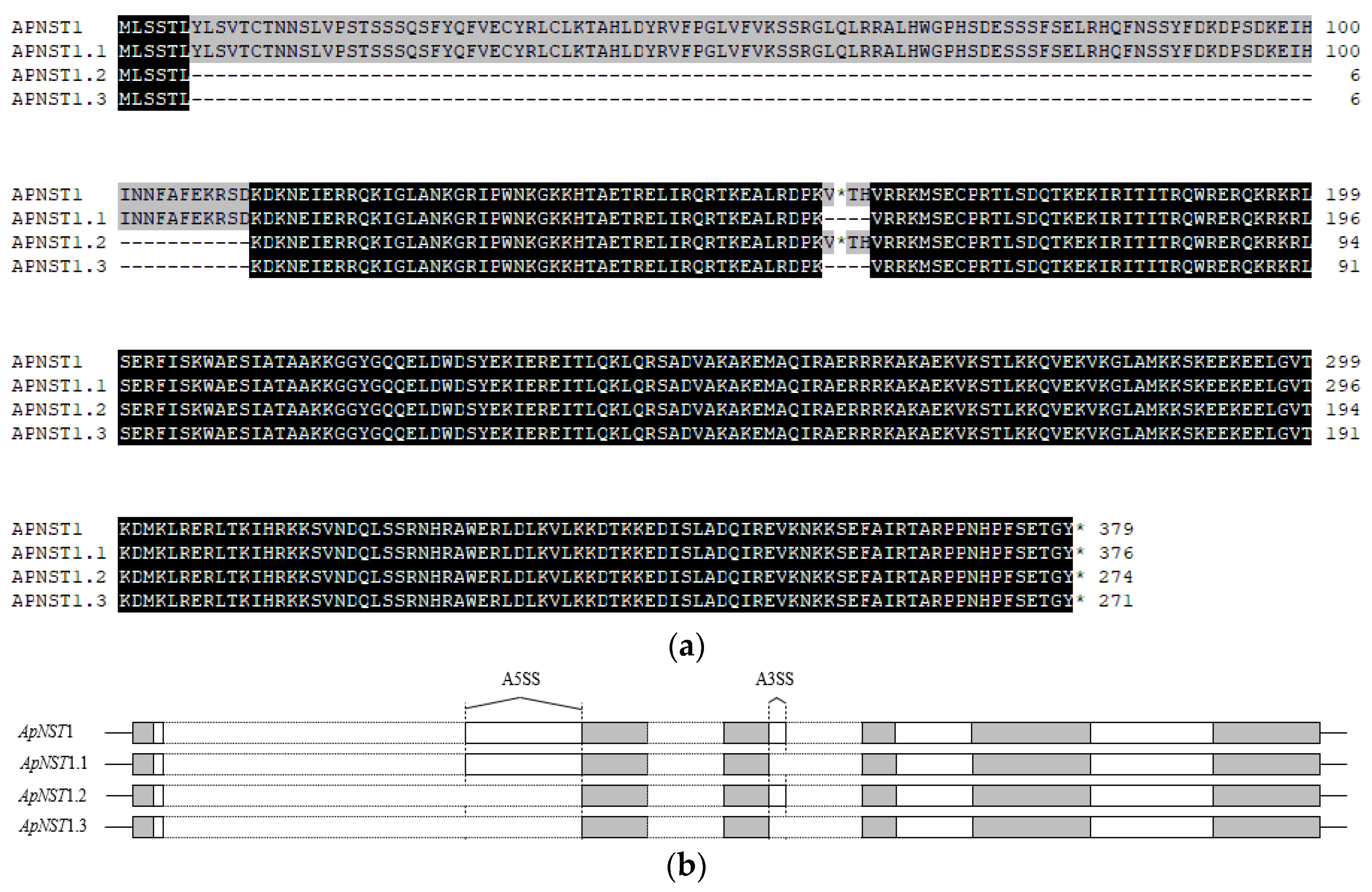
3.1. Bioinformatics Analysis of the Four Splice Variants from A. purpurea NST1
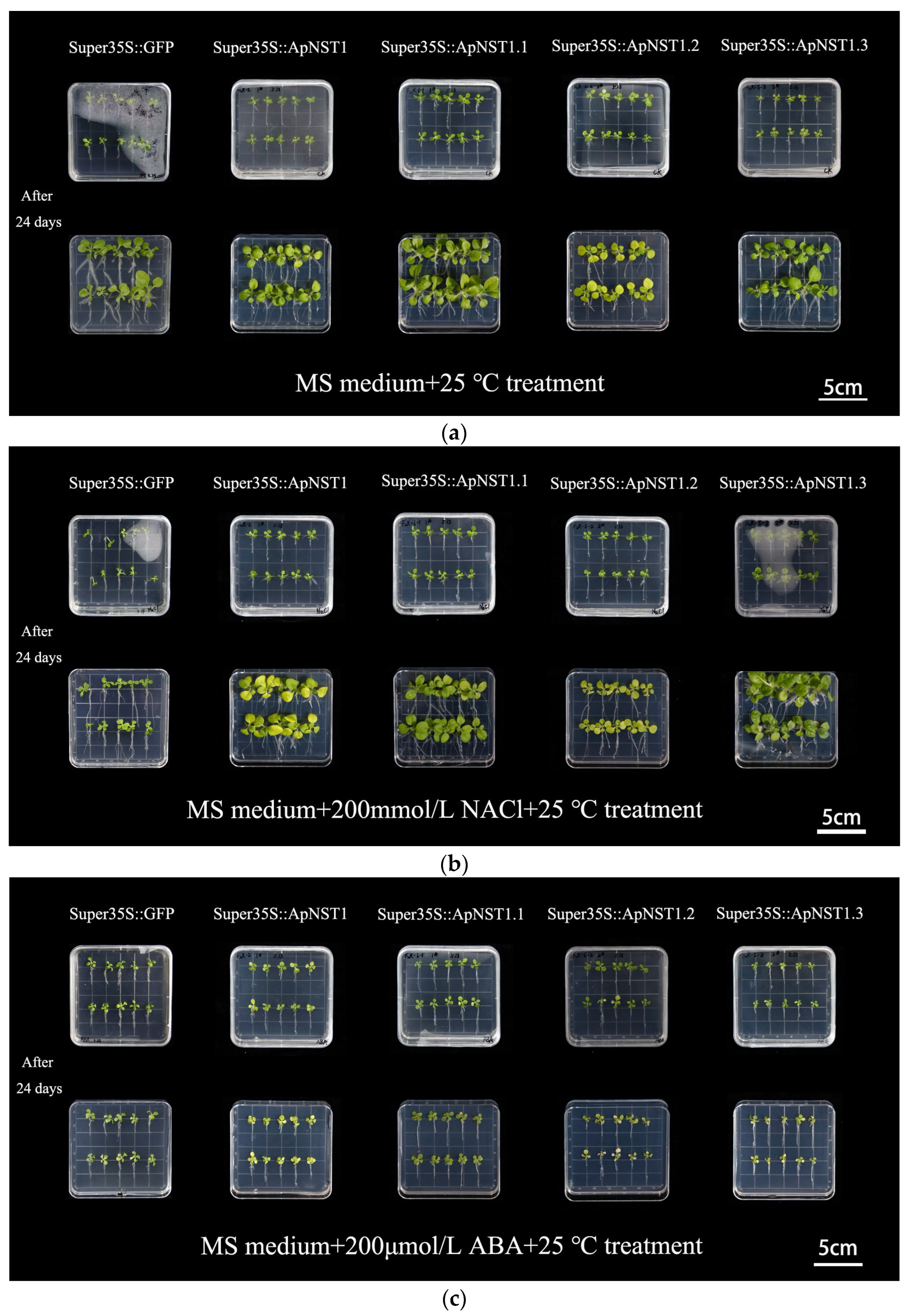
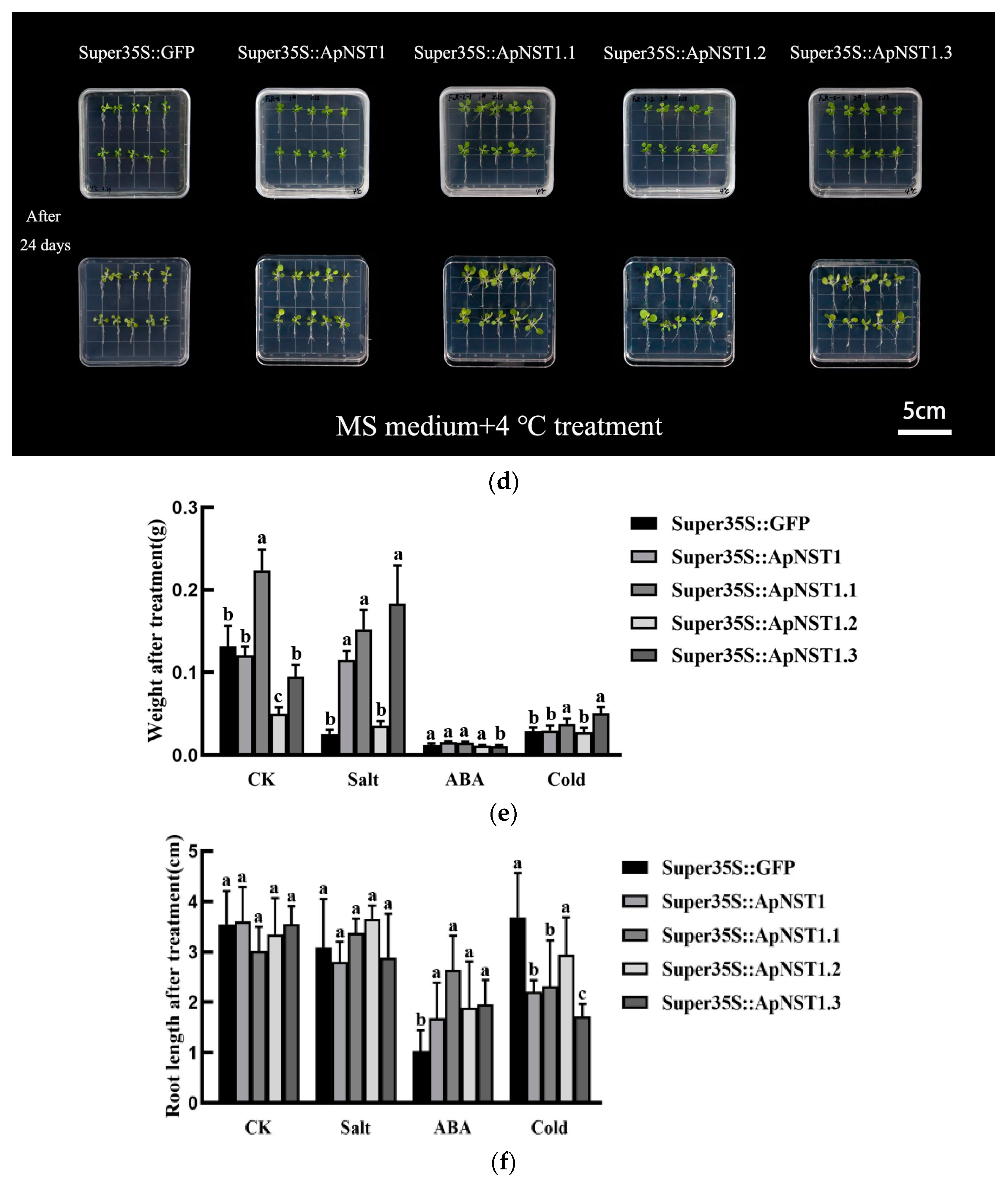
3.2. Phenotypes of Transgenic Tobacco Seedlings under Abiotic Stress
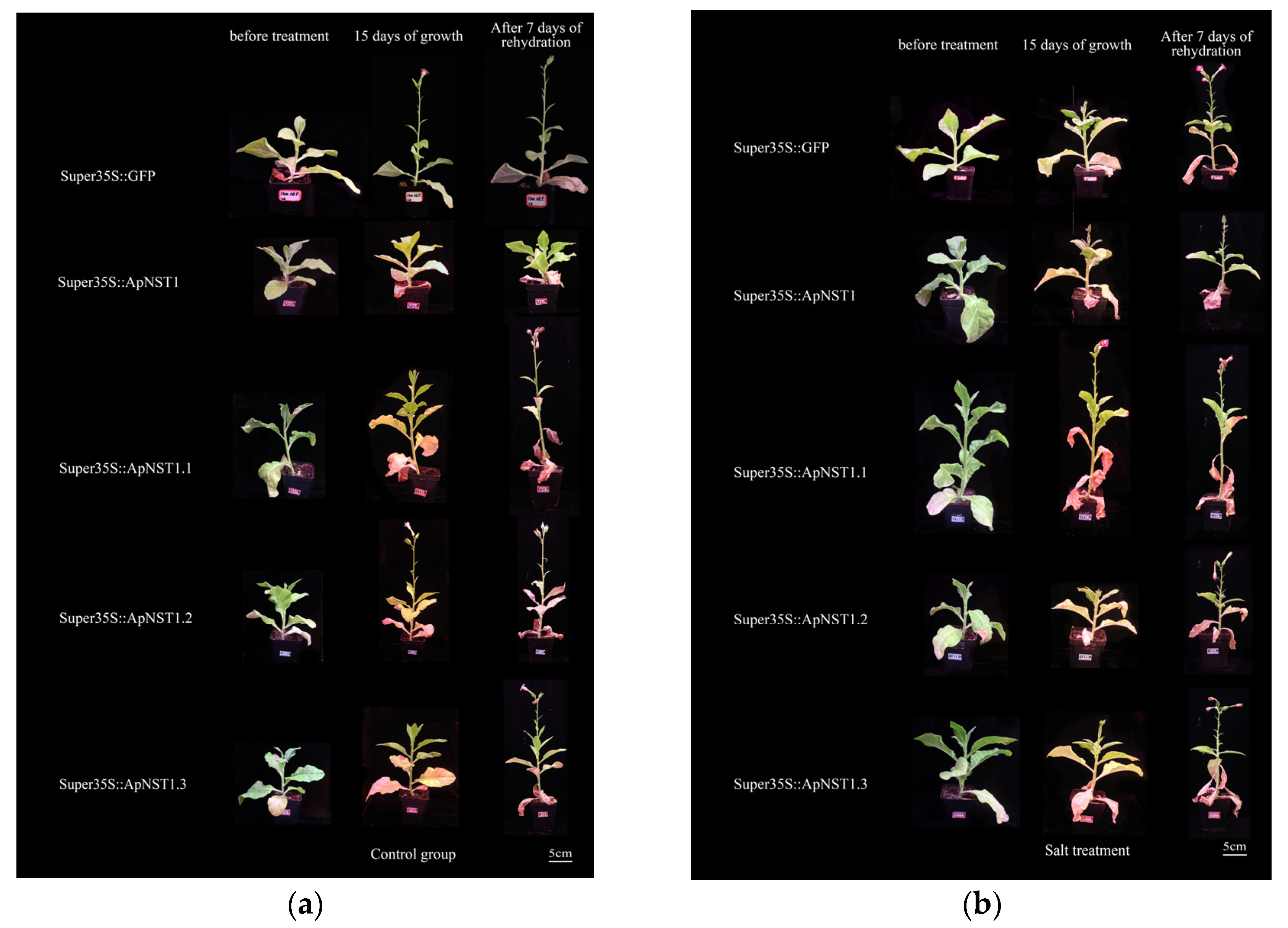
3.3. Phenotypes of Transgenic Tobacco Mature Seedlings under Abiotic Stress
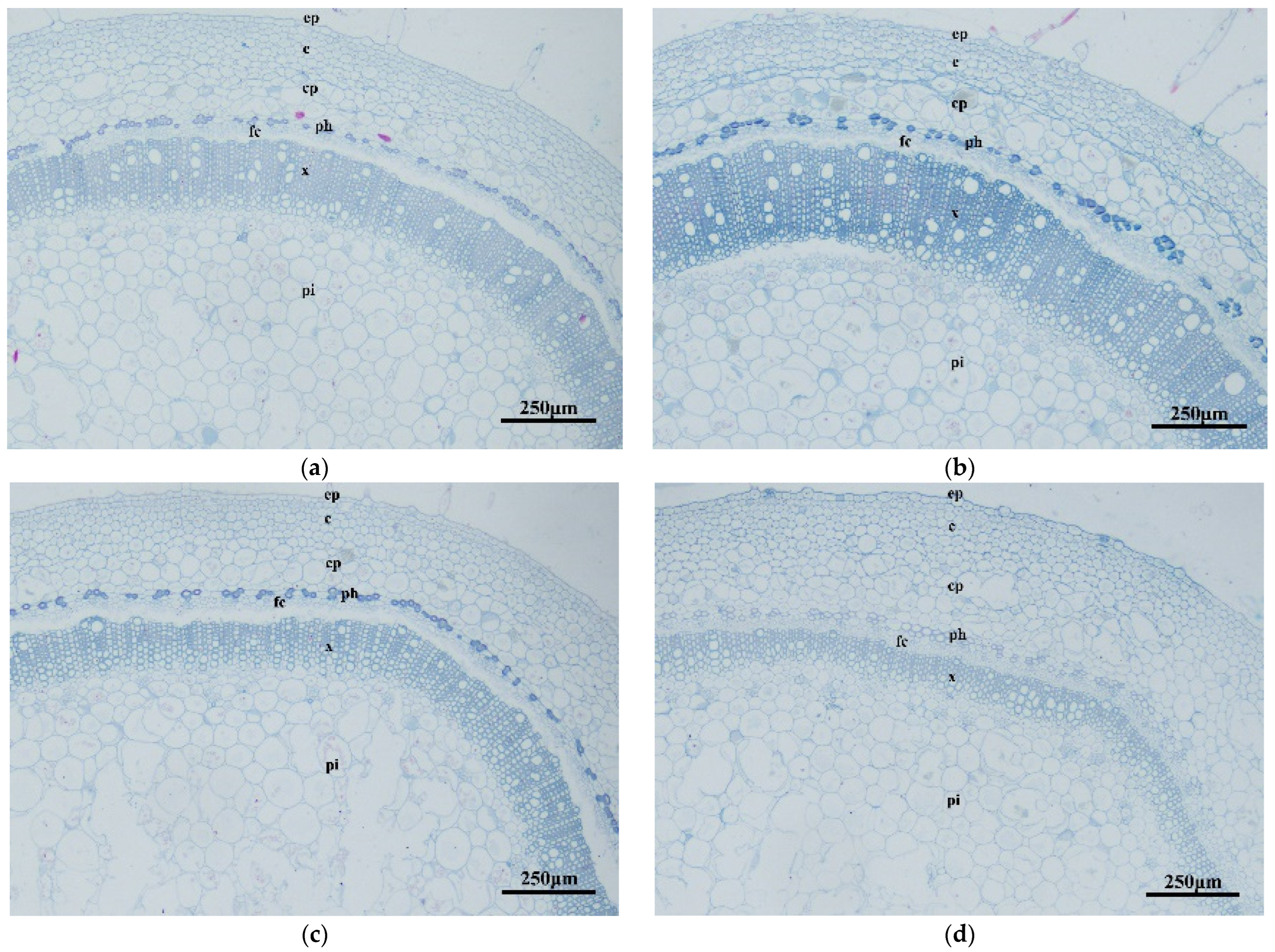
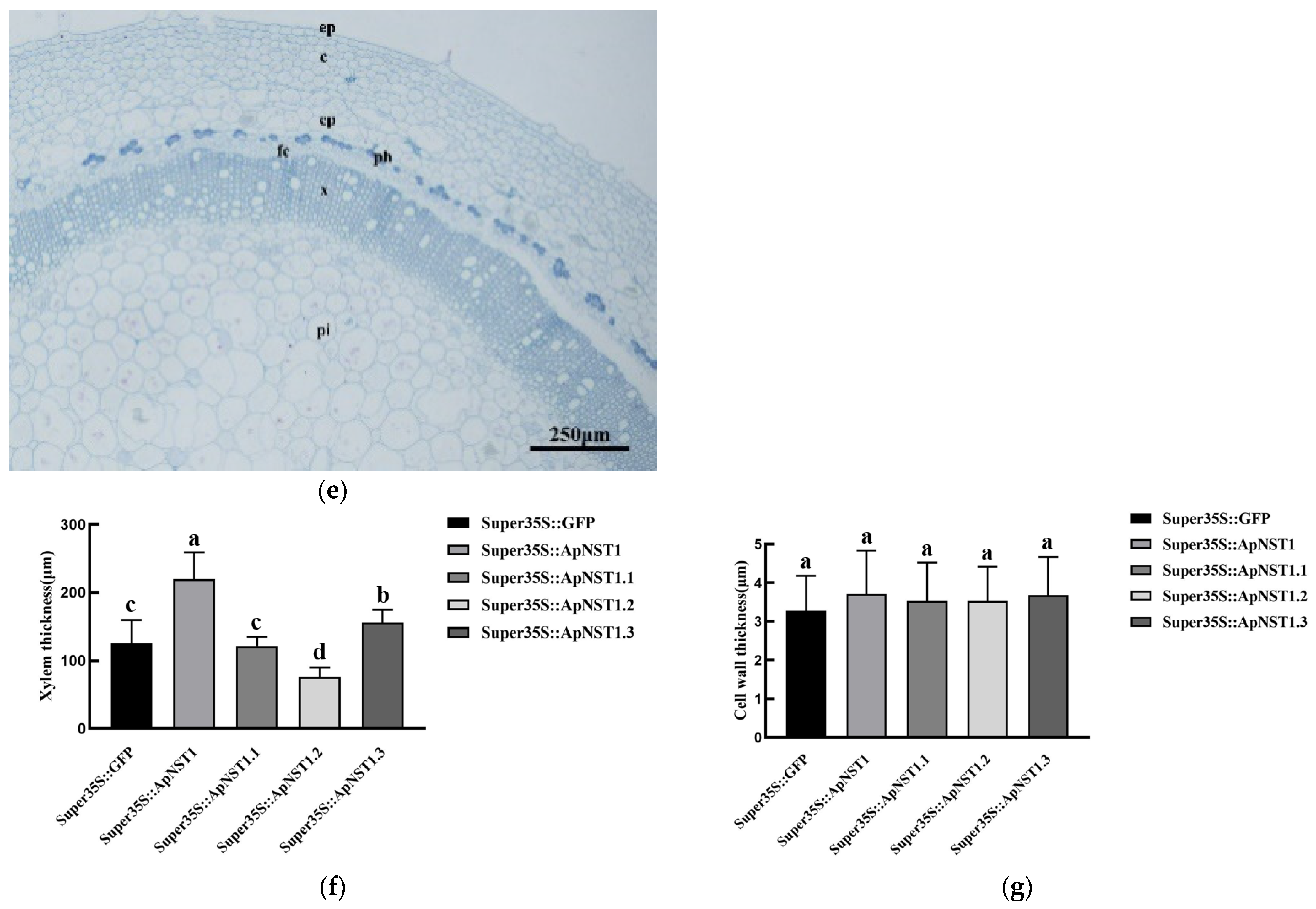
3.4. Analysis of Cross-Cut Structure of Transgenic Tobacco Stem
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Data Availability Statement
Conflicts of Interest
References
- Wu, X.; Ge, H.; Zhao, H. Callus induction and plant regeneration of Ajania purpurea. Acta Agric. Zhejiangensis 2014, 26, 335–338. [Google Scholar]
- Yang, X.; Jia, Z.; Pu, Q.; Tian, Y.; Zhu, F.; Liu, Y. ABA Mediates Plant Development and Abiotic Stress via Alternative Splicing. Int. J. Mol. Sci. 2022, 23, 3796. [Google Scholar] [CrossRef] [PubMed]
- Liu, C.; Yu, H.S.; Rao, X.L.; Li, L.G.; Dixon, R.A. Abscisic acid regulates secondary cell-wall formation and lignin deposition in Arabidopsis thaliana through phosphorylation of NST1. Proc. Natl. Acad. Sci. USA 2021, 118, e2010911118. [Google Scholar] [CrossRef] [PubMed]
- Zhang, L.; Zhang, H.W.; Yang, S.S. Cytosolic TaGAPC2 Enhances Tolerance to Drought Stress in Transgenic Arabidopsis Plants. Int. J. Mol. Sci. 2020, 21, 7499. [Google Scholar] [CrossRef]
- Lu, M.; Sun, Q.; Zhang, D.; Wang, T.; Pan, J. Identification of 7 stress-related NAC transcription factor members in maize (Zea mays L.) and characterization of the expression pattern of these genes. Biochem. Biophys. Res. Commun. 2015, 462, 144–150. [Google Scholar] [CrossRef]
- Nuruzzaman, M.; Sharoni, A.M.; Kikuchi, S. Roles of NAC transcription factors in the regulation of biotic and abiotic stress responses in plants. Front. Microbiol. 2013, 4, 248. [Google Scholar] [CrossRef]
- Souer, E.; van Houwelingen, A.; Kloos, D.; Mol, J.; Koes, R. The no apical meristem gene of Petunia is required for pattern formation in embryos and flowers and is expressed at meristem and primordia boundaries. Cell 1996, 85, 159–170. [Google Scholar] [CrossRef]
- Fang, Y.; You, J.; Xie, K.; Xie, W.; Xiong, L. Systematic sequence analysis and identification of tissue-specific or stress-responsive genes of NAC transcription factor family in rice. Mol. Genet. Genom. MGG 2008, 280, 547–563. [Google Scholar] [CrossRef]
- Ju, Y.L.; Yue, X.F.; Min, Z.; Wang, X.H.; Fang, Y.L.; Zhang, J.X. VvNAC17, a novel stress-responsive grapevine (Vitis vinifera L.) NAC transcription factor, increases sensitivity to abscisic acid and enhances salinity, freezing, and drought tolerance in transgenic Arabidopsis. Plant Physiol. Biochem. 2020, 146, 98–111. [Google Scholar] [CrossRef]
- Seok, H.; Woo, D.; Linh Vu, N.; Tran, H.T.; Tarte, V.N.; Mehdi, S.M.M.; Lee, S.; Moon, Y. Arabidopsis AtNAP functions as a negative regulator via repression of AREB1 in salt stress response. Planta 2017, 245, 329–341. [Google Scholar] [CrossRef]
- Yuan, X.; Wang, H.; Cai, J.; Bi, Y.; Li, D.; Song, F. Rice NAC transcription factor ONAC066 functions as a positive regulator of drought and oxidative stress response. BMC Plant Biol. 2019, 19, 278. [Google Scholar] [CrossRef] [PubMed]
- Liu, W.; Mei, Z.; Yu, L.; Gu, T.; Li, Z.; Zou, Q.; Zhang, S.; Fang, H.; Wang, Y.; Zhang, Z.; et al. The ABA-induced NAC transcription factor MdNAC1 interacts with a bZIP-type transcription factor to promote anthocyanin synthesis in red-fleshed apples. Hortic. Res. 2023, 10, uhad049. [Google Scholar] [CrossRef] [PubMed]
- Zhao, J.; Wu, Q.; Wu, H.; Wang, A.; Wang, X.; Li, C.; Zhao, H.; Wu, Q. FtNAC31, a Tartary buckwheat NAC transcription factor, enhances salt and drought tolerance in transgenic Arabidopsis. Plant Physiol. Biochem. 2022, 191, 20–33. [Google Scholar] [CrossRef]
- Xi, Y.; Ling, Q.; Zhou, Y.; Liu, X.; Qian, Y. ZmNAC074, a maize stress-responsive NAC transcription factor, confers heat stress tolerance in transgenic Arabidopsis. Front. Plant Sci. 2022, 13, 986628. [Google Scholar] [CrossRef]
- Takasaki, H.; Maruyama, K.; Kidokoro, S.; Ito, Y.; Fujita, Y.; Shinozaki, K.; Yamaguchi-Shinozaki, K.; Nakashima, K. The abiotic stress-responsive NAC-type transcription factor OsNAC5 regulates stress-inducible genes and stress tolerance in rice. Mol. Genet. Genom. MGG 2010, 284, 173–183. [Google Scholar] [CrossRef] [PubMed]
- Hu, H.; You, J.; Fang, Y.; Zhu, X.; Qi, Z.; Xiong, L. Characterization of transcription factor gene SNAC2 conferring cold and salt tolerance in rice. Plant Mol. Biol. 2008, 67, 169–181. [Google Scholar] [CrossRef]
- Ren, Y.; Huang, Z.; Jiang, H.; Wang, Z.; Wu, F.; Xiong, Y.; Yao, J. A heat stress responsive NAC transcription factor heterodimer plays key roles in rice grain filling. J. Exp. Bot. 2021, 72, 2947–2964. [Google Scholar] [CrossRef]
- Gonzali, S.; Perata, P. Fruit Colour and Novel Mechanisms of Genetic Regulation of Pigment Production in Tomato Fruits. Horticulturae 2021, 7, 259. [Google Scholar] [CrossRef]
- Thirumalaikumar, V.P.; Devkar, V.; Mehterov, N.; Ali, S.; Ozgur, R.; Turkan, I.; Mueller-Roeber, B.; Balazadeh, S. NAC transcription factor JUNGBRUNNEN1 enhances drought tolerance in tomato. Plant Biotechnol. J. 2018, 16, 354–366. [Google Scholar] [CrossRef]
- Zhou, H.; Sheng, Y.; Qiu, K.L.; Ren, F.; Shi, P.; Xie, Q.M.; Guo, J.Y.; Pan, H.F.; Zhang, J.Y. Improved Annotation of the Peach (Prunus persica) Genome and Identification of Tissue- or Development Stage-Specific Alternative Splicing through the Integration of Iso-Seq and RNA-Seq Data. Horticulturae 2023, 9, 175. [Google Scholar] [CrossRef]
- Filichkin, S.; Priest, H.D.; Megraw, M.; Mockler, T.C. Alternative splicing in plants: Directing traffic at the crossroads of adaptation and environmental stress. Curr. Opin. Plant Biol. 2015, 24, 125–135. [Google Scholar] [CrossRef]
- Ling, Y.; Mahfouz, M.M.; Zhou, S. Pre-mRNA alternative splicing as a modulator for heat stress response in plants. Trends Plant Sci. 2021, 26, 1153–1170. [Google Scholar] [CrossRef]
- Kornblihtt, A.R. A Long Noncoding Way to Alternative Splicing in Plant Development. Dev. Cell 2014, 30, 117–119. [Google Scholar] [CrossRef][Green Version]
- Capovilla, G.; Pajoro, A.; Immink, R.G.H.; Schmid, M. Role of alternative pre-mRNA splicing in temperature signaling. Curr. Opin. Plant Biol. 2015, 27, 97–103. [Google Scholar] [CrossRef] [PubMed]
- Shikata, H.; Hanada, K.; Ushijima, T.; Nakashima, M.; Suzuki, Y.; Matsushita, T. Phytochrome controls alternative splicing to mediate light responses in Arabidopsis. Proc. Natl. Acad. Sci. USA 2014, 111, 18781–18786. [Google Scholar] [CrossRef] [PubMed]
- Thatcher, S.R.; Danilevskaya, O.N.; Meng, X.; Beatty, M.; Zastrow-Hayes, G.; Harris, C.; Van Allen, B.; Habben, J.; Li, B. Genome-Wide Analysis of Alternative Splicing during Development and Drought Stress in Maize. Plant Physiol. 2016, 170, 586–599. [Google Scholar] [CrossRef] [PubMed]
- Lee, J.H.; Ryu, H.; Chung, K.S.; Pose, D.; Kim, S.; Schmid, M.; Ahn, J.H. Regulation of Temperature-Responsive Flowering by MADS-Box Transcription Factor Repressors. Science 2013, 342, 628–632. [Google Scholar] [CrossRef]
- Hemming, M.N.; Trevaskis, B. Make hay when the sun shines: The role of MADS-box genes in temperature-dependant seasonal flowering responses. Plant Sci. Int. J. Exp. Plant Biol. 2011, 180, 447–453. [Google Scholar] [CrossRef]
- Wang, X.; Wu, F.; Xie, Q.; Wang, H.; Wang, Y.; Yue, Y.; Gahura, O.; Ma, S.; Liu, L.; Cao, Y.; et al. SKIP Is a Component of the Spliceosome Linking Alternative Splicing and the Circadian Clock in Arabidopsis. Plant Cell 2012, 24, 3278–3295. [Google Scholar] [CrossRef]
- Horsch, R.B.; Fry, J.E.; Hoffmann, N.L.; Wallroth, M.; Eichholtz, D.; Rogers, S.G.; Fraley, R.T. A simple and general method for transferring genes into plants. Science 1985, 227, 1229–1231. [Google Scholar] [CrossRef]
- Wang, H. Function and Gene Editing of Nac Transcription Factors Related to Chamomile Lignin Synthesis; Beijing Forestry University: Beijing, China, 2021. [Google Scholar]
- Punzo, P.; Grillo, S.; Batelli, G. Alternative splicing in plant abiotic stress responses. Biochem. Soc. Trans. 2020, 48, 2117–2126. [Google Scholar] [CrossRef] [PubMed]
- Grau-Bove, X.; Ruiz-Trillo, I.; Irimia, M. Origin of exon skipping-rich transcriptomes in animals driven by evolution of gene architecture. Genome Biol. 2018, 19, 135. [Google Scholar] [CrossRef] [PubMed]
- Iniguez, L.P.; Ramirez, M.; Barbazuk, W.B.; Hernandez, G. Identification and analysis of alternative splicing events in Phaseolus vulgaris and Glycine max. BMC Genom. 2017, 18, 650. [Google Scholar] [CrossRef]
- Xing, Y.; Lee, C. Alternative splicing and RNA selection pressure—Evolutionary consequences for eukaryotic genomes. Nat. Rev. Genet. 2006, 7, 499–509. [Google Scholar] [CrossRef] [PubMed]
- Smith, C.C.R.; Tittes, S.; Mendieta, J.P.; Collier-zans, E.; Rowe, H.C.; Rieseberg, L.H.; Kane, N.C. Genetics of alternative splicing evolution during sunflower domestication. Proc. Natl. Acad. Sci. USA 2018, 115, 6768–6773. [Google Scholar] [CrossRef]
- Barbosa-Morais, N.L.; Irimia, M.; Pan, Q.; Xiong, H.Y.; Gueroussov, S.; Lee, L.J.; Slobodeniuc, V.; Kutter, C.; Watt, S.; Colak, R.; et al. The Evolutionary Landscape of Alternative Splicing in Vertebrate Species. Science 2012, 338, 1587–1593. [Google Scholar] [CrossRef]
- Liu, H.; Zhou, Y.; Li, H.; Wang, T.; Zhang, J.; Ouyang, B.; Ye, Z. Molecular and functional characterization of ShNAC1, an NAC transcription factor from Solanum habrochaites. Plant Sci. 2018, 271, 9–19. [Google Scholar] [CrossRef]
- Wang, G.D.; Liu, Q.; Shang, X.T.; Chen, C.; Xu, N.; Guan, J.; Meng, Q.W. Overexpression of transcription factor SlNAC35 enhances the chilling tolerance of transgenic tomato. Biol. Plant. 2018, 62, 479–488. [Google Scholar] [CrossRef]
- Fang, S.; Shang, X.; Yao, Y.; Li, W.; Guo, W. NST- and SND-subgroup NAC proteins coordinately act to regulate secondary cell wall formation in cotton. Plant Sci. 2020, 301, 110657. [Google Scholar] [CrossRef]




Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2023 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Wang, H.; Hao, X.; Zhang, W.; Guo, Y.; Zhao, X.; Li, Y.; He, W.; Cai, S.; Song, X. Functional Verification of the Four Splice Variants from Ajania purpurea NST1 in Transgenic Tobacco. Horticulturae 2023, 9, 916. https://doi.org/10.3390/horticulturae9080916
Wang H, Hao X, Zhang W, Guo Y, Zhao X, Li Y, He W, Cai S, Song X. Functional Verification of the Four Splice Variants from Ajania purpurea NST1 in Transgenic Tobacco. Horticulturae. 2023; 9(8):916. https://doi.org/10.3390/horticulturae9080916
Chicago/Turabian StyleWang, Hai, Xueying Hao, Wenxin Zhang, Yuning Guo, Xiang Zhao, Yanxi Li, Wenting He, Shiyi Cai, and Xuebin Song. 2023. "Functional Verification of the Four Splice Variants from Ajania purpurea NST1 in Transgenic Tobacco" Horticulturae 9, no. 8: 916. https://doi.org/10.3390/horticulturae9080916
APA StyleWang, H., Hao, X., Zhang, W., Guo, Y., Zhao, X., Li, Y., He, W., Cai, S., & Song, X. (2023). Functional Verification of the Four Splice Variants from Ajania purpurea NST1 in Transgenic Tobacco. Horticulturae, 9(8), 916. https://doi.org/10.3390/horticulturae9080916






