Antimicrobial Multiresistant Phenotypes of Genetically Diverse Pseudomonas spp. Isolates Associated with Tomato Plants in Chilean Orchards
Abstract
:1. Introduction
2. Materials and Methods
2.1. Bacterial Isolation and Culture Conditions
2.2. Molecular Characterization of Bacterial Isolates
2.3. Amplification, Sequencing and Analysis of the 16S rRNA Gene
2.4. Housekeeping Gene Sequencing and Multilocus Sequence Analysis (MLSA)
2.5. Copper Tolerance
2.6. Streptomycin Susceptibility
2.7. Antimicrobial Susceptibility Test (AST)
3. Results
3.1. Isolation and Identification of Pseudomonas from Orchards
3.2. Genetic Diversity of Pseudomonas spp. Isolates
3.3. Phenotypic Tolerance to Copper and Antibiotics
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Muñoz, V.M. Boletín de Hortalizas. Diciembre 2021 [En Linea]. Available online: https://app.powerbi.com/view?r=eyJrIjoiOGQ1OTI0YzMtYzUzOC00OWM5LWFiMzEtMTQxMGNmOTEyYzU3IiwidCI6IjMzYjdmNzA3LTZlNmYtNDJkMi04ZDZmLTk4YmZmOWZiNWZhMCIsImMiOjR9 (accessed on 10 July 2022).
- Basim, H.; Basim, E.; Yilmaz, S.; Dickstein, E.R.; Jones, J.B. An Outbreak of Bacterial Speck Caused by Pseudomonas syringae pv. tomato on Tomato Transplants Grown in Commercial Seedling Companies Located in the Western Mediterranean Region of Turkey. Plant Dis. 2004, 88, 1050. [Google Scholar] [CrossRef] [PubMed]
- Ottesen, A.R.; Peña, A.G.; White, J.R.; Pettengill, J.B.; Li, C.; Allard, S.; Rideout, S.; Allard, M.; Hill, T.; Evans, P.; et al. Baseline survey of the anatomical microbial ecology of an important food plant: Solanum lycopersicum (tomato). BMC Microbiol. 2013, 13, 114. [Google Scholar] [CrossRef] [PubMed]
- Dong, C.-J.; Wang, L.-L.; Li, Q.; Shang, Q.-M. Bacterial communities in the rhizosphere, phyllosphere and endosphere of tomato plants. PLoS ONE 2019, 14, e0223847. [Google Scholar] [CrossRef]
- Toju, H.; Okayasu, K.; Notaguchi, M. Leaf-associated microbiomes of grafted tomato plants. Sci. Rep. 2019, 9, 1787. [Google Scholar] [CrossRef] [PubMed]
- Parte, A.C. LPSN—List of prokaryotic names with standing in nomenclature (Bacterio.Net), 20 years on. Int. J. Syst. Evol. Microbiol. 2018, 68, 1825–1829. [Google Scholar] [CrossRef]
- Catara, V. Pseudomonas corrugata: Plant pathogen and/or biological resource? Mol. Plant Pathol. 2007, 8, 233–244. [Google Scholar] [CrossRef]
- Jones, J.B. Pseudomonas viridiflava: Causal Agent of Bacterial Leaf Blight of Tomato. Plant Dis. 1984, 68, 341–342. [Google Scholar] [CrossRef]
- Dimartino, M.; Panebianco, S.; Vitale, A.; Castello, I.; Leonardi, C.; Cirvilleri, G.; Polizzi, G. Occurrence and Patho-genicity of Pseudomonas fluorescens and P. putida on tomato plants in Italy. J. Plant Pathol. 2011, 93, 79–87. [Google Scholar] [CrossRef]
- Pekhtereva, E.; Kornev, K.; Matveeva, E.; Polityko, V.; Budenkov, N.; Ignatov, A.; Schaad, N. Pith necrosis of tomato in Russia. Acta Hortic. 2009, 808, 251–254. [Google Scholar] [CrossRef]
- Lamichhane, J.R.; Osdaghi, E.; Behlau, F.; Köhl, J.; Jones, J.B.; Aubertot, J.-N. Thirteen decades of antimicrobial copper compounds applied in agriculture. A review. Agron. Sustain. Dev. 2018, 38, 28. [Google Scholar] [CrossRef]
- Sundin, G.W.; Wang, N. Antibiotic Resistance in Plant-Pathogenic Bacteria. Annu. Rev. Phytopathol. 2018, 56, 161–180. [Google Scholar] [CrossRef]
- Cooksey, D.A.; Azad, H.R. Accumulation of Copper and Other Metals by Copper-Resistant Plant-Pathogenic and Saprophytic Pseudomonads. Appl. Environ. Microbiol. 1992, 58, 274–278. [Google Scholar] [CrossRef]
- Yang, C.H.; Menge, J.A.; Cooksey, D.A. Role of copper resistance in competitive survival of Pseudomonas fluorescens in soil. Appl. Environ. Microbiol. 1993, 59, 580–584. [Google Scholar] [CrossRef]
- Griffin, K.; Gambley, C.; Brown, P.; Li, Y. Copper-tolerance in Pseudomonas syringae pv. tomato and Xanthomonas spp. and the control of diseases associated with these pathogens in tomato and pepper. A systematic literature review. Crop Prot. 2017, 96, 144–150. [Google Scholar] [CrossRef]
- Cameron, A.; Sarojini, V. Pseudomonas syringae pv actinidiae: Chemical control, resistance mechanisms and possible alternatives. Plant Pathol. 2013, 63, 1–11. [Google Scholar] [CrossRef]
- Xu, Y.; Luo, Q.-Q.; Zhou, M.-G. Identification and Characterization of Integron-Mediated Antibiotic Resistance in the Phytopathogen Xanthomonas oryzae pv. oryzae. PLoS ONE 2013, 8, e55962. [Google Scholar] [CrossRef]
- Cooksey, D.A. Molecular mechanisms of copper resistance and accumulation in bacteria. FEMS Microbiol. Rev. 1994, 14, 381–386. [Google Scholar] [CrossRef]
- Walsh, C.A. Molecular mechanisms that confer antibacterial drug resistance. Nature 2000, 406, 775–781. [Google Scholar] [CrossRef]
- Thanner, S.; Drissner, D.; Walsh, F. Antimicrobial Resistance in Agriculture. mBio 2016, 7, e02227-15. [Google Scholar] [CrossRef]
- Ruimy, R.; Brisabois, A.; Bernede, C.; Skurnik, D.; Barnat, S.; Arlet, G.; Momcilovic, S.; Elbaz, S.; Moury, F.; Vibet, M.-A.; et al. Organic and conventional fruits and vegetables contain equivalent counts of Gram-negative bacteria expressing resistance to antibacterial agents. Environ. Microbiol. 2010, 12, 608–615. [Google Scholar] [CrossRef]
- Allydice-Francis, K.; Brown, P.D. Diversity of Antimicrobial Resistance and Virulence Determinants in Pseudomonas aeruginosa Associated with Fresh Vegetables. Int. J. Microbiol. 2012, 2012, 426241. [Google Scholar] [CrossRef]
- Sun, Y.; Guo, G.; Tian, F.; Chen, H.; Liu, W.; Li, M.; Wang, S. Antibiotic resistance genes and bacterial community on the surfaces of five cultivars of fresh tomatoes. Ecotoxicology 2020, 30, 1550–1558. [Google Scholar] [CrossRef]
- Scaccia, N.; Vaz-Moreira, I.; Manaia, C.M. The risk of transmitting antibiotic resistance through endophytic bacteria. Trends Plant Sci. 2021, 26, 1213–1226. [Google Scholar] [CrossRef]
- Baker-Austin, C.; Wright, M.S.; Stepanauskas, R.; McArthur, J. Co-selection of antibiotic and metal resistance. Trends Microbiol. 2006, 14, 176–182. [Google Scholar] [CrossRef]
- Seiler, C.; Berendonk, T.U. Heavy metal driven co-selection of antibiotic resistance in soil and water bodies impacted by agriculture and aquaculture. Front. Microbiol. 2012, 3, 399. [Google Scholar] [CrossRef]
- Scheck, H.J.; Pscheidt, J.W.; Moore, L.W. Copper and Streptomycin Resistance in Strains of Pseudomonas syringae from Pacific Northwest Nurseries. Plant Dis. 1996, 80, 1034–1039. [Google Scholar] [CrossRef]
- Hwang, M.S.H.; Morgan, R.L.; Sarkar, S.F.; Wang, P.W.; Guttman, D.S. Phylogenetic Characterization of Virulence and Resistance Phenotypes of Pseudomonas syringae. Appl. Environ. Microbiol. 2005, 71, 5182–5191. [Google Scholar] [CrossRef] [PubMed]
- Corzo-Ariyama, H.A.; García-Heredia, A.; Heredia, N.; García, S.; León, J.; Jaykus, L.; Solís-Soto, L. Phylogroups, pathotypes, biofilm formation and antimicrobial resistance of Escherichia coli isolates in farms and packing facilities of tomato, jalapeño pepper and cantaloupe from Northern Mexico. Int. J. Food Microbiol. 2018, 290, 96–104. [Google Scholar] [CrossRef] [PubMed]
- Berg, J.; Tom-Petersen, A.; Nybroe, O. Copper amendment of agricultural soil selects for bacterial antibiotic resistance in the field. Lett. Appl. Microbiol. 2004, 40, 146–151. [Google Scholar] [CrossRef] [PubMed]
- Esterio, M.; Pérez, L.; Auger, J. Analysis of genetic diversity of Xanthomonas arboricola pv juglandis strains, in Chile. Fitopatología 2007, 43, 110–121. [Google Scholar]
- Altimira, F.; Yáñez, C.; Bravo, G.; González, M.; Rojas, L.A.; Seeger, M. Characterization of copper-resistant bacteria and bacterial communities from copper-polluted agricultural soils of central Chile. BMC Microbiol. 2012, 12, 193. [Google Scholar] [CrossRef]
- Méndez, V.; Fuentes, S.; Morgante, V.; Hernández, M.; González, M.; Moore, E.; Seeger, M. Novel hydrocarbon clastic metal-tolerant Acinetobacter and Pseudomonas strains from Aconcagua River oil-polluted soil. J. Soil Sci. Plant Nutr. 2017, 17, 1074–1087. [Google Scholar] [CrossRef]
- Arguello, J.M.; Raimunda, D.; Padilla-Benavides, T. Mechanisms of copper homeostasis in bacteria. Front. Cell. Infect. Microbiol. 2013, 3, 73. [Google Scholar] [CrossRef]
- Igbinosa, I.H.; Nwodo, U.U.; Sosa, A.; Tom, M.; Okoh, A.I. Commensal Pseudomonas Species Isolated from Wastewater and Freshwater Milieus in the Eastern Cape Province, South Africa, as Reservoir of Antibiotic Resistant Determinants. Int. J. Environ. Res. Public Health 2012, 9, 2537–2549. [Google Scholar] [CrossRef]
- Roberts, M.C.; Schwarz, S. Tetracycline and Phenicol Resistance Genes and Mechanisms: Importance for Agriculture, the Environment, and Humans. J. Environ. Qual. 2016, 45, 576–592. [Google Scholar] [CrossRef]
- Armalytė, J.; Skerniškytė, J.; Bakienė, E.; Krasauskas, R.; Šiugždinienė, R.; Kareivienė, V.; Kerzienė, S.; Klimienė, I.; Sužiedėlienė, E.; Ružauskas, M. Microbial Diversity and Antimicrobial Resistance Profile in Microbiota From Soils of Conventional and Organic Farming Systems. Front. Microbiol. 2019, 10, 892. [Google Scholar] [CrossRef]
- Stockwell, V.; Duffy, B. Use of antibiotics in plant agriculture. Rev. Sci. Et Technol. De L’oie 2012, 31, 199–210. [Google Scholar] [CrossRef]
- Monteil, C.L.; Cai, R.; Liu, H.; Llontop, M.E.M.; Leman, S.; Studholme, D.J.; Morris, C.E.; Vinatzer, B.A. Nonagricultural reservoirs contribute to emergence and evolution of Pseudomonas syringae crop pathogens. New Phytol. 2013, 199, 800–811. [Google Scholar] [CrossRef]
- King, E.O.; Ward, M.K.; Raney, D.E. Two simple media for the demonstration of pyocyanin and fluorescin. J. Lab. Clin. Med. 1954, 44, 301–307. [Google Scholar]
- Frank, J.A.; Reich, C.I.; Sharma, S.; Weisbaum, J.S.; Wilson, B.A.; Olsen, G.J. Critical Evaluation of Two Primers Commonly Used for Amplification of Bacterial 16S rRNA Genes. Appl. Environ. Microbiol. 2008, 74, 2461–2470. [Google Scholar] [CrossRef]
- Sambrook, J.; Russell, D.W. Molecular Cloning: A Laboratory Manual (3-Volume Set); Molecular Cloning: A Laboratory Manual; Cold Spring Harbor Laboratory Press: Cold Spring Harbor, NY, USA, 2001. [Google Scholar]
- Sarkar, S.F.; Guttman, D.S. Evolution of the Core Genome of Pseudomonas syringae, a Highly Clonal, Endemic Plant Pathogen. Appl. Environ. Microbiol. 2004, 70, 1999–2012. [Google Scholar] [CrossRef] [PubMed]
- Yamamoto, S.; Harayama, S. Phylogenetic relationships of Pseudomonas putida strains deduced from the nucleotide sequences of gyrB, rpoD and 16S rRNA genes. Int. J. Syst. Bacteriol. 1998, 48, 813–819. [Google Scholar] [CrossRef] [PubMed]
- Anzai, Y.; Kim, H.; Park, J.-Y.; Wakabayashi, H.; Oyaizu, H. Phylogenetic affiliation of the Pseudomonads based on 16S rRNA sequence. Int. J. Syst. Evol. Microbiol. 2000, 50, 1563–1589. [Google Scholar] [CrossRef] [PubMed]
- Singh, V.; Kumar, A.; Kumar, S. Phylogenetic analysis on 16s ribosomal DNA of Pseudomonas strains from clinical cases of animals. Progress. Res.-Int. J. 2018, 13, 92–95. [Google Scholar]
- Berge, O.; Monteil, C.; Bartoli, C.; Chandeysson, C.; Guilbaud, C.; Sands, D.C.; Morris, C.E. A User’s Guide to a Data Base of the Diversity of Pseudomonas syringae and Its Application to Classifying Strains in This Phylogenetic Complex. PLoS ONE 2014, 9, e105547. [Google Scholar] [CrossRef]
- Flores, O.; Prince, C.; Nuñez, M.; Vallejos, A.; Mardones, C.; Yáñez, C.; Besoain, X.; Bastías, R. Genetic and Phenotypic Characterization of Indole-Producing Isolates of Pseudomonas syringae pv. actinidiae Obtained From Chilean Kiwifruit Orchards. Front. Microbiol. 2018, 9, 1907. [Google Scholar] [CrossRef]
- van de Peer, Y.; Salemi, M. Phylogenetic Inference Based on Distance Methods. In The Phylogenetic Handbook; Cambridge University Press: Cambridge, UK, 2012. [Google Scholar]
- Abbasi, P.A.; Khabbaz, S.E.; Weselowski, B.; Zhang, L. Occurrence of copper-resistant strains and a shift in Xanthomonas spp. causing tomato bacterial spot in Ontario. Can. J. Microbiol. 2015, 61, 753–761. [Google Scholar] [CrossRef]
- Nakajima, M.; Goto, M.; Hibi, T. Similarity between Copper Resistance Genes from Pseudomonas syringae pv. actinidiae and P. syringae pv. tomato. J. Gen. Plant Pathol. 2002, 68, 68–74. [Google Scholar] [CrossRef]
- Valenzuela, M.; Méndez, V.; Montenegro, I.; Besoain, X.; Seeger, M. Streptomycin resistance in Clavibacter michiganensis subsp. michiganensis strains from Chile is related to an rpsL gene mutation. Plant Pathol. 2018, 68, 426–433. [Google Scholar] [CrossRef]
- Sundin, G.W.; Bender, C.L. Ecological and genetic analysis of copper and streptomycin resistance in Pseudomonas syringae pv. syringae. Appl. Environ. Microbiol. 1993, 59, 1018–1024. [Google Scholar] [CrossRef]
- Clinical and Laboratory Standards Institute. Performance Standards for Antimicrobial Susceptibility Testing; Twenty-Second Informational Supplement; Scientific Research: Wuhan, China, 2012; Volume 32. [Google Scholar]
- Jorgensen, J.H.; Ferraro, M.J. Antimicrobial Susceptibility Testing: A Review of General Principles and Contemporary Practices overview of commonly used susceptibility testing methods. Clin. Infect. Dis. 2009, 49, 1749–1755. [Google Scholar] [CrossRef]
- Khezri, S.; Rahimian, H.; Ahangaran, A.; Mohammadi, M. Comparisons of Iranian strains of Pseudomonas syringae pv syringae from various hosts with different methods. Int. J. Agric. Biol. 2010, 12, 106–110. [Google Scholar]
- Sarker, M.R.; Islam, K.N.; Huri, H.Z.; Rahman, M.; Imam, H.; Hosen, B.; Mohammad, N.; Sarker, Z.I. Studies of the Impact of Occupational Exposure of Pharmaceutical Workers on the Development of Antimicrobial Drug Resistance. J. Occup. Health 2014, 56, 260–270. [Google Scholar] [CrossRef]
- Estay, P.; Bruna, A. Insectos, Ácaros y Enfermedades Asociadas al Tomate en Chile; Colección Libros INIA; Instituto de Investigaciones Agropecuarias: Santiago, Chile, 2002; Volume 7. [Google Scholar]
- Garrido-Sanz, D.; Arrebola, E.; Martínez-Granero, F.; García-Méndez, S.; Muriel, C.; Blanco-Romero, E.; Martín, M.; Rivilla, R.; Redondo-Nieto, M. Classification of Isolates from the Pseudomonas fluorescens Complex into Phylogenomic Groups Based in Group-Specific Markers. Front. Microbiol. 2017, 8, 413. [Google Scholar] [CrossRef]
- Haque, M.; Mosharaf, K.; Khatun, M.; Haque, A.; Biswas, S.; Islam, S.; Islam, M.; Shozib, H.B.; Miah, M.U.; Molla, A.H.; et al. Biofilm Producing Rhizobacteria with Multiple Plant Growth-Promoting Traits Promote Growth of Tomato Under Water-Deficit Stress. Front. Microbiol. 2020, 11, 542053. [Google Scholar] [CrossRef]
- Guo, Q.; Sun, Y.; Shi, M.; Han, X.; Jing, Y.; Li, Y.; Li, H. Pseudomonas koreensis promotes tomato growth and shows potential to induce stress tolerance via auxin and polyphenol-related pathways. Plant Soil 2021, 462, 141–158. [Google Scholar] [CrossRef]
- Couillerot, O.; Prigent-Combaret, C.; Caballero-Mellado, J.; Moënne-Loccoz, Y. Pseudomonas fluorescens and closely-related fluorescent pseudomonads as biocontrol agents of soil-borne phytopathogens. Lett. Appl. Microbiol. 2009, 48, 505–512. [Google Scholar] [CrossRef]
- Oueslati, M.; Mulet, M.; Zouaoui, M.; Chandeysson, C.; Lalucat, J.; Hajlaoui, M.R.; Berge, O.; García-Valdés, E.; Sadfi-Zouaoui, N. Diversity of pathogenic Pseudomonas isolated from citrus in Tunisia. AMB Express 2020, 10, 198. [Google Scholar] [CrossRef]
- Cai, R.; Lewis, J.; Yan, S.; Liu, H.; Clarke, C.R.; Campanile, F.; Almeida, N.; Studholme, D.; Lindeberg, M.; Schneider, D.; et al. The Plant Pathogen Pseudomonas syringae pv. tomato Is Genetically Monomorphic and under Strong Selection to Evade Tomato Immunity. PLOS Pathog. 2011, 7, e1002130. [Google Scholar] [CrossRef]
- Almeida, N.F.; Yan, S.; Lindeberg, M.; Studholme, D.J.; Schneider, D.J.; Condon, B.; Liu, H.; Viana, C.J.; Warren, A.; Evans, C.; et al. A Draft genome sequence of Pseudomonas syringae pv tomato T1 reveals a type iii effector repertoire significantly divergent from that of Pseudomonas syringae pv tomato DC3000. Mol. Plant-Microbe Interact. MPMI 2009, 22, 52–62. [Google Scholar] [CrossRef]
- Economou, V.; Gousia, P. Agriculture and food animals as a source of antimicrobial-resistant bacteria. Infect. Drug Resist. 2015, 8, 49–61. [Google Scholar] [CrossRef]
- Compeau’t Boutros, G.; Al-Achi, B.J.; Platsouka, E.; Levy, S.B. Survival of rifampin-resistant mutants of Pseudomonas fluorescens and Pseudomonas putida in soil systems. Appl. Environ. Microbiol. 1988, 54, 2432–2438. [Google Scholar] [CrossRef]
- Leclercq, R.; Cantón, R.; Brown, D.F.J.; Giske, C.G.; Heisig, P.; MacGowan, A.P.; Mouton, J.W.; Nordmann, P.; Rodloff, A.C.; Rossolini, G.M.; et al. EUCAST expert rules in antimicrobial susceptibility testing. Clin. Microbiol. Infect. 2013, 19, 141–160. [Google Scholar] [CrossRef]
- Estepa, V.; Rojo-Bezares, B.; Torres, C.; Sáenz, Y. Genetic Lineages and Antimicrobial Resistance in Pseudomonas spp. Isolates Recovered from Food Samples. Foodborne Pathog. Dis. 2015, 12, 486–491. [Google Scholar] [CrossRef]
- Zhou, Y.; Xu, Y.-B.; Xu, J.-X.; Zhang, X.-H.; Xu, S.-H.; Du, Q.-P. Combined Toxic Effects of Heavy Metals and Antibiotics on a Pseudomonas fluorescens Strain ZY2 Isolated from Swine Wastewater. Int. J. Mol. Sci. 2015, 16, 2839–2850. [Google Scholar] [CrossRef]
- Quintieri, L.; Fanelli, F.; Caputo, L. Antibiotic Resistant Pseudomonas Spp. Spoilers in Fresh Dairy Products: An Underestimated Risk and the Control Strategies. Foods 2019, 8, 372. [Google Scholar] [CrossRef]
- Mann, A.; Nehra, K.; Rana, J.; Dahiya, T. Antibiotic resistance in agriculture: Perspectives on upcoming strategies to overcome upsurge in resistance. Curr. Res. Microb. Sci. 2021, 2, 100030. [Google Scholar] [CrossRef]
- Buttimer, C.; McAuliffe, O.; Ross, R.P.; Hill, C.; O’Mahony, J.; Coffey, A. Bacteriophages and Bacterial Plant Diseases. Front. Microbiol. 2017, 8, 34. [Google Scholar] [CrossRef]
- Oniciuc, E.A.; Likotrafiti, E.; Alvarez-Molina, A.; Prieto, M.; Santos, J.A.; Alvarez-Ordóñez, A. The Present and Future of Whole Genome Sequencing (WGS) and Whole Metagenome Sequencing (WMS) for Surveillance of Antimicrobial Resistant Microorganisms and Antimicrobial Resistance Genes across the Food Chain. Genes 2018, 9, 268. [Google Scholar] [CrossRef]
- Bender, C.L.; Cooksey, D.A. Indigenous plasmids in Pseudomonas syringae pv. tomato: Conjugative transfer and role in copper resistance. J. Bacteriol. 1986, 165, 534–541. [Google Scholar] [CrossRef]
- Nongkhlaw, F.M.W.; Joshi, S.R. Horizontal Gene Transfer of the Non-ribosomal Peptide Synthetase Gene Among Endophytic and Epiphytic Bacteria Associated with Ethnomedicinal Plants. Curr. Microbiol. 2016, 72, 1–11. [Google Scholar] [CrossRef] [PubMed]
- Karmakar, R.; Bindiya, S.; Hariprasad, P. Convergent evolution in bacteria from multiple origins under antibiotic and heavy metal stress, and endophytic conditions of host plant. Sci. Total Environ. 2019, 650, 858–867. [Google Scholar] [CrossRef] [PubMed]
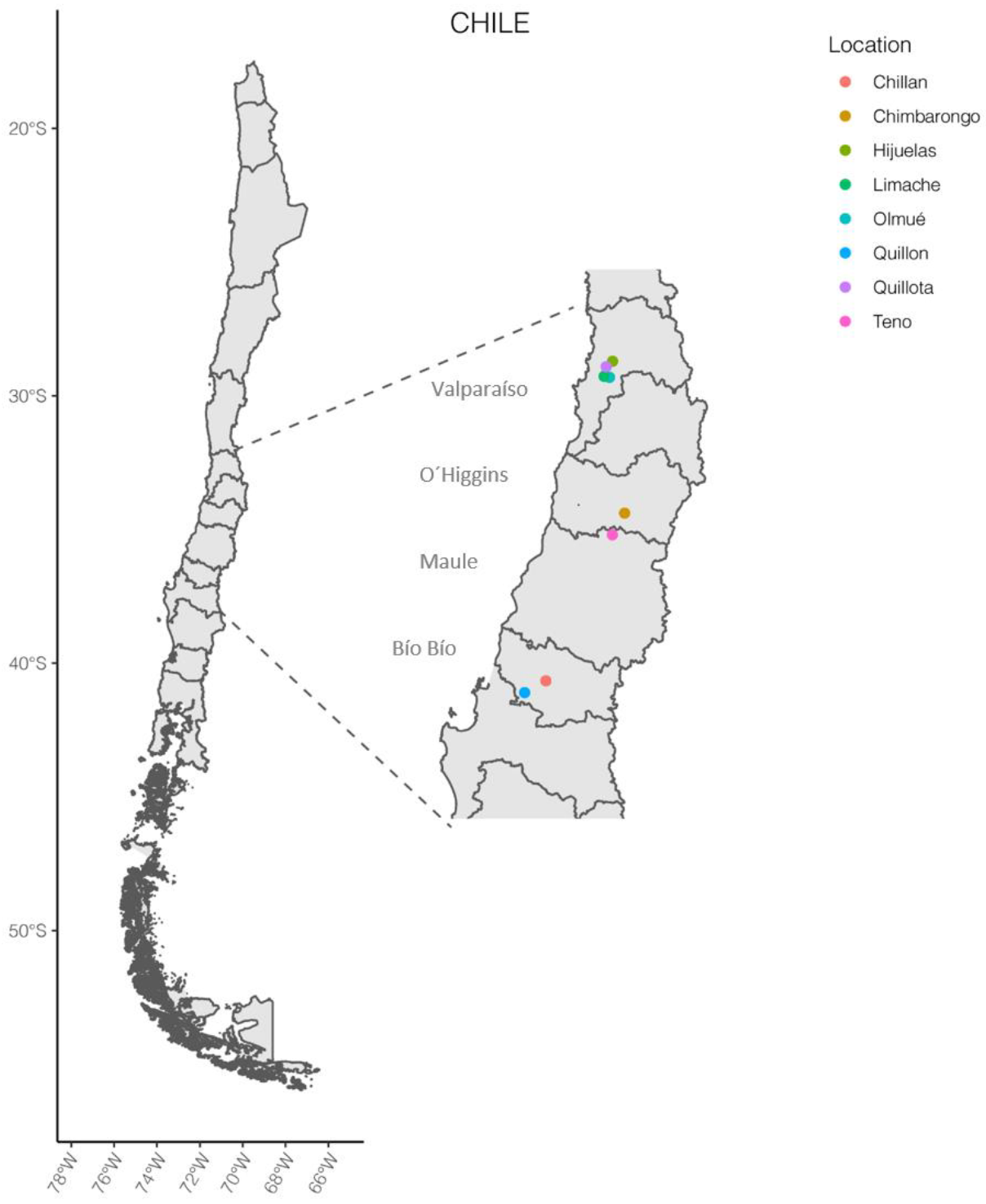


| Strain | Sample Origin | Plant Health Appearance 2,3 | Cultivation Modality | Región 1/Location | Isolation Date |
|---|---|---|---|---|---|
| pA2.4 | tomato plant | with signs of illness | greenhouse | V/Olmué | August 2018 |
| pA2.5 | tomato plant | with signs of illness | greenhouse | V/Olmué | August 2018 |
| pA2.6 | tomato plant | with signs of illness | greenhouse | V/Olmué | August 2018 |
| pAI.1 | tomato plant | with signs of illness | greenhouse | V/Olmué | August 2018 |
| pAII.1 | tomato plant | with signs of illness | greenhouse | V/Olmué | August 2018 |
| pAII.2 | tomato plant | with signs of illness | greenhouse | V/Olmué | August 2018 |
| pFL13 | tomato plant debris | recently dead | greenhouse | V/Limache | July 2018 |
| pFL17 | tomato plant debris | recently dead | greenhouse | V/Limache | July 2018 |
| pFL2 | tomato plant | healthy | greenhouse | V/Limache | July 2018 |
| pFL3 | tomato plant | healthy | greenhouse | V/Limache | July 2018 |
| pFL4 | tomato plant | healthy | greenhouse | V/Limache | July 2018 |
| pFL5 | tomato plant | healthy | greenhouse | V/Limache | July 2018 |
| pFL8 | tomato plant | healthy | greenhouse | V/Limache | July 2018 |
| pJV.8r | tomato plant | with signs of illness | greenhouse | V/Limache | August 2018 |
| pJV.9r | tomato plant | with signs of illness | greenhouse | V/Limache | August 2018 |
| pJ.1 | tomato plant | with signs of illness | greenhouse | V/Limache | August 2018 |
| pJ3.2 | tomato plant | with signs of illness | greenhouse | V/Limache | August 2018 |
| pJ4.1 | tomato plant | with signs of illness | greenhouse | V/Limache | August 2018 |
| pJ5.1 | tomato plant | with signs of illness | greenhouse | V/Limache | August 2018 |
| pJ5.2 | tomato plant | with signs of illness | greenhouse | V/Limache | August 2018 |
| pJ4.2 | tomato plant | with signs of illness | greenhouse | V/Limache | August 2018 |
| pJ6.3 | tomato plant | with signs of illness | greenhouse | V/Limache | August 2018 |
| p1.12 | tomato seedling | healthy | nursery | V/Hijuelas | August 2018 |
| p10.3 | tomato seedling | healthy | nursery | V/Hijuelas | August 2018 |
| p2.2 | tomato seedling | healthy | nursery | V/Hijuelas | August 2018 |
| p4.1 | tomato seedling | healthy | nursery | V/Hijuelas | August 2018 |
| p4.8 | tomato seedling | healthy | nursery | V/Hijuelas | August 2018 |
| p9.7 | tomato seedling | healthy | nursery | V/Hijuelas | August 2018 |
| p9.9 | tomato seedling | healthy | nursery | V/Hijuelas | August 2018 |
| p4.3 | tomato seedling | with signs of illness | nursery | V/Hijuelas | August 2018 |
| pAZ.24 | tomato plant | healthy | open field | VII/Teno | December 2018 |
| pAZ.25 | tomato plant | healthy | open field | VII/Teno | December 2018 |
| pAZ.26 | tomato plant | healthy | open field | VII/Teno | December 2018 |
| p27Ch | tomato plant | with signs of illness | open field | VIII/Chillán | April 2018 |
| p29Ch | tomato plant | with signs of illness | open field | VIII/Chillán | April 2018 |
| p42Ch | tomato plant | with signs of illness | open field | VIII/Chillán | April 2018 |
| pH7.13 | tomato plant | healthy | greenhouse | VIII/Quillón | December 2018 |
| pH7.17 | tomato plant | healthy | greenhouse | VIII/Quillón | December 2018 |
| pH6.6 | tomato plant | healthy | greenhouse | VIII/Quillón | December 2018 |
| pH7.10 | tomato plant | healthy | greenhouse | VIII/Quillón | December 2018 |
| pHm27 | tomato plant | healthy | greenhouse | VIII/Quillón | December 2018 |
| pH7.14 | tomato plant | healthy | greenhouse | VIII/Quillón | December 2018 |
| pHm28 | tomato plant | healthy | greenhouse | VIII/Quillón | December 2018 |
| pH7.18 | tomato plant | healthy | greenhouse | VIII/Quillón | December 2018 |
| pH7.9 | tomato plant | healthy | greenhouse | VIII/Quillón | December 2018 |
| pH6.2 | tomato plant | healthy | greenhouse | VIII/Quillón | December 2018 |
| pH6.5 | tomato plant | healthy | greenhouse | VIII/Quillón | December 2018 |
| p22 | tomato seedling | with signs of illness | nursery | V/Quillota | November 2020 |
| p23 | tomato seedling | with signs of illness | nursery | V/Hijuelas | November 2020 |
| p24 | tomato seedling | with signs of illness | nursery | V/Quillota | November 2020 |
| p26 | tomato seedling | with signs of illness | nursery | V/Hijuelas | March 2021 |
| p27 | tomato seedling | with signs of illness | nursery | V/Hijuelas | March 2021 |
| p28 | tomato seedling | with signs of illness | nursery | V/Limache | March 2021 |
| pT11 | tomato seedling | with signs of illness | nursery | VI/Chimbarongo | August 2018 |
| pLO2.3 | tobacco seedling | with signs of illness | nursery | VI/Chimbarongo | October 2018 |
| pLO4.2 | tobacco seedling | with signs of illness | nursery | VI/Chimbarongo | October 2018 |
| pLO5.2 | cucumber seedling | healthy | nursery | VI/Chimbarongo | October 2018 |
| pl1.1 | lettuce seedling | healthy | nursery | VI/Chimbarongo | August 2018 |
| pl3.12 | lettuce seedling | healthy | nursery | VI/Chimbarongo | August 2018 |
| pl1.2 | lettuce seedling | with signs of illness | nursery | VI/Chimbarongo | August 2018 |
| pl1.3 | lettuce seedling | with signs of illness | nursery | VI/Chimbarongo | August 2018 |
| pJS4.2 | soil | - | greenhouse | V/Limache | August 2018 |
| pJS5.5 | soil | - | greenhouse | V/Limache | August 2018 |
| pAg2 | irrigation water | - | greenhouse | VII/Teno | October 2018 |
| Primer Name | Sequence (5′-3′) | Target Gene | Tm (°C) 1 |
|---|---|---|---|
| 27F | AGAGTTTGATCMTGGCTCAG | 16SrRNA | 58 |
| 1492R | TACGGYTACCTTGTTACGACTT | ||
| cts-Fp | AGTTGATCATCGAGGGCGCWGCC | cts | 56 |
| cts-Rp | TGATCGGTTTGATCTCGCACGG | ||
| acn-Fp | ACATCCCGCTGCACGCYCTGGCC | acn | 60 |
| acn-Rp | GTGGTGTCCTGGGAACCGACGGTG | ||
| pgi-Fp | TGCAGGACTTCAGCATGCGCGAAGC | pgi | 60 |
| pgi-Rp | CGAGCCGCCCTGSGCCAGGTACCAG | ||
| rpoD-Fp | AAGGCGARATCGAAATCGCCAAGCG | rpoD | 63 |
| rpoD-Rps | GGAACWKGCGCAGGAAGTCGGCACG |
| Species | Strain Name | Country | Access Number |
|---|---|---|---|
| Pseudomonas fluorescens | FDAARGOS 1088 | Germany | NZ_CP068151.1 |
| Pseudomonas syringae pv. Syringae | B728a | USA | NC_007005.1 |
| Pseudomonas syringae pv. tomato | DC3000 | UK | NC_004578.1 |
| Pseudomonas syringae pv. tomato | T1 | Canada | NZ_ABSM00000000 |
| Pseudomonas viridiflava | CFBP 1590 | France | NZ_LT855380.1 |
| Pseudomonas putida | KT2440 | Japan | NC_002947.4 |
| Pseudomonas koreensis | BS3658 | Korea | LT629687.1 |
| Pseudomonas gessardii | BS2982 | France | FNKR01000003 |
| Pseudomonas azotoformans | LMG21611 | Japan | LT629702 |
| Escherichia coli | K-12 | USA | U00096 |
| Isolates | Species Identification | Identity (%) | GenBank Accession Number 1 | |
|---|---|---|---|---|
| Isolates matching with single Pseudomonas species | p22, p23, p24, p26, p27, p28, pJV.8r, pJ.1, pJ3.2, pJ5.1, pJ5.2, pH6.5, pH6.6, pH7.9, pH7.10, pH7.13, pH7.14, pH7.17, pH7.18 | Pseudomonas syringae | 99.5–100 | MW138064 |
| pAZ.24, pAZ.25, pAZ.26, pH6.2, pFL4, pFL5 | Pseudomonas viridiflava | 100 | OK091003 | |
| p27Ch, p29Ch, p42Ch, pLO4.2, pLO5.2, p1.12, p2.2 | Pseudomonas koreensis | 99.9–100 | MZ707723 | |
| pJ6.3, p10.3, pLO2.3 | Pseudomonas azotoformans | 100 | MW221357 | |
| pA2.4, pA2.5, pA2.6 | Pseudomonas gessardii | 100 | MT889683 | |
| pHm27 | Pseudomonas putida | 100 | MZ209185 | |
| pJV.9r | Pseudomonas marginalis | 100 | OK086045 | |
| pHm28 | Pseudomonas fluorescens | 99.3 | KJ590506 | |
| Isolates matching with at least two Pseudomonas species | pFL2, pFL3, pFL13, pFL17, pJ4.2, pl1.3, pAII.1, pAII.2 | Pseudomonas paralactis | 100 | MZ674188 |
| Pseudomonas putida | 100 | MZ497031 | ||
| Pseudomonas gessardii | 100 | MZ452411 | ||
| pFL8, p4.8, p9.7, p9.9 | Pseudomonas fluorescens | 100 | MZ503692 | |
| Pseudomonas koreensis | 100 | MW524107 | ||
| pAI.1, pT11 | Pseudomonas putida | 99.9 | OK083428 | |
| Pseudomonas koreensis | 99.9 | MZ965051 | ||
| Pseudomonas fluorescens | 99.9 | MZ914652 | ||
| pJS5.5 | Pseudomonas punonensis | 99.9 | NR_109583 | |
| Pseudomonas argentinensis | 99.9 | MZ853950 | ||
| Pseudomonas sp. | 99.9 | MW033798 | ||
| pl1.1 | Pseudomonas sp. | 100 | MT354167 | |
| Pseudomonas koreensis | 100 | MK790616 | ||
| pJS4.2 | Pseudomonas brenneri | 100 | MZ914419 | |
| Pseudomonas fluorescens | 100 | MW295497 | ||
| Isolates matching with undefined Pseudomonas species | pl3.12 | Pseudomonas sp. | 100 | MZ825296 |
| pl1.2 | Pseudomonas sp. | 100 | MZ758888 | |
| p4.3 | Pseudomonas sp. | 100 | MW930799 | |
| pAg2 | Pseudomonas sp. | 100 | JN899567 | |
| pJ4.1 | Pseudomonas sp. | 99.1 | MW089209 |
| MIC (µg/mL) | AST Test 1 | ||||||
|---|---|---|---|---|---|---|---|
| Isolate | Cu | Streptomycin | Ampicillin | Chloramphenicol | Gentamicin | Tetracycline | Rifampin |
| pA2.4 | 64 | 15 | R | R | S | S | R |
| pA2.5 | 32 | 4 | R | S | MS | S | R |
| pA2.6 | 64 | 250 | MS | S | S | S | R |
| pAI.1 | 32 | 6 | S | S | S | S | R |
| pAII.1 | 32 | 100 | R | R | S | S | R |
| pAII.2 | 64 | 50 | R | R | S | S | R |
| pFL13 | 32 | 100 | R | R | S | S | R |
| pFL17 | 64 | 250 | R | R | S | S | R |
| pFL2 | 100 | 500 | R | R | S | S | R |
| pFL3 | 80 | 500 | R | R | S | S | R |
| pFL4 | 32 | 4 | R | S | S | S | R |
| pFL5 | 32 | 4 | R | MS | S | S | R |
| pFL8 | 32 | 15 | R | R | S | MS | R |
| pJV.8r | 32 | 100 | MS | S | S | MS | R |
| pJV.9r | 100 | 250 | R | MS | S | R | R |
| pJ.1 | 64 | 50 | S | MS | S | S | R |
| pJ3.2 | 64 | 15 | R | R | S | S | R |
| pJ4.1 | 64 | 50 | R | R | S | S | R |
| pJ5.1 | 32 | 100 | R | R | S | S | R |
| pJ5.2 | 64 | 250 | R | S | S | S | R |
| pJ4.2 | 100 | 250 | R | R | S | S | R |
| pJ6.3 | 80 | 100 | R | R | S | S | R |
| p1.12 | 80 | 12 | R | R | S | S | MS |
| p10.3 | 64 | 250 | R | R | S | S | R |
| p2.2 | 80 | 15 | R | R | S | S | R |
| p4.1 | 32 | 12 | R | R | S | S | R |
| p4.8 | 64 | 15 | R | R | S | S | R |
| p9.7 | 32 | 15 | R | R | S | MS | R |
| p9.9 | 32 | 15 | R | R | S | S | R |
| p4.3 | 64 | 4 | R | S | S | S | MS |
| pAZ.24 | 64 | 4 | R | MS | S | S | R |
| pAZ.25 | 32 | 2 | R | MS | S | S | R |
| pAZ.26 | 64 | 15 | R | MS | S | S | R |
| p27Ch | 32 | 15 | R | R | S | S | R |
| p29Ch | 32 | 15 | R | R | S | S | R |
| p42Ch | 64 | 15 | R | R | S | MS | R |
| pH7.13 | 64 | 15 | R | S | S | S | R |
| pH7.17 | 100 | 2 | R | S | S | S | R |
| pH6.6 | 32 | 4 | R | MS | S | S | R |
| pH7.10 | 80 | 6 | R | S | S | S | R |
| pHm27 | 100 | 15 | R | R | S | S | R |
| pH7.14 | 100 | 2 | R | S | S | S | R |
| pHm28 | 80 | 15 | R | R | S | S | R |
| pH7.18 | 100 | 4 | R | S | S | S | R |
| pH7.9 | 80 | 4 | R | S | S | S | R |
| pH6.2 | 32 | 2 | R | S | S | S | R |
| pH6.5 | 32 | 10 | R | S | S | S | R |
| p22 | 64 | 250 | MS | S | S | S | R |
| p23 | 64 | 250 | R | S | S | S | R |
| p24 | 32 | 500 | R | S | S | S | R |
| p26 | 64 | 250 | MS | S | S | S | R |
| p27 | 64 | 250 | MS | S | S | S | R |
| p28 | 64 | 250 | R | S | S | S | R |
| pT11 | 80 | 500 | R | R | S | MS | R |
| pLO2.3 | 80 | 100 | R | R | S | S | R |
| pLO4.2 | 100 | 100 | R | R | S | S | R |
| pLO5.2 | 64 | 20 | R | R | S | S | R |
| pl1.1 | 100 | 500 | R | R | S | S | R |
| pl3.12 | 64 | 2 | MS | S | S | S | R |
| pl1.2 | 64 | 100 | R | R | S | S | R |
| pl1.3 | 64 | 100 | R | R | S | S | R |
| pJS4.2 | 64 | 20 | R | R | S | S | R |
| pJS5.5 | 64 | 15 | MS | S | S | S | S |
| pAg2 | 64 | 100 | R | R | S | S | R |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Córdova, P.; Rivera-González, J.P.; Rojas-Martínez, V.; Villarreal, P.; Zamorano, A.; Fiore, N.; San Martín, D.; Vera, F.; Gálvez, E.; Romero, J.; et al. Antimicrobial Multiresistant Phenotypes of Genetically Diverse Pseudomonas spp. Isolates Associated with Tomato Plants in Chilean Orchards. Horticulturae 2022, 8, 750. https://doi.org/10.3390/horticulturae8080750
Córdova P, Rivera-González JP, Rojas-Martínez V, Villarreal P, Zamorano A, Fiore N, San Martín D, Vera F, Gálvez E, Romero J, et al. Antimicrobial Multiresistant Phenotypes of Genetically Diverse Pseudomonas spp. Isolates Associated with Tomato Plants in Chilean Orchards. Horticulturae. 2022; 8(8):750. https://doi.org/10.3390/horticulturae8080750
Chicago/Turabian StyleCórdova, Pamela, Juan Pablo Rivera-González, Victoria Rojas-Martínez, Pablo Villarreal, Alan Zamorano, Nicola Fiore, Daniel San Martín, Francisca Vera, Eduardo Gálvez, Jaime Romero, and et al. 2022. "Antimicrobial Multiresistant Phenotypes of Genetically Diverse Pseudomonas spp. Isolates Associated with Tomato Plants in Chilean Orchards" Horticulturae 8, no. 8: 750. https://doi.org/10.3390/horticulturae8080750
APA StyleCórdova, P., Rivera-González, J. P., Rojas-Martínez, V., Villarreal, P., Zamorano, A., Fiore, N., San Martín, D., Vera, F., Gálvez, E., Romero, J., Barrueto, J., Ilabaca-Díaz, C., & Higuera, G. (2022). Antimicrobial Multiresistant Phenotypes of Genetically Diverse Pseudomonas spp. Isolates Associated with Tomato Plants in Chilean Orchards. Horticulturae, 8(8), 750. https://doi.org/10.3390/horticulturae8080750






