Abstract
Clubroot is a devastating disease caused by the protist Plasmodiophora brassicae Woronin. After root hair colonization, the clubroot pathogen induces clubs that block water uptake, leading to dehydration and death. The study of the severity of plant diseases is very important. It allows us to characterize the level of resistance of plant germplasm and to classify the virulence of pathogen strains or isolates. Lately, the use of learning machines and automatization has expanded to plant pathology. Fast, reliable and unbiased methods are always necessary, and with clubroot disease indexing this is not different. From this perspective, we discuss why this is the case and how we could achieve this long overdue task for clubroot disease.
1. Clubroot, an Introduction
In recent years, clubroot has become one of the most devastating diseases affecting the family Brassicaceae worldwide. This disease is caused by the soil-borne obligate parasite Plasmodiophora brassicae [1]. The susceptible plant hosts develop clubs on their roots as a result of P. brassicae infection, leading to dehydration and death of the infected plant [1]. Clubroot disease usually leads to approximately 10–15% yield losses on a global scale and could exceed 30% under disease-inducing environmental conditions [2,3]. Control of clubroot disease has proven to be very difficult to achieve, mainly because the resting spores released into the soil by the pathogen are resistant to most of the fungicides available on the market, and the fact that they can survive on the soil for many years [1]. To this day, the best alternative is the use of resistant genotypes, but after one or two years of using certain clubroot-resistant crops, resistance has been broken by different P. brassicae pathotypes [4]. A very important step to identify which crop variety is the best for each field is the determination of the virulence of the different isolates in susceptible and clubroot-resistant germplasm. This relies on the determination of disease index (DI) through a very laborious observational and subjective process [4,5,6,7,8,9]. Disease indexing would benefit from digitalization, which not only would make it easier and faster, but also reproducible across different laboratories. From this perspective, we discuss why this is the right time for the digitalization of clubroot disease indexing and how we could be able to achieve it successfully.
2. Clubroot Disease Index and Plasmodiophora brassicae Pathotyping
The management of clubroot disease relies on the use of clubroot-resistant (CR) varieties. However, the genetic resistance to clubroot is highly vulnerable to pathotype shifts. Currently, the use of host differential systems along with population genetics provides insights into the pathogen identity [4]. The most commonly employed differentials to identify P. brassicae pathotypes are Somé [5], Williams [6], European Clubroot Differential (ECD) [7] and the Canadian Clubroot Differential (CCD) [8]. To date, more than 30 P. brassicae pathotypes have been identified in Canada using the CCD [8]. Although this differential system has been effective for detecting predominant pathotypes of P. brassicae in Canadian fields and to study the virulence of single spore isolates (SSI), it does not discriminate between virulent and avirulent strains of P. brassicae against the resistance in CR varieties or cultivars. This has been addressed by the novel differential system SCD (Sinitic Clubroot Differential), which theoretically has the potential to detect around 250 pathotypes and has been successful in different isolates from Brassica crops in China and Korea [9].
Disease indexing is a key part of the classification of P. brassicae pathotypes, allowing us to evaluate the virulence of the pathogen. Usually, after 35–45 days post inoculation of the pathogen the disease symptoms are assessed. The commonly accepted practice is ranking the root for the severity of the disease by the human eye. The scoring system for the galls in the roots follows a score of 0 to 3, where 0—no symptoms, 1—few small clubs on lateral roots, 2—small clubs on the main root and larger clubs on lateral roots and 3—large galls on both main root and lateral roots.
Although ranking the infected roots via visual estimation by trained personnel is the current way of calculating the disease index of clubroot, it is inefficient in terms of labor and cost requirements and could be subject to human biases as well. In addition, the assessments can be influenced by temporal variations as well as those acquired from different assessors. Therefore, the foregoing drawbacks demand an objective visualization technique with the highest accuracy and reproducibility.
3. Moving to the Digital World—Which Are the Best Alternatives?
Lately, digital imaging systems are being adopted across many scientific disciplines for rapid pathotyping and disease indexing in plant science [10]. The digitalization of root morphology and architecture is often achieved by obtaining digital root images followed by analyses using software such as WinRhizo, ROOTEDGE, RootSystemAnalyser, GLO-RIA, RootGraph, RhizoScan and others [11]. These software are broadly grouped as semi-automated construction, fully automated reconstruction and fully automated phene construction, depending on the architecture of the roots. A comprehensive description of the foregoing software packages can be found in the following online resource (www.plant-image-analysis.org) (accessed on 1 July 2021). These software aid in assessing the number of roots, number of root tips, depth, width, depth–to-width ratio, areas, angles and other topological features [12,13,14].
Selecting the appropriate growing conditions of the sample population is essential in selecting the suitable digital software, because obtaining the right images for analysis is a trade-off between the growth environment and its throughput. Therefore, choosing the right growth conditions is challenging as there are many methods available (Table 1). When selecting the method, care should be taken to recreate as closely as possible the same environment in which the plants are infected in the field. The P. brassicae infections are conditioned by the transition of resting spores into motile spores, zoospores, which are chemotactically attracted by the host roots under water-saturated conditions [1]. In fact, hydroponic systems have been already successfully applied to grow Brassica spp. and to infect the plants with the clubroot pathogen resting spores [15], but several modifications should be performed in order to evaluate the infected roots with any of the software mentioned above.

Table 1.
Plant growth conditions and pros/cons for further root analyses.
4. Conclusions
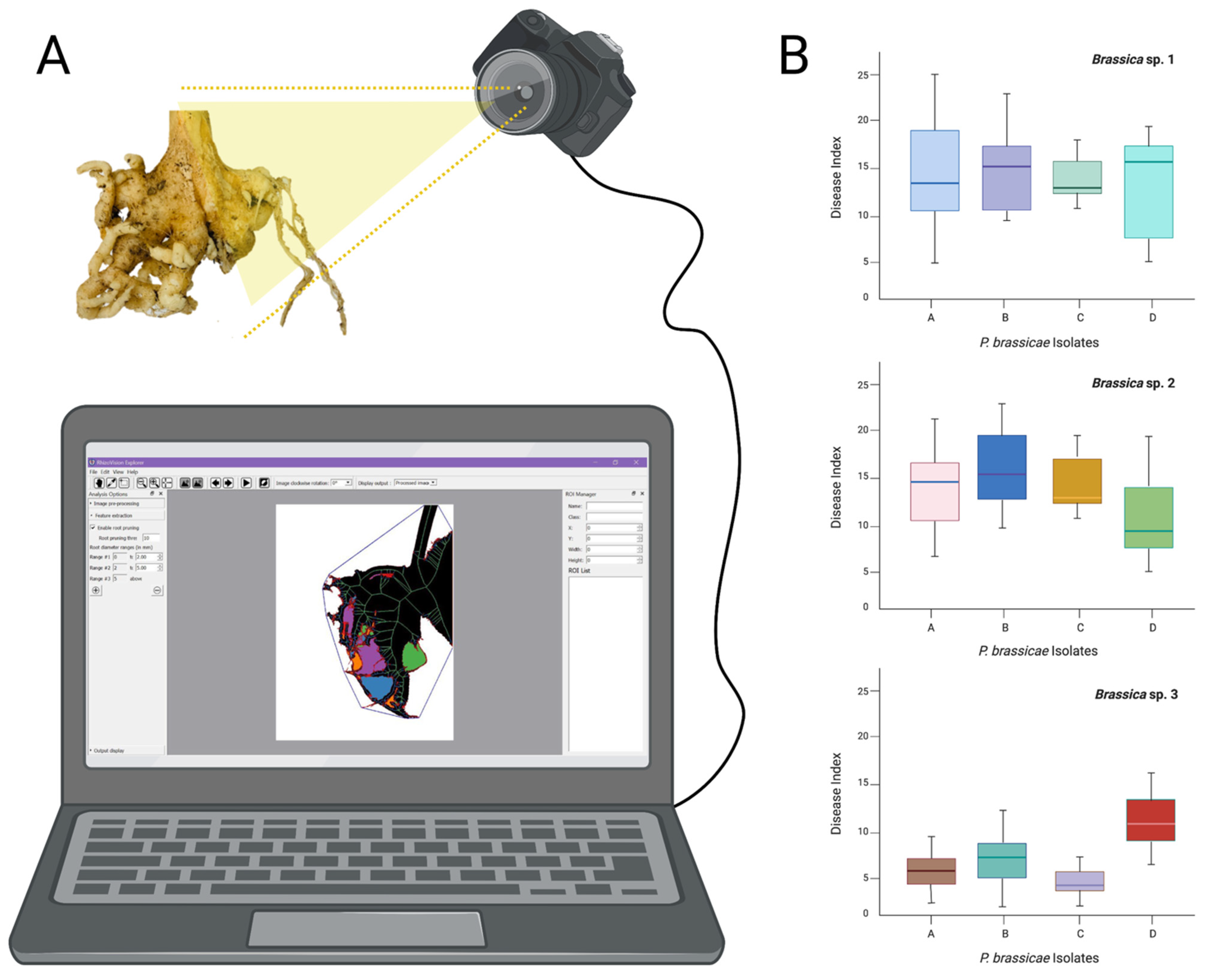
Climate change is impacting clubroot distribution and new geographic areas are becoming affected by the disease, along with new and uncharacterized P. brassicae pathotypes with increased virulence towards resistant canola cultivars [21]. There is a long way to go before fully automatized clubroot disease indexing (Figure 1), but we believe that it is and will be, in the near future, extremely necessary.
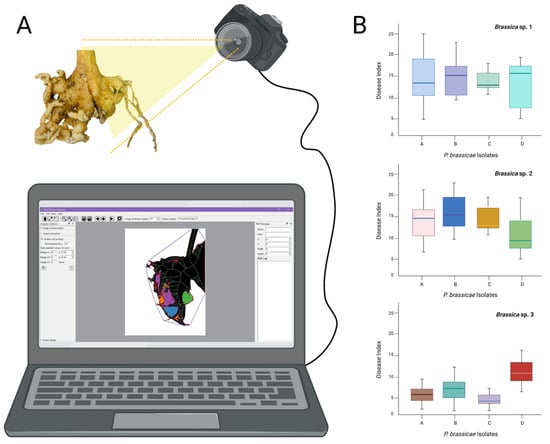
Figure 1.
Schematic representation of root galls digitalization. (A), analysis of roots using a suitable software and disease index for P. brassicae isolates infecting different canola genotypes. (B), the graphics presented are hypothetical, made with three hypothetical Brassica spp. (1 to 3) and four hypothetical clubroot pathogen isolates (A to D), used to represent how we could present the clubroot disease index calculated by the software.
Author Contributions
Conceptualization, R.S. and E.P.-L.; writing—original draft preparation, R.S. and E.P.-L.; writing—review and editing, R.S. and E.P.-L.; supervision, E.P.-L.; funding acquisition, E.P.-L. All authors have read and agreed to the published version of the manuscript.
Funding
This research was funded by Canola Council of Canada and Western Grains Research Foundation, grant number 2021.04.
Institutional Review Board Statement
Not applicable.
Informed Consent Statement
Not applicable.
Conflicts of Interest
The authors declare no conflict of interest.
References
- Dixon, G.R. The biology of Plasmodiophora brassicae Wor.–A review of recent advances. Acta Hortic. 2006, 706, 271–282. [Google Scholar] [CrossRef]
- Dixon, G.R. The occurrence and economic impact of Plasmodiophora brassicae and clubroot disease. J. Plant Growth Regul. 2009, 28, 194–202. [Google Scholar] [CrossRef]
- Strelkov, S.E.; Hwang, S.F.; Manolii, V.; Turnbull, G.D.; Fredua-Agyeman, R.; Hollman, K.; Kaus, S. Characterization of clubroot (Plasmodiophora brassicae) from canola (Brassica napus) in the Peace Country of Alberta, Canada. Can. J. Plant Pathol. 2020, 43, 155–161. [Google Scholar] [CrossRef]
- Strelkov, S.E.; Hwang, S.F.; Howard, R.J.; Hartman, M.; Turkington, T.K. Progress towards the sustainable management of clubroot [Plasmodiophora brassicae] of canola in the Canadian prairies. Prairie Soils Crop. 2011, 4, 114–121. [Google Scholar] [CrossRef]
- Somé, A.; Manzanares, M.J.; Laurens, F.; Baron, F.; Thomas, G.; Rouxel, F. Variation for virulence on Brassica napus L. amongst Plasmodiophora brassicae collections from France and derived single-spore isolates. Plant Pathol. 1996, 45, 432–439. [Google Scholar] [CrossRef]
- Williams, P.H. A system for the determination of races of Plasmodiophora brassicae that infect cabbage and rutabaga. Phytopathology 1966, 56, 624–626. [Google Scholar]
- Buczacki, S.T.; Toxopeus, H.; Mattusch, P.; Johnston, T.D.; Dixon, G.R.; Hobolth, L.A. Study of physiologic specialization in Plasmodiophora brassicae: Proposals for attempted rationalization through an international approach. Trans. Br. Mycol. Soc. 1975, 65, 295–303. [Google Scholar] [CrossRef]
- Strelkov, S.E.; Hwang, S.F.; Manolii, V.P.; Cao, T.; Fredua-Agyeman, R.; Harding, M.W.; Peng, G.; Gossen, B.D.; Mcdonald, M.R.; Feindel, D. Virulence and pathotype classification of Plasmodiophora brassicae populations collected from clubroot resistant canola (Brassica napus) in Canada. Can. J. Plant Pathol. 2018, 40, 284–298. [Google Scholar] [CrossRef]
- Pang, W.; Liang, Y.; Zhan, Z.; Li, X.; Piao, Z. Development of a Sinitic Clubroot Differential Set for the Pathotype Classification of Plasmodiophora brassicae. Front. Plant Sci. 2020, 11, 1360. [Google Scholar] [CrossRef]
- Griffiths, M. A 3D Print Repository for Plant Phenomics. Plant Phenomics 2020. [Google Scholar] [CrossRef]
- Lobet, G.; Draye, X. Novel scanning procedure enabling the vectorization of entire rhizotron-grown root systems. Plant Methods 2013, 9, 1. [Google Scholar] [CrossRef] [Green Version]
- Himmelbauer, M.L. Estimating length, average diameter and surface area of roots using two different image analyses systems. Plant Soil 2004, 260, 111–120. [Google Scholar] [CrossRef]
- Mattupalli, C.; Seethepalli, A.; York, L.M.; Young, C.A. Digital imaging to evaluate root system architectural changes associated with soil biotic factors. Phytobiomes J. 2019, 3, 102–111. [Google Scholar] [CrossRef] [Green Version]
- Kimura, K.; Kikuchi, S.; Yamasaki, S. Accurate root length measurement by image analysis. Plant Soil 1999, 216, 117–127. [Google Scholar] [CrossRef]
- Lebreton, A.; Labbé, C.; De Ronne, M.; Xue, A.G.; Marchand, G.; Bélanger, R.R. Development of a simple hydroponic assay to study vertical and horizontal resistance of soybean and pathotypes of Phytophthora sojae. Plant Dis. 2018, 102, 114–123. [Google Scholar] [CrossRef] [Green Version]
- Wagner, R.E.; Wilkinson, H.T. An aeroponics system for investigating disease development on soybean taproots infected with Phytophthora sojae. Plant Dis. 1992, 76, 610–614. [Google Scholar] [CrossRef]
- Trachsel, S.; Kaeppler, S.M.; Brown, K.M.; Lynch, J.P. Shovelomics: High throughput phenotyping of maize (Zea mays L.) root architecture in the field. Plant Soil 2011, 341, 75–87. [Google Scholar] [CrossRef]
- Cai, G.; Vanderborght, J.; Klotzsche, A.; van der Kruk, J.; Neumann, J.; Hermes, J.N.; Vereecken, H. Construction of minirhizotron facilities for investigating root zone processes. Vadose Zone J. 2016, 15, vzj2016.05.0043. [Google Scholar] [CrossRef] [Green Version]
- Auer, S. A costum-made hydroponic culture system to study plant roots during root infection with Plasmodiophora brassicae. Protocols.io 2021. [Google Scholar] [CrossRef]
- Hund, A.; Trachsel, S.; Stamp, P. Growth of axile and lateral roots of maize: I development of a phenotying platform. Plant Soil 2009, 325, 335–349. [Google Scholar] [CrossRef]
- Botero, A.; García, C.; Gossen, B.D.; Strelkov, S.E.; Todd, C.D.; Bonham-Smith, P.C.; Pérez-López, E. Clubroot disease in Latin America: Distribution and management strategies. Plant Pathol. 2019, 68, 827–833. [Google Scholar] [CrossRef] [Green Version]
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).