High-Throughput SSR Marker Development and the Analysis of Genetic Diversity in Capsicum frutescens
Abstract
:1. Introduction
2. Materials and Methods
2.1. Plant Materials and DNA Extraction
2.2. Source of Genic Sequences, SSR Identification and Primer Design
2.3. PCR Amplification and Polyacrylamide Gel Electrophoresis
2.4. Data Statistics and Analysis Results
3. Results
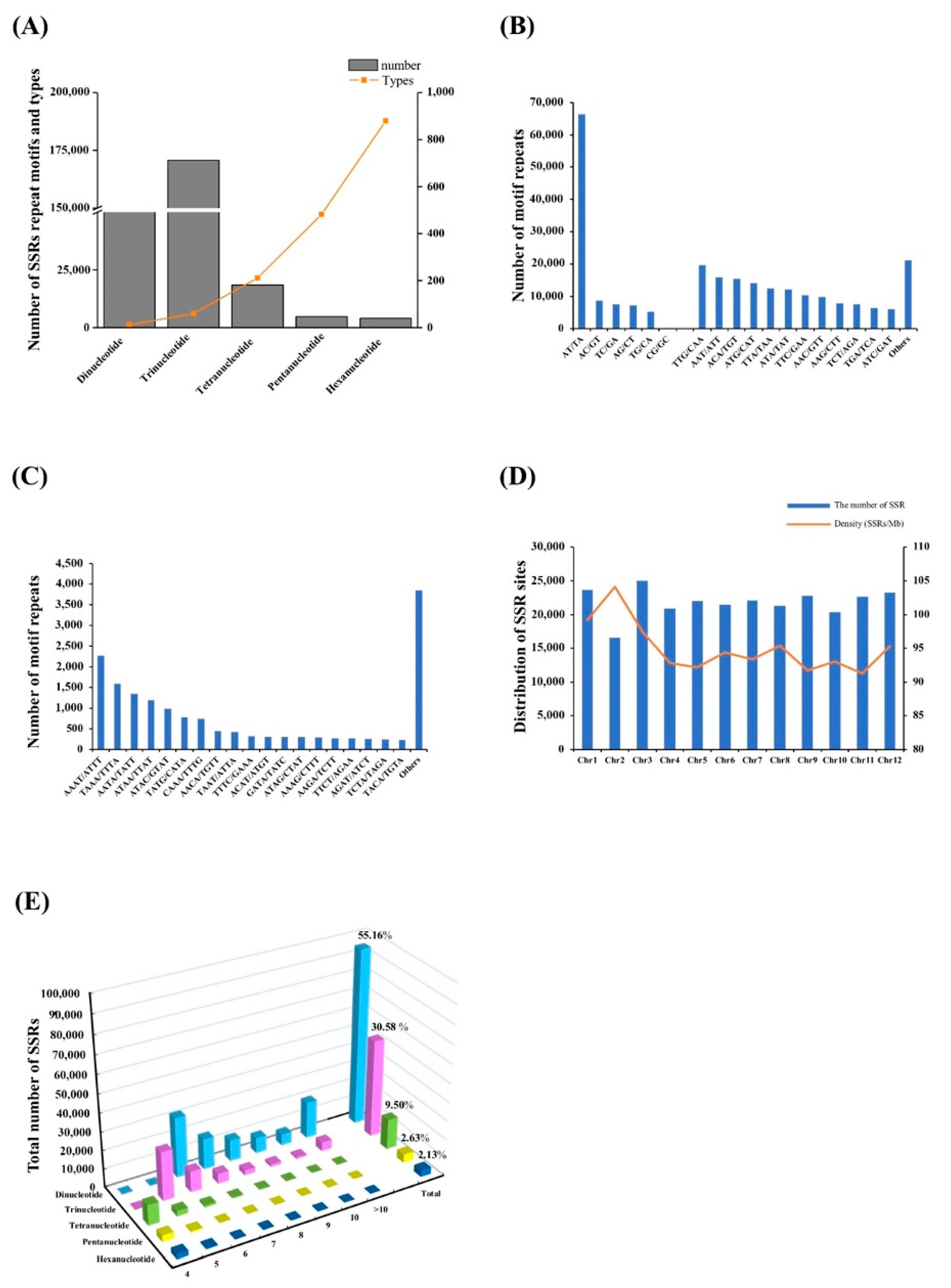
3.1. Distribution of the SSRs in the Capsicum Frutescens Genome
3.2. Analysis of the SSR Repeat Motif Types and Frequencies
3.3. Primer Design of the Pepper Plant Genomic SSR Markers
3.4. Polymorphism Analysis with SSR
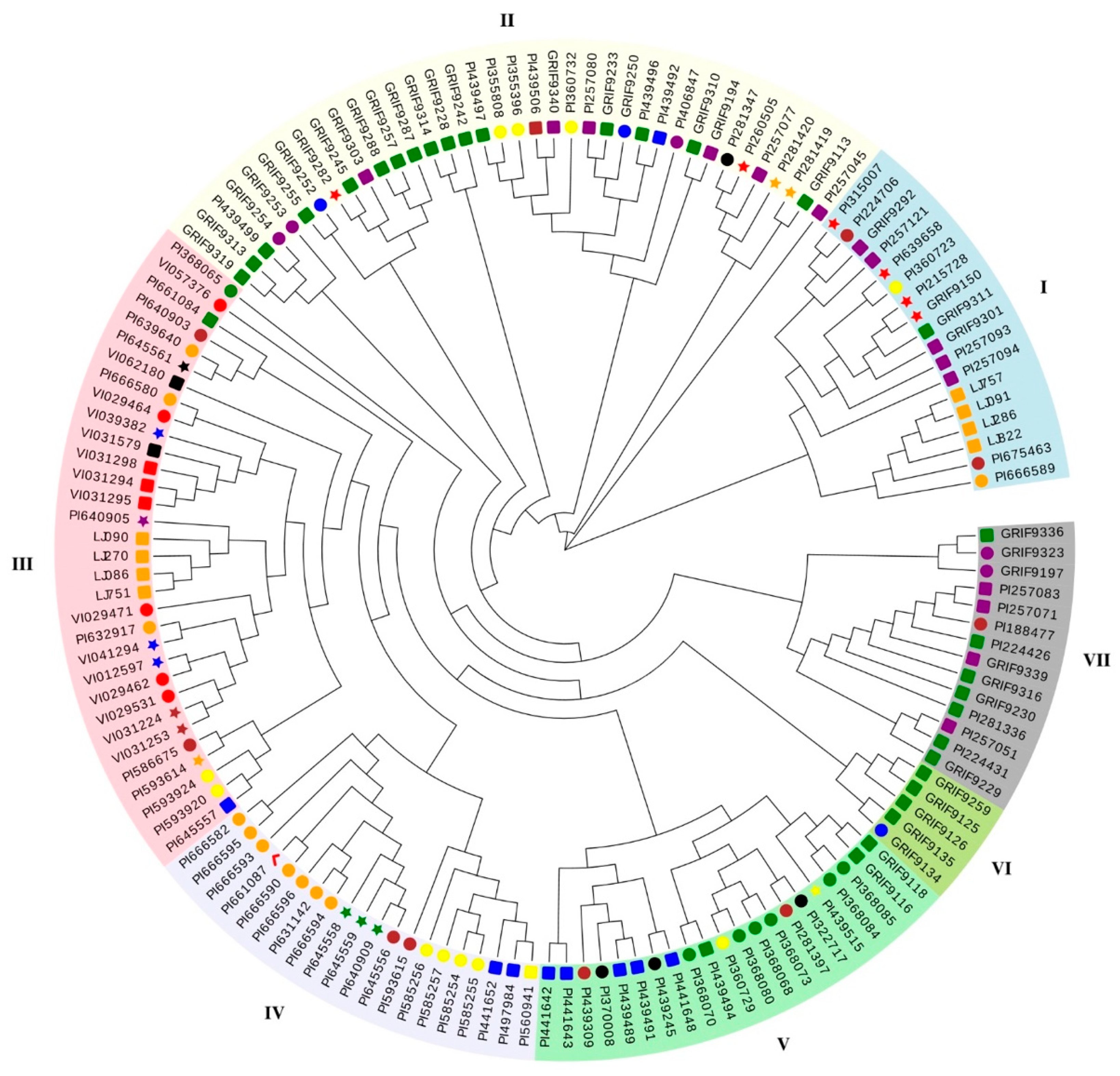
3.5. Genetic Diversity Analysis
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Moscone, E.A.; Scaldaferro, M.; Grabiele, M.; Cecchini, N.; García, Y.S.; Jarret, R.; Daviña, J.; Ducasse, D.; Barboza, G.; Ehrendorfer, F. The evolution of chili peppers (Capsicum-solanaceae): A cytogenetic perspective. Acta Hortic. 2007, 745, 137–169. [Google Scholar] [CrossRef]
- Olmstead, R.G.; Bohs, L.; Migid, H.A.; Santiago-Valentin, E.; Garcia, V.F.; Collier, S.M. A molecular phylogeny of the solanaceae. TAXON 2008, 57, 1159–1181. [Google Scholar] [CrossRef]
- Qin, C.; Yu, C.; Shen, Y.; Fang, X.; Chen, L.; Min, J.; Cheng, J.; Zhao, S.; Xu, M.; Luo, Y.; et al. Whole-genome sequencing of cultivated and wild peppers provides insights into Capsicum domestication and specialization. Proc. Natl. Acad. Sci. USA 2014, 111, 5135–5140. [Google Scholar] [CrossRef] [Green Version]
- Nee, M. Peppers, the domesticated Capsicums. Brittonia 1985, 37, 218. [Google Scholar] [CrossRef]
- Chen, X.; Zhou, K.; Zong, H.; Fang, R. Genetic diversity of Capsicum frutescens in China as revealed by SRAP and SSR markers. Acta Bot. Boreali-Occident. Sin. 2012, 32, 2201–2205. [Google Scholar]
- Zhao, H.; Cao, Y.; Zhang, Z.; Zhang, B.; Bai, R.; Zhao, Y.; Wang, L. Analysis and evaluation of genetic diversity of pepper (Capsicum spp.) Core Germplasm Resources in China. China Veg. 2018, 1, 25–34. [Google Scholar]
- Jia, H.; Wei, X.; Yao, Q.; Yuan, Y.; Wang, Z.; Jiang, J.; Yang, S.; Zhao, Y.; Wang, B.; Zhang, X. Genetic diversity analysis of Capsicum genus germplasm resources using SSR markers. Mol. Plant Breed. 2017, 15, 353–363. [Google Scholar]
- Deng, M.; Wen, J.; Zhu, H.; Zou, X. The hottest pepper variety in China. Genet. Resour. Crop Evol. 2009, 56, 605–608. [Google Scholar] [CrossRef]
- Liu, S.; Li, W.; Wu, Y.; Chen, C.; Lei, J. De Novo Transcriptome Assembly in Chili Pepper (Capsicum frutescens) to Identify Genes Involved in the Biosynthesis of Capsaicinoids. PLoS ONE 2013, 8, e48156. [Google Scholar] [CrossRef] [Green Version]
- Wei, J.; Zheng, J.; Yu, J.; Zhao, D.; Cheng, Y.; Ruan, M.; Ye, Q.; Yao, Z.; Wang, R.; Zhou, G.; et al. Production and identification of interspecific hybrids between pepper (Capsicum annuum L.) and the wild relative (Capsicum frutescens L.). J. Agric. Sci. Technol. 2019, 21, 761–769. [Google Scholar]
- Wei, J.; Li, J.; Yu, J.; Cheng, Y.; Ruan, M.; Ye, Q.; Yao, Z.; Wang, R.; Zhou, G.; Deng, M.; et al. Construction of high-density bin map and QTL mapping of horticultural traits from an interspecific cross between Capsicum annuum and Chinese wild Capsicum frutescens. Biotechnol. Biotechnol. Equip. 2020, 34, 549–561. [Google Scholar] [CrossRef]
- Tautz, D.; Renz, M. Simple sequences are ubiquitous repetitive components of eukaryotic genomes. Nucleic Acids Res. 1984, 12, 4127–4138. [Google Scholar] [CrossRef] [Green Version]
- Tóth, G.; Gáspári, Z.; Jurka, J. Microsatellites in different eukaryotic genomes: Survey and analysis. Genome Res. 2000, 10, 967–981. [Google Scholar] [CrossRef] [Green Version]
- Powell, W.; Machray, G.C.; Provan, J. Polymorphism revealed by simple sequence repeats. Trends Plant Sci. 1996, 1, 215–222. [Google Scholar] [CrossRef]
- Chen, C.; Zhou, P.; Choi, Y.A.; Huang, S.; Gmitter, F.G. Mining and characterizing microsatellites from citrus ESTs. Theor. Appl. Genet. 2006, 112, 1248–1257. [Google Scholar] [CrossRef]
- Cheng, Z.; Huang, H. SSR fingerprinting Chinese peach cultivars and landraces (Prunus persica) and analysis of their genetic relationships. Sci. Hortic. 2008, 120, 188–193. [Google Scholar] [CrossRef]
- Mccouch, S.; Teytelman, L.; Xu, Y.; Lobos, K.; Clare, K.; Walton, M.; Fu, B.; Maghirang, R.; Li, Z.; Xing, Y.; et al. Development and mapping of 2240 new SSR markers for rice (Oryza sativa L.). DNA Res. 2002, 9, 199–207. [Google Scholar] [CrossRef]
- Liu, S.; Li, W.; Long, D.; Hu, C.; Zhang, J. Development and characterization of genomic and expressed SSRs in citrus by genome-wide analysis. PLoS ONE 2013, 8, e75149. [Google Scholar] [CrossRef]
- Zhu, H.; Song, P.; Dal-Hoe, K.; Guo, L.; Li, Y.; Sun, S.; Weng, Y.; Yang, L. Genome wide characterization of simple sequence repeats in watermelon genome and their application in comparative mapping and genetic diversity analysis. BMC Genom. 2016, 17, 557–583. [Google Scholar] [CrossRef] [Green Version]
- Tan, L.; Wang, L.; Wei, K.; Zhang, C.; Wu, L.; Qi, G.; Cheng, H.; Zhang, Q.; Cui, Q.; Liang, J. Floral transcriptome sequencing for SSR marker development and linkage map construction in the tea plant (Camellia sinensis). PLoS ONE 2013, 8, e81611. [Google Scholar] [CrossRef] [Green Version]
- Bali, S.; Mamgain, A.; Raina, S.N.; Yadava, S.K.; Bhat, V.; Das, S.; Pradhan, A.K.; Goel, S. Construction of a genetic linkage map and mapping of drought tolerance trait in Indian beveragial tea. Mol. Breed. 2015, 35, 1–20. [Google Scholar] [CrossRef]
- Li, L.; He, W.; Ma, L.; Liu, P.; Xu, H.; Xu, J.; Zheng, X. Construction Chinese cabbage (Brassica rapa L.) core collection and its EST-SSR finger print database by EST-SSR molecular markers. Genom. Appl. Biol. 2009, 28, 76–88. [Google Scholar]
- Sarao, N.K.; Vikal, Y.; Singh, K.; Joshi, M.A.; Sharma, R.C. SSR marker-based DNA fingerprinting and cultivar identification of rice (Oryza sativa L.) in Punjab state of India. Plant Genet. Resour. 2009, 8, 42–44. [Google Scholar] [CrossRef]
- Tan, L.; Peng, M.; Xu, L.; Wang, L.; Chen, S.; Zou, Y.; Qi, G.; Cheng, H. Fingerprinting 128 Chinese clonal tea cultivars using SSR markers provides new insights into their pedigree relationships. Tree Genet. Genomes 2015, 11, 1–12. [Google Scholar] [CrossRef]
- Liu, S.; Liu, H.; Wu, A.; Hou, Y.; An, Y.; Wei, C. Construction of fingerprinting for tea plant (Camellia sinensis) accessions using new genomic SSR markers. Mol. Breed. 2017, 37, 1–14. [Google Scholar] [CrossRef]
- Ercisli, S.; Ipek, A.; Barut, E. SSR marker-based DNA fingerprinting and cultivar identification of olives (Olea europaea). Biochem. Genet. 2011, 49, 555–561. [Google Scholar] [CrossRef]
- Hameed, U.; Pan, Y.B.; Muhammad, K.; Afghan, S.; Iqbal, J. Use of simple sequence repeat markers for DNA fingerprinting and diversity analysis of sugarcane (Saccharum spp.) cultivars resistant and susceptible to red rot. Genet. Mol. Res. 2012, 11, 1195–1204. [Google Scholar] [CrossRef] [PubMed]
- Lanteri, S.; Portis, E.; Acquadro, A.; Mauro, R.; Mauromicale, G. Morphology and SSR fingerprinting of newly developed Cynara cardunculus genotypes exploitable as ornamentals. Euphytica 2012, 184, 311–321. [Google Scholar] [CrossRef]
- Zhebentyayeva, T.; Reighard, G.; Gorina, V.; Abbott, A. Simple sequence repeat (SSR) analysis for assessment of genetic variability in apricot germplasm. Theor. Appl. Genet. 2003, 106, 435–444. [Google Scholar] [CrossRef]
- Grover, A.; Sharma, P.C. Development and use of molecular markers: Past and present. Crit. Rev. Biotechnol. 2016, 36, 290–302. [Google Scholar] [CrossRef]
- Gu, X.; Cao, Y.; Zhang, Z.; Zhang, B.; Zhao, H.; Zhang, X.; Wang, H.; Li, X.; Wang, L. Genetic diversity and population structure analysis of Capsicum germplasm accessions. J. Integr. Agric. 2019, 18, 1312–1320. [Google Scholar] [CrossRef]
- Zhang, X.; Zhang, Z.; Gu, X.; Mao, S.; Li, X.; Chadoeuf, J.; Palloix, A.; Wang, L.; Zhang, B. Genetic diversity of pepper (Capsicum spp.) germplasm resources in China reflects selection for cultivar types and spatial distribution. J. Integr. Agric. 2016, 15, 1991–2001. [Google Scholar] [CrossRef]
- Li, Y.; Zhao, H.; Wang, Y.; Jiang, J.; Meng, X.; Wei, X.; Li, J. Genetic diversity analysis of 169 accessions of Capsicum. J. Henan Agric. Sci. 2018, 47, 91–97. [Google Scholar]
- Murray, M.G.; Thompson, W.F. Rapid isolation of high molecular weight plant DNA. Nucleic Acids Res. 1980, 8, 4321–4325. [Google Scholar] [CrossRef] [Green Version]
- Thiel, T.; Michalek, W.; Varshney, R.; Graner, A. Exploiting EST databases for the development and characterization of gene-derived SSR-markers in barley (Hordeum vulgare L.). Theor. Appl. Genet. 2003, 106, 411–422. [Google Scholar] [CrossRef]
- Yeh, F.; Boyle, T.B. Population genetic analysis of co-dominant and dominant markers and quantitative traits. Belg. J. Bot. 1997, 129, 157. [Google Scholar]
- Liu, K.; Muse, S.V. PowerMarker: An integrated analysis environment for genetic marker analysis. Bioinformatics 2005, 21, 2128–2129. [Google Scholar] [CrossRef] [Green Version]
- Nei, M. Estimation of average heterozygosity and genetic distance from a small number of individuals. Genetics 1978, 89, 583–590. [Google Scholar] [CrossRef]
- Kumar, S.; Stecher, G.; Tamura, K. MEGA7: Molecular evolutionary genetics analysis version 7.0 for bigger datasets. Mol. Biol. Evol. 2016, 33, 1870–1874. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Zhao, D.; Yang, J.; Yang, S.; Kenji, K.; Luo, J. Genetic diversity and domestication origin of tea plant Camellia taliensis (Theaceae) as revealed by microsatellite markers. BMC Plant Biol. 2014, 14, 14. [Google Scholar] [CrossRef] [Green Version]
- Göl, Ş.; Göktay, M.; Allmer, J.; Doğanlar, S.; Frary, A. Newly developed SSR markers reveal genetic diversity and geographical clustering in spinach (Spinacia oleracea). Mol. Genet. Genom. 2017, 292, 847–855. [Google Scholar] [CrossRef] [PubMed]
- Wu, J.; Cheng, F.; Cai, C.; Zhong, Y.; Jie, X. Association mapping for floral traits in cultivated Paeonia rockii based on SSR markers. Mol. Genet. Genom. 2017, 292, 187–200. [Google Scholar] [CrossRef] [PubMed]
- Wang, D.; Cao, L.; Gao, J. Data mining of simple sequence repeats in Codonopsis pilosula transcriptome. Chin. Tradit. Herb. Drugs 2014, 45, 2390–2394. [Google Scholar]
- Yu, Y.; Tian, D.; Pan, X.; Jin, L.; Ge, Y. Mining and developing SSR molecular markers based on transcriptome sequences of Anthurium. Mol. Plant Breed. 2015, 13, 1349–1354. [Google Scholar]
- Liu, S.; An, Y.; Li, F.; Li, S.; Liu, L.; Zhou, Q.; Zhao, S.; Wei, C. Genome-wide identification of simple sequence repeats and development of polymorphic SSR markers for genetic studies in tea plant (Camellia sinensis). Mol. Breed. 2018, 38, 1–13. [Google Scholar] [CrossRef]
- Mun, J.; Kim, D.; Choi, H.; Gish, J.; Debéllé, F.; Mudge, J.; Denny, R.; Endré, G.; Saurat, O.; Dudez, A.; et al. Distribution of microsatellites in the genome of Medicago truncatula: A resource of genetic markers that integrate genetic and physical maps. Genetics 2006, 172, 2541–2555. [Google Scholar] [CrossRef] [Green Version]
- Jaillon, O.; Aury, J.; Noel, B.; Policriti, A.; Clépet, C.; Casagrande, A.; Choisne, N.; Aubourg, S.; Vitulo, N.; Jubin, C.; et al. The grapevine genome sequence suggests ancestral hexaploidization in major angiosperm phyla. Nature 2007, 449, 463–467. [Google Scholar]
- Cavagnaro, P.F.; Senalik, D.A.; Yang, L.M.; Simon, P.W.; Harkins, T.T.; Kodira, C.D.; Huang, S.W.; Weng, Y.Q. Genome-wide characterization of simple sequence repeats in cucumber (Cucumis sativus L.). BMC Genom. 2010, 11, 569–587. [Google Scholar] [CrossRef] [Green Version]
- Dong, Q.; Wang, X.; Zhao, M.; Song, C.; Ge, A.; Wang, J. Development of EST-derived SSR markers and their application in strawberry genetic diversity analysis. Sci. Agric. Sin. 2011, 44, 3603–3612. [Google Scholar]
- Xu, J.; Liu, L.; Xu, Y.; Chen, C.; Rong, T.; Ali, F.; Zhou, S.; Wu, F.; Liu, Y.; Wang, J.; et al. Development and characterization of simple sequence repeat markers providing genome-wide coverage and high resolution in maize. DNA Res. 2013, 20, 497–509. [Google Scholar] [CrossRef] [Green Version]
- Gong, L.; Cheng, Y.; Gan, X.; Chen, Y.; Nie, F.; Zhang, L.; Shi, L.; Song, Y.; Guo, Z.; Wang, F. Mining EST-SSRs based on potato transcriptome and analyzing their polymorphism. Mol. Plant Breed. 2015, 13, 1535–1544. [Google Scholar]
- Fang, Z.; Ye, X.; Zhou, D.; Jiang, C.; Pan, S. Analysis on SSR information in ‘Furongli’ plum transcriptome and development of molecular markers in Prunus salicina Lindl. J. Fruit Sci. 2016, 33, 416–424. [Google Scholar]
- Zhang, Y.; Wei, X.; Wang, X.; Pan, S.; Ye, X. SSR loci in transcriptome of Moringa oleifera lam. Fujian J. Agric. Sci. 2017, 32, 955–958. [Google Scholar]
- Zhang, Z.; Deng, Y.; Tan, J.; Hu, S.; Yu, J.; Xue, Q. A Genome-wide microsatellite polymorphism database for the indica and japonica Rice. DNA Res. 2007, 14, 37–45. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Li, M.; Zhang, S.; Deng, P.; Hou, X.; Wang, J. Analysis on SSR information in transcriptome of onion and the polymorphism. Acta Hortic. Sin. 2015, 42, 1103–1111. [Google Scholar]
- Yue, C.; Chen, C.; Guo, F.; Li, H.; Sun, H.; Pei, D.; Ma, X.; Chen, F.; Yang, H.; Li, Q. Data mining of simple sequence repeats in transcriptome sequences of mongolia medicinal plant artemisia frigida willd. J. Agric. Sci. Technol. 2016, 18, 31–43. [Google Scholar]
- Liu, Z.; Yang, Y.; Sun, J.; Liu, Z.; Cao, Z. Purity identification of Rela No.2 pepper and genetic diversity analysis of excellent pepper inbred lines. Chin. J. Trop. Crop. 2014, 35, 847–853. [Google Scholar]
- Rivera, A.; Monteagudo, A.B.; Igartua, E.; Taboada, A.; García-Ulloa, A.; Pomar, F.; Riveiro-Leira, M.; Silvar, C. Assessing genetic and phenotypic diversity in pepper (Capsicum annuum L.) landraces from North-West Spain. Sci. Hortic. 2016, 203, 1–11. [Google Scholar] [CrossRef] [Green Version]
- Yuan, X.; Zhou, K.; Wu, Y.; Fang, R.; Chen, X. Genetic Diversity and population structure analysis of pepper core collections. Mol. Plant Breed. 2019, 17, 3090–3104. [Google Scholar]
- Uncu, A.T. Genome-wide identification of simple sequence repeat (SSR) markers in Capsicum chinense Jacq. with high potential for use in pepper introgression breeding. Biologia 2019, 74, 119–126. [Google Scholar] [CrossRef]
- Yao, M.; Ma, C.; Qiao, T.; Jin, J.; Chen, L. Diversity distribution and population structure of tea germplasms in China revealed by EST-SSR markers. Tree Genet. Genomes 2012, 8, 205–220. [Google Scholar] [CrossRef]
- Nicolaï, M.; Cantet, M.; Lefebvre, V.; Sage-Palloix, A.M.; Palloix, A. Genotyping a large collection of pepper (Capsicum spp.) with SSR loci brings new evidence for the wild origin of cultivated C. annuum and the structuring of genetic diversity by human selection of cultivar types. Genet. Resour. Crop Evol. 2013, 60, 2375–2390. [Google Scholar] [CrossRef]
- Baba, V.Y.; Rocha, K.; Gomes, G.P.; Ruas, C.F.; Ruas, P.; Rodrigues, R.; Gonçalves, L. Genetic diversity of Capsicum chinense accessions based on fruit morphological characterization and AFLP markers. Genet. Resour. Crop Evol. 2015, 63, 1371–1381. [Google Scholar] [CrossRef]
- Lee, H.; Ro, N.; Jeong, H.; Kwon, J.; Jo, J.; Ha, Y.; Jung, A.; Han, J.; Venkatesh, J.; Kang, B. Genetic diversity and population structure analysis to construct a core collection from a large Capsicum germplasm. BMC Genet. 2016, 17, 142–154. [Google Scholar] [CrossRef] [Green Version]
- Luo, Y.; Li, J.; Li, M. Analysis of genetic diversity of Capsicum germplasm resources by using SSR markers. Biotechnol. Bull. 2006, S1, 337–341. [Google Scholar]

| No. | Primers | Forward Primer Sequence (5′-3′) | Reverse Primer Sequence (5′-3′) | Repeat Unit | Repeat Number |
|---|---|---|---|---|---|
| 1 | Chr1SSR8 | GACTATAGCAAACATGTCCCCAG | CCAGATTTTGTGGATCCTATTGA | TCA | 4 |
| 2 | Chr1SSR12 | ATTCAAAGAGGGCATGATGTAGA | TGTCAACTTAGTGGGTGGGTTAG | GGTAGG | 6 |
| 3 | Chr1SSR18 | TTAGTGTTGTCAAAATACCCGTG | CAATAAACATATCACACGTGCAAC | ATAG | 6 |
| 4 | Chr2SSR12 | GAAAGATCCGTCTAAGCAAACAA | AAATCTCATGATTTTCGGTGATG | AAT | 4 |
| 5 | Chr2SSR14 | TGAATTTCGGAGTGTTACGTAGAG | GCGAAGGTGAGTCTGTTCAATTA | TAA | 4 |
| 6 | Chr2SSR15 | AATTTGAAGAGCGTGCATAAAAA | ACCATACCATACCATGGAAACAA | ATGGT | 4 |
| 7 | Chr3SSR5 | ATAAAATAGACCCCACCCACTTC | GGGCTGCTGTACTAAAGAAGAAGA | TTC | 4 |
| 8 | Chr3SSR6 | CTCTCTAGAATGAAAAGTGCCGA | TTGTCGATTTGTTCTTCTTCCAT | CAC | 4 |
| 9 | Chr3SSR13 | ACCCTAAAAGCTATGGAGTTGCT | CCCACCATTCTTCTAAACTTTCC | AC | 6 |
| 10 | Chr4SSR11 | CCTACAAGTGAGGTCTGAGGAGA | CAGCAAGTTGGAGAAACTAATGC | ACCCT | 4 |
| 11 | Chr4SSR14 | AGTGGAAAGTGCTGTTACGATGT | GCGTGATAATTTTTCCACAAGAA | TTTA | 5 |
| 12 | Chr4SSR20 | AAAACAACACGACACACAGCTTA | TATATTTTTCTTGGGAACGAGCA | TATAT | 6 |
| 13 | Chr5SSR13 | TAAAACCTTGTCACATGTACCCC | AAATGAATAAATCCTCTGCATGG | TGCA | 4 |
| 14 | Chr5SSR14 | TAAGTTGTTCGAGAAATCAGACG | CTGTCTCGTATTGAAGGGTGTTT | CGGTGA | 4 |
| 15 | Chr5SSR16 | CACTGTGAGAGCAACTTTCTGTG | CACTATTTTCTCATGCACTTTACCA | ACGGGC | 4 |
| 16 | Chr6SSR12 | TTCGACCTCCGTATCACTATCAG | CGTACTCTATCGCTTGTTGCTTT | ACTC | 7 |
| 17 | Chr6SSR17 | TCTGAAAAATCCTCGGCTAAAGT | ACTGTCCCACCTTACATCCCTAT | AAAAG | 5 |
| 18 | Chr6SSR19 | CAGAGGCAGTTAGGTAGTAGCGA | TGCTAAACCCACCTTCAGTCTTA | GGATTC | 4 |
| 19 | Chr6SSR20 | TCTGCTTTCCCACAGTATCTCTC | TCAACAGATAAGCGTCAAGTGAA | TCACCA | 4 |
| 20 | Chr7SSR2 | CTGCTTAAAAAGTTGAAGATGAGAA | TTGATACGCTAATAAAATTGTTGAAA | AAT | 4 |
| 21 | Chr7SSR15 | GCCTGGCATGTTTTGTATTTCTA | TTTGGTGACCGACAATATAAAGG | ATT | 4 |
| 22 | Chr8SSR6 | TGAGTCAAGAAAACTTGCAGAAA | TTTGAAAATAATTGAAGTTCCGC | AAAGCA | 4 |
| 23 | Chr8SSR13 | TAGATGTTGAACCCCTATTGGAA | GGTAGAGGGTAGAGTGTACGCAG | TACCAC | 4 |
| 24 | Chr8SSR15 | GGTGTCATGCGTAAGCTCATAGT | GGTGTCATGCGTAAGCTCATAGT | GAGTTG | 4 |
| 25 | Chr9SSR16 | CCCCACCGATGAATTTAGTAGA | TGATGATGTGTCATGGTGTATGA | CCCCCA | 4 |
| 26 | Chr9SSR17 | ATTCCCCATATCGAAACTTCTTC | AAATCAAGGAGACGATTGTTGAA | CTGGTG | 5 |
| 27 | Chr10SSR12 | ACGAGAGTTTGCTTTCTTTTCCT | TCAGAGGTAGAGGTATGGACTGC | AAC | 4 |
| 28 | Chr10SSR14 | TTTTTCAGGCTTTTGTGTATGTAAA | CGGAAACAACCTCTCTACTTCAG | CAA | 5 |
| 29 | Chr11SSR4 | AATTTTCAACAACAAACTCCACG | CAGCAGTGAGGATGAAAAAGTTTA | GAG | 4 |
| 30 | Chr11SSR14 | TTTTAACTTTTTGTAATTTCGTGTCA | ATCCAATTTTGTTAGGCCTATTG | ATCAAA | 4 |
| 31 | Chr11SSR16 | ACGAGTGAACACGTACATGAAAA | GGTAGGAGCTAGGAGTAGTGACG | TTA | 6 |
| 32 | Chr12SSR10 | AAAAGGGACTTGTTTTCCTATCA | AAATGGGAATGCGATTATCTAAA | AAAAAT | 4 |
| 33 | Chr12SSR19 | TCTTTCCTTCGGATTAAGTTTCC | GAGGCAAGAAATAAGAGATGCCT | CAAATC | 4 |
| Primer | Allele Size | Na | Ne | MAF | Ho | He | I | PIC | Gene Diversity |
|---|---|---|---|---|---|---|---|---|---|
| Chr1SSR8 | 134 | 2.0000 | 1.0615 | 0.9701 | 0.0000 | 0.0584 | 0.1342 | 0.0562 | 0.0579 |
| Chr1SSR12 | 294 | 3.0000 | 1.2132 | 0.9048 | 0.1837 | 0.1763 | 0.3680 | 0.1665 | 0.1757 |
| Chr1SSR18 | 278 | 4.0000 | 2.7999 | 0.4532 | 0.0360 | 0.6452 | 1.1545 | 0.5758 | 0.6428 |
| Chr2SSR12 | 286 | 2.0000 | 1.1104 | 0.9476 | 0.0350 | 0.0997 | 0.2057 | 0.0945 | 0.0994 |
| Chr2SSR14 | 294 | 2.0000 | 1.1764 | 0.9184 | 0.0272 | 0.1504 | 0.2827 | 0.1387 | 0.1499 |
| Chr2SSR15 | 278 | 3.0000 | 1.3743 | 0.8417 | 0.2014 | 0.2733 | 0.4998 | 0.2452 | 0.2723 |
| Chr3SSR5 | 286 | 2.0000 | 1.1259 | 0.9406 | 0.1189 | 0.1122 | 0.2254 | 0.1056 | 0.1118 |
| Chr3SSR6 | 290 | 2.0000 | 1.3729 | 0.8379 | 0.0345 | 0.2725 | 0.4431 | 0.2347 | 0.2716 |
| Chr3SSR13 | 272 | 3.0000 | 1.4543 | 0.8162 | 0.2059 | 0.3135 | 0.5784 | 0.2837 | 0.3124 |
| Chr4SSR11 | 290 | 2.0000 | 1.1012 | 0.9517 | 0.0000 | 0.0922 | 0.1934 | 0.0877 | 0.0919 |
| Chr4SSR14 | 204 | 3.0000 | 2.1276 | 0.5392 | 0.0588 | 0.5326 | 0.8242 | 0.4252 | 0.5300 |
| Chr4SSR20 | 292 | 6.0000 | 2.3599 | 0.5822 | 0.0137 | 0.5782 | 1.0946 | 0.5175 | 0.5763 |
| Chr5SSR13 | 282 | 2.0000 | 1.4302 | 0.8156 | 0.3404 | 0.3019 | 0.4780 | 0.2556 | 0.3008 |
| Chr5SSR14 | 278 | 2.0000 | 1.0516 | 0.9748 | 0.0216 | 0.0493 | 0.1176 | 0.0479 | 0.0491 |
| Chr5SSR16 | 292 | 3.0000 | 2.1992 | 0.5068 | 0.9863 | 0.5472 | 0.8578 | 0.4426 | 0.5453 |
| Chr6SSR12 | 248 | 2.0000 | 1.9541 | 0.5766 | 0.0081 | 0.4902 | 0.6814 | 0.3691 | 0.4883 |
| Chr6SSR17 | 288 | 3.0000 | 1.1656 | 0.9236 | 0.0000 | 0.1426 | 0.2931 | 0.1338 | 0.1421 |
| Chr6SSR19 | 294 | 3.0000 | 1.3276 | 0.8571 | 0.0136 | 0.2476 | 0.4375 | 0.2195 | 0.2467 |
| Chr6SSR20 | 280 | 2.0000 | 1.0894 | 0.9571 | 0.0857 | 0.0823 | 0.1769 | 0.0787 | 0.0820 |
| Chr7SSR2 | 282 | 2.0000 | 1.0288 | 0.9858 | 0.0000 | 0.0281 | 0.0744 | 0.0276 | 0.0280 |
| Chr7SSR15 | 294 | 4.0000 | 3.6226 | 0.3231 | 0.0476 | 0.7264 | 1.3273 | 0.6718 | 0.7240 |
| Chr8SSR6 | 288 | 2.0000 | 1.5732 | 0.7604 | 0.0486 | 0.3656 | 0.5506 | 0.2980 | 0.3644 |
| Chr8SSR13 | 294 | 3.0000 | 1.4308 | 0.8265 | 0.1361 | 0.3021 | 0.5775 | 0.2795 | 0.3011 |
| Chr8SSR15 | 286 | 4.0000 | 3.4282 | 0.4021 | 0.0699 | 0.7108 | 1.3023 | 0.6566 | 0.7083 |
| Chr9SSR16 | 250 | 2.0000 | 1.9678 | 0.5640 | 0.0560 | 0.4938 | 0.6849 | 0.3709 | 0.4918 |
| Chr9SSR17 | 294 | 3.0000 | 1.6031 | 0.7721 | 0.0748 | 0.3775 | 0.6873 | 0.3430 | 0.3762 |
| Chr10SSR12 | 270 | 4.0000 | 3.5014 | 0.3556 | 0.0000 | 0.7171 | 1.3058 | 0.6606 | 0.7144 |
| Chr10SSR14 | 292 | 2.0000 | 1.5026 | 0.7877 | 0.0411 | 0.3356 | 0.5170 | 0.2785 | 0.3345 |
| Chr11SSR4 | 292 | 3.0000 | 1.3060 | 0.8699 | 0.0000 | 0.2351 | 0.4723 | 0.2206 | 0.2343 |
| Chr11SSR14 | 76 | 2.0000 | 1.2321 | 0.8947 | 0.0000 | 0.1909 | 0.3365 | 0.1706 | 0.1884 |
| Chr11SSR16 | 294 | 2.0000 | 1.7306 | 0.6973 | 0.0884 | 0.4236 | 0.6132 | 0.3331 | 0.4222 |
| Chr12SSR10 | 278 | 3.0000 | 2.5179 | 0.4928 | 0.1007 | 0.6050 | 0.9921 | 0.5226 | 0.6028 |
| Chr12SSR19 | 294 | 4.0000 | 1.3399 | 0.8571 | 0.2313 | 0.2546 | 0.5140 | 0.2366 | 0.2537 |
| Mean | 271 | 2.7576 | 1.7055 | 0.7547 | 0.0989 | 0.3313 | 0.5758 | 0.2893 | 0.3300 |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Zhong, Y.; Cheng, Y.; Ruan, M.; Ye, Q.; Wang, R.; Yao, Z.; Zhou, G.; Liu, J.; Yu, J.; Wan, H. High-Throughput SSR Marker Development and the Analysis of Genetic Diversity in Capsicum frutescens. Horticulturae 2021, 7, 187. https://doi.org/10.3390/horticulturae7070187
Zhong Y, Cheng Y, Ruan M, Ye Q, Wang R, Yao Z, Zhou G, Liu J, Yu J, Wan H. High-Throughput SSR Marker Development and the Analysis of Genetic Diversity in Capsicum frutescens. Horticulturae. 2021; 7(7):187. https://doi.org/10.3390/horticulturae7070187
Chicago/Turabian StyleZhong, Yangmin, Yuan Cheng, Meiying Ruan, Qingjing Ye, Rongqing Wang, Zhuping Yao, Guozhi Zhou, Jia Liu, Jiahong Yu, and Hongjian Wan. 2021. "High-Throughput SSR Marker Development and the Analysis of Genetic Diversity in Capsicum frutescens" Horticulturae 7, no. 7: 187. https://doi.org/10.3390/horticulturae7070187
APA StyleZhong, Y., Cheng, Y., Ruan, M., Ye, Q., Wang, R., Yao, Z., Zhou, G., Liu, J., Yu, J., & Wan, H. (2021). High-Throughput SSR Marker Development and the Analysis of Genetic Diversity in Capsicum frutescens. Horticulturae, 7(7), 187. https://doi.org/10.3390/horticulturae7070187






