Changes in the Metabolite Profile during Micropropagation of Normal and Somaclonal Variants of Banana Musa AAA cv. Williams
Abstract
1. Introduction
2. Materials and Methods
2.1. Plant Material
2.2. In Vitro Establishment of Banana Plants
2.3. Phenotype Analysis
2.4. Metabolite Profiling
2.5. Data Analysis
3. Results
3.1. Phenotype Heritability
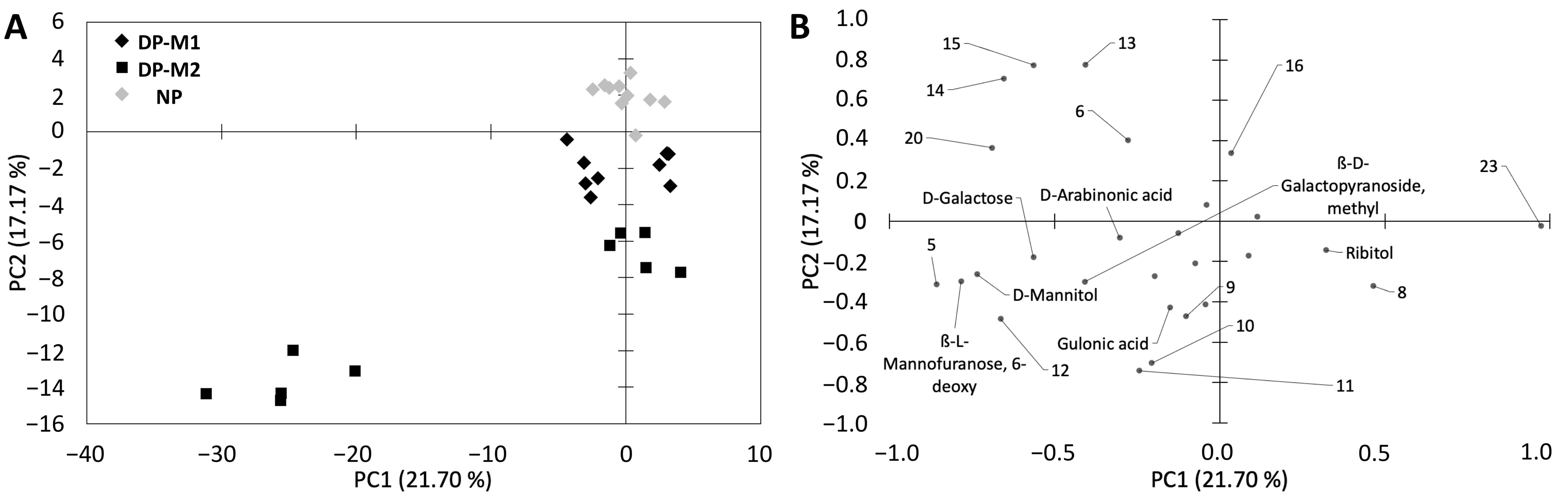
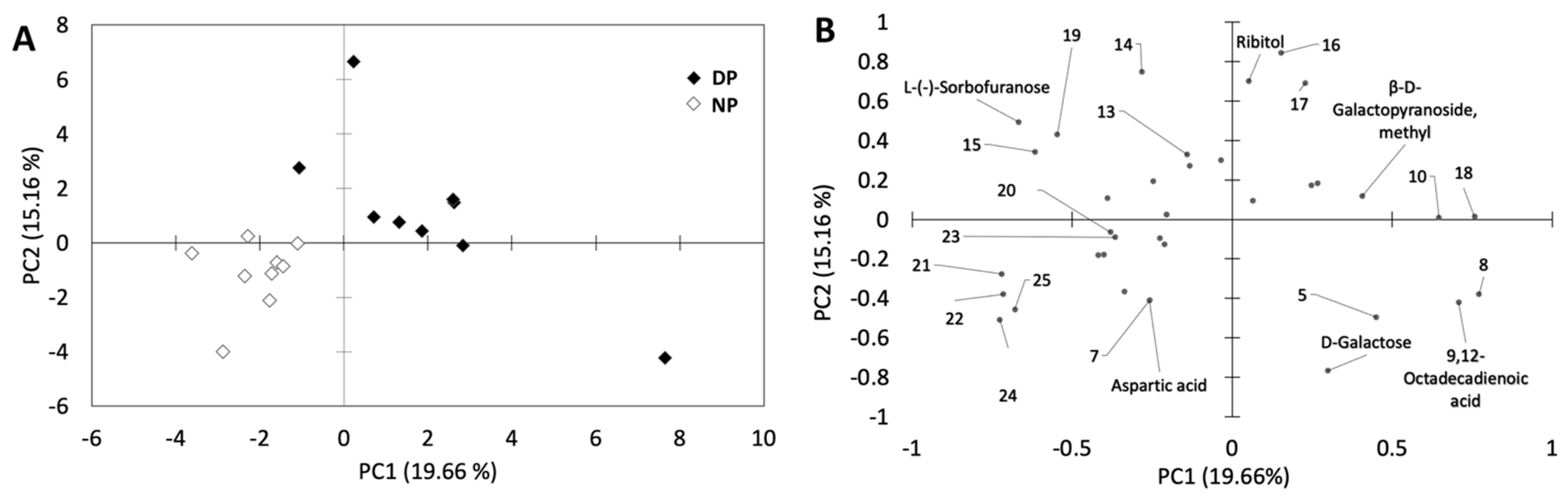
3.2. Overall Metabolite Profiling
3.3. Individual Metabolites
4. Discussion
4.1. Phenotype Analysis
4.2. Overall Metabolite Profile
4.3. Differentially Accumulated Carboxylic Acids
4.4. Differentially Accumulated Monosaccharides and Plant Cell Wall Components
4.5. Differentially Accumulated Aminated Metabolites
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Jain, S.M.; Priyadarshan, P.M. Breeding Plantation Tree Crops: Tropical Species; Springer: Berlin/Heidelberg, Germany, 2009. [Google Scholar]
- Dale, J.; James, A.; Paul, J.-Y.; Khanna, H.; Smith, M.; Peraza-Echeverria, S.; Garcia-Bastidas, F.; Kema, G.; Waterhouse, P.; Mengersen, K.; et al. Transgenic Cavendish bananas with resistance to Fusarium wilt tropical race 4. Nat. Commun. 2017, 8, 1–8. [Google Scholar] [CrossRef]
- Hamill, S. Processes, costs and traits of plants produced in tissue culture must be considered to develop effective crop production systems. Acta Hortic. 2016, 1113, 85–92. [Google Scholar] [CrossRef]
- Govindaraju, S.; Arulselvi, P.I. Effect of cytokinin combined elicitors (L-phenylalanine, salicylic acid and chitosan) on in vitro propagation, secondary metabolites and molecular characterization of medicinal herb–Coleus aromaticus Benth (L). J. Saudi Soc. Agric. Sci. 2018, 17, 435–444. [Google Scholar] [CrossRef]
- Valledor, L.; Hasbun, R.; Meijón, M.; Rodríguez, J.L.; Santamaría, E.; Viejo, M.; Berdasco, M.; Feito, I.; Fraga, M.F.; Canal, M.J.; et al. Involvement of DNA methylation in tree development and micropropagation. Plant Cell Tissue Organ Cult. (PCTOC) 2007, 91, 75–86. [Google Scholar] [CrossRef]
- Sahijram, L.; Soneji, J.R.; Bollamma, K.T. Analyzing somaclonal variation in micropropagated bananas (Musa spp.). Vitr. Cell. Dev. Biol. Anim. 2003, 39, 551–556. [Google Scholar] [CrossRef]
- Abdellatif, K.F.; Hegazy, A.E.; Aboshama, H.M.; Emara, H.A.; El-Shahed, A.A. Morphological and molecular characterization of somaclonal variations in tissue culture-derived banana plants. J. Genet. Eng. Biotechnol. 2012, 10, 47–53. [Google Scholar] [CrossRef]
- Israeli, Y.; Reuveni, O.; Lahav, E. Qualitative aspects of somaclonal variations in banana propagated by in vitro techniques. Sci. Hortic. 1991, 48, 71–88. [Google Scholar] [CrossRef]
- Jain, S.M. Tissue culture-derived variation in crop improvement. Euphytica 2001, 118, 153–166. [Google Scholar] [CrossRef]
- Bairu, M.W.; Fennell, C.W.; Van Staden, J. The effect of plant growth regulators on somaclonal variation in Cavendish banana (Musa AAA cv. ’Zelig’). Sci. Hortic. 2006, 108, 347–351. [Google Scholar] [CrossRef]
- Côte, F.X.; Sandoval-Fernández, J.A.; Marie, P.; Erik, A. Variations in micropropagated bananas and plantains: Literature survey. Fruits 1993, 48, 15–23. [Google Scholar]
- De Langhe, E.; Vrydaghs, L.; de Maret, P.; Perrier, X.; Denham, T. Why Bananas Matter: An introduction to the history of banana domestication. Ethnobot. Res. Appl. 2009, 7, 165–177. [Google Scholar] [CrossRef]
- Hřibová, E.; Čížková, J.; Christelová, P.; Taudien, S.; De Langhe, E.; Doležel, J. The ITS1-5.8S-ITS2 sequence region in the musaceae: Structure, diversity and use in molecular phylogeny. PLoS ONE 2011, 6, e17863. [Google Scholar] [CrossRef]
- Hapsari, L.; Azrianingsih, R.; Arumingtyas, E.L. Genetic variability and relationship of banana cultivars (musa l.) from East Java, Indonesia based on the internal transcribed spacer region nrdna sequences. J. Trop. Biol. Conserv. 2018, 15, 101–120. [Google Scholar]
- Cote, F.; Sandoval-Fernández, J.A.; Marie, P.; Erik, A. Phenotypic variation in micropropagated bananas and plantains. Variación fenotípica de bananos y plátanos micropropagados. CORBANA 1998, 23, 177–198. [Google Scholar]
- Sheidai, M.; Aminpoor, H.; Noormohammadi, Z.; Farahani, F. RAPD analysis of somaclonal variation in banana (Musa acuminate L.) cultivar Valery. Acta Biol. Szeged. 2008, 52, 307–311. [Google Scholar]
- Choudhary, D.; Kajla, S.; Poonia, A.K.; Brar, B.; Surekha; Duhan, J.S. Molecular assessment of genetic stability using ISSR and RAPD markers in in vitro multiplied copies of commercial banana cv. Robusta. Indian J. Biotechnol. 2015, 14, 420–424. [Google Scholar]
- Khatab, I.; Youssef, M. Micropropagation and Assessment of Genetic Stability of Musa sp. cv. Williams using RAPD and SRAP Markers. Egypt. J. Bot. 2018, 58, 371–380. [Google Scholar] [CrossRef]
- Noceda, C.; Vargas, A.; Roels, S.; Cejas, I.; Santamaría, E.; Escalona, M.; DeBergh, P.; Rodríguez, R.; Sandoval, J.; Canal, M.J.; et al. Field performance and (epi)genetic profile of plantain (Musa AAB) clone ‘CEMSA ¾’ plants micropropagated by temporary immersion systems. Sci. Hortic. 2012, 146, 65–75. [Google Scholar] [CrossRef]
- Ray, T.; Dutta, I.; Saha, P.; Das, S.; Roy, S. Genetic stability of three economically important micropropagated banana (Musa spp.) cultivars of lower Indo-Gangetic plains, as assessed by RAPD and ISSR markers. Plant Cell Tissue Organ Cult. (PCTOC) 2006, 85, 11–21. [Google Scholar] [CrossRef]
- Aremu, A.O.; Plačková, L.; Bairu, M.W.; Novák, O.; Szüčová, L.; Doležal, K.; Finnie, J.F.; Van Staden, J. Endogenous cytokinin profiles of tissue-cultured and acclimatized ‘Williams’ bananas subjected to different aromatic cytokinin treatments. Plant Sci. 2014, 214, 88–98. [Google Scholar] [CrossRef] [PubMed]
- Dhanapal, S.; Sathish, D.; Satheesh, P.M. Efficiency of Rapd, Ssr and Issr Markers in Evaluating the Genetic Fidelity for Micropropogated Musa Accuminata Plant Exposed To Coal Extracted Humic Acid and Commercially Available Products. Int. J. Agric. Sci. Res. 2014, 4, 77–86. [Google Scholar]
- Nandhakumar, N.; Kumar, K.; Sudhakar, D.; Soorianathasundaram, K. Plant regeneration, developmental pattern and genetic fidelity of somatic embryogenesis derived Musa spp. J. Genet. Eng. Biotechnol. 2018, 16, 587–598. [Google Scholar] [CrossRef] [PubMed]
- Harrison, B.R.; Wang, L.; Gajda, E.; Hoffman, E.V.; Chung, B.Y.; Pletcher, S.D.; Raftery, D.; Promislow, D.E.L. The metabolome as a link in the genotype-phenotype map for peroxide resistance in the fruit fly, Drosophila melanogaster. BMC Genom. 2020, 21, 341. [Google Scholar] [CrossRef] [PubMed]
- Cevallos-Cevallos, J.M.; Reyes-De-Corcuera, J.I.; Etxeberria, E.; Danyluk, M.D.; Rodrick, G.E. Metabolomic analysis in food science: A review. Trends Food Sci. Technol. 2009, 20, 557–566. [Google Scholar] [CrossRef]
- Cevallos-Cevallos, J.M.; García-Torres, R.; Etxeberria, E.; Reyes-De-Corcuera, J.I. GC-MS Analysis of Headspace and Liquid Extracts for Metabolomic Differentiation of Citrus Huanglongbing and Zinc Deficiency in Leaves of ‘Valencia’ Sweet Orange from Commercial Groves. Phytochem. Anal. 2010, 22, 236–246. [Google Scholar] [CrossRef] [PubMed]
- Mais, E.; Alolga, R.N.; Wang, S.-L.; Linus, L.O.; Yin, X.; Qi, L.-W. A comparative UPLC-Q/TOF-MS-based metabolomics approach for distinguishing Zingiber officinale Roscoe of two geographical origins. Food Chem. 2018, 240, 239–244. [Google Scholar] [CrossRef]
- Nam, K.-H.; Kim, Y.-J.; Moon, Y.S.; Pack, I.-S.; Kim, C.-G. Salinity affects metabolomic profiles of different trophic levels in a food chain. Sci. Total. Environ. 2017, 599–600, 198–206. [Google Scholar] [CrossRef]
- Mohandas, S.; Ravishankar, K.V. Banana: Genomics and Transgenic Approaches for Genetic Improvement; Springer: Berlin/Heidelberg, Germany, 2016; pp. 1–346. [Google Scholar] [CrossRef]
- Abdelrahman, M.; Burritt, D.J.; Tran, L.-S.P.; Adbelrahman, M. The use of metabolomic quantitative trait locus mapping and osmotic adjustment traits for the improvement of crop yields under environmental stresses. Semin. Cell Dev. Biol. 2018, 83, 86–94. [Google Scholar] [CrossRef] [PubMed]
- Hong, J.; Yang, L.; Zhang, D.; Shi, J. Plant Metabolomics: An Indispensable System Biology Tool for Plant Science. Int. J. Mol. Sci. 2016, 17, 767. [Google Scholar] [CrossRef]
- Cevallos-Cevallos, J.M.; Jines, C.; Maridueña-Zavala, M.G.; Molina-Miranda, M.J.; Ochoa, D.E.; Flores-Cedeno, J.A. GC-MS metabolite profiling for specific detection of dwarf somaclonal variation in banana plants. Appl. Plant Sci. 2018, 6, e01194. [Google Scholar] [CrossRef] [PubMed]
- Murashige, T.; Skoog, F. A Revised Medium for Rapid Growth and Bio Assays with Tobacco Tissue Cultures. Physiol. Plant. 1962, 15, 473–497. [Google Scholar] [CrossRef]
- Vuylsteke, D.R. Shoot-Tip Culture for the Propagation, Conservation and Exchange of Musa Germplasm; International Institute of Tropical Agriculture: Ibadan, Nigeria, 1989. [Google Scholar]
- Maridueña-Zavala, M.G.; Freire-Peñaherrera, A.; Cevallos-Cevallos, J.M.; Peralta, E.L. GC-MS metabolite profiling of Phytophthora infestans resistant to metalaxyl. Eur. J. Plant Pathol. 2017, 149, 563–574. [Google Scholar] [CrossRef]
- Pluskal, T.; Castillo, S.; Villar-Briones, A.; Orešič, M. MZmine 2: Modular framework for processing, visualizing, and analyzing mass spectrometry-based molecular profile data. BMC Bioinform. 2010, 11, 395. [Google Scholar] [CrossRef] [PubMed]
- Neumann, G.; Römheld, V. Root excretion of carboxylic acids and protons in phosphorus-deficient plants. Plant Soil 1999, 211, 121–130. [Google Scholar] [CrossRef]
- Aguiar, N.O.; Olivares, F.L.; Novotny, E.H.; Canellas, L.P. Changes in metabolic profiling of sugarcane leaves induced by endophytic diazotrophic bacteria and humic acids. PeerJ 2018, 6, e5445. [Google Scholar] [CrossRef] [PubMed]
- Li, P.; Wang, A.; Du, W.; Mao, L.; Wei, Z.; Wang, S.; Yuan, H.; Ji, R.; Zhao, L. Insight into the interaction between Fe-based nanomaterials and maize (Zea mays) plants at metabolic level. Sci. Total Environ. 2020, 738, 139795. [Google Scholar] [CrossRef]
- Stephens, C.; Christen, B.; Fuchs, T.; Sundaram, V.; Watanabe, K.; Jenal, U. Genetic Analysis of a Novel Pathway for d-Xylose Metabolism in Caulobacter crescentus. J. Bacteriol. 2006, 189, 2181–2185. [Google Scholar] [CrossRef]
- Klalvraa, S.; Gregersen, N. In vitro studies on the oxidation of medium-chain dicarboxylic acids in rat liver. ScienceDirect 1986, 876, 515–525. [Google Scholar]
- Krishna, H.; Alizadeh, M.; Singh, D.; Singh, U.; Chauhan, N.; Eftekhari, M.; Sadh, R.K. Somaclonal variations and their applications in horticultural crops improvement. 3 Biotech 2016, 6, 1–18. [Google Scholar] [CrossRef]
- Riggs, J.W.; Rockwell, N.C.; Cavales, P.C.; Callis, J. Identification of the plant ribokinase and discovery of a role for Arabidopsis Ribokinase in nucleoside metabolism. J. Biol. Chem. 2016, 291, 22572–22582. [Google Scholar] [CrossRef] [PubMed]
- Ishida, Y.; Kakibuchi, K.; Kudo, R.; Izumori, K.; Tajima, S.; Akimitsu, K.; Tanaka, K. Effects of rare sugars on growth and disease occurrence in head lettuce. Acta Hortic. 2012, 927, 929–934. [Google Scholar] [CrossRef]
- Vázquez, A.M.; Linacero, R. Stress and Somaclonal Variation. In Plant Developmental Biology—Biotechnological Perspectives; Springer International Publishing: Berlin/Heidelberg, Germany, 2010; Volume 2, pp. 45–64. [Google Scholar]
- Hossain, A.; Yamaguchi, F.; Matsuo, T.; Tsukamoto, I.; Toyoda, Y.; Ogawa, M.; Nagata, Y.; Tokuda, M. Rare sugar d-allulose: Potential role and therapeutic monitoring in maintaining obesity and type 2 diabetes mellitus. Pharmacol. Ther. 2015, 155, 49–59. [Google Scholar] [CrossRef]
- Ciereszko, I. Regulatory roles of sugars in plant growth and development. Acta Soc. Bot. Pol. 2018, 87, 1–13. [Google Scholar] [CrossRef]
- Mazelis, M. Amino Acid Catabolism. In Amino Acids and Derivatives; Academic Press: Cambridge, MA, USA, 1980; pp. 541–567. [Google Scholar]
- Sassaki, G.L.; de Souza, L.M. Mass Spectrometry Strategies for Structural Analysis of Carbohydrates and Glycoconjugates. In Tandem Mass Spectrometry—Molecular Characterization; InTech: London, UK, 2013. [Google Scholar]
- Scheller, H.V.; Ulvskov, P. Hemicelluloses. Annu. Rev. Plant Biol. 2010, 61, 263–289. [Google Scholar] [CrossRef]
- Buchanan, B.B.; Gruissem, W.; Jones, R.L. Biochemistry and Molecular Biology of Plants; John Wiley & Sons: Hoboken, NY, USA, 2000. [Google Scholar]
- Cohen, E.; Merzendorfer, H. Biologically-Inspired Systems Extracellular Sugar-Based Biopolymers Matrices. Available online: http://www.springer.com/series/8430 (accessed on 21 January 2021).
- Mathews, C.K.; Holde, K.E.; Appling, D.R. Biochemistry; Pearson: London, UK, 2013. [Google Scholar]
- Bindschedler, L.V.; Tuerck, J.; Maunders, M.; Ruel, K.; Petit-Conil, M.; Danoun, S.; Boudet, A.-M.; Joseleau, J.-P.; Bolwell, G.P. Modification of hemicellulose content by antisense down-regulation of UDP-glucuronate decarboxylase in tobacco and its consequences for cellulose extractability. Phytochemistry 2007, 68, 2635–2648. [Google Scholar] [CrossRef] [PubMed]
- Yang, Y.; Kang, L.; Wu, R.; Chen, Y.; Lu, C. Genome-wide identification and characterization of UDP-glucose dehydrogenase family genes in moso bamboo and functional analysis of PeUGDH4 in hemicellulose synthesis. Sci. Rep. 2020, 10, 1–13. [Google Scholar] [CrossRef] [PubMed]
- Xiao, C.; Anderson, C.T. Roles of pectin in biomass yield and processing for biofuels. Front. Plant Sci. 2013, 4, 67. [Google Scholar] [CrossRef] [PubMed]
- Cosgrove, D.J. Growth of the plant cell wall. Nat. Rev. Mol. Cell Biol. 2005, 6, 850–861. [Google Scholar] [CrossRef]
- Karp, A. Mechanisms of Somaclonal Variation. Biotechnol. Biotechnol. Equip. 1993, 7, 20–25. [Google Scholar] [CrossRef]
- Mathur, J. Cell shape development in plants. Trends Plant Sci. 2004, 9, 583–590. [Google Scholar] [CrossRef]
- Vogt, T. Phenylpropanoid Biosynthesis. Mol. Plant 2010, 3, 2–20. [Google Scholar] [CrossRef]
- Bais, H.P.; Vepachedu, R.; Gilroy, S.; Callaway, R.M.; Vivanco, J.M. Allelopathy and Exotic Plant Invasion: From Molecules and Genes to Species Interactions. Science 2003, 301, 1377–1380. [Google Scholar] [CrossRef]
- He, M.; He, C.-Q.; Ding, N.-Z. Abiotic Stresses: General Defenses of Land Plants and Chances for Engineering Multistress Tolerance. Front. Plant Sci. 2018, 9, 1771. [Google Scholar] [CrossRef]
- Masclaux-Daubresse, C.; Reisdorf-Cren, M.; Pageau, K.; Lelandais, M.; Grandjean, O.; Kronenberger, J.; Valadier, M.H.; Feraud, M.; Jouglet, T.; Suzuki, A. Glutamine synthetase-glutamate synthase pathway and glutamate dehydrogenase play distinct roles in the sink-source nitrogen cycle in tobacco. Plant Physiol. 2006, 140, 444–456. [Google Scholar] [CrossRef]
- Van Hengel, A.J.; Tadesse, Z.; Immerzeel, P.; Schols, H.; Van Kammen, A.; De Vries, S.C. N-Acetylglucosamine and Glucosamine-Containing Arabinogalactan Proteins Control Somatic Embryogenesis. Plant Physiol. 2001, 125, 1880–1890. [Google Scholar] [CrossRef] [PubMed]
- Yang, T.; Echols, M.; Martin, A.; Bar-Peled, M. Identification and characterization of a strict and a promiscuous N-acetylglucosamine-1-P uridylyltransferase in Arabidopsis. Biochem. J. 2010, 430, 275–284. [Google Scholar] [CrossRef]
- Bull, A.T. Chemical Composition of Wild-type and Mutant Aspergillus nidulans Cell Walls. The Nature of Polysaccharide and Melanin Constituents. J. Gen. Microbiol. 1970, 63, 75–94. [Google Scholar] [CrossRef]
- Freeze, H.H.; Lomis, W. Isolation and Characterization of a Component of the Surface Sheath of Dictyostelium discoideum. J. Biol. Chem. 1977, 10, 820–824. [Google Scholar] [CrossRef]
- Niemetz, R.; Karcher, U.; Kandler, O.; Tindall, B.J.; Konig, H. The cell wall polymer of the extremely halophilic archaeon Natronococcus occultus. JBIC J. Biol. Inorg. Chem. 1997, 249, 905–911. [Google Scholar] [CrossRef] [PubMed]
- Vroh-Bi, I.; Anagbogu, C.; Nnadi, S.; Tenkouano, A. Genomic characterization of natural and somaclonal variations in bananas (Musa spp.). Plant Mol. Biol. Rep. 2011, 29, 440–448. [Google Scholar] [CrossRef]


| No. | METABOLITE | CLASS | Log2 FC M1DP-M1NP | Log2 FC M2DP-M1NP |
|---|---|---|---|---|
| Proliferation Phase | ||||
| 1 | Sebacic acid | Dicarboxylic acid | 3.520 | n.s. |
| 2 | 2,3,4,5-Tetrahydroxypentanoic acid-1,4-lactone | Carboxylic acid | 2.304 | n.s. |
| 3 | D-Ribofuranose | Monosaccharide | 2.226 | n.s. |
| 4 | D-Glucose | Monosaccharide | −0.995 | n.s. |
| 5 | L-(-)-Sorbose | Monosaccharide | −0.908 | n.s. |
| 6 | β-L-Arabinopyranose | Monosaccharide | −0.918 | −6.252 |
| 7 | α-D-Galactopyranose | Monosaccharide | −1.479 | −1.013 |
| 8 | Phenylalanine | Aminoacid | UDP | n.s. |
| Rooting Phase | ||||
| 9 | Propanoic acid | Carboxilic acid | 3.666 | NS |
| 10 | 2-Keto-D-gluconic acid | Carboxilic acid | 2.694 | 1.898 |
| 11 | 1,2,3-Propanetricarboxylic acid, 2-hydroxi (Citric acid) | Tricarboxylic acid | 0.954 | 0.944 |
| 12 | D-Psicofuranose | Monosaccharide | 0.907 | UDP |
| 13 | D-Galactose, 2-deoxy | Monosaccharide | −1.084 | −0.620 |
| 14 | D-Fructose | Monosaccharide | −1.076 | −1.510 |
| 15 | D-(-)-Fructofuranose | Monosaccharide | −1.246 | NS |
| 6 | β-L-Arabinopyranose | Monosaccharide | −1.264 | UDP |
| Acclimatization Phase II | ||||
| 16 | β-D-Galactofuranose | Monosaccharide | UNP | n.a. |
| 17 | Gulonic acid | Carboxilic acid | UNP | n.a. |
| 18 | D-(+)-Galacturonic acid | Monosaccharide | 2.852 | n.a. |
| 19 | N-Acetyl glucosamine | Amino sugar | −0.755 | n.a. |
| 20 | D-Galactose, 2-amino-2-deoxy (D-Galactosamine) | Amino sugar | −0.969 | n.a. |
| 21 | Mannose | Monosaccharide | −2.307 | n.a. |
| 22 | L-6 deoxy-Galactopyranose (L-Fucopyranose) | Monosaccharide | −3.993 | n.a. |
| 23 | L-Glutamine | Aminoacid | UDP | n.a. |
| 24 | α-D-Glucopyranoside, methyl | Monosaccharid | UDP | n.a. |
| 25 | Uridine | Nucleoside | UDP | n.a. |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Carrera, F.P.; Noceda, C.; Maridueña-Zavala, M.G.; García, J.A.; Ruiz-Barzola, O.; Cevallos-Cevallos, J.M. Changes in the Metabolite Profile during Micropropagation of Normal and Somaclonal Variants of Banana Musa AAA cv. Williams. Horticulturae 2021, 7, 39. https://doi.org/10.3390/horticulturae7030039
Carrera FP, Noceda C, Maridueña-Zavala MG, García JA, Ruiz-Barzola O, Cevallos-Cevallos JM. Changes in the Metabolite Profile during Micropropagation of Normal and Somaclonal Variants of Banana Musa AAA cv. Williams. Horticulturae. 2021; 7(3):39. https://doi.org/10.3390/horticulturae7030039
Chicago/Turabian StyleCarrera, Fredy P., Carlos Noceda, María G. Maridueña-Zavala, José A. García, Omar Ruiz-Barzola, and Juan M. Cevallos-Cevallos. 2021. "Changes in the Metabolite Profile during Micropropagation of Normal and Somaclonal Variants of Banana Musa AAA cv. Williams" Horticulturae 7, no. 3: 39. https://doi.org/10.3390/horticulturae7030039
APA StyleCarrera, F. P., Noceda, C., Maridueña-Zavala, M. G., García, J. A., Ruiz-Barzola, O., & Cevallos-Cevallos, J. M. (2021). Changes in the Metabolite Profile during Micropropagation of Normal and Somaclonal Variants of Banana Musa AAA cv. Williams. Horticulturae, 7(3), 39. https://doi.org/10.3390/horticulturae7030039







