De novo Transcriptome Assembly and Comprehensive Annotation of Two Tree Tomato Cultivars (Solanum betaceum Cav.) with Different Fruit Color
Abstract
1. Introduction
2. Materials and Methods
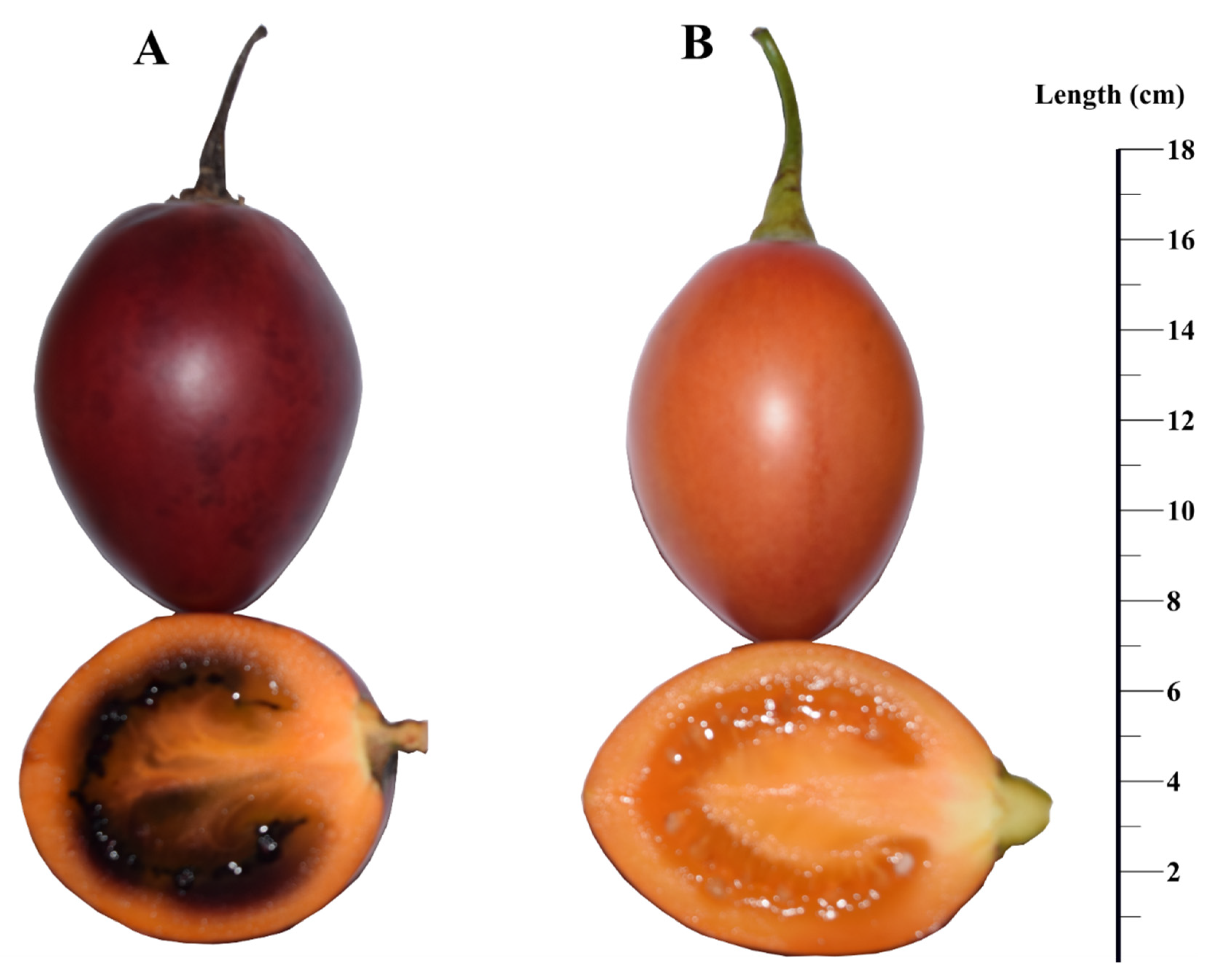
2.1. Plant Material
2.2. RNA Extraction, Library Construction and RNA Sequencing
2.3. DNA Sequence Processing and De Novo Transcriptome Assembly
2.4. Structural and Functional Annotation
2.5. Single-Nucleotide Variations (SNVs)
3. Results
3.1. Transcriptome Sequencing and Assembly
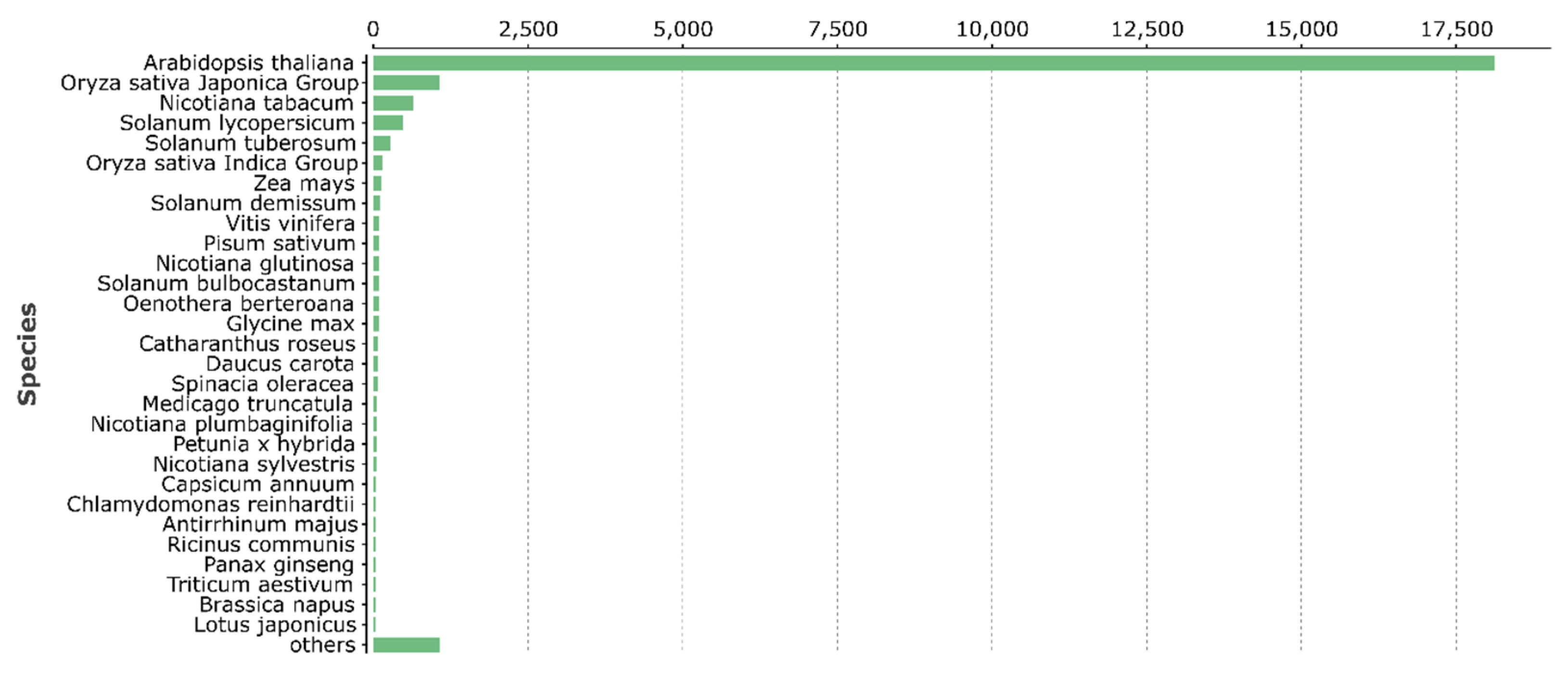
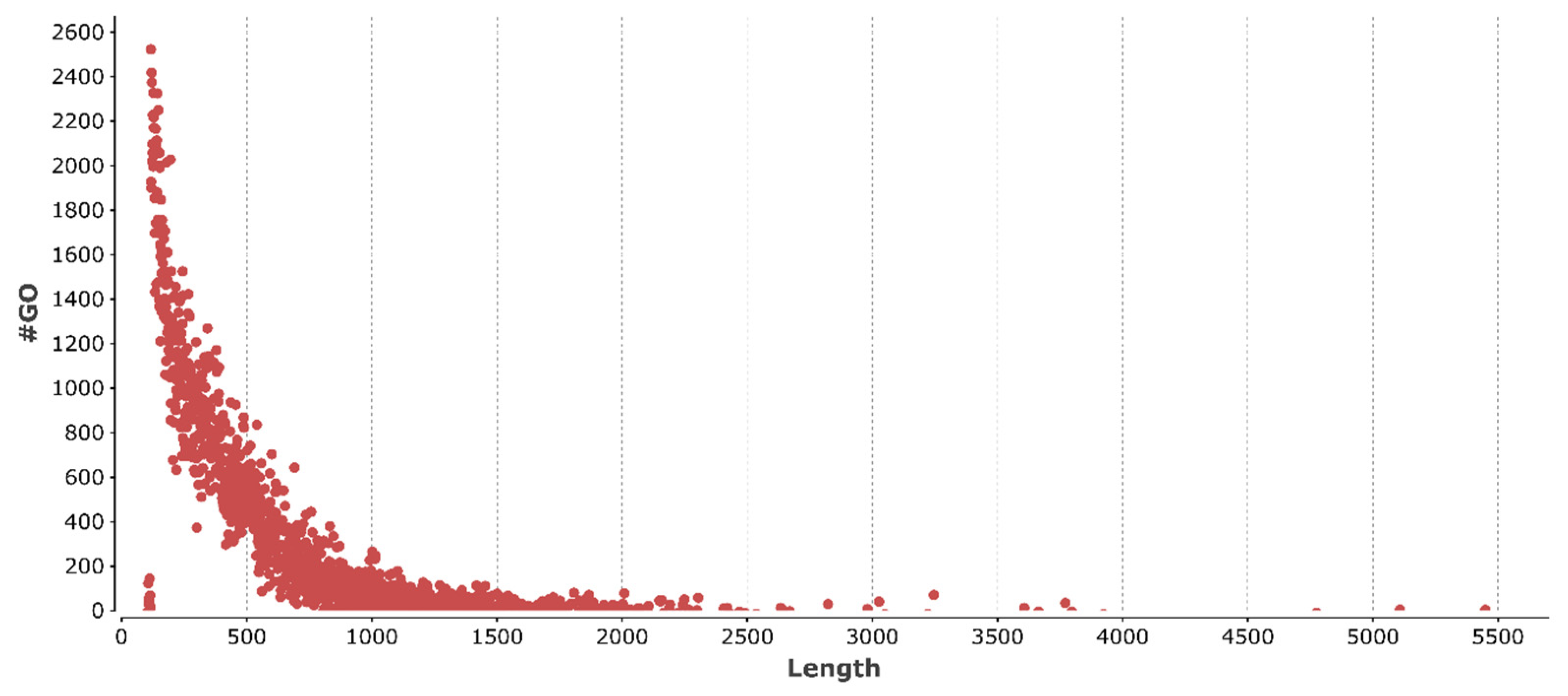
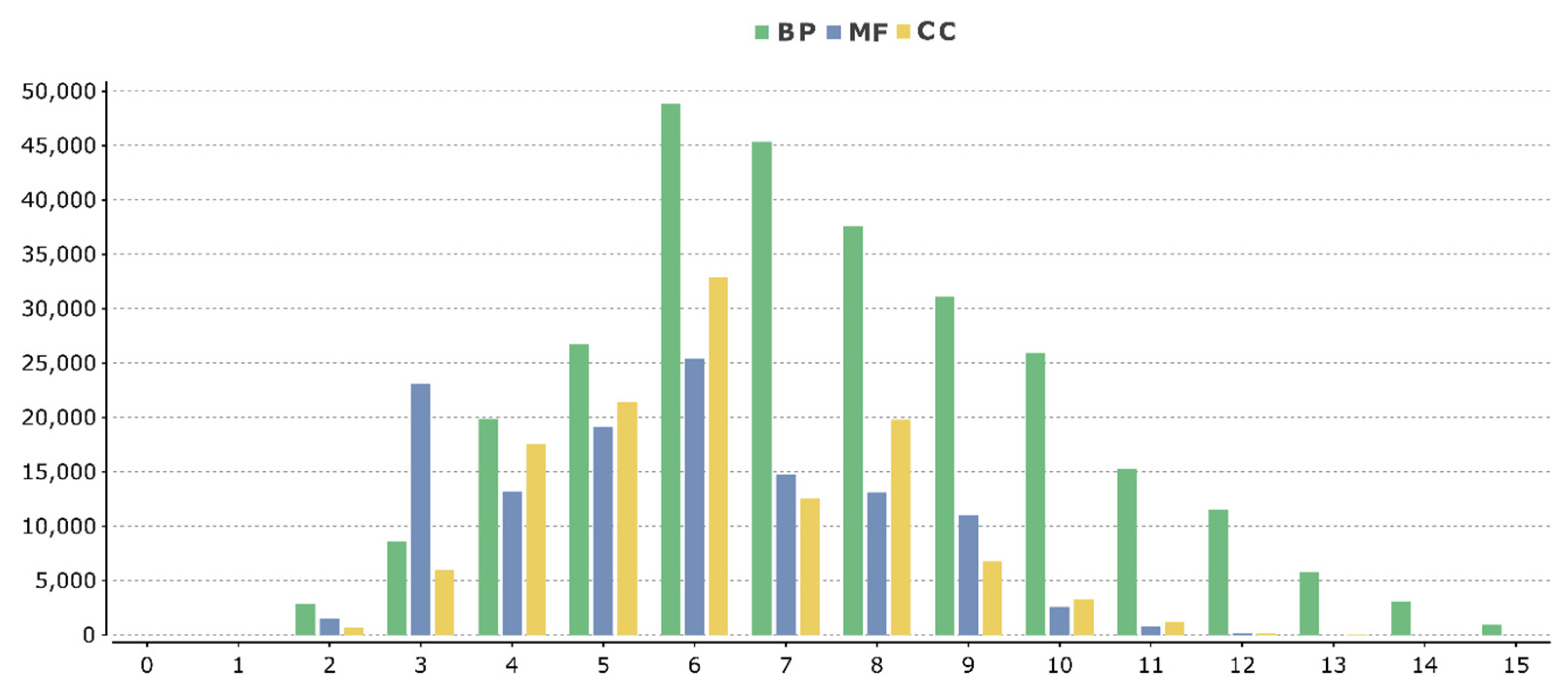
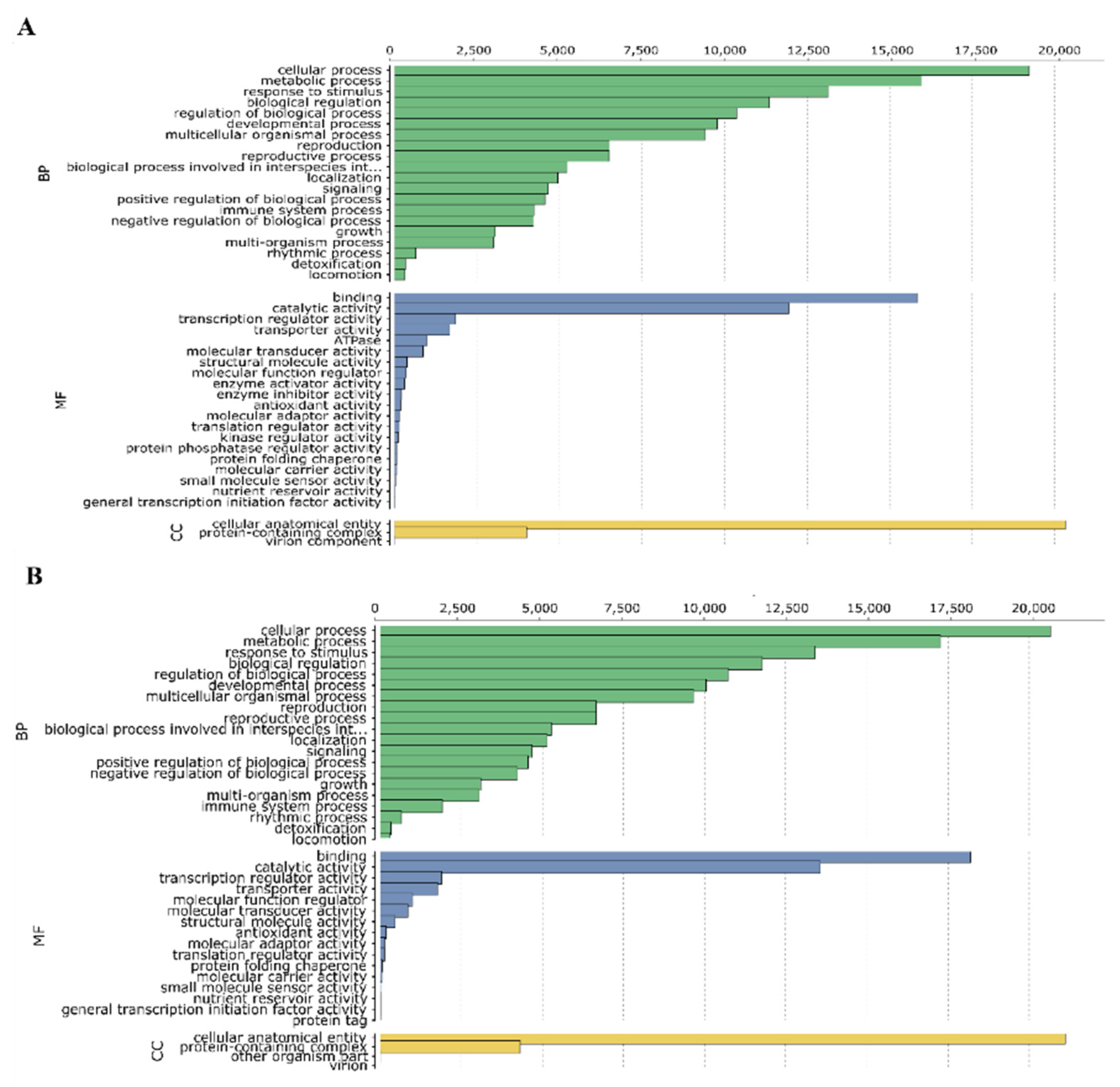
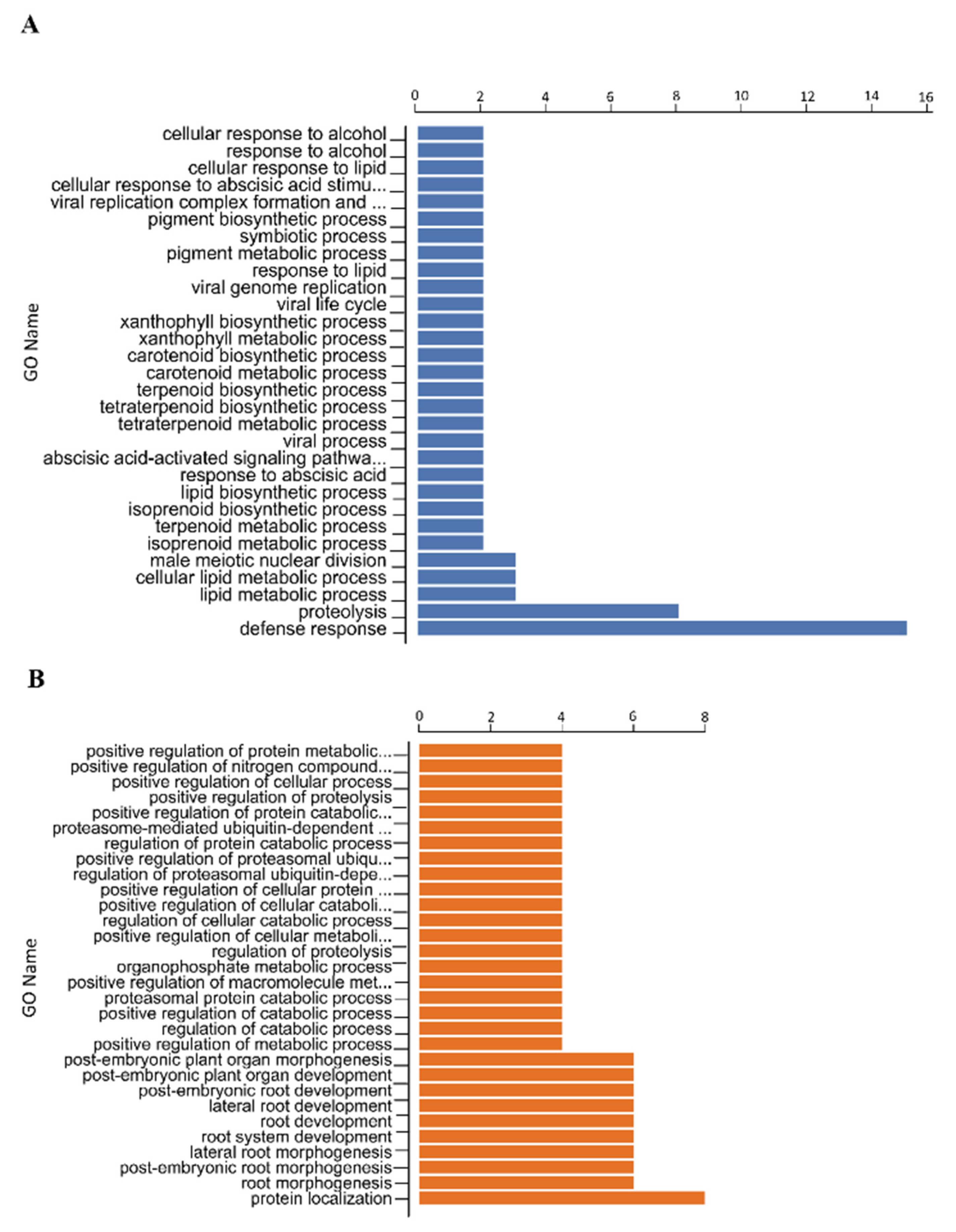
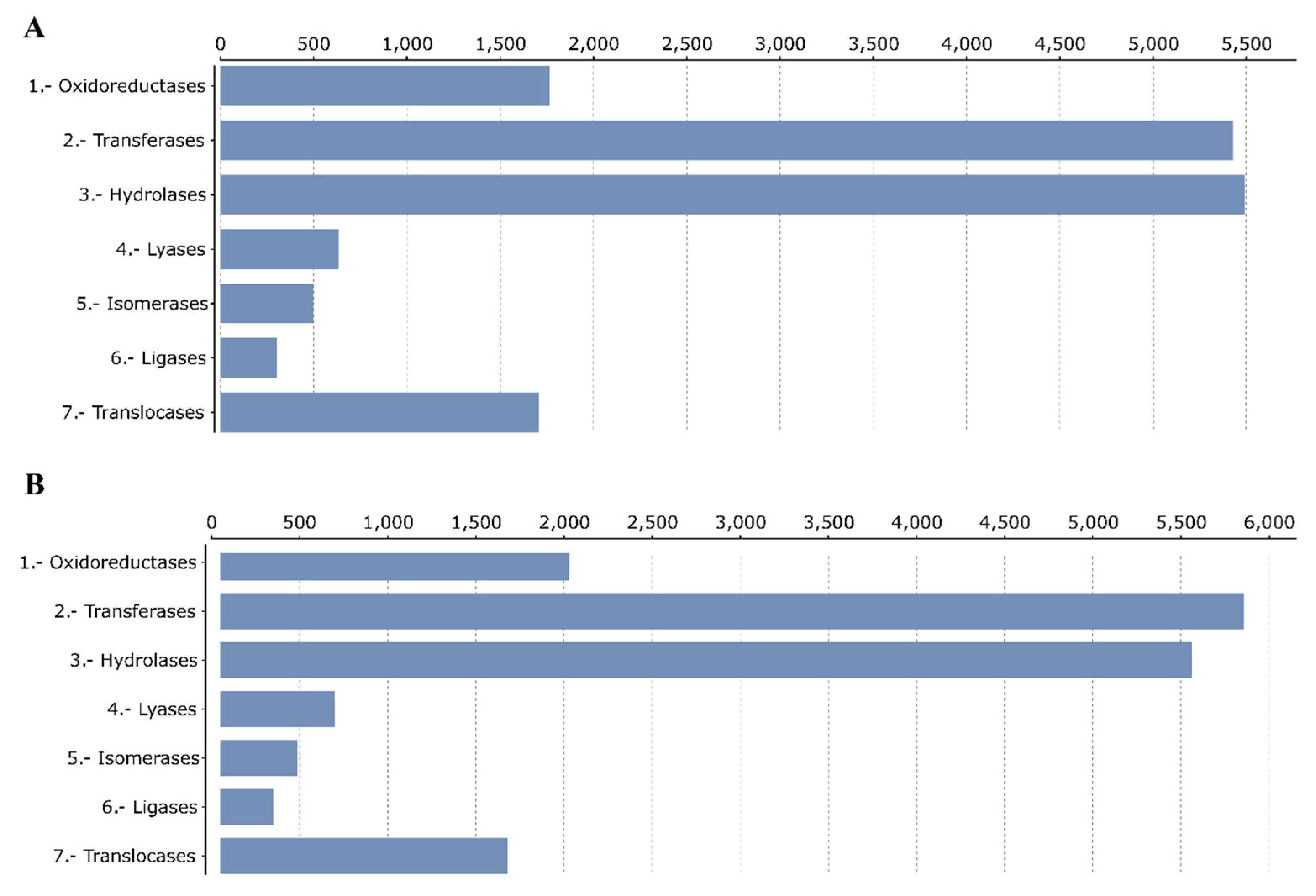
3.2. Structural and Functional Annotation
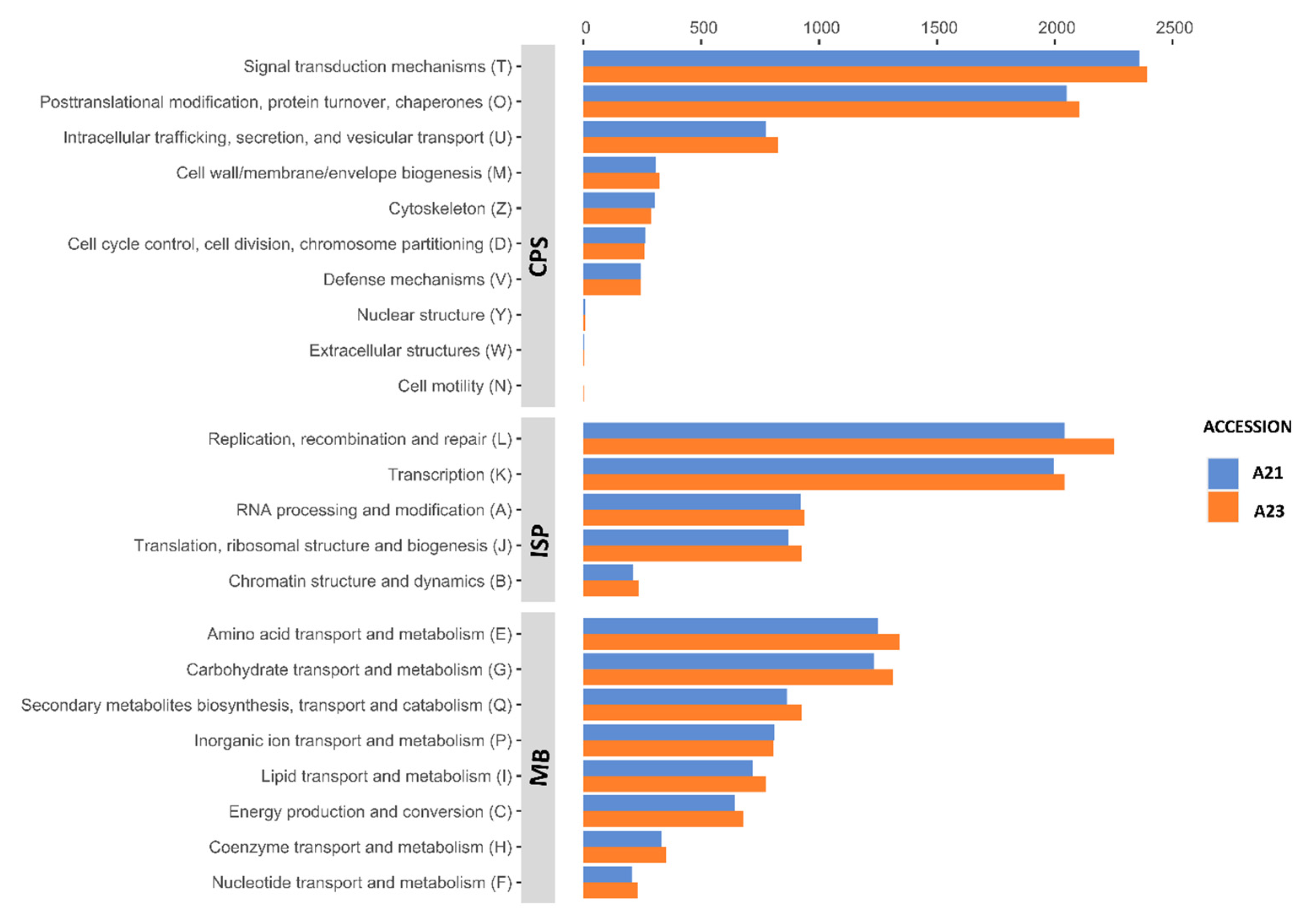
3.3. COG Classification
3.4. Identification and Characterizacion of SNVs
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Duarte, O.; Paull, R.E. Solanaceae. In Exotic Fruits and Nuts of the New World, 1st ed.; Duarte, O., Paull, R.E., Eds.; CABI: Wallingford, UK, 2015; pp. 136–144. [Google Scholar]
- Orqueda, M.E.; Zampini, I.C.; Torres, S.; Alberto, M.R.; Ramos, L.L.P.; Schmeda-Hirschmann, G.; Isla, M.I. Chemical and functional characterization of skin, pulp and seed powder from the Argentine native fruit mistol (Ziziphus mistol). Effects of phenolic fractions on key enzymes involved in metabolic syndrome and oxidative stress. J. Funct. Foods 2017, 37, 531–540. [Google Scholar] [CrossRef]
- Särkinen, T.; Bohs, L.; Olmstead, R.G.; Knapp, S. A phylogenetic framework for evolutionary study of the nightshades (Solanaceae): A dated 1000-tip tree. BMC Evol. Biol. 2013, 13, 214. [Google Scholar] [CrossRef]
- Ramírez, F.; Kallarackal, J. Tree tomato (Solanum betaceum Cav.) reproductive physiology: A review. Sci. Hortic. 2019, 248, 206–215. [Google Scholar] [CrossRef]
- Chen, X.; Quek, S.Y.; Fedrizzi, B.; Kilmartin, P.A. Characterization of free and glycosidically bound volatile compounds from tamarillo (Solanum betaceum Cav.) with considerations on hydrolysis strategies and incubation time. LWT 2020, 124, 109178. [Google Scholar] [CrossRef]
- Acosta-Quezada, P.G.; Martínez-Laborde, J.B.; Prohens, J. Variation among tree tomato (Solanum betaceum Cav.) accessions from different cultivar groups: Implications for conservation of genetic resources and breeding. Genet. Resour. Crop Evol. 2011, 58, 943–960. [Google Scholar] [CrossRef]
- Diep, T.T.; Rush, E.C.; Yoo, M.J.Y. Tamarillo (Solanum betaceum Cav.): A review of physicochemical and bioactive properties and potential applications. Food Rev. Int. 2020, 6, 1–25. [Google Scholar] [CrossRef]
- Diep, T.T.; Pook, C.; Rush, E.C.; Yoo, M.J.Y. Quantification of carotenoids, α-tocopherol, and ascorbic acid in amber, mulligan, and laird’s large cultivars of New Zealand tamarillos (Solanum betaceum Cav.). Foods 2020, 9, 769. [Google Scholar] [CrossRef]
- Mertz, C.; Brat, P.; Caris-Veyrat, C.; Gunata, Z. Characterization and thermal lability of carotenoids and vitamin C of tamarillo fruit (Solanum betaceum Cav.). Food Chem. 2010, 119, 653–659. [Google Scholar] [CrossRef]
- Acosta-Quezada, P.G.; Vilanova, S.; Martínez-Laborde, J.B.; Prohens, J. Genetic diversity and relationships in accessions from different cultivar groups and origins in the tree tomato (Solanum betaceum Cav.). Euphytica 2012, 187, 87–97. [Google Scholar] [CrossRef]
- Huang, X.; Chen, X.-G.; Armbruster, P.A. Comparative performance of transcriptome assembly methods for non-model organisms. BMC Genom. 2016, 17, 523. [Google Scholar] [CrossRef]
- Ward, J.A.; Ponnala, L.; Weber, C.A. Strategies for transcriptome analysis in nonmodel plants. Am. J. Bot. 2012, 99, 267–276. [Google Scholar] [CrossRef] [PubMed]
- Xia, Z.; Xu, H.; Zhai, J.; Li, D.; Luo, H.; He, C.; Huang, X. RNA-Seq analysis and de novo transcriptome assembly of Hevea brasiliensis. Plant Mol. Biol. 2011, 77, 299–308. [Google Scholar] [CrossRef]
- Herraiz, F.J.; Blanca, J.; Ziarsolo, P.; Gramazio, P.; Plazas, M.; Anderson, G.J.; Prohens, J.; Vilanova, S. The first de novo transcriptome of pepino (Solanum muricatum): Assembly, comprehensive analysis and comparison with the closely related species S. caripense, potato and tomato. BMC Genom. 2016, 17, 321. [Google Scholar] [CrossRef]
- Ranil, R.H.G.; Niran, H.M.L.; Plazas, M.; Fonseka, R.M.; Fonseka, H.H.; Vilanova, S.; Andújar, I.; Gramazio, P.; Fita, A.; Prohens, J. Improving seed germination of the eggplant rootstock Solanum torvum by testing multiple factors using an orthogonal array design. Sci. Hortic. 2015, 193, 174–181. [Google Scholar] [CrossRef]
- Andrews, S. FastQC: A Quality Control Tool for High Throughput Sequence Data. 2010. Available online: http://www.bioinformatics.babraham.ac.uk/projects/fastqc (accessed on 18 July 2020).
- Bolger, A.M.; Lohse, M.; Usadel, B. Trimmomatic: A flexible trimmer for Illumina sequence data. Bioinformatics 2014, 30, 2114–2120. [Google Scholar] [CrossRef]
- Grabherr, M.G.; Haas, B.J.; Yassour, M.; Levin, J.; A Thompson, D.; Amit, I.; Adiconis, X.; Fan, L.; Raychowdhury, R.; Zeng, Q.; et al. Full-length transcriptome assembly from RNA-Seq data without a reference genome. Nat. Biotechnol. 2011, 29, 644–652. [Google Scholar] [CrossRef] [PubMed]
- Fu, L.; Niu, B.; Zhu, Z.; Wu, S.; Li, W. CD-HIT: Accelerated for clustering the next-generation sequencing data. Bioinformatics 2012, 28, 3150–3152. [Google Scholar] [CrossRef]
- Langmead, B.; Salzberg, S.L. Fast gapped-read alignment with Bowtie 2. Nat. Methods 2012, 9, 357–359. [Google Scholar] [CrossRef]
- Simão, F.A.; Waterhouse, R.M.; Ioannidis, P.; Kriventseva, E.V.; Zdobnov, E.M. BUSCO: Assessing genome assembly and annotation completeness with single-copy orthologs. Bioinformatics 2015, 31, 3210–3212. [Google Scholar] [CrossRef]
- Waterhouse, R.M.; Seppey, M.; Simao, F.A.; Manni, M.; Ioannidis, P.; Klioutchnikov, G.; Kriventseva, E.V.; Zdobnov, E.M. BUSCO applications from quality assessments to gene prediction and phylogenomics. Mol. Biol. Evol. 2018, 35, 543–548. [Google Scholar] [CrossRef] [PubMed]
- Götz, S.; Garcia-Gomez, J.M.; Terol, J.; Williams, T.; Nagaraj, S.H.; Nueda, M.J.; Robles, M.; Talón, M.; Dopazo, J.; Conesa, A. High-throughput functional annotation and data mining with the Blast2GO suite. Nucleic Acids Res. 2008, 36, 3420–3435. [Google Scholar] [CrossRef] [PubMed]
- Bryant, D.M.; Johnson, K.; DiTommaso, T.; Tickle, T.; Couger, M.B.; Payzin-Dogru, D.; Lee, T.J.; Leigh, N.; Kuo, T.-H.; Davis, F.G.; et al. A tissue-mapped axolotl de novo transcriptome enables identification of limb regeneration factors. Cell Rep. 2017, 18, 762–776. [Google Scholar] [CrossRef]
- Wheeler, T.J.; Eddy, S.R. Nhmmer: DNA homology search with profile HMMs. Bioinformatics 2013, 29, 2487–2489. [Google Scholar] [CrossRef] [PubMed]
- Petersen, T.N.; Brunak, S.; Von Heijne, G.; Nielsen, H. SignalP 4.0: Discriminating signal peptides from transmembrane regions. Nat. Methods 2011, 8, 785–786. [Google Scholar] [CrossRef]
- Huerta-Cepas, J.; Szklarczyk, D.; Heller, D.; Hernández-Plaza, A.; Forslund, S.K.; Cook, H.V.; Mende, D.R.; Letunic, I.; Rattei, T.; Jensen, L.J.; et al. EggNOG 5.0: A hierarchical, functionally and phylogenetically annotated orthology resource based on 5090 organisms and 2502 viruses. Nucleic Acids Res. 2019, 47, D309–D314. [Google Scholar] [CrossRef]
- Kanehisa, M.; Goto, S. KEGG: Kyoto encyclopedia of genes and genomes. Nucleic Acids Res. 2000, 28, 27–30. [Google Scholar] [CrossRef]
- Li, H.; Durbin, R. Fast and accurate short read alignment with Burrows-Wheeler transform. Bioinformatics 2009, 25, 1754–1760. [Google Scholar] [CrossRef] [PubMed]
- Hosmani, P.S.; Flores-Gonzales, M.; van de Geest, H.; Maumus, F.; Bakker, L.V.; Schijlen, E.; van Haarst, J.; Cordewener, J.; Sanchez-Perez, G.; Peters, S.; et al. An Improved de Novo Assembly and Annotation of the Tomato Reference Genome Using Single-Molecule Sequencing, Hi-C Proximity Ligation and Optical Maps. BioRxiv under Review. Available online: https://www.biorxiv.org/content/10.1101/767764v1 (accessed on 18 July 2020).
- Pham, G.M.; Hamilton, J.P.; Wood, J.C.; Burke, J.T.; Zhao, H.; Vaillancourt, B.; Ou, S.; Jiang, J.; Buell, C.R. Construction of a chromosome-scale long-read reference genome assembly for potato. GigaScience 2020, 9, giaa100. [Google Scholar] [CrossRef]
- Li, H.; Handsaker, R.; Wysoker, A.; Fennell, T.; Ruan, J.; Homer, N.; Marth, G.; Abecasis, G.; Durbin, R. The sequence alignment/map format and samtools. Bioinformatics 2009, 25, 2078–2079. [Google Scholar] [CrossRef]
- Garrison, E.; Marth, G. Haplotype-Based Variant Detection from Short-Read Sequencing. arXiv 2012, arXiv:1207.3907. Available online: http://arXiv.org/abs/1207.3907 (accessed on 18 July 2020).
- Cingolani, P.; Platts, A.; Wang, L.L.; Coon, M.; Nguyen, T.; Wang, L.; Land, S.J.; Lu, X.; Ruden, D.M. A program for annotating and predicting the effects of single nucleotide polymorphisms, SnpEff: SNPs in the genome of Drosophila melanogaster strain w1118; iso-2; iso-3. Fly 2012, 6, 80–92. [Google Scholar] [CrossRef] [PubMed]
- Barabaschi, D.; Tondelli, A.; Desiderio, F.; Volante, A.; Vaccino, P.; Valè, G.; Cattivelli, L. Next generation breeding. Plant Sci. 2015, 242, 3–13. [Google Scholar] [CrossRef] [PubMed]
- Enciso-Rodriguez, F.; Osorio-Guarín, J.A.; Garzón-Martínez, G.A.; Delgadillo-Duran, P.; Barrero, L.S. Optimization of the genotyping-by-sequencing SNP calling for diversity analysis in cape gooseberry (Physalis peruviana L.) and related taxa. PLoS ONE 2020, 15, e0238383. [Google Scholar] [CrossRef]
- Ma, X.; Vaistij, F.E.; Li, Y.; van Rensburg, W.S.J.; Harvey, S.; Bairu, M.W.; Venter, S.L.; Mavengahama, S.; Ning, Z.; Graham, I.A.; et al. A chromosome-level Amaranthus cruentus genome assembly highlights gene family evolution and biosynthetic gene clusters that may underpin the nutritional value of this traditional crop. Plant J. 2021, 107, 613–628. [Google Scholar] [CrossRef] [PubMed]
- Moon, K.-B.; Ahn, D.-J.; Park, J.-S.; Jung, W.Y.; Cho, H.S.; Kim, H.-R.; Jeon, J.-H.; Park, Y.-I.; Kim, H.-S. Transcriptome profiling and characterization of drought-tolerant potato plant (Solanum tuberosum L.). Mol. Cells 2018, 41, 979–992. [Google Scholar] [CrossRef]
- Scarano, D.; Rao, R.; Corrado, G. In Silico identification and annotation of noncoding RNAs by RNA-seq and de novo assembly of the transcriptome of tomato fruits. PLoS ONE 2017, 12, e0171504. [Google Scholar] [CrossRef] [PubMed]
- Yang, X.; Liu, F.; Zhang, Y.; Wang, L.; Cheng, Y.-F. Cold-responsive miRNAs and their target genes in the wild eggplant species Solanum aculeatissimum. BMC Genom. 2017, 18, 1–13. [Google Scholar] [CrossRef]
- Zuluaga, A.P.; Solé, M.; Lu, H.; Góngora-Castillo, E.; Vaillancourt, B.; Coll, N.; Buell, C.R.; Valls, M. Transcriptome responses to Ralstonia solanacearum infection in the roots of the wild potato Solanum commersonii. BMC Genom. 2015, 16, 1–16. [Google Scholar] [CrossRef]
- Petek, M.; Zagorščak, M.; Ramšak, Ž.; Sanders, S.; Tomaž, Š.; Tseng, E.; Zouine, M.; Coll, A.; Gruden, K. Cultivar-specific transcriptome and pan-transcriptome reconstruction of tetraploid potato. Sci. Data 2020, 7, 1–15. [Google Scholar] [CrossRef]
- Stam, R.; Nosenko, T.; Hörger, A.C.; Stephan, W.; Seidel, M.; Kuhn, J.M.M.; Haberer, G.; Tellier, A. The de novo reference genome and transcriptome assemblies of the wild tomato species Solanum chilense highlights birth and death of NLR genes between tomato species. G3 Genes Genomes Genet. 2019, 9, 3933–3941. [Google Scholar] [CrossRef] [PubMed]
- Hu, H.; Scheben, A.; Edwards, D. Advances in integrating genomics and bioinformatics in the plant breeding pipeline. Agriculture 2018, 8, 75. [Google Scholar] [CrossRef]
- Aversano, R.; Contaldi, F.; Ercolano, M.R.; Grosso, V.; Iorizzo, M.; Tatino, F.; Xumerle, L.; Molin, A.D.; Avanzato, C.; Ferrarini, A.; et al. The Solanum commersonii genome sequence provides insights into adaptation to stress conditions and genome evolution of wild potato relatives. Plant Cell 2015, 27, 954–968. [Google Scholar] [CrossRef]
- Gramazio, P.; Blanca, J.; Ziarsolo, P.; Herraiz, F.J.; Plazas, M.; Prohens, J.; Vilanova, S. Transcriptome analysis and molecular marker discovery in Solanum incanum and S. aethiopicum, two close relatives of the common eggplant (Solanum melongena) with interest for breeding. BMC Genom. 2016, 17, 1–17. [Google Scholar] [CrossRef]
- Wang, X.; Gao, L.; Jiao, C.; Stravoravdis, S.; Hosmani, P.S.; Saha, S.; Zhang, J.; Mainiero, S.; Strickler, S.R.; Catala, C.; et al. Genome of Solanum pimpinellifolium provides insights into structural variants during tomato breeding. Nat. Commun. 2020, 11, 1–11. [Google Scholar] [CrossRef] [PubMed]
- Lateef, A.; Prabhudas, S.K.; Natarajan, P. RNA sequencing and de novo assembly of Solanum trilobatum leaf transcriptome to identify putative transcripts for major metabolic pathways. Sci. Rep. 2018, 8, 1–13. [Google Scholar] [CrossRef]
- Wu, L.; Du, G.; Bao, R.; Li, Z.; Gong, Y.; Liu, F. De novo assembly and discovery of genes involved in the response of Solanum sisymbriifolium to Verticillium dahlia. Physiol. Mol. Biol. Plants 2019, 25, 1009–1027. [Google Scholar] [CrossRef] [PubMed]
- Kanehisa, M.; Araki, M.; Goto, S.; Hattori, M.; Hirakawa, M.; Itoh, M.; Katayama, T.; Kawashima, S.; Okuda, S.; Tokimatsu, T.; et al. KEGG for linking genomes to life and the environment. Nucleic Acids Res. 2008, 36, D480–D484. [Google Scholar] [CrossRef]
- Karasawa, M.M.G.; Mohan, C. Fruits as prospective reserves of bioactive compounds: A review. Nat. Prod. Bioprospect. 2018, 8, 335–346. [Google Scholar] [CrossRef]
- Britton, G. Carotenoid research: History and new perspectives for chemistry in biological systems. Biochim. Biophys. Acta Mol. Cell Biol. Lipids 2020, 1865, 158699. [Google Scholar] [CrossRef]
- Nabavi, S.M.; Šamec, D.; Tomczyk, M.; Milella, L.; Russo, D.; Habtemariam, S.; Suntar, I.; Rastrelli, L.; Daglia, M.; Xiao, J.; et al. Flavonoid biosynthetic pathways in plants: Versatile targets for metabolic engineering. Biotechnol. Adv. 2020, 38, 107316. [Google Scholar] [CrossRef] [PubMed]
- Acosta-Quezada, P.G.; Raigón, M.D.; Riofrío-Cuenca, T.; García-Martínez, M.D.; Plazas, M.; Burneo, J.I.; Figueroa, J.G.; Vilanova, S.; Prohens, J. Diversity for chemical composition in a collection of different varietal types of tree tomato (Solanum betaceum Cav.), an Andean exotic fruit. Food Chem. 2015, 169, 327–335. [Google Scholar] [CrossRef]
- Vasco, C.; Avila, J.; Ruales, J.; Svanberg, U.; Kamal-Eldin, A. Physical and chemical characteristics of golden-yellow and purple-red varieties of tamarillo fruit (Solanum betaceum Cav.). Int. J. Food Sci. Nutr. 2009, 60, 278–288. [Google Scholar] [CrossRef] [PubMed]
- Ye, J.; Hu, T.; Yang, C.; Li, H.; Yang, M.; Ijaz, R.; Ye, Z.; Zhang, Y. Transcriptome profiling of tomato fruit development reveals transcription factors associated with ascorbic acid, carotenoid and flavonoid biosynthesis. PLoS ONE 2015, 10, e0130885. [Google Scholar] [CrossRef] [PubMed]
- Hamilton, J.P.; Hansey, C.N.; Whitty, B.R.; Stoffel, K.; Massa, A.N.; Van Deynze, A.; De Jong, W.S.; Douches, D.S.; Buell, C.R. Single nucleotide polymorphism discovery in elite north American potato germplasm. BMC Genom. 2011, 12, 302. [Google Scholar] [CrossRef]
- Gramazio, P.; Yan, H.; Hasing, T.; Vilanova, S.; Prohens, J.; Bombarely, A. Whole-genome resequencing of seven eggplant (Solanum melongena) and one wild relative (S. incanum) accessions provides new insights and breeding tools for eggplant enhancement. Front. Plant Sci. 2019, 10, 1220. [Google Scholar] [CrossRef] [PubMed]
- Olmstead, R.G.; Bohs, L.; Migid, H.A.; Santiago-Valentin, E.; Garcia, V.F.; Collier, S.M. A molecular phylogeny of the Solanaceae. Taxon 2008, 57, 1159–1181. [Google Scholar] [CrossRef]
- Isaacson, T.; Ronen, G.; Zamir, D.; Hirschberg, J. Cloning of tangerine from tomato reveals a carotenoid isomerase essential for the production of β-carotene and xanthophylls in plants. Plant Cell 2002, 14, 333–342. [Google Scholar] [CrossRef]
- Livingstone, K.; Anderson, S. Patterns of variation in the evolution of carotenoid biosynthetic pathway enzymes of higher plants. J. Hered. 2009, 100, 754–761. [Google Scholar] [CrossRef]
- Hunt, R.C.; Simhadri, V.L.; Iandoli, M.; Sauna, Z.E.; Kimchi-Sarfaty, C. Exposing synonymous mutations. Trends Genet. 2014, 30, 308–321. [Google Scholar] [CrossRef]
- Gu, W.; Wang, X.; Zhai, C.; Xie, X.; Zhou, T. Selection on synonymous sites for increased accessibility around miRNA binding sites in plants. Mol. Biol. Evol. 2020, 29, 3037–3044. [Google Scholar] [CrossRef][Green Version]
- He, J.; Zhao, X.; Laroche, A.; Lu, Z.-X.; Liu, H.K.; Li, Z. Genotyping-by-sequencing (GBS), An ultimate marker-assisted selection (MAS) tool to accelerate plant breeding. Front. Plant Sci. 2014, 5, 484. [Google Scholar] [CrossRef] [PubMed]

| Statistics | Accessions | |
|---|---|---|
| A21 | A23 | |
| Total raw reads | 100,919,310 | 113,802,281 |
| Total raw reads data size (Gb) | 14.68 | 15.84 |
| G/C (%) | 42.2 | 42.2 |
| Total clean reads | 38,411,167 | 54,474,055 |
| Total clean reads data size (Gb) | 4.25 | 5.97 |
| Number of transcripts | 174,252 | 194,417 |
| Total nucleotide length | 148,352,996 | 165,074,290 |
| Average transcript length | 851.37 | 849.07 |
| Maximum transcript length | 17,046 | 16,865 |
| N50 | 1494 | 1503 |
| G/C (%) | 38.8 | 38.6 |
| Overall alignment rate (%) | 99.09 | 99.21 |
| BUSCO (%) | 98.4 | 98.8 |
| Statistics | Accessions | |
|---|---|---|
| A21 | A23 | |
| Predicted ORFs | 27,441 | 28,336 |
| Predicted proteins | 34,636 | 36,224 |
| sprot_Top_BLASTX_hit | 57,422 | 60,772 |
| sprot_Top_BLASTP_hit | 24,311 | 25,054 |
| Pfam | 22,954 | 23,637 |
| SignalP | 1623 | 1745 |
| TmHMM | 6899 | 7216 |
| GO terms | 196,800 | 204,090 |
| EC numbers | 15,828 | 16,668 |
| Kegg | 14,035 | 14,540 |
| Statistics | SNPs | MNP | INDELs | MIXED | Total SNVs |
|---|---|---|---|---|---|
| SNVs intraspecific variations | |||||
| A21 | 14,837 | 3283 | 767 | 227 | 19,117 |
| A23 | 38,183 | 8213 | 2391 | 726 | 49,530 |
| SNVs interspecific variations | |||||
| A21 and S. tuberosum | 619,626 | 174,982 | 28,2835 | 23,115 | 1,973,023 |
| A23 and S. tuberosum | 805,997 | 242,484 | 42,142 | 36,352 | |
| A21 and S. lycopersicum | 624,503 | 194,857 | 23,407 | 20,788 | 1,809,264 |
| A23 and S. lycopersicum | 684,775 | 218,205 | 27,102 | 24,627 |
| Chromosome | Species | |
|---|---|---|
| S. tuberosum | S. lycopersicum | |
| 1 | 259,267 | 237,496 |
| 2 | 200,827 | 90,558 |
| 3 | 205,561 | 92,045 |
| 4 | 176,113 | 78,104 |
| 5 | 133,442 | 59,508 |
| 6 | 164,694 | 72,583 |
| 7 | 152,942 | 67,774 |
| 8 | 140,397 | 62,569 |
| 9 | 148,610 | 64,651 |
| 10 | 130,071 | 58,183 |
| 11 | 133,138 | 59,040 |
| 12 | 127,595 | 54,634 |
| Statistics | A21 | A23 | Total SNVs |
|---|---|---|---|
| CRTISO | 233 | 245 | 478 |
| NCED1 | 174 | 198 | 372 |
| Lcy-e | 91 | 103 | 194 |
| NSY | 79 | 85 | 164 |
| OR | 139 | 201 | 340 |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Pacheco, J.; Vilanova, S.; Grillo-Risco, R.; García-García, F.; Prohens, J.; Gramazio, P. De novo Transcriptome Assembly and Comprehensive Annotation of Two Tree Tomato Cultivars (Solanum betaceum Cav.) with Different Fruit Color. Horticulturae 2021, 7, 431. https://doi.org/10.3390/horticulturae7110431
Pacheco J, Vilanova S, Grillo-Risco R, García-García F, Prohens J, Gramazio P. De novo Transcriptome Assembly and Comprehensive Annotation of Two Tree Tomato Cultivars (Solanum betaceum Cav.) with Different Fruit Color. Horticulturae. 2021; 7(11):431. https://doi.org/10.3390/horticulturae7110431
Chicago/Turabian StylePacheco, Juan, Santiago Vilanova, Rubén Grillo-Risco, Francisco García-García, Jaime Prohens, and Pietro Gramazio. 2021. "De novo Transcriptome Assembly and Comprehensive Annotation of Two Tree Tomato Cultivars (Solanum betaceum Cav.) with Different Fruit Color" Horticulturae 7, no. 11: 431. https://doi.org/10.3390/horticulturae7110431
APA StylePacheco, J., Vilanova, S., Grillo-Risco, R., García-García, F., Prohens, J., & Gramazio, P. (2021). De novo Transcriptome Assembly and Comprehensive Annotation of Two Tree Tomato Cultivars (Solanum betaceum Cav.) with Different Fruit Color. Horticulturae, 7(11), 431. https://doi.org/10.3390/horticulturae7110431








