Insight into the Phylogenetic Relationships and Evolutionary History of Indocalamus (Bambusoideae) Through Comparative Analyses of Plastomes
Abstract
1. Introduction
2. Materials and Methods
2.1. Sampling, DNA Extraction, Sequencing, Assembly, and Annotation
2.2. Phylogenetic Analyses
2.3. Estimation of Divergence Time
2.4. Plastome Structure Comparison and Sequence Divergence Analyses
2.5. Codon Usage Analyses
2.6. Repeats and SSR Analyses
3. Results
3.1. Sampling, Sequencing, and Assembly
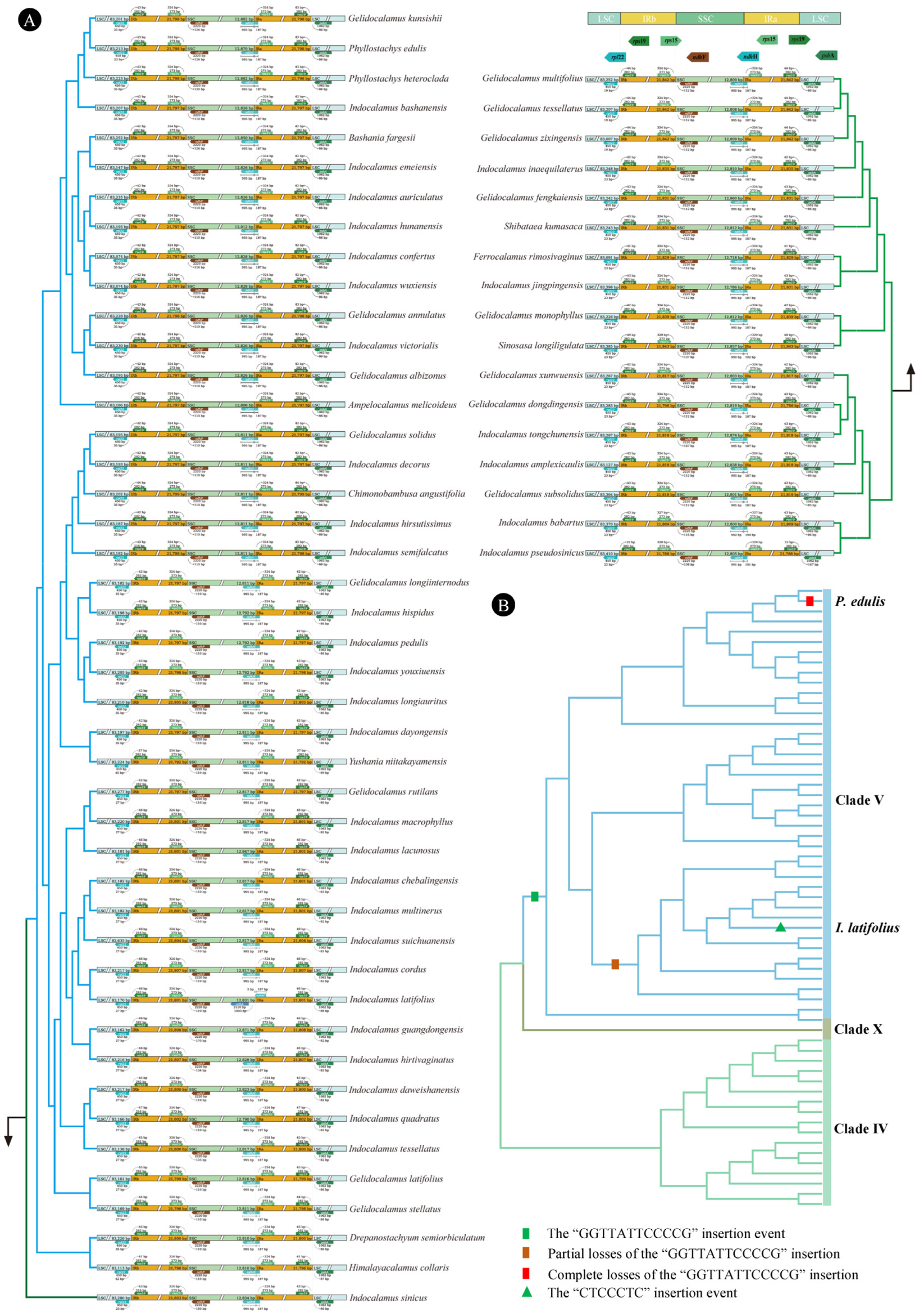
3.2. Phylogenetic Reconstruction
3.3. Divergence Time Estimation


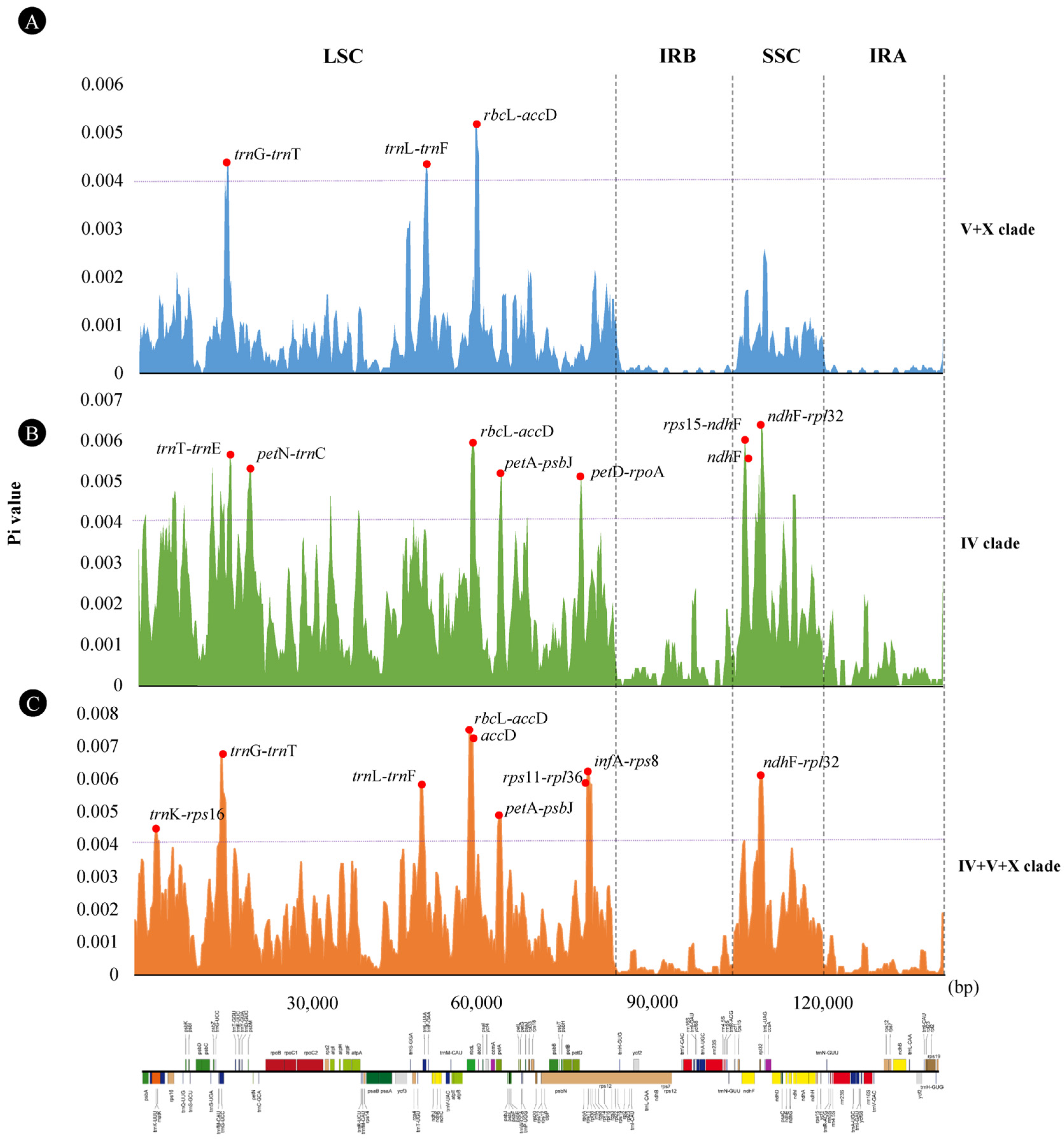
3.4. Genome Structural Variations and Sequence Divergences
3.5. Codon Usage Bias
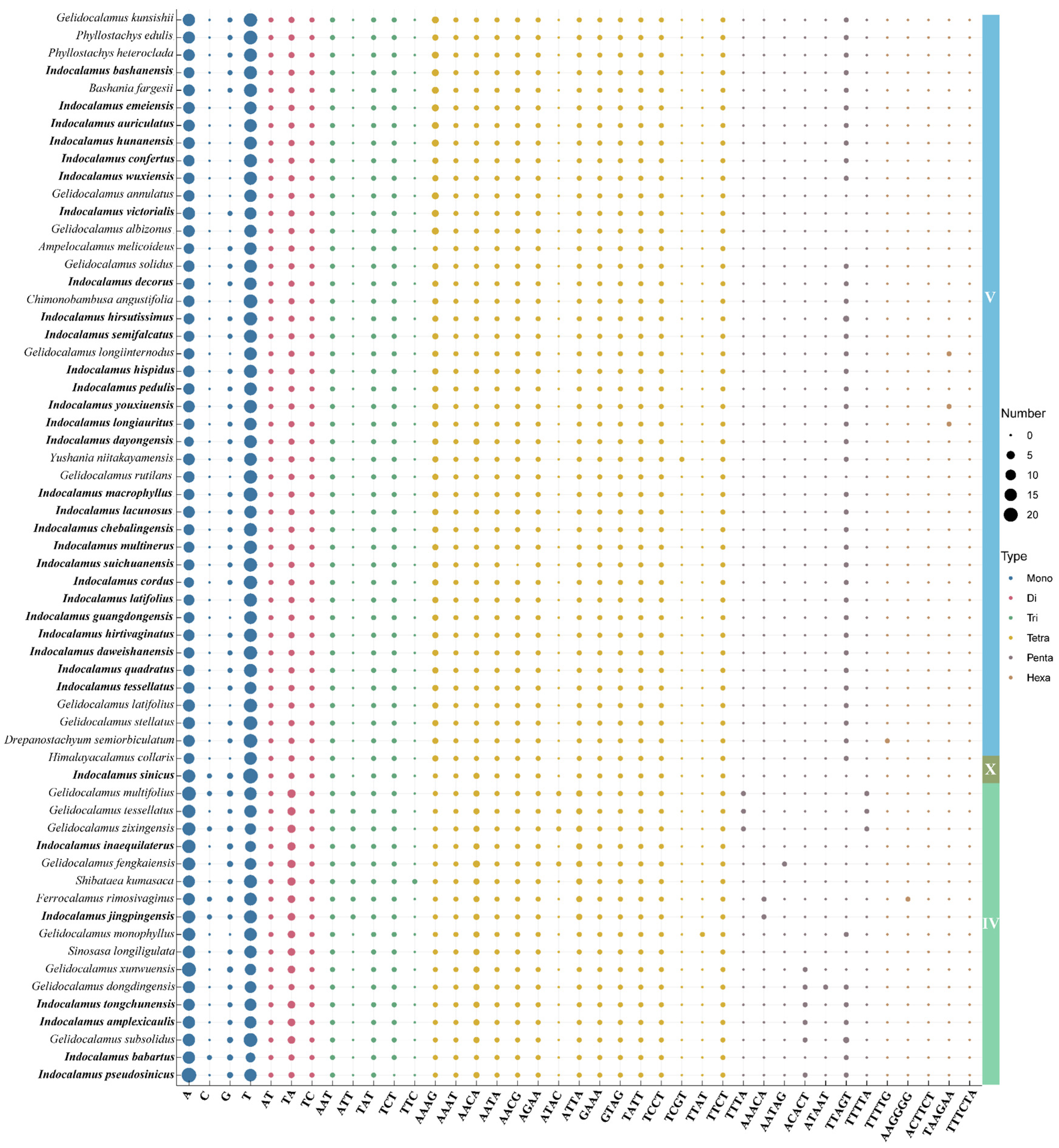
3.6. SSRs and Long Repeats Analysis
4. Discussion
4.1. Re-Evaluating the Circumscription of Indocalamus
4.2. Rapid Adaptive Radiation Evolution of Indocalamus and Its Related Genera
4.3. Evolution of the Plastomes and DNA Barcode Identification in Indocalamus
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Janzen, D.H. Why bamboos wait so long to flower. Annu. Rev. Ecol. Syst. 1976, 7, 347–391. [Google Scholar] [CrossRef]
- Zheng, X.; Lin, S.; Fu, H.; Wan, Y.; Ding, Y. The Bamboo Flowering Cycle Sheds Light on Flowering Diversity. Front. Plant Sci. 2020, 11, 381. [Google Scholar] [CrossRef]
- Soreng, R.J.; Peterson, P.M.; Romaschenko, K.; Davidse, G.; Zuloaga, F.O.; Judziewicz, E.J.; Filgueiras, T.S.; Davis, J.I.; Morrone, O. A Worldwide Phylogenetic Classification of the Poaceae (Gramineae). J. Syst. Evol. 2015, 53, 117–137. [Google Scholar] [CrossRef]
- Peng, S.; Yang, H.Q.; Li, D.Z. Highly heterogeneous generic delimitation within the temperate bamboo clade (Poaceae: Bambusoideae): Evidence from GBSSI and ITS sequences. Taxon 2008, 57, 799–810. [Google Scholar] [CrossRef]
- Zeng, C.X.; Zhang, Y.X.; Triplett, J.K.; Yang, J.B.; Li, D.Z. Large multi-locus plastid phylogeny of the tribe Arundinarieae (Poaceae: Bambusoideae) reveals ten major lineages and low rate of molecular divergence. Mol. Phylogenetics Evol. 2010, 56, 821–839. [Google Scholar] [CrossRef] [PubMed]
- Ma, P.F.; Vorontsova, M.S.; Nanjarisoa, O.P.; Razanatsoa, J.; Guo, Z.H.; Haevermans, T.; Li, D.Z. Negative correlation between rates of molecular evolution and flowering cycles in temperate woody bamboos revealed by plastid phylogenomics. BMC Plant Biol. 2017, 17, 260. [Google Scholar] [CrossRef] [PubMed]
- Zhou, Y.; Li, W.W.; Zhang, Y.Q.; Xing, X.C.; Zhang, J.Q.; Ren, Y. Extensive reticulate evolution within Fargesia (s.l.) (Bambusoideae: Poaceae) and its allies: Evidence from multiple nuclear markers. Mol. Phylogenetics Evol. 2020, 149, 106842. [Google Scholar] [CrossRef]
- Nakai, T. Two new genera of Bambusaceae, with special remarks on the related genera growing in eastern Asia. J. Arnold Arbor. 1925, 6, 145–153. [Google Scholar]
- Chao, Q.S.; Chu, Z.D.; Hsiung, W.Y. A revision of some genera and species of Chinese bamboos. Acta Phytotax. Sin 1980, 18, 20–36. [Google Scholar]
- Yang, Y.L. A revision of the genus Indocalamus of Bambusoideae from the world (I). J. Nanjing Univ. Nat. Sci. 1987, 23, 453–462. [Google Scholar]
- Yang, Y.L. The endemic genus of China: Geographical distribution of Indocalamus Nakai. J. Plant Resour. Environ. 1993, 2, 41–44. [Google Scholar]
- Wang, Z.P.; Stapleton, C. Indocalamus. In Flora of China; Wu, Z.Y., Raven, P.H., Hong, D.Y., Eds.; Science Press: Beijing, China; The Missouri Botanical Garden Press: St. Louis, MO, USA, 2006; Volume 22, pp. 135–143. [Google Scholar]
- Vorontsova, M.S.; Clark, L.G.; Dransfield, J.; Govaerts, R.; Baker, W.J. World Checklist of Bamboos and Rattans; Science Press: Beijing, China, 2016; pp. 106–110. [Google Scholar]
- Zhao, H.R.; Yang, Y.L. Bambusoideae. In Flora Reipublicae Popularis Sinicae; Keng, P.C., Wang, Z.P., Eds.; Science Press: Beijing, China, 1996; Volume 9, pp. 676–704. [Google Scholar]
- Yi, T.P.; Yang, L. A new species of Indocalamus Nakai from western Sichuan of China. J. Bamboo Res. 2004, 23, 13–15. [Google Scholar]
- Keng, P.C.; Hsueh, J.R. Ferrocalamus Hsueh & Keng f., a new bamboo genus in China. J. Bamboo Res. 1982, 1, 137–139. [Google Scholar]
- Yang, Y.L.; Zhao, H.R. Taxonomic Revision of Species of Indocalamus (Bambusoideae) in the World (II). J. Nanjing Univ. Nat. Sci. 1990, 26, 282–290. [Google Scholar]
- Qin, Q.M.; Tong, Y.H.; Zheng, X.R.; Ni, J.B.; Xia, N.H. Sinosasa (Poaceae: Bambusoideae), a new genus from China. Taxon 2020, 70, 27–47. [Google Scholar] [CrossRef]
- Hodkinson, T.R.; Ní Chonghaile, G.; Sungkaew, S.; Chase, M.W.; Salamin, N.; Stapleton, C.M.A. Phylogenetic analyses of plastid and nuclear DNA sequences indicate a rapid late Miocene radiation of the temperate bamboo tribe Arundinarieae (Poaceae, Bambusoideae). Plant Ecol. Divers. 2010, 3, 109–120. [Google Scholar] [CrossRef]
- Attigala, L.; Triplett, J.K.; Kathriarachchi, H.S.; Clark, L.G. A new genus and a major temperate bamboo lineage of the Arundinarieae (Poaceae: Bambusoideae) from Sri Lanka based on a multi-locus plastid phylogeny. Phytotaxa 2014, 174, 187–205. [Google Scholar] [CrossRef]
- Triplett, J.K.; Clark, L.G. Phylogeny of the temperate Bamboos (Poaceae: Bambusoideae: Bambuseae) with an emphasis on Arundinaria and allies. Syst. Biol. 2010, 35, 102–120. [Google Scholar] [CrossRef]
- Zhang, Y.X.; Zeng, C.X.; Li, D.Z. Complex evolution in Arundinarieae (Poaceae: Bambusoideae): Incongruence between plastid and nuclear GBSSI gene phylogenies. Mol. Phylogenetics Evol. 2012, 63, 777–797. [Google Scholar] [CrossRef]
- Yang, H.M.; Zhang, Y.X.; Yang, J.B.; Li, D.Z. The monophyly of Chimonocalamus and conflicting gene trees in Arundinarieae (Poaceae: Bambusoideae) inferred from four plastid and two nuclear markers. Mol. Phylogenetics Evol. 2013, 68, 340–356. [Google Scholar] [CrossRef]
- Ma, P.F.; Zhang, Y.X.; Zeng, C.X.; Guo, Z.H.; Li, D.Z. Chloroplast phylogenomic analyses resolve deep-level relationships of an intractable bamboo tribe Arundinarieae (Poaceae). Syst. Biol. 2014, 63, 933–950. [Google Scholar] [CrossRef]
- Zhang, Y.X.; Ma, P.F.; Li, D.Z. A new genus of temperate woody bamboos (Poaceae, Bambusoideae, Arundinarieae) from a limestone montane area of China. PhytoKeys 2018, 109, 67–76. [Google Scholar] [CrossRef] [PubMed]
- Zhang, Y.X.; Guo, C.; Li, D.Z. A new subtribal classification of Arundinarieae (Poaceae, Bambusoideae) with the description of a new genus. Plant Divers. 2020, 42, 127–134. [Google Scholar] [CrossRef] [PubMed]
- Guo, C.; Ma, P.F.; Yang, G.Q.; Ye, X.Y.; Guo, Y.; Liu, J.X.; Liu, Y.L.; Eaton, D.A.R.; Guo, Z.H.; Li, D.Z. Parallel ddRAD and genome skimming analyses reveal a radiative and reticulate evolutionary history of the temperate bamboos. Syst. Biol. 2021, 70, 756–773. [Google Scholar] [CrossRef] [PubMed]
- Murray, M.G.; Thompson, W.F. Rapid isolation of high molecular weight plant DNA. Nucleic Acids Res. 1980, 8, 4321–4326. [Google Scholar] [CrossRef]
- Chen, S.F.; Zhou, Y.Q.; Chen, Y.R.; Gu, J. fastp: An ultra-fast all-in-one fastq preprocessor. Bioinformatics 2018, 34, i884–i890. [Google Scholar] [CrossRef]
- Jin, J.J.; Yu, W.B.; Yang, J.B.; Song, Y.; dePamphilis, C.W.; Yi, T.S.; Li, D.Z. GetOrganelle: A fast and versatile toolkit for accurate de novo assembly of organelle genomes. Genome Biol. 2020, 21, 241. [Google Scholar] [CrossRef]
- Wick, R.R.; Schultz, M.B.; Zobel, J.; Holt, K.E. Bandage: Interactive visualization of de novo genome assemblies. Bioinformatics 2015, 31, 3350–3352. [Google Scholar] [CrossRef]
- Kearse, M.; Moir, R.; Wilson, A.; Stones-Havas, S.; Cheung, M.; Sturrock, S.; Buxton, S.; Cooper, A.; Markowitz, S.; Duran, C.; et al. Geneious Basic: An integrated and extendable desktop software platform for the organization and analysis of sequence data. Bioinformatics 2012, 28, 1647–1649. [Google Scholar] [CrossRef]
- Shi, L.; Chen, H.; Jiang, M.; Wang, L.; Wu, X.; Huang, L.; Liu, C. CPGAVAS2, an integrated plastome sequence annotator and analyzer. Nucleic Acids Res. 2019, 47, W65–W73. [Google Scholar] [CrossRef]
- Zheng, S.; Poczai, P.; Hyvönen, J.; Tang, J.; Amiryousefi, A. Chloroplot: An online program for the versatile plotting of organelle genomes. Front. Genet. 2020, 11, 576124. [Google Scholar] [CrossRef]
- Katoh, K.; Standley, D.M. MAFFT multiple sequence alignment software version 7: Improvements in performance and usability. Mol. Biol. Evol. 2013, 30, 772–780. [Google Scholar] [CrossRef] [PubMed]
- Capella-Gutiérrez, S.; Silla-Martínez, J.M.; Gabaldón, T. trimAl: A tool for automated alignment trimming in large-scale phylogenetic analyses. Bioinformatics 2009, 25, 1972–1973. [Google Scholar] [CrossRef] [PubMed]
- Nguyen, L.T.; Schmidt, H.A.; von Haeseler, A.; Minh, B.Q. IQ-TREE: A fast and effective stochastic algorithm for estimating maximum-likelihood phylogenies. Mol. Biol. Evol. 2015, 32, 268–274. [Google Scholar] [CrossRef] [PubMed]
- Kalyaanamoorthy, S.; Minh, B.Q.; Wong, T.K.F.; von Haeseler, A.; Jermiin, L.S. ModelFinder: Fast model selection for accurate phylogenetic estimates. Nat. Methods 2017, 14, 587–589. [Google Scholar] [CrossRef]
- Ronquist, F.; Teslenko, M.; van der Mark, P.; Ayres, D.L.; Darling, A.; Höhna, S.; Larget, B.; Liu, L.; Suchard, M.A.; Huelsenbeck, J.P. MrBayes 3.2: Efficient bayesian phylogenetic inference and model choice across a large model space. Syst. Biol. 2012, 61, 539–542. [Google Scholar] [CrossRef]
- Bouckaert, R.; Vaughan, T.G.; Barido-Sottani, J.; Duchêne, S.; Fourment, M.; Gavryushkina, A.; Heled, J.; Jones, G.; Kühnert, D.; De Maio, N.; et al. BEAST 2.5: An advanced software platform for bayesian evolutionary analysis. PLoS Comput. Biol. 2019, 15, e1006650. [Google Scholar] [CrossRef]
- Wang, C.K.; Guo, R.; Guo, C.C.; Yang, G.Y.; Zhang, W.G. Gelidocalamus zixingensis (Poaceae, Bambusoideae, Arundinarieae), a new species from southern China revealed by morphological and molecular evidence. PhytoKeys 2023, 218, 29–45. [Google Scholar] [CrossRef]
- Strömberg, C.A.E. Evolution of grasses and grassland ecosystems. Annu. Rev. Earth Planet. Sci. 2011, 39, 517–544. [Google Scholar] [CrossRef]
- Li, H.; Guo, Q.; Xu, L.; Gao, H.; Liu, L.; Zhou, X. CPJSdraw: Analysis and visualization of junction sites of chloroplast genomes. PeerJ 2023, 11, e15326. [Google Scholar] [CrossRef]
- Librado, P.; Rozas, J. DnaSP v5: A software for comprehensive analysis of DNA polymorphism data. Bioinformatics 2009, 25, 1451–1452. [Google Scholar] [CrossRef]
- Zhang, D.; Gao, F.; Jakovlić, I.; Zou, H.; Zhang, J.; Li, W.X.; Wang, G.T. PhyloSuite: An integrated and scalable desktop platform for streamlined molecular sequence data management and evolutionary phylogenetics studies. Mol. Ecol. Resour. 2019, 20, 348–355. [Google Scholar] [CrossRef] [PubMed]
- CondoW. Available online: https://codonw.sourceforge.net/culong.html#CodonW (accessed on 16 May 2025).
- Chen, C.J.; Chen, H.; Zhang, Y.; Thomas, H.R.; Frank, M.H.; He, Y.H.; Xia, R. TBtools: An integrative toolkit developed for interactive analyses of big biological data. Mol. Plant 2020, 13, 1194–1202. [Google Scholar] [CrossRef] [PubMed]
- Beier, S.; Thiel, T.; Münch, T.; Scholz, U.; Mascher, M.; Valencia, A. MISA-web: A web server for microsatellite prediction. Bioinformatics 2017, 33, 2583–2585. [Google Scholar] [CrossRef] [PubMed]
- Kurtz, S.; Choudhuri, J.V.; Ohlebusch, E.; Schleiermacher, C.; Stoye, J.; Giegerich, R. REPuter: The manifold applications of repeat analysis on a genomic scale. Nucleic Acids Res. 2001, 29, 4633–4642. [Google Scholar] [CrossRef]
- Li, Y.L.; Guo, R.; Zhang, H.J.; Yi, S.R.; Yang, G.Y.; Zhang, W.G. Gelidocalamus albozonatus (Poaceae, Bambusoideae), a new species from the southeast of Chongqing, China, and analysis of the morphological diversity in the core group of Gelidocalamus. PhytoKeys 2023, 236, 17–27. [Google Scholar] [CrossRef]
- Zhang, X.Z.; Zeng, C.X.; Ma, P.F.; Haevermans, T.; Zhang, Y.X.; Zhang, L.N.; Guo, Z.H.; Li, D.Z. Multi-locus plastid phylogenetic biogeography supports the Asian hypothesis of the temperate woody bamboos (Poaceae: Bambusoideae). Mol. Phylogenetics Evol. 2016, 96, 118–129. [Google Scholar] [CrossRef]
- Christin, P.A.; Spriggs, E.; Osborne, C.P.; Stromberg, C.A.E.; Salamin, N.; Edwards, E.J. Molecular dating, evolutionary rates, and the age of the grasses. Syst. Biol. 2013, 63, 153–165. [Google Scholar] [CrossRef]
- Bouchenak-Khelladi, Y.; Verboom, G.A.; Savolainen, V.; Hodkinson, T.R. Biogeography of the grasses (Poaceae): A phylogenetic approach to reveal evolutionary history in geographical space and geological time. Bot. J. Linn. Soc. 2010, 162, 543–557. [Google Scholar] [CrossRef]
- Janssens, S.B.; Knox, E.B.; Huysmans, S.; Smets, E.F.; Merckx, V.S.F.T. Rapid radiation of Impatiens (Balsaminaceae) during Pliocene and Pleistocene: Result of a global climate change. Mol. Phylogenetics Evol. 2009, 52, 806–824. [Google Scholar] [CrossRef]
- Hampe, A.; Jump, A.S. Climate relicts: Past, present, future. Annu. Rev. Ecol. Evol. Syst. 2011, 42, 313–333. [Google Scholar] [CrossRef]
- Zou, X.H.; Yang, Z.; Doyle, J.J.; Ge, S. Multilocus estimation of divergence times and ancestral effective population sizes of Oryza species and implications for the rapid diversification of the genus. New Phytol. 2013, 198, 1155–1164. [Google Scholar] [CrossRef] [PubMed]
- Zhisheng, A.; Kutzbach, J.E.; Prell, W.L.; Porter, S.C. Evolution of Asian monsoons and phased uplift of the Himalaya—Tibetan plateau since Late Miocene times. Nature 2001, 411, 62–66. [Google Scholar] [CrossRef] [PubMed]
- Zheng, H.B.; Powell, C.M.; Rea, D.K.; Wang, J.L.; Wang, P.X. Late Miocene and mid-Pliocene enhancement of the East Asian monsoon as viewed from the land and sea. Glob. Planet. Change 2004, 41, 147–155. [Google Scholar] [CrossRef]
- Guo, C.; Guo, Z.H.; Li, D.Z. Phylogenomic analyses reveal intractable evolutionary history of a temperate bamboo genus (Poaceae: Bambusoideae). Plant Divers. 2019, 41, 213–219. [Google Scholar] [CrossRef]
- Verma, D.; Daniell, H. Chloroplast vector systems for biotechnology applications. Plant Physiol. 2007, 145, 1129–1143. [Google Scholar] [CrossRef]
- Li, Y.H.; Ren, Y.K.; Zhao, X.H.; Liu, J.; Han, B.; Wang, C.B. Research progress on chloroplast genome of major Gramineous crops. Biotechnol. Bull. 2020, 36, 112–121. [Google Scholar] [CrossRef]
- Tu, X.D.; Xin, Y.X.; Fu, H.H.; Zhou, C.Y.; Liu, Q.L.; Tang, X.H.; Zou, L.H.; Liu, Z.J.; Chen, S.P.; Lin, W.J.; et al. The complete mitochondrial genome of Castanopsis carlesii and Castanea henryi reveals the rearrangement and size differences of mitochondrial DNA molecules. BMC Plant Biol. 2024, 24, 988. [Google Scholar] [CrossRef]
- Asaf, S.; Khan, A.L.; Khan, M.A.; Shahzad, R.; Lubna; Kang, S.M.; Al-Harrasi, A.; Al-Rawahi, A.; Lee, I.J. Complete chloroplast genome sequence and comparative analysis of loblolly pine (Pinus taeda L.) with related species. PLoS ONE 2018, 13, e0192966. [Google Scholar] [CrossRef]
- Li, J.P.; Qin, Z.; Guo, C.C.; Zhang, W.G.; Yang, G.Y. Codon bias in the chloroplast genome of Gelidocalamus tessellatus. J. Bamboo Res. 2019, 38, 79–87. [Google Scholar] [CrossRef]
- Zhang, Y.J.; Li, D.Z. Advances in phylogenomics based on complete chloroplast genomes. Plant Divers. 2011, 33, 365–375. [Google Scholar]
- Cai, Z.M.; Zhang, Y.X.; Zhang, L.N.; Li, D.Z. Testing four candidate barcoding markers in temperate woody bamboos (Poaceae: Bambusoideae). J. Syst. Evol. 2012, 50, 527–539. [Google Scholar] [CrossRef]
- Zhang, Y.X.; Xu, Y.X.; Ma, P.F.; Li, D.Z. Screening of chloroplast DNA barcodes in Bambusoideae. J. Plant Taxon. Resour. 2013, 35, 743–750. [Google Scholar]
- Meena, R.K.; Negi, N.; Uniyal, N.; Shamoon, A.; Bhandari, M.S.; Pandey, S.; Negi, R.K.; Sharma, R.; Ginwal, H.S. Chloroplast-based DNA barcode analysis indicates high discriminatory potential of matK locus in Himalayan temperate bamboos. 3 Biotech 2020, 10, 534. [Google Scholar] [CrossRef] [PubMed]
- Ji, Y.; Liu, C.; Yang, Z.; Yang, L.; He, Z.; Wang, H.; Yang, J.; Yi, T. Testing and using complete plastomes and ribosomal DNA sequences as the next generation DNA barcodes in Panax (Araliaceae). Mol. Ecol. Resour. 2019, 19, 1333–1445. [Google Scholar] [CrossRef] [PubMed]
- Dev, S.A.; Sijimol, K.; Prathibha, P.S.; Sreekumar, V.B.; Muralidharan, E.M. DNA barcoding as a valuable molecular tool for the certification of planting materials in bamboo. 3 Biotech 2020, 10, 59. [Google Scholar] [CrossRef]
- Fu, C.N.; Mo, Z.Q.; Yang, J.B.; Cai, J.; Ye, L.J.; Zou, J.Y.; Qin, H.T.; Zheng, W.; Hollingsworth, P.M.; Li, D.Z.; et al. Testing genome skimming for species discrimination in the large and taxonomically difficult genus Rhododendron. Mol. Ecol. Resour. 2022, 22, 404–414. [Google Scholar] [CrossRef]
- Lv, S.Y.; Ye, X.Y.; Li, Z.H.; Ma, P.F.; Li, D.Z. Testing complete plastomes and nuclear ribosomal DNA sequences for species identification in a taxonomically difficult bamboo genus Fargesia. Plant Divers. 2022, 45, 147–155. [Google Scholar] [CrossRef]


Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2025 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Wang, C.; Li, Y.; Cui, L.; Wang, J.; Wang, M.; Guo, C.; Yang, G.; Gao, L.; Zhang, W. Insight into the Phylogenetic Relationships and Evolutionary History of Indocalamus (Bambusoideae) Through Comparative Analyses of Plastomes. Horticulturae 2025, 11, 1018. https://doi.org/10.3390/horticulturae11091018
Wang C, Li Y, Cui L, Wang J, Wang M, Guo C, Yang G, Gao L, Zhang W. Insight into the Phylogenetic Relationships and Evolutionary History of Indocalamus (Bambusoideae) Through Comparative Analyses of Plastomes. Horticulturae. 2025; 11(9):1018. https://doi.org/10.3390/horticulturae11091018
Chicago/Turabian StyleWang, Chengkun, Yonglong Li, Ling Cui, Jianqing Wang, Meixia Wang, Chunce Guo, Guangyao Yang, Liqin Gao, and Wengen Zhang. 2025. "Insight into the Phylogenetic Relationships and Evolutionary History of Indocalamus (Bambusoideae) Through Comparative Analyses of Plastomes" Horticulturae 11, no. 9: 1018. https://doi.org/10.3390/horticulturae11091018
APA StyleWang, C., Li, Y., Cui, L., Wang, J., Wang, M., Guo, C., Yang, G., Gao, L., & Zhang, W. (2025). Insight into the Phylogenetic Relationships and Evolutionary History of Indocalamus (Bambusoideae) Through Comparative Analyses of Plastomes. Horticulturae, 11(9), 1018. https://doi.org/10.3390/horticulturae11091018




