Reference Genes in Plant–Pathogen Interaction: A Bibliometric Analysis
Abstract
1. Introduction
2. Relevant Studies on Reference Gene Stability for RT-qPCR Normalization in Plant Pathogen Interactions
2.1. Main Keywords in the Analyzed Studies
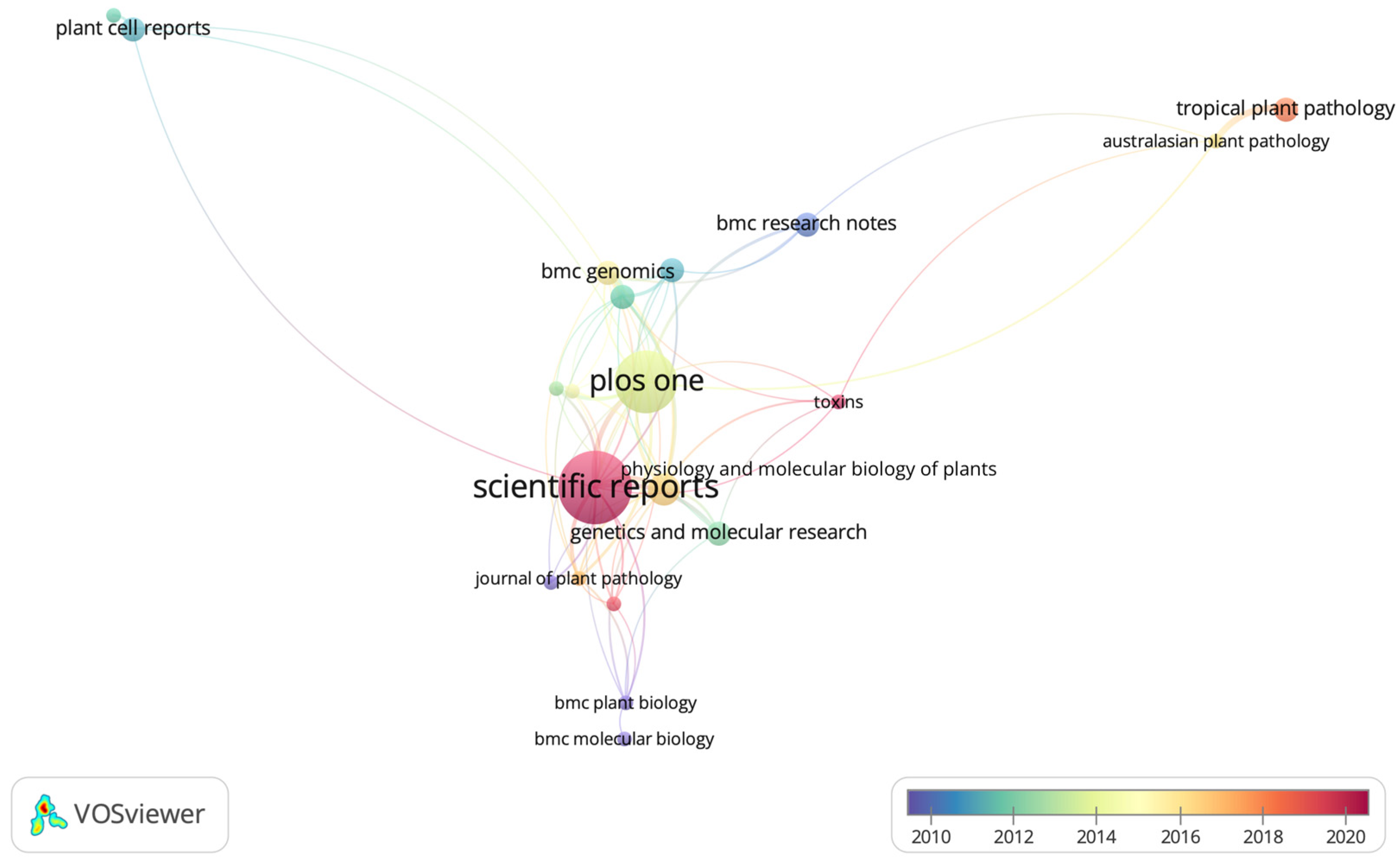
2.2. Network Visualization of Academic Journals in the Analyzed Studies
3. Number of Reference Genes per Study
4. Software Packages Used for Normalization Strategy
5. Most Used and Stable Reference Genes
6. Conclusions
Supplementary Materials
Author Contributions
Funding
Data Availability Statement
Conflicts of Interest
References
- Kumari, R.; Kumar, A.; Kumar, P.; Jakhar, D.S. Advances in plant-pathogen interactions in terms of biochemical and molecular aspects. In Plant-Microbe Interaction-Recent Advances in Molecular and Biochemical Approaches; Swapnil, P., Meena, M., Harish, Marwal, A., Vijayalakshmi, S., Zehra, A., Eds.; Elsevier Inc.: Cambridge, UK, 2023; Volume 1, pp. 111–122. [Google Scholar] [CrossRef]
- Saraiva, M.; Ściślak, M.E.; Ascurra, Y.T.; Ferrando, T.M.; Zic, N.; Henard, C.; van West, P.; Trusch, F.; Vleeshouwers, V.G. The molecular dialog between oomycete effectors and their plant and animal hosts. Fungal Biol. Rev. 2023, 43, 100289. [Google Scholar] [CrossRef]
- Wieczorek, K.; Denis, E.; Lynch, O.; and Osek, J. Molecular characterization and antibiotic resistance profiling of Campylobacter isolated from cattle in polish slaughterhouses. Food Microbiol. 2013, 34, 130–136. [Google Scholar] [CrossRef]
- Boava, L.P.; Laia, M.L.; Jacob, T.R.; Dabbas, K.M.; Goncalves, J.F.; Ferro, J.A.; Ferro, M.I.; and Furtado, E.L. Selection of endogenous genes for gene expression studiesin Eucalyptus under biotic (Puccinia psidii) and abiotic (acibenzolar-S-methyl) stresses using RT-qPCR. BMC Res. Notes 2010, 3, 43. [Google Scholar] [CrossRef]
- Kong, Q.; Yuan, J.; Gao, L.; Zhao, S.; Jiang, W.; Huang, Y.; Bie, Z. Identification of suitable reference genes for gene expression normalization in qRT-PCR analysis in watermelon. PLoS ONE 2014, 9, e90612. [Google Scholar] [CrossRef]
- Petriccione, M.; Mastrobuoni, F.; Zampella, L.; Scortichini, M. Reference gene selection for normalization of RT-qPCR gene expression data from Actinidia deliciosa leaves infected with Pseudomonas syringae pv. actinidiae. Sci. Rep. 2015, 5, 16961. [Google Scholar] [CrossRef] [PubMed]
- Bunnik, E.M.; Le Roch, K.G. An introduction to functional genomics and systems biology. Adv. Wound Care 2013, 2, 490–498. [Google Scholar] [CrossRef]
- Pfaffl, M.W. Relative quantification. In Real-Time PCR; Dorak, M.T., Ed.; Taylor & Francis: New York, NY, USA, 2006; pp. 63–82. [Google Scholar]
- Kundu, A.; Patel, A.; Pal, A. Defining reference genes for qPCR normalization to study biotic and abiotic stress responses in Vigna mungo. Plant Cell Rep. 2013, 32, 1647–1658. [Google Scholar] [CrossRef]
- Petriccione, M.; Magri, A.; Gaudiano, M.; Scortichini, M. Selection and validation of reference genes for gene expression studies in Xanthomonas arboricola pv. juglandis subjected to abiotic stress. Plant Pathol. 2021, 70, 1455–1466. [Google Scholar] [CrossRef]
- Chapman, J.R.; Waldenström, J. With reference to reference genes: A systematic review of endogenous controls in gene expression studies. PLoS ONE 2015, 10, e0141853. [Google Scholar] [CrossRef]
- Lal, P.; Tiwari, R.K.; Kumar, A.; Altaf, M.A.; Alsahli, A.A.; Lal, M.K.; Kumar, R. Bibliometric analysis of real-time PCR-based pathogen detection in plant protection research: A comprehensive study. Front. Plant Sci. 2023, 14, 1129714. [Google Scholar] [CrossRef] [PubMed]
- Derveaux, S.; Vandesompele, J.; Hellemans, J. How to do successful gene expression analysis using real-time PCR. Methods 2010, 50, 227–230. [Google Scholar] [CrossRef] [PubMed]
- Orrego, A.; Gavilán, M.C.; Arévalos, A.; Ortíz, B.; Gaete Humada, B.; Pineda-Fretez, A.; Romero-Rodríguez, M.C.; Flores Giubi, M.E.; Kohli, M.M.; Iehisa, J.C.M. Identification of reference genes and their validation for gene expression analysis in phytopathogenic fungus Macrophomina phaseolina. PLoS ONE 2022, 17, e0272603. [Google Scholar] [CrossRef]
- Bustin, S.A.; Benes, V.; Garson, J.A.; Hellemans, J.; Huggett, J.; Kubista, M.; Mueller, R.; Nolan, T.; Pfaffl, M.W.; Shipley, G.L.; et al. The MIQE guidelines: Minimum information for publication of quantitative real-time PCR experiments. Clin. Chem. 2009, 55, 611–622. [Google Scholar] [CrossRef] [PubMed]
- Bustin, S.A.; Benes, V.; Garson, J.; Hellemans, J.; Huggett, J.; Kubista, M.; Mueller, R.; Nolan, T.; Pfaffl, M.W.; Shipley, G.; et al. The need for transparency and good practices in the qPCR literature. Nat. Methods 2013, 10, 1063–1067. [Google Scholar] [CrossRef]
- Selim, M.; Legay, S.; Berkelmann-Löhnertz, B.; Langen, G.; Kogel, K.H.; Evers, D. Identification of suitable reference genes for real-time RT-PCR normalization in the grapevine-downy mildew pathosystem. Plant Cell Rep. 2012, 31, 205–216. [Google Scholar] [CrossRef] [PubMed]
- Radonić, A.; Thulke, S.; Mackay, I.M.; Landt, O.; Siegert, W.; Nitsche, A. Guideline to reference gene selection for quantitative real-time PCR. Biochem. Biophys. Res. Commun. 2004, 313, 856–862. [Google Scholar] [CrossRef]
- Kalushkov, P.; and Hodek, I. The effects of thirteen species of aphids on some life history parameters of the ladybird Coccinella septempunctata. Biol. Control 2004, 49, 21–32. [Google Scholar] [CrossRef]
- Couto, D.; Zipfel, C. Regulation of pattern recognition receptor signalling in plants. Nat. Rev. Immunol. 2016, 16, 537–552. [Google Scholar] [CrossRef]
- Tang, Y.; Bao, X.; Liu, K.; Wang, J.; Zhang, J.; Feng, Y.; Wang, Y.; Lin, L.; Feng, J.; Li, C. Genome-wide identification and expression profiling of the auxin response factor (ARF) gene family in physic nut. PLoS ONE 2018, 13, e0201024. [Google Scholar] [CrossRef]
- Andersen, E.J.; Ali, S.; Byamukama, E.; Yen, Y.; Nepal, M.P. Disease resistance mechanisms in plants. Genes 2018, 9, 339. [Google Scholar] [CrossRef]
- Wang, M.R.; Bi, W.L.; Bettoni, J.C.; Zhang, D.; Volk, G.M.; Wang, Q.C. Shoot tip cryotherapy for plant pathogen eradication. Plant Pathol. 2022, 71, 1241–1254. [Google Scholar] [CrossRef]
- Backes, A.; Charton, S.; Planchon, S.; Esmaeel, Q.; Sergeant, K.; Hausman, J.F.; Renaut, J.; Barka, E.A.; Jacquard, C.; Guerriero, G. Gene expression and metabolite analysis in barley inoculated with net blotch fungus and plant growth-promoting rhizobacteria. Plant Physiol. Biochem. 2021, 168, 488–500. [Google Scholar] [CrossRef]
- Pandit, M.A.; Kumar, J.; Gulati, S.; Bhandari, N.; Mehta, P.; Katyal, R.; Rawat, C.D.; Mishra, V.; Kaur, J. Major Biological Control Strategies for Plant Pathogens. Pathogens 2022, 1, 273. [Google Scholar] [CrossRef]
- Sowa, S.; Toporowska, J.; Paczos-Grzęda, E. Identification of reference genes for gene expression assessment in Avena sativa under biotic stress triggered by Blumeria graminis. Sci. Rep. 2024, 14, 30006. [Google Scholar] [CrossRef] [PubMed]
- Sowa, S.; Sozoniuk, M.; Toporowska, J.; Kowalczyk, K.; Paczos-Grzęda, E. Reference genes expression stability in Avena sativa L. during compatible and incompatible interactions with Puccinia graminis. Sci. Rep. 2022, 12, 18369. [Google Scholar] [CrossRef]
- Sowa, S.; Sozoniuk, M.; Toporowska, J.; Kowalczyk, K.; Paczos-Grzęda, E. Validation of reference genes as an internal control for studying Avena sativa-Puccinia coronata interaction by RT-qPCR. Sci. Rep. 2022, 12, 14601. [Google Scholar] [CrossRef] [PubMed]
- Zhu, H.; Ma, Y.; Guo, Q. Expression stability of internal reference gene in response to Trichoderma polysporum infection in Avena fatua L. Curr. Genet. 2021, 67, 909–918. [Google Scholar] [CrossRef]
- Qi, J.; Yu, S.; Zhang, F.; Shen, X.; Zhao, X.; Yu, Y.; Zhang, D. Reference gene selection for real-time quantitative polymerase chain reaction of mRNA transcript levels in Chinese cabbage (Brassica rapa L. ssp. pekinensis). Plant Mol. Biol. Rep. 2010, 28, 597–604. [Google Scholar] [CrossRef]
- Zhu, X.; Li, X.; Chen, W.; Chen, J.; Lu, W.; Chen, L.; Fu, D. Evaluation of new reference genes in papaya for accurate transcript normalization under different experimental conditions. PLoS ONE 2012, 7, e44405. [Google Scholar] [CrossRef]
- Mafra, V.; Kubo, K.S.; Alves-Ferreira, M.; Ribeiro-Alves, M.; Stuart, R.M.; Boava, L.P.; Rodrigues, C.M.; Machado, M.A. Reference genes for accurate transcript normalization in citrus genotypes under different experimental conditions. PLoS ONE 2012, 7, e31263. [Google Scholar] [CrossRef]
- Maximo, H.J.; Dalio, R.J.; Rodrigues, C.M.; Breton, M.C.; Machado, M.A. Reference genes for RT-qPCR analysis in Citrus and Poncirus infected by zoospores of Phytophthora parasitica. Trop. Plant Path. 2017, 42, 76–85. [Google Scholar] [CrossRef]
- Figueiredo, A.; Loureiro, A.; Batista, D.; Monteiro, F.; Várzea, V.; Pais, M.S.; Gichuru, E.K.; Silva, M.C. Validation of reference genes for normalization of qPCR gene expression data from Coffea spp. hypocotyls inoculated with Colletotrichum kahawae. BMC Res. Notes 2013, 6, 388. [Google Scholar] [CrossRef]
- Barsalobres-Cavallari, C.F.; Severino, F.E.; Maluf, M.P.; Maia, I.G. Identification of suitable internal control genes for expression studies in Coffea arabica under different experimental conditions. BMC Mol. Biol. 2009, 10, 1. [Google Scholar] [CrossRef] [PubMed]
- Kwan, Y.M.; Meon, S.; Ho, C.L.; Wong, M.Y. Selection of reference genes for quantitative real-time PCR normalization in Ganoderma-infected oil palm (Elaeis guineensis) seedlings. Australas. Plant Pathol. 2016, 45, 261–268. [Google Scholar] [CrossRef]
- Olbrich, M.; Gerstner, E.; Welzl, G.; Fleischmann, F.; Oßwald, W.; Bahnweg, G.; Ernst, D. Quantification of mRNAs and housekeeping gene selection for quantitative real-time RT-PCR normalization in European beech (Fagus sylvatica L.) during abiotic and biotic stress. Z. Naturforsch. C 2008, 63, 574–582. [Google Scholar] [CrossRef][Green Version]
- Bansal, R.; Mittapelly, P.; Cassone, B.J.; Mamidala, P.; Redinbaugh, M.G.; Michel, A. Recommended reference genes for quantitative PCR analysis in soybean have variable stabilities during diverse biotic stresses. PLoS ONE 2015, 10, e0134890. [Google Scholar] [CrossRef] [PubMed]
- Hirschburger, D.; Müller, M.; Voegele, R.T.; Link, T. Reference genes in the pathosystem Phakopsora pachyrhizi/soybean suitable for normalization in transcript profiling. Int. J. Mol. Sci. 2015, 16, 23057–23075. [Google Scholar] [CrossRef] [PubMed]
- Štajner, N.; Cregeen, S.; Javornik, B. Evaluation of reference genes for RT-qPCR expression studies in hop (Humulus lupulus L.) during infection with vascular pathogen Verticillium albo-atrum. PLoS ONE 2013, 8, e68228. [Google Scholar] [CrossRef]
- Saha, G.C.; Vandemark, G.J. Stability of expression of reference genes among different lentil (Lens culinaris) genotypes subjected to cold stress, white mold disease, and Aphanomyces root rot. Plant Mol. Biol. Rep. 2013, 31, 1109–1115. [Google Scholar] [CrossRef]
- Prakash, H.; Shukla, P.; Ramesha, A.; Arunakumar, G.S.; Doss, S.G.; Ponnuvel, K.M. Evaluation of reference genes for accurate normalization of qPCR data under biotic stresses in mulberry (Morus indica L.). Sci. Hortic. 2024, 323, 112507. [Google Scholar] [CrossRef]
- Rego, E.C.S.; Pinheiro, T.D.M.; Antonino, J.D.; Alves, G.S.C.; Cotta, M.G.; Fonseca, F.C.D.A.; Miller, R.N.G. Stable reference genes for RT-qPCR analysis of gene expression in the Musa acuminata-Pseudocercospora musae interaction. Sci. Rep. 2019, 9, 14592. [Google Scholar] [CrossRef]
- de Castro Costa, É.; Bastos, L.S.; Gomes, T.G.; Miller, R.N.G. Reference genes for RT-qPCR analysis in Musa acuminata genotypes contrasting in resistance to Fusarium oxysporum f. sp. cubense subtropical race 4. Sci. Rep. 2024, 14, 16578. [Google Scholar] [CrossRef]
- Chen, L.; Zhong, H.Y.; Kuang, J.F.; Li, J.G.; Lu, W.J.; Chen, J.Y. Validation of reference genes for RT-qPCR studies of gene expression in banana fruit under different experimental conditions. Planta 2011, 234, 377–390. [Google Scholar] [CrossRef] [PubMed]
- Zhang, L.; Liu, L.; Cheng, P.; Shen, H.; Rong, B.; Liu, W.; Yu, G. Identification and validation of reference genes for RT-qPCR analysis in banana (Musa spp.) under Fusarium wilt resistance induction conditions. J. Phytopathol. 2017, 165, 746–754. [Google Scholar] [CrossRef]
- Hashemipetroudi, S.H.; Ghorbani, H.; Rostami, M.; Rezaei, A.; Goodwin, P.H.; Kuhlmann, M. Selection of reference genes for RT-qPCR analysis of rice with Rhizoctonia solani infection and biocontrol PGPR/KSi application. Mol. Biol. Rep. 2023, 50, 4225–4237. [Google Scholar] [CrossRef] [PubMed]
- Bevitori, R.; Oliveira, M.B.; Grossi-de-Sá, M.F.; Lanna, A.C.; da Silveira, R.D.; Petrofeza, S. Selection of optimized candidate reference genes for qRT-PCR normalization in rice (Oryza sativa L.) during Magnaporthe oryzae infection and drought. Genet. Mol. Res. 2014, 13, 9795–9805. [Google Scholar] [CrossRef]
- Narsai, R.; Ivanova, A.; Ng, S.; Whelan, J. Defining reference genes in Oryza sativa using organ, development, biotic and abiotic transcriptome datasets. BMC Plant Biol. 2010, 10, 56. [Google Scholar] [CrossRef]
- Borges, A.; Tsai, S.M.; Caldas, D.G.G. Validation of reference genes for RT-qPCR normalization in common bean during biotic and abiotic stresses. Plant Cell Rep. 2012, 31, 827–838. [Google Scholar] [CrossRef]
- Umadevi, P.; Suraby, E.J.; Anandaraj, M.; Nepolean, T. Identification of stable reference gene for transcript normalization in black pepper-Phytophthora capsici pathosystem. Physiol. Mol. Biol. Plants 2019, 25, 945–952. [Google Scholar] [CrossRef]
- Saha, G.C.; Vandemark, G.J. Evaluation of expression stability of candidate references genes among green and yellow pea cultivars (Pisum sativum L.) subjected to abiotic and biotic stress. Am. J. Plant Sci. 2012, 3, 235–242. [Google Scholar] [CrossRef][Green Version]
- Klie, M.; Debener, T. Identification of superior reference genes for data normalisation of expression studies via quantitative PCR in hybrid roses (Rosa hybrida). BMC Res. Notes 2011, 4, 518. [Google Scholar] [CrossRef]
- Huang, N.; Ling, H.; Liu, F.; Su, Y.; Su, W.; Mao, H.; Zhang, X.; Wang, L.; Chen, R.; Que, Y. Identification and evaluation of PCR reference genes for host and pathogen in sugarcane-Sporisorium scitamineum interaction system. BMC Genom. 2018, 19, 479. [Google Scholar] [CrossRef]
- Wei, L.; Miao, H.; Zhao, R.; Han, X.; Zhang, T.; Zhang, H. Identification and testing of reference genes for Sesame gene expression analysis by quantitative real-time PCR. Planta 2013, 237, 873–889. [Google Scholar] [CrossRef] [PubMed]
- Bai, S.; Wang, X.; Guo, M.; Cheng, G.; Khan, A.; Yao, W.; Gao, Y.; Li, J. Selection and Evaluation of Reference Genes for Quantitative Real-Time PCR in Tomato (Solanum lycopersicum L.) Inoculated with Oidium neolycopersici. Agronomy 2022, 12, 3171. [Google Scholar] [CrossRef]
- Pinheiro, T.T.; Litholdo, C.G., Jr.; Sereno, M.L.; Leal, G.A., Jr.; Albuquerque, P.S.B.D.; Figueira, A. Establishing references for gene expression analyses by RT-qPCR in Theobroma cacao tissues. Genet. Mol. Res. 2011, 10, 3291–3305. [Google Scholar] [CrossRef]
- Prasad, P.; Savadi, S.; Bhardwaj, S.C.; Gangwar, O.P.; Khan, H.; Kumar, S. Evaluation of candidate reference genes for normalization of RT-qPCR data in wheat during pre-haustorial stages of Puccinia triticina infection. Trop. Plant Path. 2020, 45, 96–101. [Google Scholar] [CrossRef]
- Scholtz, J.J.; Visser, B. Reference gene selection for qPCR gene expression analysis of rust-infected wheat. Physiol. Mol. Plant Pathol. 2013, 81, 22–25. [Google Scholar] [CrossRef]
- Jose, S.; Abbey, J.; Jaakola, L.; Percival, D. Selection and validation of reliable reference genes for gene expression studies from Monilinia vaccinii-corymbosi infected wild blueberry phenotypes. Sci. Rep. 2020, 10, 11688. [Google Scholar] [CrossRef] [PubMed]
- Monteiro, F.; Sebastiana, M.; Pais, M.S.; Figueiredo, A. Reference gene selection and validation for the early responses to downy mildew infection in susceptible and resistant Vitis vinifera cultivars. PLoS ONE 2013, 8, e72998. [Google Scholar] [CrossRef] [PubMed]
- Alves Oliveira, D.; Tang, J.D.; Warburton, M.L. Reference gene selection for RT-qPCR analysis in Maize Kernels inoculated with Aspergillus flavus. Toxins 2021, 13, 386. [Google Scholar] [CrossRef]
- Joseph, J.T.; Poolakkalody, N.J.; Shah, J.M. Screening internal controls for expression analyses involving numerous treatments by combining statistical methods with reference gene selection tools. Physiol. Mol. Biol. Plants 2019, 25, 289–301. [Google Scholar] [CrossRef] [PubMed]
- Pombo, M.A.; Ramos, R.N.; Zheng, Y.; Fei, Z.; Martin, G.B.; Rosli, H.G. Transcriptome-based identification and validation of reference genes for plant-bacteria interaction studies using Nicotiana benthamiana. Sci. Rep. 2019, 9, 1632. [Google Scholar] [CrossRef]
- Sagun, C.M.; Grandmottet, F.; Suachaowna, N.; Sujipuli, K.; Ratanasut, K. Validation of suitable reference genes for normalization of quantitative reverse transcriptase-polymerase chain reaction in rice infected by Xanthomonas oryzae pv. oryzae. Plant Gene 2020, 21, 100217. [Google Scholar] [CrossRef]
- Albuquerque, G.M.R.; Fonseca, F.C.A.; Boiteux, L.S.; Borges, R.C.F.; Miller, R.N.G.; Lopes, C.A.; Souza, E.B.; Fonseca, M.E.N. Stability analysis of reference genes for RT-qPCR assays involving compatible and incompatible Ralstonia solanacearum-tomato ‘Hawaii 7996’ interactions. Sci. Rep. 2021, 11, 18719. [Google Scholar] [CrossRef]
- Ghareeb, H.; Bozsó, Z.; Ott, P.G.; Wydra, K. Silicon and Ralstonia solanacearum modulate expression stability of housekeeping genes in tomato. Physiol. Mol. Plant Pathol. 2011, 75, 176–179. [Google Scholar] [CrossRef]
- Pombo, M.A.; Zheng, Y.; Fei, Z.; Martin, G.B.; Rosli, H.G. Use of RNA-seq data to identify and validate RT-qPCR reference genes for studying the tomato-Pseudomonas pathosystem. Sci. Rep. 2017, 7, 44905. [Google Scholar] [CrossRef]
- De Spiegelaere, W.; Dern-Wieloch, J.; Weigel, R.; Schumacher, V.; Schorle, H.; Nettersheim, D.; Bergmann, M.; Brehm, R.; Kliesch, S.; Vandekerckhove, L.; et al. Reference gene validation for RT-qPCR, a note on different available software packages. PLoS ONE 2015, 10, e0122515. [Google Scholar] [CrossRef] [PubMed]
- Vandesompele, J.; De Preter, K.; Pattyn, F.; Poppe, B.; Van Roy, N.; De Paepe, A.; Speleman, F. Accurate normalization of real-time quantitative RT-PCR data by geometric averaging of multiple internal control genes. Genome Biol. 2002, 3, research0034-1. [Google Scholar] [CrossRef]
- Andersen, C.L.; Jensen, J.L.; Orntoft, T.F. Normalization of real-time quantitative reverse transcription-PCR data: A model-based variance estimation approach to identify genes suited for normalization, applied to bladder and colon cancer data sets. Cancer Res. 2004, 64, 5245–5250. [Google Scholar] [CrossRef]
- Pfaffl, M.W.; Tichopad, A.; Prgomet, C.; Neuvians, T.P. Determination of stable housekeeping genes, differentially regulated target genes and sample integrity: BestKeeper–Excel-based tool using pair-wise correlations. Biotechnol. Lett. 2004, 26, 509–515. [Google Scholar] [CrossRef]
- Silver, N.; Best, S.; Jiang, J.; Thein, S.L. Selection of housekeeping genes for gene expression studies in human reticulocytes using real-time PCR. BMC Mol. Biol. 2006, 7, 33. [Google Scholar] [CrossRef] [PubMed]
- Available online: https://github.com/fulxie/RefFinder (accessed on 15 November 2025).
- Duan, M.; Wang, J.; Zhang, X.; Yang, H.; Wang, H.; Qiu, Y.; Song, J.; Guo, Y.; Li, X. Identification of Optimal Reference Genes for Expression Analysis in Radish (Raphanus sativus L.) and Its Relatives Based on Expression Stability. Front. Plant Sci. 2017, 8, 1605. [Google Scholar] [CrossRef]
- Xie, F.; Wang, J.; Zhang, B. RefFinder: A web-based tool for comprehensively analyzing and identifying reference genes. Funct. Integr. Genomics 2023, 23, 125. [Google Scholar] [CrossRef]
- Auler, P.A.; Benitez, L.C.; do Amaral, M.N.; Vighi, I.L.; Dos Santos Rodrigues, G.; da Maia, L.C.; Braga, E.J. Evaluation of stability and validation of reference genes for RT-qPCR expression studies in rice plants under water deficit. J. Appl. Genet. 2017, 58, 163–177. [Google Scholar] [CrossRef]
- Sun, C.W.; Griffen, S.; Callis, J. A model for the evolution of polyubiquitin genes from the study of Arabidopsis thaliana ecotypes. Plant Mol. Biol. 1997, 34, 745–758. [Google Scholar] [CrossRef] [PubMed]
- Hernandez-Garcia, C.M.; Martinelli, A.P.; Bouchard, R.A.; Finer, J.J. A soybean (Glycine max) polyubiquitin promoter gives strong constitutive expression in transgenic soybean. Plant Cell Rep. 2009, 28, 837–849. [Google Scholar] [CrossRef]
- Binet, M.N.; Weil, J.H.; Tessier, L.H. Structure and expression of sunflower ubiquitin genes. Plant Mol. Biol. 1991, 17, 395–407. [Google Scholar] [CrossRef] [PubMed]
- Callis, J.; Raasch, J.A.; Vierstra, R.D. Ubiquitin extension proteins of Arabidopsis thaliana. Structure, localization, and expression of their promoters in transgenic tobacco. J. Biol. Chem. 1990, 265, 12486–12493. [Google Scholar] [CrossRef] [PubMed]
- Gao, W.; Zhang, L.; Zhang, Y.; Zhang, P.; Shahinnia, F.; Chen, T.; Yang, D. Genome-wide identification and expression analysis of the UBC gene family in wheat (Triticum aestivum L.). BMC Plant Biol. 2024, 24, 341. [Google Scholar] [CrossRef]
- Jeon, E.H.; Pak, J.H.; Kim, M.J.; Kim, H.J.; Shin, S.H.; Lee, J.H.; Kim, D.H.; Oh, J.S.; Oh, B.J.; Jung, H.W.; et al. Ectopic expression of ubiquitin-conjugating enzyme gene from wild rice, OgUBC1, confers resistance against UV-B radiation and Botrytis infection in Arabidopsis thaliana. Biochem. Biophys. Res. Commun. 2012, 427, 309–314. [Google Scholar] [CrossRef]
- Pickart, C.M. Mechanisms underlying ubiquitination. Annu. Rev. Biochem. 2001, 70, 503–533. [Google Scholar] [CrossRef]
- Kim, S.C.; Guo, L.; Wang, X. Nuclear moonlighting of cytosolic glyceraldehyde-3-phosphate dehydrogenase regulates Arabidopsis response to heat stress. Nat. Commun. 2020, 11, 3439. [Google Scholar] [CrossRef]
- Zhang, H.; Zhao, Y.; Zhou, D.-X. Rice NAD+ dependent histone deacetylase OsSRT1 represses glycolysis function of GAPDH as a transcriptional activator of glycolytic genes. Nucleic Acids Res. 2017, 45, 12241–12255. [Google Scholar] [CrossRef] [PubMed]
- Zaffagnini, M.; Fermani, S.; Costa, A.; Lemaire, S.; Trost, P. Plant cytoplasmic GAPDH: Redox post-translational modifications and moonlighting properties. Front. Plant Sci. 2013, 4, 450. [Google Scholar] [CrossRef]
- McDowell, J.M.; Huang, S.; McKinney, E.C.; Yong-Qiang, A.; Meaghed, R.B. Structure evolution of the actin gene family in Arabidopsis thaliana. Genetics 1996, 142, 587–602. [Google Scholar] [CrossRef]
- Eleftheriou, E.P.; Palevitz, B.A. The effect of cytochalasin D on preprophase band organization in root tip cells of Allium. J. Cell Sci. 1992, 103, 989–998. [Google Scholar] [CrossRef]
- Wernicke, W.; Jung, G. Role of cytoskeleton in cell shaping of developing mesophyll of wheat (Triticum aestivum L.). Eur. J. Cell Biol. 1992, 57, 88–94. [Google Scholar] [PubMed]
- Šlajcherová, K.; Fišerová, J.; Fischer, L.; Schwarzerová, K. Multiple Actin Isotypes in Plants: Diverse Genes for Diverse Roles? Front. Plant Sci. 2012, 3, 226. [Google Scholar] [CrossRef] [PubMed]
- Radchuk, V.V. The transcriptome of the tubulin gene family in plants. In The Plant Cytoskeleton: A Key Tool for Agro-Biotechnology, 2nd ed.; Blume, Y.B., Baird, W.V., Yemets, A.I., Breviario, D., Eds.; Springer: Dordrecht, The Netherland, 2008; pp. 219–241. [Google Scholar] [CrossRef]
- Silflow, C.D.; Oppenheimer, D.G.; Kopozak, S.D.; Ploense, S.E.; Ludwig, S.R.; Haas, N.; Peter Snustad, D. Plant tubulin genes: Structure and differential expression during development. Dev. Genet. 1987, 8, 435–460. [Google Scholar] [CrossRef]
- Gardiner, J.; Overall, R.; Marc, J. Plant microtubule cytoskeleton complexity: Microtubule arrays as fractals. J. Exp. Bot. 2012, 63, 635–642. [Google Scholar] [CrossRef][Green Version]
- Dutt, S.; Parkash, J.; Mehra, R.; Sharma, N.; Singh, B.; Raigond, P.; Joshi, A.; Chopra, S.; Singh, B.P. Translation initiation in plants: Roles and implications beyond protein synthesis. Biol. Plant. 2015, 59, 401–412. [Google Scholar] [CrossRef]
- Shopan, J.; Liu, C.; Hu, Z.; Zhang, M.; Yang, J. Identification of eukaryotic translation initiation factors and the temperature-dependent nature of Turnip mosaic virus epidemics in allopolyploid Brassica juncea. 3Biotech 2020, 10, 75. [Google Scholar] [CrossRef]
- Raabe, K.; Honys, D.; Michailidis, C. The role of eukaryotic initiation factor 3 in plant translation regulation. Plant Physiol. Biochem. 2019, 145, 75–83. [Google Scholar] [CrossRef] [PubMed]
- Dinkova, T.D.; Martinez-Castilla, L.; Cruz-Espíndola, M.A. The Diversification of eIF4E Family Members in Plants and Their Role in the Plant-Virus Interaction. In Evolution of the Protein Synthesis Machinery and Its Regulation; Hernández, G., Jagus, R., Eds.; Springer International Publishing: Cham, Switzerland, 2016; pp. 187–205. [Google Scholar] [CrossRef]
- Cocco, E.; Farci, D.; Guadalupi, G.; Manconi, B.; Maxia, A.; Piano, D. The elongation factor 1-alpha as storage reserve and environmental sensor in Nicotiana tabacum L. seeds. Plant Sci. 2024, 345, 112113. [Google Scholar] [CrossRef] [PubMed]
- Xu, L.; Zhang, L.; Liu, Y.; Sod, B.; Li, M.; Yang, T.; Gao, T.; Yang, Q.; Long, R. Overexpression of the elongation factor MtEF1A1 promotes salt stress tolerance in Arabidopsis thaliana and Medicago truncatula. BMC Plant Biol. 2023, 23, 138. [Google Scholar] [CrossRef]
- Fu, J.; Momčilović, I.; Prasad, P.V.V. Roles of protein synthesis elongation factor EF-TU in heat tolerance in plants. J. Bot. 2012, 2012, 835836. [Google Scholar] [CrossRef]
- Cottage, A.; Edwards, Y.J.; Elgar, G. SAND, a new protein family: From nucleic acid to protein structure and function prediction. Comp. Funct. Genom. 2001, 2, 226–235. [Google Scholar] [CrossRef]
- Carles, C.C.; Fletcher, J.C. Missing links between histones and RNA Pol II arising from SAND? Epigenetics 2010, 5, 381–385. [Google Scholar] [CrossRef]
- Francois, M.; Donovan, P.; Fontaine, F. Modulating transcription factor activity: Interfering with protein-protein interaction networks. Semin. Cell Dev. Biol. 2020, 99, 12–19. [Google Scholar] [CrossRef]



| Plant | Pathogen | Reference |
|---|---|---|
| Fungi and Oomycetes | ||
| Avena sativa | Blumeria graminis f.sp. avenae Puccinia graminis f.sp. avenae Puccinia coronata f.sp. avenae | [26] [27] [28] |
| Avena fatua | Trichoderma polysporum | [29] |
| Brassica rapa | Peronospora parasitica (Hyaloperonospora parasitica) | [30] |
| Carica papaya | Colletotrichum gloeosporioides | [31] |
| Citrus × clementina | Alternaria alternata Phytophthora parasitica (Phytophthora nicotianae) | [32] |
| Citrus reshni | Alternaria alternata Phytophthora parasitica (Phytophthora nicotianae) | [32] |
| Citrus reticulata | Alternaria alternata Phytophthora parasitica (Phytophthora nicotianae) | [32] |
| Citrus sinensis | Alternaria alternata Phytophthora parasitica (Phytophthora nicotianae) | [32] |
| Citrus sunki | Phytophthora parasitica (Phytophthora nicotianae) Alternaria alternata | [33] [32] |
| Coffea arabica | Colletotrichum kahawae Hemileia vastatrix | [34] [35] |
| Cucumis melo | Fusarium oxysporum f.sp. melonis | [5] |
| Elaeis guineensis | Ganoderma boninense | [36] |
| Eucalyptus grandis × Eucalyptus urophylla | Puccinia psidii (Austropuccinia psidii) | [4] |
| Fagus sylvatica | Phytophthora citricola | [37] |
| Glycine max | Microsphaera diffusa (Erysiphe diffusa) Phakopsora pachyrhizi | [38] [39] |
| Humulus lupulus | Verticillium alboatrum | [40] |
| Lens culinaris | Sclerotinia sclerotiorum Aphanomyces euteiches | [41] |
| Morus indica | Fusarium solani Lasiodiplodia theobromae Phyllactinia corylea (Phyllactinia guttata) | [42] |
| Musa acuminata | Pseudocercospora musae Fusarium oxysporum f.sp. cubense Colletotrichum musae | [43] [44] [45] |
| Musa spp. | Fusarium oxysporum f.sp. cubense | [46] |
| Oryza sativa | Rhizoctonia solani Magnaporthe oryzae (Pyricularia oryzae) Magnaporthe grisea (Pyricularia grisea) | [47] [48] [49] |
| Phaseolus vulgaris | Colletotrichum lindemuthianum | [50] |
| Piper nigrum | Phytophthora capsici | [51] |
| Pisum sativum | Sclerotinia sclerotiorum | [52] |
| Poncirus trifoliata | Phytophthora parasitica (Phytophthora nicotianae) Alternaria alternata | [33] [32] |
| Rosa hybrida | Diplocarpon rosae | [53] |
| Saccharum officinarum | Sporisorium scitamineum | [54] |
| Sesamum indicum | Fusarium oxysporum | [55] |
| Solanum lycopersicum | Oidium neolycopersici (Erysiphe neolycopersici) | [56] |
| Theobroma cacao | Moniliophthora perniciosa | [57] |
| Triticum aestivum | Puccinia triticina (Puccinia recondita) Puccinia striiformis Puccinia graminis f.sp. tritici | [58] [59] |
| Vaccinium angustifolium f. nigrum | Monilinia vaccinii-corymbosi | [60] |
| Vaccinium myrtilloides | Monilinia vaccinii-corymbosi | [60] |
| Vitis vinifera | Plasmopara viticola | [17,61] |
| Zea mays | Aspergillus flavus | [62] |
| Bacteria | ||
| Actinidia deliciosa | Pseudomonas syringae pv. actinidiae | [6] |
| Arabidopsis thaliana | Agrobacterium tumefaciens | [63] |
| Citrus clementina | Xylella fastidiosa Candidatus Liberibacter asiaticus | [32] |
| Citrus reshni | Xylella fastidiosa Candidatus Liberibacter asiaticus | [32] |
| Citrus reticulata | Xylella fastidiosa Candidatus Liberibacter asiaticus | [32] |
| Citrus sinensis | Xylella fastidiosa Candidatus Liberibacter asiaticus | [32] |
| Citrus sunki | Xylella fastidiosa Candidatus Liberibacter asiaticus | [32] |
| Nicotiana benthamiana | Pseudomonas fluorescens | [64] |
| Oryza sativa | Xantomonas oryzae pv. oryzae Xantomonas oryzae pv. oryzicola | [65] [49] |
| Poncirus trifoliata | Xylella fastidiosa Candidatus Liberibacter asiaticus | [32] |
| Solanum lycopersicum | Ralstonia solanacearum Xantomonas syringae pv. tomato Pseudomonas syringae pv. tomato | [66] [67] [68] |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2025 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Lizzio, A.; Battaglia, V.; Lahoz, E.; Reverberi, M.; Petriccione, M. Reference Genes in Plant–Pathogen Interaction: A Bibliometric Analysis. Horticulturae 2025, 11, 1416. https://doi.org/10.3390/horticulturae11121416
Lizzio A, Battaglia V, Lahoz E, Reverberi M, Petriccione M. Reference Genes in Plant–Pathogen Interaction: A Bibliometric Analysis. Horticulturae. 2025; 11(12):1416. https://doi.org/10.3390/horticulturae11121416
Chicago/Turabian StyleLizzio, Agata, Valerio Battaglia, Ernesto Lahoz, Massimo Reverberi, and Milena Petriccione. 2025. "Reference Genes in Plant–Pathogen Interaction: A Bibliometric Analysis" Horticulturae 11, no. 12: 1416. https://doi.org/10.3390/horticulturae11121416
APA StyleLizzio, A., Battaglia, V., Lahoz, E., Reverberi, M., & Petriccione, M. (2025). Reference Genes in Plant–Pathogen Interaction: A Bibliometric Analysis. Horticulturae, 11(12), 1416. https://doi.org/10.3390/horticulturae11121416






