Novel Propagation Strategy of Saccharomyces cerevisiae for Enhanced Xylose Metabolism during Fermentation on Softwood Hydrolysate
Abstract
1. Introduction
2. Materials and Methods
2.1. Yeast Strain and Maintenance
2.2. Yeast Pre-Culture
2.3. Fermentation on Defined Media
2.4. Fermentation on Hydrolysate
2.5. Cell Propagation on Ethanol and Glycerol
2.6. Stoichiometry
2.7. Analytical Methods
3. Results and Discussion
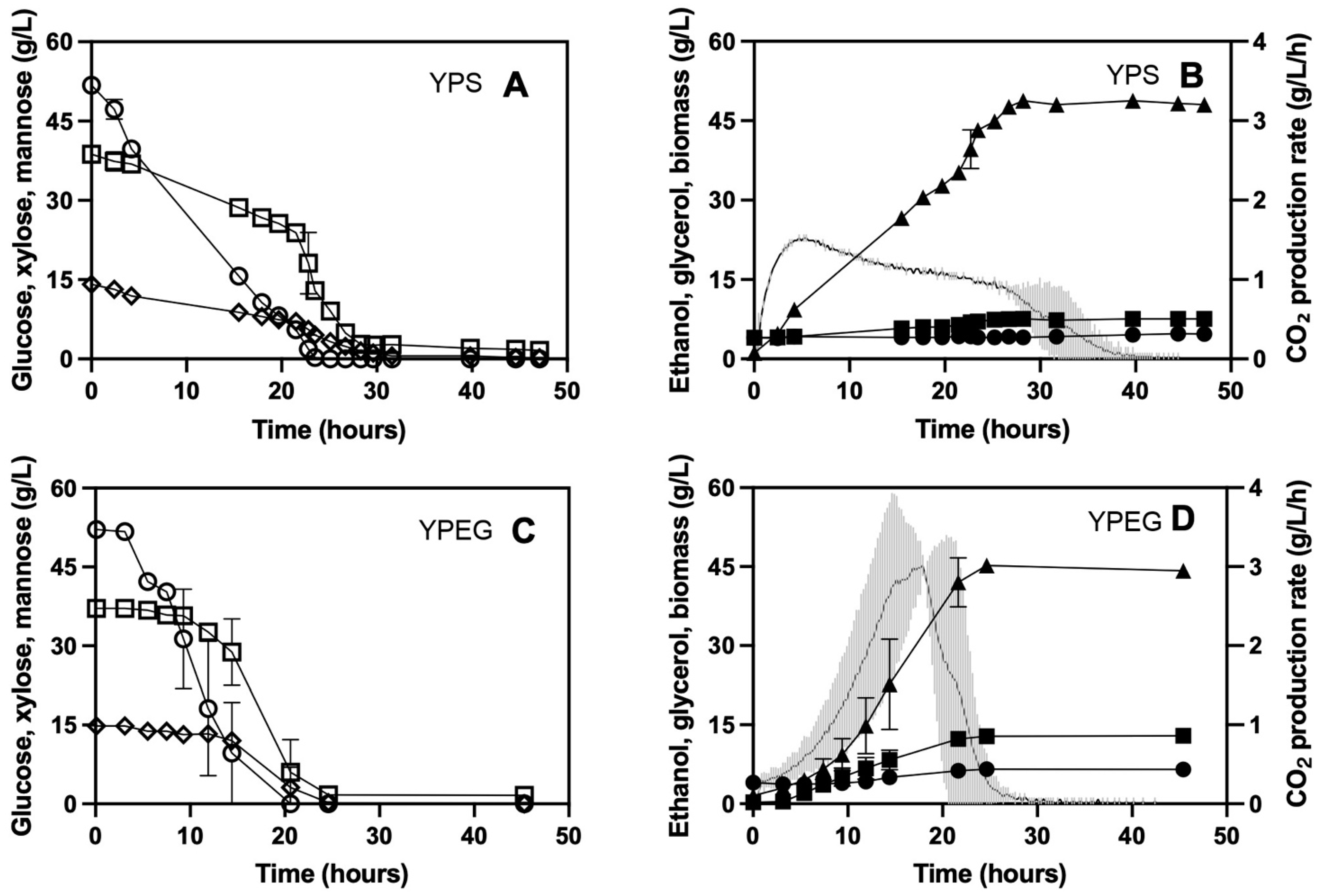
3.1. Fermentation on Defined Glucose/Xylose Media
3.2. Fermentation on Softwood Hydrolysate
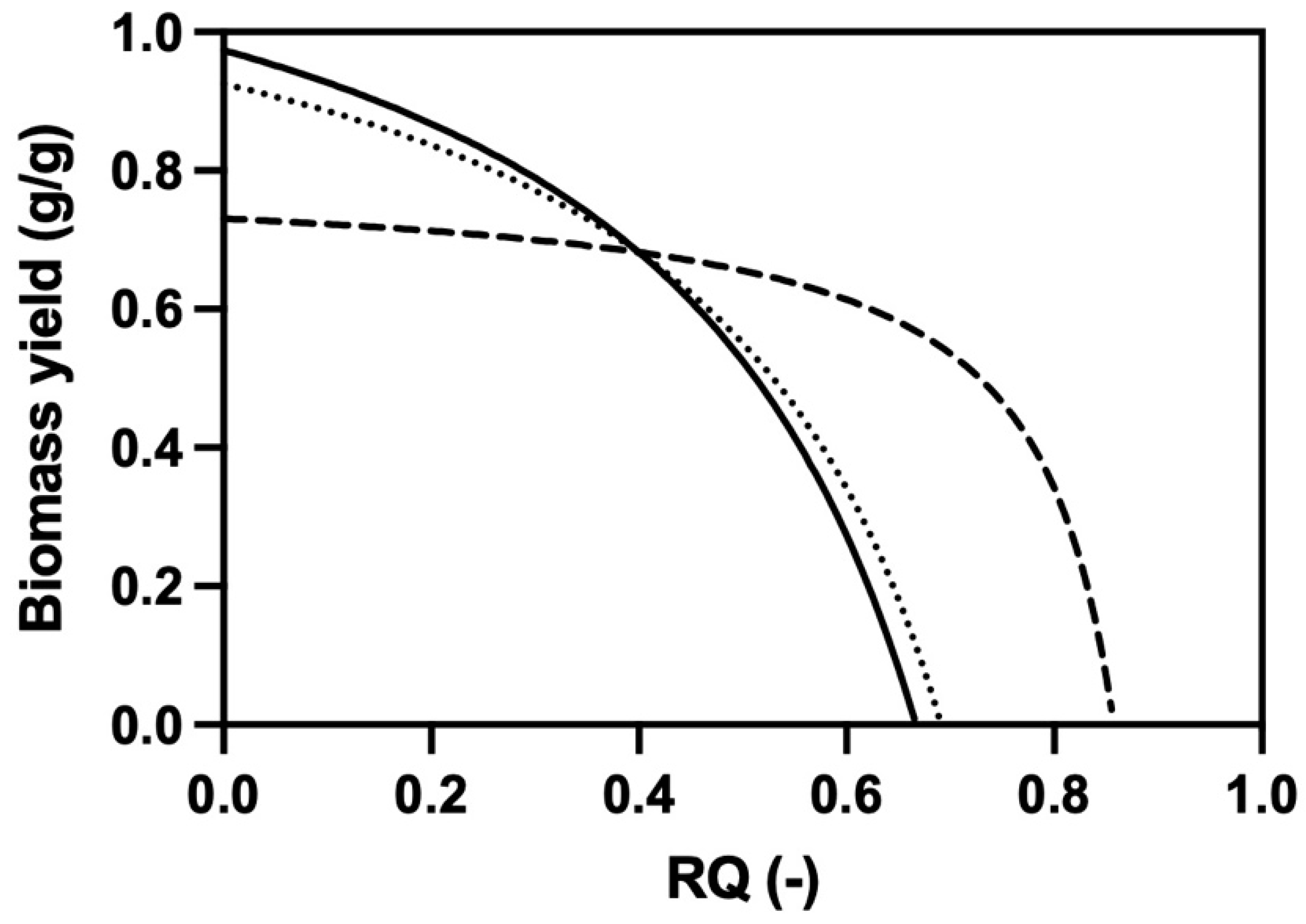
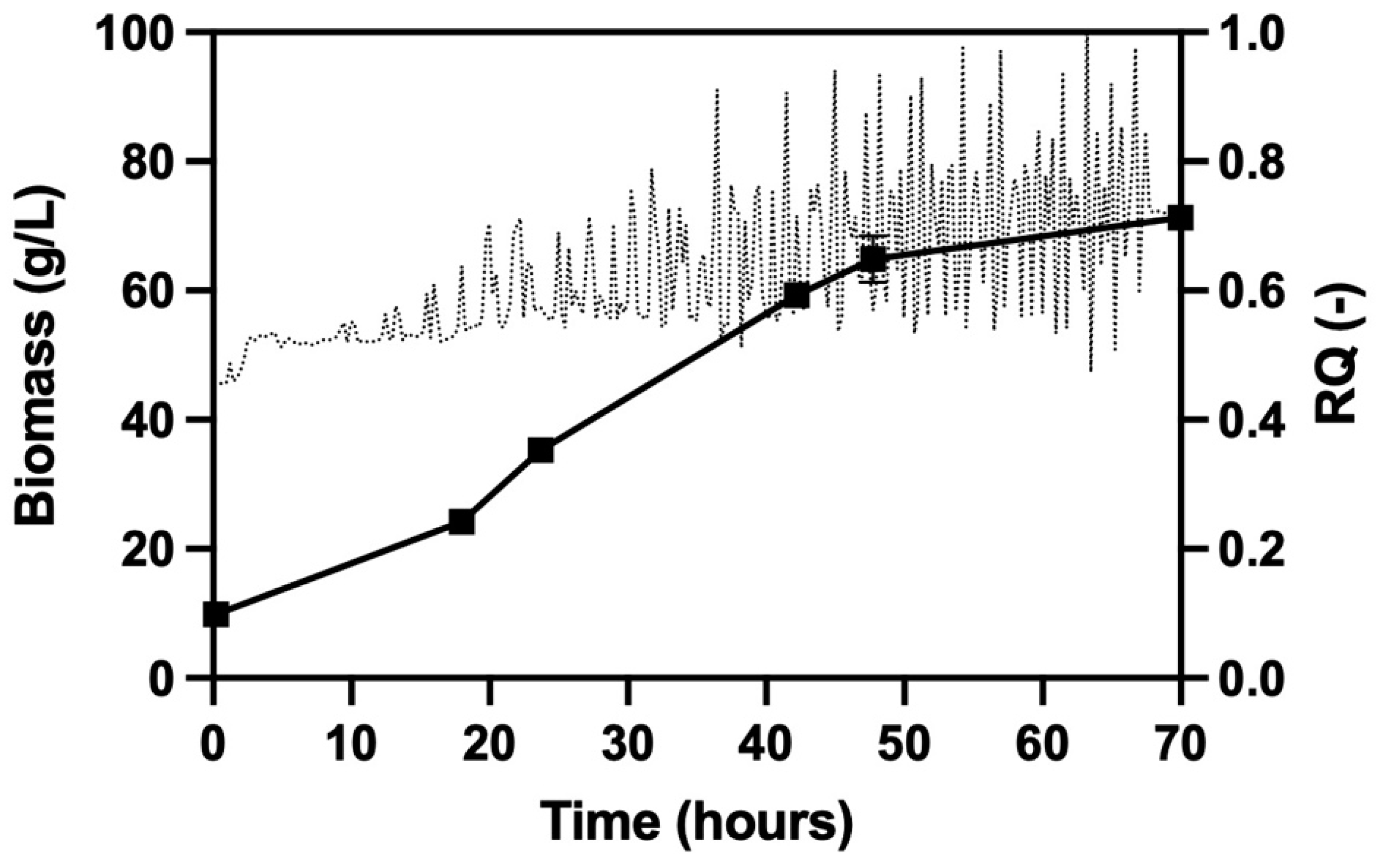
3.3. Development of a Novel Propagation Protocol on Ethanol and Glycerol
4. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Sharma, B.; Larroche, C.; Dussap, C. Comprehensive assessment of 2G bioethanol production. Bioresour. Technol. 2020, 313, 123630. [Google Scholar] [CrossRef] [PubMed]
- European Union 2020. Energy Policy Review; International Energy Agency: France, Paris, 2020. [Google Scholar]
- Directive (EU). 2018/2001 of the European Parliament and of the Council on the promotion of the use of energy from renewable resources (recast). Off. J. Eur. Union 2018, L 328/82, 125–127. [Google Scholar]
- European Union 2020. Trends and projections in Europe 2020. In Tracking Progress Towards Europe’s Climate and Energy Targets; No 13/2020; European Environment Agency: Luxemburg, 2020. [Google Scholar]
- Biofuels Digest. Available online: https://www.biofuelsdigest.com/bdigest/2021/08/25/the-fuel-economy-rule-an-epa-that-wont-see-hear-or-speak-to-the-facts/ (accessed on 29 September 2021).
- Agarwal, A.K. Biofuels (alcohols and biodiesel) applications as fuels for internal combustion engines. Prog. Energy Combust. Sci. 2007, 33, 233–271. [Google Scholar] [CrossRef]
- Del Pecchia, M.; Pessina, V.; Berni, F.; D’Adamo, A.; Fontanesi, S. Gasoline-Ethanol blend formulation to mimic laminar flame speed and auto-ignition quality in automotive engines. Fuel 2020, 264, 116741. [Google Scholar] [CrossRef]
- Ayodele, B.V.; Alsaffar, M.A.; Mustapa, S.I. An overview of integration opportunities for sustainable bioethanol production from first-And second-Generation sugar-Based feedstocks. J. Clean. Prod. 2020, 245, 118857. [Google Scholar] [CrossRef]
- Callegari, A.; Bolognesi, S.; Cecconet, D.; Capodaglio, A.G. Production technologies, current role, and future prospects of biofuels feedstocks: A state-Of-The-Art review. Crit. Rev. Environ. Sci. Technol. 2020, 50, 384–436. [Google Scholar] [CrossRef]
- Saini, J.K.; Saini, R.; Tewari, L. Lignocellulosic agriculture wastes as biomass feedstocks for second-generation bioethanol production: Concepts and recent developments. 3 Biotech 2015, 5, 337–353. [Google Scholar] [CrossRef] [PubMed]
- Balan, V. Current Challenges in Commercially Producing Biofuels from Lignocellulosic Biomass. ISRN Biotechnol. 2014, 2014, 463074. [Google Scholar] [CrossRef]
- Pasin, T.M.; de Almeida, P.Z.; de Almeida Scarcella, A.S.; da Conceição Infante, J.; de Teixeira de Moraes Polizeli de Lourdes, M. Bioconversion of Agro-Industrial Residues to Second-Generation Bioethanol. In Biorefinery of Alternative Resources: Targeting Green Fuels and Platform Chemicals; Nanda, S., Vo, D.N., Sarangi, P.K., Eds.; Springer: Singapore, 2020; pp. 23–47. [Google Scholar]
- Kwak, S.; Jin, Y. Production of fuels and chemicals from xylose by engineered Saccharomyces cerevisiae: A review and per-spective. Microb. Cell Fact. 2017, 16, 82. [Google Scholar] [CrossRef] [PubMed]
- Favaro, L.; Jansen, T.; van Zyl, W.H. Exploring industrial and natural Saccharomyces cerevisiae strains for the bio-based economy from biomass: The case of bioethanol. Crit. Rev. Biotechnol. 2019, 39, 800–816. [Google Scholar]
- Kim, S.R.; Park, Y.; Jin, Y.; Seo, J. Strain engineering of Saccharomyces cerevisiae for enhanced xylose metabolism. Biotechnol. Adv. 2013, 31, 851–861. [Google Scholar] [CrossRef]
- Vilela, L.; de Araujo, V.; Paredes, R.; Bon, E.; Torres, F.; Neves, B.; Eleutherio, E. Enhanced xylose fermentation and ethanol production by engineered Saccharomyces cerevisiae strain. AMB Express 2015, 5, 1–7. [Google Scholar] [CrossRef] [PubMed]
- Hoang Nguyen Tran, P.; Ko, J.K.; Gong, G.; Um, Y.; Lee, S. Improved simultaneous co-fermentation of glucose and xylose by Saccharomyces cerevisiae for efficient lignocellulosic biorefinery. Biotechnol. Biofuels 2020, 13, 12. [Google Scholar] [CrossRef] [PubMed]
- Subtil, T.; Boles, E. Competition between pentoses and glucose during uptake and catabolism in recombinant Saccharomyces cerevisiae. Biotechnol. Biofuels 2012, 5, 14. [Google Scholar] [CrossRef]
- Bergdahl, B.; Sandström, A.G.; Borgström, C.; Boonyawan, T.; van Niel, E.W.J.; Gorwa-Grauslund, M.F. Engineering Yeast Hexokinase 2 for Improved Tolerance Toward Xylose-Induced Inactivation. PLoS ONE 2013, 8, e75055. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Salusjärvi, L.; Kankainen, M.; Soliymani, R.; Pitkänen, J.; Penttilä, M.; Ruohonen, L. Regulation of xylose metabolism in re-combinant Saccharomyces cerevisiae. Microb. Cell Fact. 2008, 7, 18. [Google Scholar] [CrossRef]
- Alff-Tuomala, S.; Salusjärvi, L.; Barth, D.; Oja, M.; Penttilä, M.; Pitkänen, J.; Ruohonen, L.; Jouhten, P. Xylose-Induced dynamic effects on metabolism and gene expression in engineered Saccharomyces cerevisiae in anaerobic glucose-Xylose cultures. Appl. Microbiol. Biotechnol. 2016, 100, 969–985. [Google Scholar] [CrossRef]
- Jin, Y.-S.; Laplaza, J.M.; Jeffries, T.W. Saccharomyces cerevisiae Engineered for Xylose Metabolism Exhibits a Respiratory Response. Appl. Environ. Microbiol. 2004, 70, 6816–6825. [Google Scholar] [CrossRef] [PubMed]
- Rose, A.H.; Vijayalakshmi, G. Baker’s Yeast. In Biochemistry and Genetics of Yeast; Bacila, M., Horecker, B.L., Stoppani, A.O.M., Eds.; Academic Press: New York, NY, USA, 1978; Volume 2, pp. 385–387. [Google Scholar]
- Ingledew, W.M. The Alcohol Textbook, 5th ed.; Nottingham Univ. Press: Nottingham, UK, 2009; pp. 152–156. [Google Scholar]
- Ochoa-Estopier, A.; Lesage, J.; Gorret, N.; Guillouet, S.E. Kinetic analysis of a Saccharomyces cerevisiae strain adapted for im-proved growth on glycerol: Implications for the development of yeast bioprocesses on glycerol. Bioresour. Technol. 2011, 102, 1521–1527. [Google Scholar] [CrossRef] [PubMed]
- Beudecher, R.F.; van Dam, H.W.; van der Plaat, J.B.; Vellega, K. Developments in baker’s yeast production. In Yeast Biotech-nology and Biocatalysis; Verachtert, H., de Mot, R., Eds.; Marcel Dekker: New York, NY, USA, 1990; pp. 103–146. [Google Scholar]
- Jørgensen, H.; Olsson, L.; Rønnow, B.; Palmqvist, E. Fed-Batch cultivation of baker’s yeast followed by nitrogen or carbon starvation: Effects on fermentaNive capacity and content of trehalose and glycogen. Appl. Microbiol. Biotechnol. 2002, 59, 310–317. [Google Scholar]
- Doran, P.M. Bioprocess. In Engineering Principles; Academic Press: London, UK, 1995; pp. 75–79. [Google Scholar]
- Heyman, B.; Tulke, H.; Putri, S.P.; Fukusaki, E.; Büchs, J. Online monitoring of the respiratory quotient reveals metabolic phases during microaerobic 2,3-butanediol production with Bacillus licheniformis. Eng. Life Sci. 2020, 20, 133–144. [Google Scholar] [CrossRef]
- Tippmann, S.; Scalcinati, G.; Siewers, V.; Nielsen, J. Production of farnesene and santalene by Saccharomyces cerevisiae using fed-Batch cultivations with RQ-controlled feed. Biotechnol. Bioeng. 2016, 113, 72–81. [Google Scholar] [CrossRef] [PubMed]
- Li, X.; Yu, C.; Yao, J.; Wang, Z.; Lu, S. An Online Respiratory Quotient-Feedback Strategy of Feeding Yeast Extract for Efficient Arachidonic Acid Production by Mortierella alpina. Front. Bioeng. Biotechnol. 2018, 5, 83. [Google Scholar] [CrossRef]
- Nielsen, F.; Tomás-Pejó, E.; Olsson, L.; Wallberg, O. Short-term adaptation during propagation improves the performance of xylose-fermenting Saccharomyces cerevisiae in simultaneous saccharification and co-fermentation. Biotechnol. Biofuels 2015, 8, 219. [Google Scholar] [CrossRef]
- Van Dijk, M.; Erdei, B.; Galbe, M.; Nygård, Y.; Olsson, L. Strain-dependent variance in short-term adaptation effects of two xylose-fermenting strains of Saccharomyces cerevisiae. Bioresour. Technol. 2019, 292, 121922. [Google Scholar] [CrossRef] [PubMed]
- Alkasrawi, M.; Rudolf, A.; Lidén, G.; Zacchi, G. Influence of strain and cultivation procedure on the performance of simul-taneous saccharification and fermentation of steam pretreated spruce. Enzyme Microb. Tech. 2006, 38, 279–286. [Google Scholar] [CrossRef]
- Zhang, K.; Wells, P.; Liang, Y.; Love, J.; Parker, D.A.; Botella, C. Effect of diluted hydrolysate as yeast propagation medium on ethanol production. Bioresour. Technol. 2019, 271, 1–8. [Google Scholar] [CrossRef] [PubMed]
- Li, B.; Yuan, Y. Transcriptome shifts in response to furfural and acetic acid in Saccharomyces cerevisiae. Appl. Microbiol. Biotechnol. 2010, 86, 1915–1924. [Google Scholar] [CrossRef]
- Viegas, C.A.; Sa-Correia, I. Activation of plasma membrane ATPase of Saccharomyces cerevisiae by octanoic acid. J. Gen. Microbiol. 1991, 137, 645–651. [Google Scholar] [CrossRef] [PubMed]
- Verstrepen, K.J.; Iserentant, D.; Malcorps, P.; Derdelinckx, G.; Van Dijck, P.; Winderickx, J.; Pretorius, I.S.; Thevelein, J.M.; Delvaux, F.R. Glucose and sucrose: Hazardous fast-food for industrial yeast? Trends Biotechnol. 2004, 22, 531–537. [Google Scholar] [CrossRef]
- Randez-Gil, F.; Córcoles-Sáez, I.; Prieto, J.A. Genetic and Phenotypic Characteristics of Baker’s Yeast: Relevance to Baking. Annu. Rev. Food Sci. Technol. 2013, 4, 191–214. [Google Scholar] [CrossRef]
- Verduyn, C.; Stouthamer, A.H.; Scheffers, W.A.; Van Dijken, J.P. A theoretical evaluation of growth yields of yeasts. Antonie van Leeuwenhoek 1991, 59, 49–63. [Google Scholar] [CrossRef] [PubMed]
- Stanley, D.; Bandara, A.; Fraser, S.; Chambers, P.J.; Stanley, G.A. The ethanol stress response and ethanol tolerance of Sac-charomyces cerevisiae. J. Appl. Microbiol. 2010, 109, 13–24. [Google Scholar] [CrossRef] [PubMed]
- Welsh, D.T. Ecological significance of compatible solute accumulation by micro-organisms: From single cells to global cli-mate. FEMS Microbiol. Rev. 2000, 24, 263–290. [Google Scholar] [CrossRef] [PubMed]
- Klein, M.; Islam, Z.; Knudsen, P.B.; Carrillo, M.; Swinnen, S.; Workman, M.; Nevoigt, E. The expression of glycerol facilitators from various yeast species improves growth on glycerol of Saccharomyces cerevisiae. Metab. Eng. Commun. 2016, 3, 252–257. [Google Scholar] [CrossRef]
- Rolland, F.; de Winde, J.H.; Lemaire, K.; Boles, E.; Thevelein, J.M.; Winderickx, J. Glucose-induced cAMP signalling in yeast requires both a G-protein coupled receptor system for extracellular glucose detection and a separable hexose kinase-dependent sensing process. Mol. Microbiol. 2000, 38, 348–358. [Google Scholar] [CrossRef]
- Saini, P.; Beniwal, A.; Kokkiligadda, A.; Vij, S. Response and tolerance of yeast to changing environmental stress during ethanol fermentation. Process. Biochem. 2018, 72, 1–12. [Google Scholar] [CrossRef]
- Microorganism Expressing Xylose Isomerase. U.S. Patent WO/2010/070549, December 2010.
- Microorganism Expressing Aldose-1-Epimerase. U.S. Patent WO/2010/001363, July 2009.
- Verduyn, C.; Postma, E.; Scheffers, W.A.; Van Dijken, J.P. Effect of benzoic acid on metabolic fluxes in yeasts: A continu-ous-culture study on the regulation of respiration and alcoholic fermentation. Yeast 1992, 8, 501–517. [Google Scholar] [CrossRef]
- Verduyn, C. Physiology of yeasts in relation to biomass yields. Antonie van Leeuwenhoek 1991, 60, 325–353. [Google Scholar] [CrossRef] [PubMed]
- Watteeuw, C.M.; Armiger, W.B.; Ristroph, D.L.; Humphrey, A.E. Production of single cell protein from ethanol by fed-batch process. Biotechnol. Bioeng. 1979, 21, 1221–1237. [Google Scholar] [CrossRef]
- Ruiz, H.A.; Silva, D.P.; Ruzene, D.S.; Lima, L.F.; Vicente, A.A.; Teixeira, J.A. Bioethanol production from hydrothermal pre-treated wheat straw by a flocculating Saccharomyces cerevisiae strain–Effect of process conditions. Fuel 2012, 95, 528–536. [Google Scholar] [CrossRef]
- Matsushika, A.; Goshima, T.; Hoshino, T. Transcription analysis of recombinant industrial and laboratory Saccharomyces cerevisiae strains reveals the molecular basis for fermentation of glucose and xylose. Microb. Cell Factories 2014, 13, 16. [Google Scholar] [CrossRef] [PubMed]
- Shen, Y.; Hou, J.; Bao, X. Enhanced xylose fermentation capacity related to an altered glucose sensing and repression network in a recombinant Saccharomyces cerevisiae. Bioengineered 2013, 4, 435–437. [Google Scholar] [CrossRef] [PubMed]
- Papapetridis, I.; Verhoeven, M.D.; Wiersma, S.J.; Gouiaan, M.; van Maris, A.J.A.; Pronk, J.T. Laboratory evolution for forced glucose-Xylose co-Consumption enables identification of mutations that improve mixed-Sugar fermentation by xy-Lose-Fermenting Saccharomyces cerevisiae. FEMS Yeast Res. 2018, 18, foy056. [Google Scholar] [CrossRef] [PubMed]
- Runquist, D.; Hahn-Hägerdal, B.; Bettiga, M. Increased expression of the oxidative pentose phosphate pathway and gluco-neogenesis in anaerobically growing xylose-Utilizing Saccharomyces cerevisiae. Microb. Cell Fact. 2009, 8, 49. [Google Scholar] [CrossRef] [PubMed]
- Zeng, W.-Y.; Tang, Y.-Q.; Gou, M.; Xia, Z.-Y.; Kida, K. Transcriptomes of a xylose-utilizing industrial flocculating Saccharomyces cerevisiae strain cultured in media containing different sugar sources. AMB Express 2016, 6, 1–13. [Google Scholar] [CrossRef] [PubMed]
- Mittelman, K.; Barkai, N. The Genetic Requirements for Pentose Fermentation in Budding Yeast. G3 Genes Genomes Genet. 2017, 7, 1743–1752. [Google Scholar] [CrossRef][Green Version]
- Van Dijken, J.P.; Scheffers, W.A. Redox balances in the metabolism of sugars by yeasts. FEMS Microbiol. Lett. 1986, 32, 199–224. [Google Scholar] [CrossRef]
- Daran-Lapujade, P.; Jansen, M.L.A.; Daran, J.; van Gulik, W.; de Winde, J.H.; Pronk, J.T. Role of Transcriptional Regulation in Controlling Fluxes in Central Carbon Metabolism of Saccharomyces cerevisiae. A chemostat culture study. J. Biol. Chem. 2004, 279, 9125–9138. [Google Scholar] [CrossRef] [PubMed]
- Knudsen, J.D.; Rønnow, B. Extended fed-batch fermentation of a C5/C6 optimised yeast strain on wheat straw hydrolysate using an online refractive index sensor to measure the relative fermentation rate. Sci. Rep. 2020, 10, 6705. [Google Scholar] [CrossRef] [PubMed]
- Margono, M.; Kaavessina, M.; Khushairi Mohd Zahari, M.A.; Hisyam, A. Continuous Bioethanol Production Using Uncontrolled Process in a Laboratory Scale of Integrated Aerobic–Anaerobic Baffled Reactor. Period. Polytech. Chem. Eng. 2019, 64, 172–178. [Google Scholar] [CrossRef]
- Beckner, M.L.; Ivey, M.; Phister, T.G. Microbial contamination of fuel ethanol fermentations. Lett. Appl. Microbiol. 2011, 53, 387–394. [Google Scholar] [CrossRef]
- Lane, S.; Xu, H.; Oh, E.J.; Kim, H.; Lesmana, A.; Jeong, D.; Zhang, G.; Tsai, C.-S.; Jin, Y.-S.; Kim, S.R. Glucose repression can be alleviated by reducing glucose phosphorylation rate in Saccharomyces cerevisiae. Sci. Rep. 2018, 8, 2613. [Google Scholar] [CrossRef] [PubMed]
- Hagman, A.; Säll, T.; Compagno, C.; Piškur, J. Yeast “Make-Accumulate-Consume” Life Strategy Evolved as a Multi-Step Process That Predates the Whole Genome Duplication. PLoS ONE 2013, 8, e68734. [Google Scholar] [CrossRef]
- Pirt, S.J. The maintenance energy of bacteria in growing cultures. Proc. R. Soc. Lond. Ser. B Boil. Sci. 1965, 163, 224–231. [Google Scholar] [CrossRef]
- Vos, T.; Hakkaart, X.D.V.; de Hulster, A.F.; van Maris, A.J.A.; Pronk, J.T.; Daran-Lapujade, P.A.S. Maintenance-energy re-Quirements and robustness of Saccharomyces cerevisiae at aerobic near-Zero specific growth rates. Microb. Cell Factories 2016, 15, 111. [Google Scholar] [CrossRef] [PubMed]
- Raj, A.E.; Kumar, H.S.S.; Kumar, S.U.; Misra, M.C.; Ghildyal, N.P.; Karanth, N.G. High-Cell-Density Fermentation of Re-combinant Saccharomyces cerevisiae Using Glycerol. Biotechnol. Prog. 2002, 18, 1130–1132. [Google Scholar]
- Ferreira, A.D.; Mussatto, S.I.; Cadete, R.M.; Rosa, C.A.; Silva, S.S. Ethanol production by a new pentose-fermenting yeast strain, Scheffersomyces stipitis UFMG-IMH 43.2, isolated from the Brazilian forest. Yeast 2011, 28, 547–554. [Google Scholar] [CrossRef]
- Sperotto, G.; Stasiak, L.G.; Godoi, J.P.M.G.; Gabiatti, N.C.; de Souza, S.S. A review of culture media for bacterial cellulose production: Complex, chemically defined and minimal media modulations. Cellulose 2021, 28, 2649–2673. [Google Scholar] [CrossRef]
- Lorenz, E.; Schmacht, M.; Stahl, U.; Senz, M. Enhanced incorporation yield of cysteine for glutathione overproduction by fed-batch fermentation of Saccharomyces cerevisiae. J. Biotechnol. 2015, 216, 131–139. [Google Scholar] [CrossRef]
- Raghavendran, V.; Webb, J.P.; Cartron, M.L.; Springthorpe, V.; Larson, T.R.; Hines, M.; Mohammed, H.; Zimmerman, W.B.; Poole, R.K.; Green, J. A microbubble-sparged yeast propagation–fermentation process for bioethanol production. Biotechnol. Biofuels 2020, 13, 1–104. [Google Scholar] [CrossRef]
- Shiloach, J.; Fass, R. Growing E. coli to high cell density—A historical perspective on method development. Biotechnol. Adv. 2005, 23, 345–357. [Google Scholar] [CrossRef]
- Hanotu, J.; Kong, D.; Zimmerman, W.B. Intensification of yeast production with microbubbles. Food Bioprod. Process. 2016, 100, 424–431. [Google Scholar] [CrossRef]
- Devi, T.T.; Kumar, B. Mass transfer and power characteristics of stirred tank with Rushton and curved blade impeller. Eng. Sci. Technol. Int. J. 2017, 20, 730–737. [Google Scholar] [CrossRef]
- Schaepe, S.; Kuprijanov, A.; Sieblist, C.; Jenzsch, M.; Simutis, R.; Lübbert, A. kLa of stirred tank bioreactors revisited. J. Biotechnol. 2013, 168, 576–583. [Google Scholar] [CrossRef] [PubMed]
- Aßkamp, M.R.; Klein, M.; Nevoigt, E. Saccharomyces cerevisiae exhibiting a modified route for uptake and catabolism of glycerol forms significant amounts of ethanol from this carbon source considered as ‘non-fermentable’. Biotechnol. Biofuels 2019, 12, 257. [Google Scholar] [CrossRef] [PubMed]
- Lages, F.; Lucas, C. Contribution to the physiological characterization of glycerol active uptake in Saccharomyces cerevisiae. Biochim. Biophys. Acta 1997, 1322, 8–18. [Google Scholar]
- Sutherland, F.C.W.; Lages, F.; Lucas, C.; Luyten, K.; Albertyn, J.; Hohmann, S.; Prior, B.A.; Kilian, S.G. Characteristics of Fps1-dependent and -independent glycerol transport in Saccharomyces cerevisiae. J. Bacteriol. 1997, 179, 7790–7795. [Google Scholar] [CrossRef][Green Version]
| Media | Preculture | Max. Substrate Degradation Rate (g/L/h) | Final Ethanol Titre (g/L) | Ethanol Yield (g/g) | Glycerol Yield (g/g) |
|---|---|---|---|---|---|
| Glucose | YPS | 4.75 ± 0.04 | 45.13 ± 1.07 | 0.39 ± 0.01 | 0.09 ± 0.00 |
| YPEG | 4.73 ± 0.10 | 44.57 ± 0.24 | 0.38 ± 0.00 | 0.12 ± 0.00 | |
| Xylose | YPS | 1.62 ± 0.18 | 46.74 ± 1.18 | 0.39 ± 0.00 | 0.07 ± 0.00 |
| YPEG | 2.54 ± 0.11 | 45.81 ± 0.64 | 0.38 ± 0.01 | 0.10 ± 0.00 | |
| Glucose–Xylose | YPS | 3.19 ± 0.23 | 46.10 ± 0.31 | 0.40 ± 0.00 | 0.11 ± 0.00 |
| YPEG | 4.14 ± 0.02 | 45.45 ± 0.43 | 0.39 ± 0.00 | 0.12 ± 0.00 |
| Preculture | Ethanol Productivity (g/L/h) | Ethanol Yield (g/g) | Glycerol Yield (g/g) | Biomass Yield (g/g) |
|---|---|---|---|---|
| YPS | 1.19 ± 0.02 | 0.47 ± 0.01 | 0.03 ± 0.00 | 0.00 ± 0.00 |
| YPEG | 1.47 ± 0.01 | 0.45 ± 0.01 | 0.13 ± 0.00 | 0.03 ± 0.00 |
| Process Parameter | Value ± Std |
|---|---|
| Average biomass yield (g/g) | 0.39 ± 0.00 |
| Percent biomass yield (%) 1 | 47.8 ± 0.64 |
| Average volumetric productivity (g/L/h) | 1.03 ± 0.05 |
| Final biomass concentration (g/L) | 71.3 ± 0.91 |
| Ethanol consumed (g/L) | 174 ± 28.9 |
| Glycerol consumed (g/L) | 43.2 ± 7.14 |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Dobrescu, A.C.; Veras, H.C.T.; Varrone, C.; Knudsen, J.D. Novel Propagation Strategy of Saccharomyces cerevisiae for Enhanced Xylose Metabolism during Fermentation on Softwood Hydrolysate. Fermentation 2021, 7, 288. https://doi.org/10.3390/fermentation7040288
Dobrescu AC, Veras HCT, Varrone C, Knudsen JD. Novel Propagation Strategy of Saccharomyces cerevisiae for Enhanced Xylose Metabolism during Fermentation on Softwood Hydrolysate. Fermentation. 2021; 7(4):288. https://doi.org/10.3390/fermentation7040288
Chicago/Turabian StyleDobrescu, Andreea Cristina, Henrique César Teixeira Veras, Cristiano Varrone, and Jan Dines Knudsen. 2021. "Novel Propagation Strategy of Saccharomyces cerevisiae for Enhanced Xylose Metabolism during Fermentation on Softwood Hydrolysate" Fermentation 7, no. 4: 288. https://doi.org/10.3390/fermentation7040288
APA StyleDobrescu, A. C., Veras, H. C. T., Varrone, C., & Knudsen, J. D. (2021). Novel Propagation Strategy of Saccharomyces cerevisiae for Enhanced Xylose Metabolism during Fermentation on Softwood Hydrolysate. Fermentation, 7(4), 288. https://doi.org/10.3390/fermentation7040288







