Grape and Wine Metabolomics to Develop New Insights Using Untargeted and Targeted Approaches
Abstract
:1. Introduction
2. Advancements in Metabolomics as an Emerging Tool within the Last Decade
2.1. Development of Sensitive and Reproducible Separation and Detection Techniques
2.2. Advancements in Data Analysis Pipelines
3. Application of Metabolomics in Grape and Wine Research: State of the Art
3.1. Untargeted Metabolomics As A Hypothesis-Generating Tool in Grape and Wine Science
3.2. Study of Terroir, Authenticity and Originality of Grapes and Wines Using a Metabolomics Approach
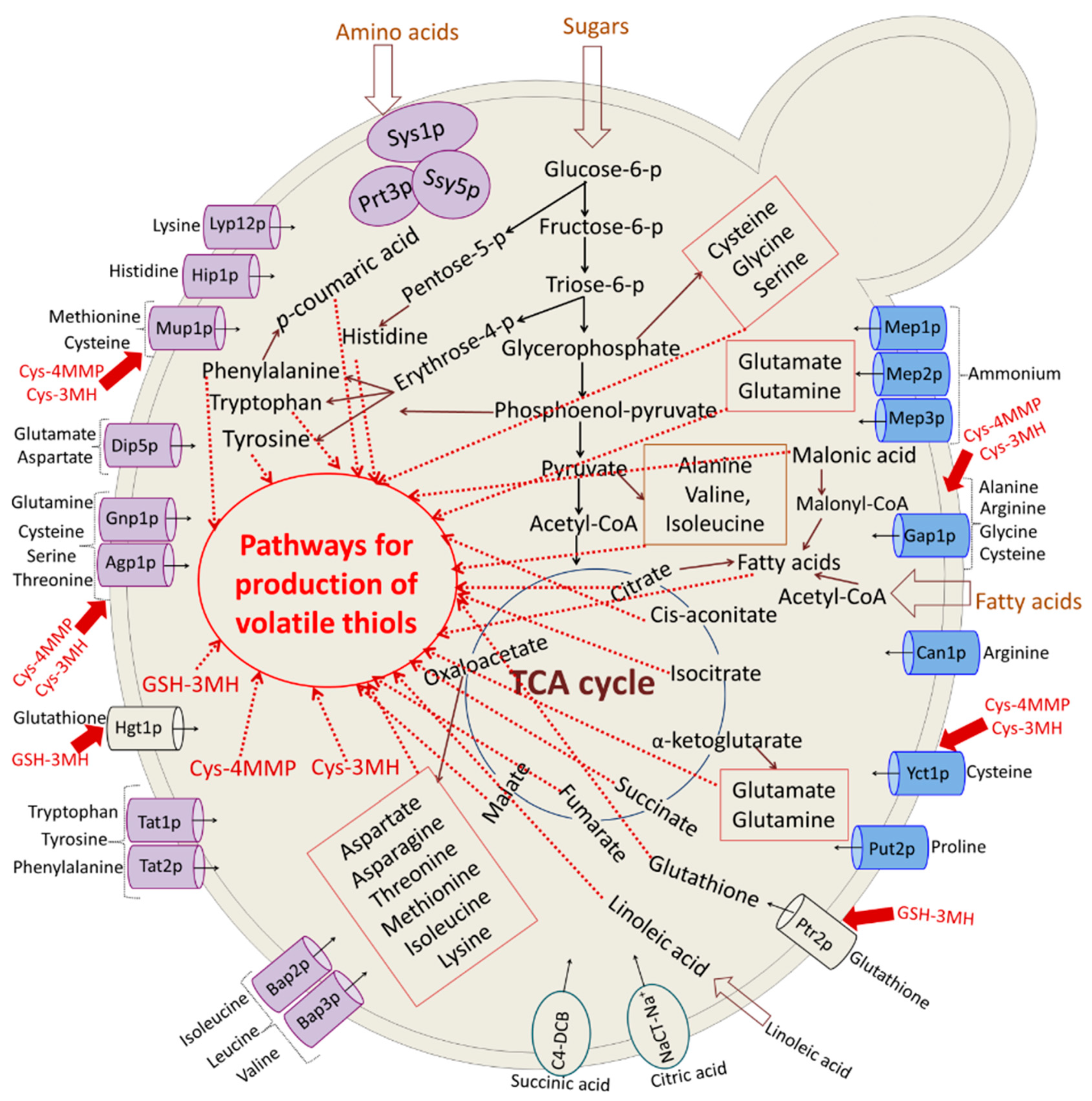
3.3. Study of Yeast Metabolism and Aroma Compound Development during Wine Making
3.4. Combination of Metabolomics and Transcriptomics to Unravel New Knowledge
3.5. Application of Metabolomics to Study Grape Growth Developments and Grape Vine Disease
4. Future Perspectives
5. Conclusions
Funding
Conflicts of Interest
References
- Webb, A.D. Applications of gas chromatography in studying the aromatic qualities of wines. Qual. Plant. Mater. Veg. 1964, 11, 234–243. [Google Scholar] [CrossRef]
- Noble, A.C.; Flath, R.A.; Forrey, R.R. Wine headspace analysis. Reproducibility and application to varietal classification. J. Agric. Food Chem. 1980, 28, 346–353. [Google Scholar] [CrossRef]
- Guth, H. Quantitation and sensory studies of character impact odorants of different white wine varieties. J. Agric. Food Chem. 1997, 45, 3027–3032. [Google Scholar] [CrossRef]
- Mestres, M.; Busto, O.; Guasch, J. Headspace solid-phase microextraction analysis of volatile sulphides and disulphides in wine aroma. J. Chromatogr. A 1998, 808, 211–218. [Google Scholar] [CrossRef]
- Joslyn, M.A.; Dunn, R. Acid metabolism of wine yeast. I. The relation of volatile acid formation to alcoholic fermentation. J. Am. Chem. Soc. 1938, 60, 1137–1141. [Google Scholar] [CrossRef]
- Nelson, R.R.; Acree, T.E.; Butts, R.M. Isolation and identification of volatiles Catawba wine. J. Agric. Food Chem. 1978, 26, 1188–1190. [Google Scholar] [CrossRef]
- Cobb, C.S.; Bursey, M.M. Comparison of extracting solvents for typical volatile components of eastern wines in model aqueous-alcoholic systems. J. Agric. Food Chem. 1978, 26, 197–199. [Google Scholar] [CrossRef]
- Kwan, W.O.; Kowalski, B.R. Pattern recognition analysis of gas chromatographic data. Geographic classification of wines of Vitis vinifera cv. Pinot noir from France and the united states. J. Agric. Food Chem. 1980, 28, 356–359. [Google Scholar] [CrossRef]
- Mayer, K.; Busch, I. On th enzymatic determination of glyerin in the grape juice and wine. UMitteilungen Gebiete Lebensmittel Hyg. 1963, 54, 297–303. [Google Scholar]
- Caldwell, J.S. Some effects of seasonal conditions upon the chemical composition of American grape juice. J. Agric. Res. 1925, 30, 1133–1176. [Google Scholar]
- Webster, J.E.; Cross, F.B. Chemical Analysis of Grape Juices: Varietal Comparisons; American Society of Horticultural Sciences: Alexandria, VA, USA, 1936. [Google Scholar]
- Lafon-Lafourcade, S. Enzymatic methods in the analysis of musts and wines. Appl. Méthodes Enzymatiques L’anal. Mouts Vins 1978, 32, 969–974. [Google Scholar]
- Pinu, F.R. Sauvignon Blanc Metabolomics: Metabolite Profile Analysis before and after Fermentation; University of Auckland: Auckland, New Zealand, 2013. [Google Scholar]
- Oliver, S.G. Systematic functional analysis of the yeast genome. Trends Biotechnol. 1998, 16, 373–378. [Google Scholar] [CrossRef] [Green Version]
- Villas-Bôas, S.G.; Rasmussen, S.; Lane, G.A. Metabolomics or metabolite profiles? Trends Biotechnol. 2005, 23, 385–386. [Google Scholar] [CrossRef] [PubMed]
- Patti, G.J.; Yanes, O.; Siuzdak, G. Metabolomics: The apogee of the omics trilogy. Nat. Rev. Mol. Cell Biol. 2012, 13, 263–269. [Google Scholar] [CrossRef] [PubMed]
- Creydt, M.; Fischer, M. Plant metabolomics: Maximizing metabolome coverage by optimizing mobile phase additives for nontargeted mass spectrometry in positive and negative electrospray ionization mode. Anal. Chem. 2017, 89, 10474–10486. [Google Scholar] [CrossRef] [PubMed]
- Jacob, M.; Malkawi, A.; Albast, N.; Al Bougha, S.; Lopata, A.; Dasouki, M.; Rahman, A.M.A. A targeted metabolomics approach for clinical diagnosis of inborn errors of metabolism. Anal. Chim. Acta 2018, 1025, 141–153. [Google Scholar] [CrossRef] [PubMed]
- Dervishi, E.; Zhang, G.; Mandal, R.; Wishart, D.S.; Ametaj, B.N. Targeted metabolomics: New insights into pathobiology of retained placenta in dairy cows and potential risk biomarkers. Animal 2018, 12, 1050–1059. [Google Scholar] [CrossRef] [PubMed]
- Fiehn, O.; Kopka, J.; Dörmann, P.; Altmann, T.; Trethewey, R.N.; Willmitzer, L. Metabolite profiling for plant functional genomics. Nat. Biotechnol. 2000, 18, 1157–1161. [Google Scholar] [CrossRef] [PubMed]
- Nielsen, J.; Jewett, M.C. The role of metabolomics in systems biology. In Topics in Current Genetics; Springer: Berlin/Heidelberg, Germany, 2007; Volume 18, pp. 1–10. [Google Scholar]
- Sumner, L.W.; Mendes, P.; Dixon, R.A. Plant metabolomics: Large-scale phytochemistry in the functional genomics era. PhytoChem 2003, 62, 817–836. [Google Scholar] [CrossRef]
- Trethewey, R.N. Gene discovery via metabolic profiling. Cur. Opin. Biotechnol. 2001, 12, 135–138. [Google Scholar] [CrossRef]
- Bino, R.J.; Hall, R.D.; Fiehn, O.; Kopka, J.; Saito, K.; Draper, J.; Nikolau, B.J.; Mendes, P.; Roessner-Tunali, U.; Beale, M.H.; et al. Potential of metabolomics as a functional genomics tool. Trends Plant Sci. 2004, 9, 418–425. [Google Scholar] [CrossRef] [PubMed]
- Dyar, K.A.; Eckel-Mahan, K.L. Circadian metabolomics in time and space. Front. Neurosci. 2017, 11, 369. [Google Scholar] [CrossRef] [PubMed]
- Saito, K.; Matsuda, F. Metabolomics for Functional Genomics, Systems Biology, and Biotechnology. Annu. Rev. Plant Biol. 2010, 61, 463–489. [Google Scholar] [CrossRef] [PubMed]
- Dixon, R.A.; Gang, D.R.; Charlton, A.J.; Fiehn, O.; Kuiper, H.A.; Reynolds, T.L.; Tjeerdema, R.S.; Jeffery, E.H.; German, J.B.; Ridley, W.P.; et al. Perspective—Applications of metabolomics in agriculture. J. Agric. Food Chem. 2006, 54, 8984–8994. [Google Scholar] [CrossRef] [PubMed]
- Wishart, D.S. Metabolomics: Applications to food science and nutrition research. Trends Food Sci. Technol. 2008, 19, 482–493. [Google Scholar] [CrossRef]
- Wang, D.D.; Zhang, L.X.; Huang, X.R.; Wang, X.; Yang, R.N.; Mao, J.; Wang, X.F.; Wang, X.P.; Zhang, Q.; Li, P.W. Identification of nutritional components in black sesame determined by widely targeted metabolomics and traditional Chinese medicines. Molecules 2018, 23, 12. [Google Scholar] [CrossRef] [PubMed]
- Beckner Whitener, M.E.; Stanstrup, J.; Panzeri, V.; Carlin, S.; Divol, B.; Du Toit, M.; Vrhovsek, U. Untangling the wine metabolome by combining untargeted SPME–GCXGC-TOF-MS and sensory analysis to profile sauvignon blanc co-fermented with seven different yeasts. Metabolomics 2016, 12, 53. [Google Scholar] [CrossRef]
- Martins, C.; Brandao, T.; Almeida, A.; Rocha, S.M. Metabolomics strategy for the mapping of volatile exometabolome from saccharomyces spp. Widely used in the food industry based on comprehensive two-dimensional gas chromatography. J. Sep. Sci. 2017, 40, 2228–2237. [Google Scholar] [CrossRef] [PubMed]
- Billet, K.; Houille, B.; de Bernonville, T.D.; Besseau, S.; Oudin, A.; Courdavault, V.; Delanoue, G.; Guerin, L.; Clastre, M.; Giglioli-Guivarc’h, N.; et al. Field-based metabolomics of vitis vinifera l. Stems provides new insights for genotype discrimination and polyphenol metabolism structuring. Front. Plant Sci. 2018, 9, 15. [Google Scholar] [CrossRef] [PubMed]
- Flamini, R.; De Rosso, M.; De Marchi, F.; Dalla Vedova, A.; Panighel, A.; Gardiman, M.; Bavaresco, L. Study of grape metabolomics by suspect screening analysis. In IX International Symposium on Grapevine Physiology and Biotechnology; Pinto, M., Ed.; International Society for Horticultural Science: Leuven, Belgium, 2017; Volume 1157, pp. 329–335. [Google Scholar]
- Pinu, F.R.; Edwards, P.J.B.; Jouanneau, S.; Kilmartin, P.A.; Gardner, R.C.; Villas-Boas, S.G. Sauvignon blanc metabolomics: Grape juice metabolites affecting the development of varietal thiols and other aroma compounds in wines. Metabolomics 2014, 10, 556–573. [Google Scholar] [CrossRef]
- Alanon, M.E.; Perez-Coello, M.S.; Marina, M.L. Wine science in the metabolomics era. Trac-Trends Anal. Chem. 2015, 74, 1–20. [Google Scholar] [CrossRef]
- Arapitsas, P.; Ugliano, M.; Perenzoni, D.; Angeli, A.; Pangrazzi, P.; Mattivi, F. Wine metabolomics reveals new sulfonated products in bottled white wines, promoted by small amounts of oxygen. J. Chromatogr. A 2016, 1429, 155–165. [Google Scholar] [CrossRef] [PubMed]
- Cozzolino, D. Metabolomics in grape and wine: Definition, current status and future prospects. Food Anal. Meth. 2016, 9, 2986–2997. [Google Scholar] [CrossRef]
- Lloyd, N.; Johnson, D.L.; Herderich, M.J. Metabolomics approaches for resolving and harnessing chemical diversity in grapes, yeast and wine. Aust. J. Grape Wine Res. 2015, 21, 723–740. [Google Scholar] [CrossRef]
- Roullier-Gall, C.; Heinzmann, S.S.; Garcia, J.-P.; Schmitt-Kopplin, P.; Gougeon, R.D. Chemical messages from an ancient buried bottle: Metabolomics for wine archeochemistry. NPJ Sci. Food 2017, 1. [Google Scholar] [CrossRef]
- Hall, R.; Beale, M.; Fiehn, O.; Hardy, N.; Sumner, L.; Bino, R. Plant metabolomics: The missing link in functional genomics strategies. Plant Cell 2002, 14, 1437–1440. [Google Scholar] [CrossRef] [PubMed]
- Wishart, D.S.; Knox, C.; Guo, A.C.; Eisner, R.; Young, N.; Gautam, B.; Hau, D.D.; Psychogios, N.; Dong, E.; Bouatra, S.; et al. Hmdb: A knowledgebase for the human metabolome. Nucleic Acids Res. 2009, 37, D603–D610. [Google Scholar] [CrossRef] [PubMed]
- Stitt, M.; Sulpice, R.; Keurentjes, J. Metabolic networks: How to identify key components in the regulation of metabolism and growth. Plant Physiol. 2010, 152, 428–444. [Google Scholar] [CrossRef] [PubMed]
- Villas-Bôas, S.G.; Mas, S.; Åkesson, M.; Smedsgaard, J.; Nielsen, J. Mass spectrometry in metabolome analysis. Mass Spectrom. Rev. 2005, 24, 613–646. [Google Scholar] [CrossRef] [PubMed]
- Shulaev, V. Metabolomics technology and bioinformatics. Brief. Bioinform. 2006, 7, 128–139. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Goeddel, L.C.; Patti, G.J. Maximizing the value of metabolomic data. Bioanalysis 2012, 4, 2199–2201. [Google Scholar] [CrossRef] [PubMed]
- Han, J.; Datla, R.; Chan, S.; Borchers, C.H. Mass spectrometry-based technologies for high-throughput metabolomics. Bioanalysis 2009, 1, 1665–1684. [Google Scholar] [CrossRef] [PubMed]
- Mas, S.; Villas-Bôas, S.G.; Hansen, M.E.; Åkesson, M.; Nielsen, J. A comparison of direct infusion MSand GC-MS for metabolic footprinting of yeast mutants. Biotechnol. Bioeng. 2007, 96, 1014–1022. [Google Scholar] [CrossRef] [PubMed]
- Glinski, M.; Weckwerth, W. The role of mass spectrometry in plant systems biology. Mass Spectrom. Rev. 2006, 25, 173–214. [Google Scholar] [CrossRef] [PubMed]
- Pope, G.A.; MacKenzie, D.A.; Defernez, M.; Aroso, M.A.M.M.; Fuller, L.J.; Mellon, F.A.; Dunn, W.B.; Brown, M.; Goodacre, R.; Kell, D.B.; et al. Metabolic footprinting as a tool for discriminating between brewing yeasts. Yeast 2007, 24, 667–679. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Dunn, W.B. Mass Spectrometry in Systems Biology: An Introduction. In Methods in Enzymology; Daniel, J., Malkhey, V., Hans, V.W., Eds.; Academic Press: Oxford, UK, 2011; Volume 500, pp. 15–35. [Google Scholar]
- Huo, Y.Q.; Kamal, G.M.; Wang, J.; Liu, H.L.; Zhang, G.N.; Hu, Z.Y.; Anwar, F.; Du, H.Y. H-1 NMR-based metabolomics for discrimination of rice from different geographical origins of China. J. Cereal Sci. 2017, 76, 243–252. [Google Scholar] [CrossRef]
- Li, Q.Q.; Yu, Z.B.; Zhu, D.; Meng, X.H.; Pang, X.M.; Liu, Y.; Frew, R.; Chen, H.; Chen, G. The application of NMR-based milk metabolite analysis in milk authenticity identification. J. Sci. Food Agric. 2017, 97, 2875–2882. [Google Scholar] [CrossRef] [PubMed]
- Fotakis, C.; Kokkotou, K.; Zoumpoulakis, P.; Zervou, M. NMR metabolite fingerprinting in grape derived products: An overview. Food Res. Int. 2013, 54, 1184–1194. [Google Scholar] [CrossRef]
- Lee, J.E.; Lee, B.J.; Chung, J.O.; Hwang, J.A.; Lee, S.J.; Lee, C.H.; Hong, Y.S. Geographical and climatic dependencies of green tea (Camellia sinensis) metabolites: A H-1 NMRr-based metabolomics study. J. Agric. Food Chem. 2010, 58, 10582–10589. [Google Scholar] [CrossRef] [PubMed]
- Hong, Y.S. NMR-based metabolomics in wine science. Magn. Reson. Chem. 2011, 49, S13–S21. [Google Scholar] [CrossRef] [PubMed]
- Dieterle, F.; Riefke, B.; Schlotterbeck, G.; Ross, A.; Senn, H.; Amberg, A. NMR and MS methods for metabonomics. Methods Mol. Biol. 2011, 691, 385–415. [Google Scholar] [PubMed]
- Alves, E.G.; Almeida, F.D.L.; Cavalcante, R.S.; de Brito, E.S.; Cullen, P.J.; Frias, J.M.; Bourke, P.; Fernandes, F.A.N.; Rodrigues, S. H-1 NMR spectroscopy and chemometrics evaluation of non-thermal processing of orange juice. Food Chem. 2016, 204, 102–107. [Google Scholar] [CrossRef] [PubMed]
- Spraul, M.; Link, M.; Schaefer, H.; Fang, F.; Schuetz, B. Wine analysis to check quality and authenticity by fully-automated H-1-NMR. In 38th World Congress of Vine and Wine; JeanMarie, A., Ed.; EDP Sciences: Les Ulis, France, 2015; Volume 5. [Google Scholar]
- Mulas, G.; Galaffu, M.G.; Pretti, L.; Nieddu, G.; Mercenaro, L.; Tonelli, R.; Anedda, R. NMR analysis of seven selections of vermentino grape berry: Metabolites composition and development. J. Agric. Food Chem. 2011, 59, 793–802. [Google Scholar] [CrossRef] [PubMed]
- Picone, G.; Trimigno, A.; Tessarin, P.; Donnini, S.; Rombola, A.D.; Capozzi, F. H-1 nmr foodomics reveals that the biodynamic and the organic cultivation managements produce different grape berries (Vitis vinifera L. cv. Sangiovese). Food Chem. 2016, 213, 187–195. [Google Scholar] [CrossRef] [PubMed]
- Mallamace, D.; Longo, S.; Corsaro, C. Proton NMR study of extra virgin olive oil with temperature: Freezing and melting kinetics. Physica A 2018, 499, 20–27. [Google Scholar] [CrossRef]
- Ozdemir, I.S.; Dag, C.; Makuc, D.; Ertas, E.; Plavec, J.; Bekiroglu, S. Characterisation of the turkish and slovenian extra virgin olive oils by chemometric analysis of the presaturation h-1 nmr spectra. LWT-Food Sci. Technol. 2018, 92, 10–15. [Google Scholar] [CrossRef]
- Mannina, L.; Marini, F.; Antiochia, R.; Cesa, S.; Magr, A.; Capitani, D.; Sobolev, A.P. Tracing the origin of beer samples by NMR and chemometrics: Trappist beers as a case study. Electrophoresis 2016, 37, 2710–2719. [Google Scholar] [CrossRef] [PubMed]
- Sanchez-Estebanez, C.; Ferrero, S.; Alvarez, C.M.; Villafane, F.; Caballero, I.; Blanco, C.A. Nuclear magnetic resonance methodology for the analysis of regular and non-alcoholic lager beers. Food Anal. Meth. 2018, 11, 11–22. [Google Scholar] [CrossRef]
- Bingol, K.; Brüschweiler, R. Two elephants in the room: New hybrid nuclear magnetic resonance and mass spectrometry approaches for metabolomics. Curr. Opin. Clin. Nutr. Metab. Care 2015, 18, 471–477. [Google Scholar] [CrossRef] [PubMed]
- Xia, J.; Sinelnikov, I.V.; Han, B.; Wishart, D.S. Metaboanalyst 3.0-making metabolomics more meaningful. Nucleic Acids Res. 2015, 43, W251–W257. [Google Scholar] [CrossRef] [PubMed]
- Forsberg, E.M.; Huan, T.; Rinehart, D.; Benton, H.P.; Warth, B.; Hilmers, B.; Siuzdak, G. Data processing, multi-omic pathway mapping, and metabolite activity analysis using XCMS online. Nat. Protoc. 2018, 13, 633–651. [Google Scholar] [CrossRef] [PubMed]
- Spicer, R.; Salek, R.M.; Moreno, P.; Cañueto, D.; Steinbeck, C. Navigating freely-available software tools for metabolomics analysis. Metabolomics 2017, 13, 106. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Ellinger, J.J.; Chylla, R.A.; Ulrich, E.L.; Markley, J.L. Databases and software for NMR-based metabolomics. Curr. Metab. 2013, 1. [Google Scholar] [CrossRef]
- Lewis, I.A.; Shortreed, M.R.; Hegeman, A.D.; Markley, J.L. Novel NMR and MS Approaches to Metabolomics; Fan, T., Higashi, R.M., Lane, A.N., Eds.; Springer: New York, NY, USA, 2012; Volume 17, pp. 199–230. [Google Scholar]
- Aggio, R.; Villas-Bôas, S.G.; Ruggiero, K. Metab: An R package for high-throughput analysis of metabolomics data generated by GC-MS. Bioinformatics 2011, 27, 2316–2318. [Google Scholar] [CrossRef] [PubMed]
- Kirpich, A.S.; Ibarra, M.; Moskalenko, O.; Fear, J.M.; Gerken, J.; Mi, X.; Ashrafi, A.; Morse, A.M.; McIntyre, L.M. Secimtools: A suite of metabolomics data analysis tools. BMC Bioinform. 2018, 19, 151. [Google Scholar] [CrossRef] [PubMed]
- Cambiaghi, A.; Ferrario, M.; Masseroli, M. Analysis of metabolomic data: Tools, current strategies and future challenges for omics data integration. Brief. Bioinform. 2017, 18, 498–510. [Google Scholar] [CrossRef] [PubMed]
- Uppal, K.; Soltow, Q.A.; Promislow, D.E.L.; Wachtman, L.M.; Quyyumi, A.A.; Jones, D.P. Metabnet: An R package for metabolic association analysis of high-resolution metabolomics data. Front. Bioeng. Biotechnol. 2015, 3, 87. [Google Scholar] [CrossRef] [PubMed]
- Uppal, K.; Walker, D.I.; Jones, D.P. Xmsannotator: An R package for network-based annotation of high-resolution metabolomics data. Anal. Chem. 2017, 89, 1063–1067. [Google Scholar] [CrossRef] [PubMed]
- Aggio, R.B.M.; Ruggiero, K.; Villas-Bôas, S.G. Pathway activity profiling (PAPI): From the metabolite profile to the metabolic pathway activity. Bioinformatics 2010, 26, 2969–2976. [Google Scholar] [CrossRef] [PubMed]
- Helmus, J.J.; Jaroniec, C.P. Nmrglue: An open source python package for the analysis of multidimensional nmr data. J. Biomol. NMR 2013, 55, 355–367. [Google Scholar] [CrossRef] [PubMed]
- Pedregosa, F.; Weiss, R.; Brucher, M. Scikit-learn: Machine learning in python. J. Mach. Learn. Res. 2011, 12, 2825–2830. [Google Scholar]
- Warth, B.; Levin, N.; Rinehart, D.; Teijaro, J.; Benton, H.P.; Siuzdak, G. Metabolizing data in the cloud. Trends Biotechnol. 2017, 35, 481–483. [Google Scholar] [CrossRef] [PubMed]
- Chong, J.; Soufan, O.; Li, C.; Caraus, I.; Li, S.; Bourque, G.; Wishart, D.S.; Xia, J. Metaboanalyst 4.0: Towards more transparent and integrative metabolomics analysis. Nucleic Acids Res. 2018, 46, W486–W494. [Google Scholar] [CrossRef] [PubMed]
- Haug, K.; Salek, R.M.; Steinbeck, C. Global open data management in metabolomics. Curr. Opin. Chem. Biol. 2017, 36, 58–63. [Google Scholar] [CrossRef] [PubMed]
- Haug, K.; Salek, R.M.; Conesa, P.; Hastings, J.; de Matos, P.; Rijnbeek, M.; Mahendraker, T.; Williams, M.; Neumann, S.; Rocca-Serra, P.; et al. Metabolights—An open-access general-purpose repository for metabolomics studies and associated meta-data. Nucleic Acids Res 2013, 41, D781–D786. [Google Scholar] [CrossRef] [PubMed]
- Sud, M.; Fahy, E.; Cotter, D.; Azam, K.; Vadivelu, I.; Burant, C.; Edison, A.; Fiehn, O.; Higashi, R.; Nair, K.S.; et al. Metabolomics workbench: An international repository for metabolomics data and metadata, metabolite standards, protocols, tutorials and training, and analysis tools. Nucleic Acids Res. 2016, 44, D463–D470. [Google Scholar] [CrossRef] [PubMed]
- Fiehn, O.; Robertson, D.; Griffin, J.; van der Werf, M.; Nikolau, B.; Morrison, N.; Sumner, L.W.; Goodacre, R.; Hardy, N.W.; Taylor, C.; et al. The metabolomics standards initiative (MSI). Metabolomics 2007, 3, 175–178. [Google Scholar] [CrossRef]
- Salek, R.M.; Neumann, S.; Schober, D.; Hummel, J.; Billiau, K.; Kopka, J.; Correa, E.; Reijmers, T.; Rosato, A.; Tenori, L.; et al. Coordination of standards in metabolomics (cosmos): Facilitating integrated metabolomics data access. Metabolomics 2015, 11, 1587–1597. [Google Scholar] [CrossRef] [PubMed]
- Trufelli, H.; Palma, P.; Famiglini, G.; Cappiello, A. An overview of matrix effects in liquid chromatography-mass spectrometry. Mass Spectrom. Rev. 2011, 30, 491–509. [Google Scholar] [CrossRef] [PubMed]
- Cappiello, A.; Famiglini, G.; Palma, P.; Trufelli, H. Matrix effects in liquid chromatography-mass spectrometry. J. Liquid Chromatogr. Relat. Technol. 2010, 33, 1067–1081. [Google Scholar] [CrossRef]
- Weber, C.M.; Cauchi, M.; Patel, M.; Bessant, C.; Turner, C.; Britton, L.E.; Willis, C.M. Evaluation of a gas sensor array and pattern recognition for the identification of bladder cancer from urine headspace. Analyst 2011, 136, 359–364. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Klapková, E.; Uřinovská, R.; Průša, R. The influence of matrix effects on high performance liquid chromatography-mass spectrometry methods development and validation. Vliv Matricových Efektů Vývoji 2011, 19, 5–8. [Google Scholar]
- Ye, Z.; Tsao, H.; Gao, H.; Brummel, C.L. Minimizing matrix effects while preserving throughput in LC-MS/MS bioanalysis. Bioanalysis 2011, 3, 1587–1601. [Google Scholar] [CrossRef] [PubMed]
- Wang, L.Q.; Zeng, Z.L.; Su, Y.J.; Zhang, G.K.; Zhong, X.L.; Liang, Z.P.; He, L.M. Matrix effects in analysis of β-agonists with LC-MS/MS: Influence of analyte concentration, sample source, and SPE type. J. Agric. Food Chem. 2012, 60, 6359–6363. [Google Scholar] [CrossRef] [PubMed]
- Peters, F.T.; Remane, D. Aspects of matrix effects in applications of liquid chromatography-mass spectrometry to forensic and clinical toxicology—A review. Anal. Bioanal. Chem. 2012, 403, 2155–2172. [Google Scholar] [CrossRef] [PubMed]
- Jiang, H.; Cao, H.; Zhang, Y.; Fast, D.M. Systematic evaluation of supported liquid extraction in reducing matrix effect and improving extraction efficiency in LC-MS/MS based bioanalysis for 10 model pharmaceutical compounds. J. Chromatogr. B 2012, 891–892, 71–80. [Google Scholar] [CrossRef] [PubMed]
- Villas-Bôas, S.G.; Koulman, A.; Lane, G.A. Analytical Methods from the Perspective of Method Standardization; Nielsen, J., Jewett, M.C., Eds.; Springer: Berlin/Heidelberg, Germany, 2007; Volume 18, pp. 11–52. [Google Scholar]
- Ali, K.; Maltese, F.; Toepfer, R.; Choi, Y.H.; Verpoorte, R. Metabolic characterization of palatinate German white wines according to sensory attributes, varieties, and vintages using NMR spectroscopy and multivariate data analyses. J. Biomol. NMR 2011, 49, 255–266. [Google Scholar] [CrossRef] [PubMed]
- Lee, J.E.; Hwang, G.S.; Van Den Berg, F.; Lee, C.H.; Hong, Y.S. Evidence of vintage effects on grape wines using 1H NMR-based metabolomic study. Anal. Chim. Acta 2009, 648, 71–76. [Google Scholar] [CrossRef] [PubMed]
- Atanassov, I.; Hvarleva, T.; Rusanov, K.; Tsvetkov, I.; Atanassov, A. Wine metabolite profiling: Possible application in winemaking and grapevine breeding in bulgaria. Biotechnol. Biotechnol. Equip. 2009, 23, 1449–1452. [Google Scholar] [CrossRef]
- Howell, K.S.; Cozzolino, D.; Bartowsky, E.J.; Fleet, G.H.; Henschke, P.A. Metabolic profiling as a tool for revealing Saccharomyces interactions during wine fermentation. FEMS Yeast Res. 2006, 6, 91–101. [Google Scholar] [CrossRef] [PubMed]
- Tumanov, S.; Zubenko, Y.; Greven, M.; Greenwood, D.R.; Shmanai, V.; Villas-Boas, S.G. Comprehensive lipidome profiling of Sauvignon blanc grape juice. Food Chem. 2015, 180, 249–256. [Google Scholar] [CrossRef] [PubMed]
- Pinto, J.; Oliveira, A.S.; Azevedo, J.; De Freitas, V.; Lopes, P.; Roseira, I.; Cabral, M.; de Pinho, P.G. Assessment of oxidation compounds in oaked chardonnay wines: A GC-MS and H-1 NMR metabolomics approach. Food Chem. 2018, 257, 120–127. [Google Scholar] [CrossRef] [PubMed]
- Arita, K.; Honma, T.; Suzuki, S. Comprehensive and comparative lipidome analysis of vitis vinifera l. Cv. Pinot noir and japanese indigenous v. Vinifera l. Cv. Koshu grape berries. PLoS ONE 2017, 12, e0186952. [Google Scholar] [CrossRef] [PubMed]
- Tejero Rioseras, A.; Garcia Gomez, D.; Ebert, B.E.; Blank, L.M.; Ibáñez, A.J.; Sinues, P.M.L. Comprehensive real-time analysis of the yeast volatilome. Sci. Rep. 2017, 7, 14236. [Google Scholar] [CrossRef] [PubMed]
- Amann, A.; Costello, B.L.; Miekisch, W.; Schubert, J.; Buszewski, B.; Pleil, J.; Ratcliffe, N.; Risby, T. The human volatilome: Volatile organic compounds (VOCs) in exhaled breath, skin emanations, urine, feces and saliva. J. Breath Res. 2014, 8, 034001. [Google Scholar] [CrossRef] [PubMed]
- Opitz, P.; Herbarth, O. The volatilome—Investigation of volatile organic metabolites (vom) as potential tumor markers in patients with head and neck squamous cell carcinoma (hnscc). J. Otolaryngol. Head Neck Surg. 2018, 47, 42. [Google Scholar] [CrossRef] [PubMed]
- Skogerson, K.; Runnebaum, R.O.N.; Wohlgemuth, G.; De Ropp, J.; Heymann, H.; Fiehn, O. Comparison of gas chromatography-coupled time-of-flight mass spectrometry and 1H nuclear magnetic resonance spectroscopy metabolite identification in white wines from a sensory study investigating wine body. J. Agric. Food Chem. 2009, 57, 6899–6907. [Google Scholar] [CrossRef] [PubMed]
- Zott, K.; Thibon, C.; Bely, M.; Lonvaud-Funel, A.; Dubourdieu, D.; Masneuf-Pomarede, I. The grape must non-Saccharomyces microbial community: Impact on volatile thiol release. Int. J. Food Microbiol. 2011, 151, 210–215. [Google Scholar] [CrossRef] [PubMed]
- Pinu, F.R.; Edwards, P.J.B.; Gardner, R.C.; Villas-Boas, S.G. Nitrogen and carbon assimilation by Saccharomyces cerevisiae during Sauvignon blanc juice fermentation. FEMS Yeast Res. 2014, 14, 1206–1222. [Google Scholar] [CrossRef] [PubMed]
- Casu, F.; Pinu, F.R.; Fedrizzi, B.; Greenwood, D.R.; Villas-Boas, S.G. The effect of linoleic acid on the Sauvignon blanc fermentation by different wine yeast strains. FEMS Yeast Res. 2016, 16, fow050. [Google Scholar] [CrossRef] [PubMed]
- Schueuermann, C.; Steel, C.C.; Blackman, J.W.; Clark, A.C.; Schwarz, L.J.; Moraga, J.; Collado, I.G.; Schmidtke, L.M. A GC-MS untargeted metabolomics approach for the classification of chemical differences in grape juices based on fungal pathogen. Food Chem. 2019, 270, 375–384. [Google Scholar] [CrossRef] [PubMed]
- Cuadros-Inostroza, A.; Ruíz-Lara, S.; González, E.; Eckardt, A.; Willmitzer, L.; Peña-Cortés, H. Gc–ms metabolic profiling of cabernet sauvignon and merlot cultivars during grapevine berry development and network analysis reveals a stage- and cultivar-dependent connectivity of primary metabolites. Metabolomics 2016, 12, 39. [Google Scholar] [CrossRef] [PubMed]
- Theodoridis, G.; Gika, H.; Franceschi, P.; Caputi, L.; Arapitsas, P.; Scholz, M.; Masuero, D.; Wehrens, R.; Vrhovsek, U.; Mattivi, F. LC-MS based global metabolite profiling of grapes: Solvent extraction protocol optimisation. Metabolomics 2012, 8, 175–185. [Google Scholar] [CrossRef]
- Capone, D.L.; Pardon, K.H.; Cordente, A.G.; Jeffery, D.W. Identification and quantitation of 3-s-cysteinylglycinehexan-1-ol (cysgly-3-mh) in sauvignon blanc grape juice by HPLC-MS/MS. J. Agric. Food Chem. 2011, 59, 11204–11210. [Google Scholar] [CrossRef] [PubMed]
- Capone, D.L.; Black, C.A.; Jeffery, D.W. Effects on 3-mercaptohexan-1-ol precursor concentrations from prolonged storage of Sauvignon blanc grapes prior to crushing and pressing. J. Agric. Food Chem. 2012, 60, 3515–3523. [Google Scholar] [CrossRef] [PubMed]
- Arapitsas, P.; Della Corte, A.; Gika, H.; Narduzzi, L.; Mattivi, F.; Theodoridis, G. Studying the effect of storage conditions on the metabolite content of red wine using HILIC LC-MS based metabolomics. Food Chem. 2016, 197, 1331–1340. [Google Scholar] [CrossRef] [PubMed]
- Diaz, R.; Gallart-Ayala, H.; Sancho, J.V.; Nunez, O.; Zamora, T.; Martins, C.P.B.; Hernandez, F.; Hernandez-Cassou, S.; Saurina, J.; Checa, A. Told through the wine: A liquid chromatography-mass spectrometry interplatform comparison reveals the influence of the global approach on the final annotated metabolites in non-targeted metabolomics. J. Chromatogr. A 2016, 1433, 90–97. [Google Scholar] [CrossRef] [PubMed]
- Roullier-Gall, C.; Hemmler, D.; Witting, M.; Moritz, F.; Heinzmann, S.; Jeandet, P.; Gonsior, M.; Gougeon, R.; Schmitt-Kopplin, P. Metabolomics characterization of bottled wine: Impact of environmental parameters. In Abstracts of Papers of the American Chemical Society; American Chemical Society: Washington, DC, USA, 2016; Volume 252. [Google Scholar]
- Roullier-Gall, C.; Witting, M.; Gougeon, R.D.; Schmitt-Kopplin, P. High precision mass measurements for wine metabolomics. Front. Chem. 2014, 2, 102. [Google Scholar] [CrossRef] [PubMed]
- Roullier-Gall, C.; Witting, M.; Tziotis, D.; Ruf, A.; Gougeon, R.D.; Schmitt-Kopplin, P. Integrating analytical resolutions in non-targeted wine metabolomics. Tetrahedron 2015, 71, 2983–2990. [Google Scholar] [CrossRef]
- Ruocco, S.; Stefanini, M.; Stanstrup, J.; Perenzoni, D.; Mattivi, F.; Vrhovsek, U. The metabolomic profile of red non-v-vinifera genotypes. Food Res. Int. 2017, 98, 10–19. [Google Scholar] [CrossRef] [PubMed]
- Son, H.S.; Hwang, G.S.; Kim, K.M.; Ahn, H.J.; Park, W.M.; Van Den Berg, F.; Hong, Y.S.; Lee, C.H. Metabolomic studies on geographical grapes and their wines using 1H NMR analysis coupled with multivariate statistics. J. Agric. Food Chem. 2009, 57, 1481–1490. [Google Scholar] [CrossRef] [PubMed]
- Rochfort, S.; Ezernieks, V.; Bastian, S.E.P.; Downey, M.O. Sensory attributes of wine influenced by variety and berry shading discriminated by NMR metabolomics. Food Chem. 2010, 121, 1296–1304. [Google Scholar] [CrossRef]
- Hong, Y.S.; Cilindre, C.; Liger-Belair, G.; Jeandet, P.; Hertkorn, N.; Schmitt-Kopplin, P. Metabolic influence of Botrytis cinerea infection in champagne base wine. J. Agric. Food Chem. 2011, 59, 7237–7245. [Google Scholar] [CrossRef] [PubMed]
- Zhu, J.Y.; Hu, B.R.; Lu, J.; Xu, S.C. Analysis of metabolites in Cabernet sauvignon and shiraz dry red wines from Shanxi by H1 NMR spectroscopy combined with pattern recognition analysis. Open Chem. 2018, 16, 446–452. [Google Scholar] [CrossRef]
- Simó, C.; Moreno-Arribas, M.V.; Cifuentes, A. Ion-trap versus time-of-flight mass spectrometry coupled to capillary electrophoresis to analyze biogenic amines in wine. J. Chromatogr. A 2008, 1195, 150–156. [Google Scholar] [CrossRef] [PubMed]
- Vanhoenacker, G.; De Villiers, A.; Lazou, K.; De Keukeleire, D.; Sandra, P. Comparison of high-performance liquid chromatography-mass spectroscopy and capillary electrophoresis-mass spectroscopy for the analysis of phenolic compounds in diethyl ether extracts of red wines. Chromatographia 2001, 54, 309–315. [Google Scholar] [CrossRef]
- Wine-Omics. Available online: https://www.nature.com/articles/455699a (accessed on 2 October 2008).
- Arapitsas, P.; Scholz, M.; Vrhovsek, U.; Di Blasi, S.; Biondi Bartolini, A.; Masuero, D.; Perenzoni, D.; Rigo, A.; Mattivi, F. A metabolomic approach to the study of wine micro-oxygenation. PLoS ONE 2012, 7, e37783. [Google Scholar] [CrossRef] [PubMed]
- Liu, Y.Z.; Forcisi, S.; Harir, M.; Deleris-Bou, M.; Krieger-Weber, S.; Lucio, M.; Longin, C.; Degueurce, C.; Gougeon, R.D.; Schmitt-Kopplin, P.; et al. New molecular evidence of wine yeast-bacteria interaction unraveled by non-targeted exometabolomic profiling. Metabolomics 2016, 12, 16. [Google Scholar] [CrossRef]
- Conterno, L.; Aprea, E.; Franceschi, P.; Viola, R.; Vrhovsek, U. Overview of Dekkera bruxellensis behaviour in an ethanol-rich environment using untargeted and targeted metabolomic approaches. Food Res. Int. 2013, 51, 670–678. [Google Scholar] [CrossRef]
- Rocchetti, G.; Gatti, M.; Bavaresco, L.; Lucini, L. Untargeted metabolomics to investigate the phenolic composition of Chardonnay wines from different origins. J. Food Compos. Anal. 2018, 71, 87–93. [Google Scholar] [CrossRef]
- López-Rituerto, E.; Savorani, F.; Avenoza, A.; Busto, J.H.; Peregrina, J.M.; Engelsen, S.B. Investigations of la rioja terroir for wine production using 1H NMR metabolomics. J. Agric. Food Chem. 2012, 60, 3452–3461. [Google Scholar] [CrossRef] [PubMed]
- Amargianitaki, M.; Spyros, A. NMR-based metabolomics in wine quality control and authentication. Chem. Biol. Technol. Agric. 2017, 4, 9. [Google Scholar] [CrossRef]
- Cassino, C.; Tsolakis, C.; Bonello, F.; Gianotti, V.; Osella, D. Effects of area, year and climatic factors on barbera wine characteristics studied by the combination of 1H-NMR metabolomics and chemometrics. J. Wine Res. 2017, 28, 259–277. [Google Scholar] [CrossRef]
- Roullier-Gall, C.; Boutegrabet, L.; Gougeon, R.D.; Schmitt-Kopplin, P. A grape and wine chemodiversity comparison of different appellations in burgundy: Vintage vs terroir effects. Food Chem. 2014, 152, 100–107. [Google Scholar] [CrossRef] [PubMed]
- Roullier-Gall, C.; Lucio, M.; Noret, L.; Schmitt-Kopplin, P.; Gougeon, R.D. How subtle is the “terroir” effect? Chemistry-related signatures of two “climats de bourgogne”. PLoS ONE 2014, 9, e97615. [Google Scholar] [CrossRef] [PubMed]
- Bokulich, N.A.; Collins, T.S.; Masarweh, C.; Allen, G.; Heymann, H.; Ebeler, S.E.; Mills, D.A. Associations among wine grape microbiome, metabolome, and fermentation behavior suggest microbial contribution to regional wine characteristics. mBio 2016, 7. [Google Scholar] [CrossRef] [PubMed]
- Anesi, A.; Stocchero, M.; Dal Santo, S.; Commisso, M.; Zenoni, S.; Ceoldo, S.; Tornielli, G.B.; Siebert, T.E.; Herderich, M.; Pezzotti, M.; et al. Towards a scientific interpretation of the terroir concept: Plasticity of the grape berry metabolome. BMC Plant Biol. 2015, 15, 191. [Google Scholar] [CrossRef] [PubMed]
- King, E.S.; Kievit, R.L.; Curtin, C.; Swiegers, J.H.; Pretorius, I.S.; Bastian, S.E.P.; Leigh Francis, I. The effect of multiple yeasts co-inoculations on Sauvignon blanc wine aroma composition, sensory properties and consumer preference. Food Chem. 2010, 122, 618–626. [Google Scholar] [CrossRef]
- Ciani, M.; Capece, A.; Comitini, F.; Canonico, L.; Siesto, G.; Romano, P. Yeast interactions in inoculated wine fermentation. Front. Microbiol. 2016, 7, 555. [Google Scholar] [CrossRef] [PubMed]
- Pizarro, F.; Vargas, F.A.; Agosin, E. A systems biology perspective of wine fermentations. Yeast 2007, 24, 977–991. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Tumanov, S.; Pinu, F.R.; Greenwood, D.R.; Villas-Boas, S.G. The effect of free fatty acids and lipolysis on sauvignon blanc fermentation. Aust. J. Grape Wine Res. 2017, in press. [Google Scholar] [CrossRef]
- Aliverdieva, D.A.; Mamaev, D.V.; Bondarenko, D.I.; Sholtz, K.F. Properties of yeast Saccharomyces cerevisiae plasma membrane dicarboxylate transporter. Biochemistry 2006, 71, 1161–1169. [Google Scholar] [CrossRef] [PubMed]
- Crépin, L.; Nidelet, T.; Sanchez, I.; Dequin, S.; Camarasa, C. Sequential use of nitrogen compounds by yeast during wine fermentation: A model based on kinetic and regulation characteristics of nitrogen permeases. Appl. Environ. Microbiol. 2012, in press. [Google Scholar]
- Cooper, T.G. Nitrogen metabolism in Saccharomyces cerevisiae. In The Molecular Biology of the Yeast Saccharomyces: Metabolism and Gene Expression; Strathern, J.N., Jones, E.W., Broach, J.R., Eds.; Cold Spring Harbor Laboratory Press: Cold Spring Harbor, NY, USA, 1982; pp. 39–99. [Google Scholar]
- Hofman-Bang, J. Nitrogen catabolite repression in saccharomyces cerevisiae. Mol. Biotechnol. 1999, 12, 35–73. [Google Scholar] [CrossRef]
- Henscke, P.; Jiranek, V. Yeasts—Metabolism of nitrogen compounds. In Wine Microbiology, Biotechnology; Harwood Academic: Chur, Switzerland, 1993; pp. 77–164. [Google Scholar]
- Fairbairn, S.; McKinnon, A.; Musarurwa, H.T.; Ferreira, A.C.; Bauer, F.F. The impact of single amino acids on growth and volatile aroma production by Saccharomyces cerevisiae strains. Front. Microbiol. 2017, 8, 2554. [Google Scholar] [CrossRef] [PubMed]
- Peltier, E.; Bernard, M.; Trujillo, M.; Prodhomme, D.; Barbe, J.-C.; Gibon, Y.; Marullo, P. Wine yeast phenomics: A standardized fermentation method for assessing quantitative traits of Saccharomyces cerevisiae strains in enological conditions. PLoS ONE 2018, 13, e0190094. [Google Scholar] [CrossRef] [PubMed]
- Varela, C.; Torrea, D.; Schmidt, S.A.; Ancin-Azpilicueta, C.; Henschke, P.A. Effect of oxygen and lipid supplementation on the volatile composition of chemically defined medium and chardonnay wine fermented with Saccharomyces cerevisiae. Food Chem. 2012, 135, 2863–2871. [Google Scholar] [CrossRef] [PubMed]
- López-Malo, M.; Querol, A.; Guillamon, J.M. Metabolomic comparison of Saccharomyces cerevisiae and the cryotolerant species S. bayanus var. uvarum and S. kudriavzevii during wine fermentation at low temperature. PLoS ONE 2013, 8, e60135. [Google Scholar] [CrossRef] [PubMed]
- Pizarro, F.J.; Jewett, M.C.; Nielsen, J.; Agosin, E. Growth temperature exerts differential physiological and transcriptional responses in laboratory and wine strains of Saccharomyces cerevisiae. Appl. Environ. Microbiol. 2008, 74, 6358. [Google Scholar] [CrossRef] [PubMed]
- Rollero, S.; Bloem, A.; Camarasa, C.; Sanchez, I.; Ortiz-Julien, A.; Sablayrolles, J.-M.; Dequin, S.; Mouret, J.-R. Combined effects of nutrients and temperature on the production of fermentative aromas by Saccharomyces cerevisiae during wine fermentation. Appl. Microbiol. Biotechnol. 2015, 99, 2291–2304. [Google Scholar] [CrossRef] [PubMed]
- Aceituno, F.F.; Orellana, M.; Torres, J.; Mendoza, S.; Slater, A.W.; Melo, F.; Agosin, E. Oxygen response of the wine yeast Saccharomyces cerevisiae EC1118 grown under carbon-sufficient, nitrogen-limited enological conditions. Appl. Environ. Microbiol. 2012, 78, 8340. [Google Scholar] [CrossRef] [PubMed]
- Orellana, M.; Aceituno, F.F.; Slater, A.W.; Almonacid, L.I.; Melo, F.; Agosin, E. Metabolic and transcriptomic response of the wine yeast Saccharomyces cerevisiae strain ec1118 after an oxygen impulse under carbon-sufficient, nitrogen-limited fermentative conditions. FEMS Yeast Res. 2014, 14, 412–424. [Google Scholar] [CrossRef] [PubMed]
- Mendes, I.; Sanchez, I.; Franco-Duarte, R.; Camarasa, C.; Schuller, D.; Dequin, S.; Sousa, M.J. Integrating transcriptomics and metabolomics for the analysis of the aroma profiles of Saccharomyces cerevisiae strains from diverse origins. BMC Genomics 2017, 18, 13. [Google Scholar] [CrossRef] [PubMed]
- Hochberg, U.; Degu, A.; Cramer, G.R.; Rachmilevitch, S.; Fait, A. Cultivar specific metabolic changes in grapevines berry skins in relation to deficit irrigation and hydraulic behavior. Plant Physiol. Biochem. 2015, 88, 42–52. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Pinasseau, L.; Vallverdu-Queralt, A.; Verbaere, A.; Roques, M.; Meudec, E.; Le Cunff, L.; Peros, J.P.; Ageorges, A.; Sommerer, N.; Boulet, J.C.; et al. Cultivar diversity of grape skin polyphenol composition and changes in response to drought investigated by LC-MS based metabolomics. Front. Plant Sci. 2017, 8, 24. [Google Scholar] [CrossRef] [PubMed]
- Brilli, M.; Asquini, E.; Moser, M.; Bianchedi, P.L.; Perazzolli, M.; Si-Ammour, A. A multi-omics study of the grapevine-downy mildew (Plasmopara viticola) pathosystem unveils a complex protein coding- and noncoding-based arms race during infection. Sci. Rep. 2018, 8, 757. [Google Scholar] [CrossRef] [PubMed]
- Benheim, D.; Rochfort, S.; Ezernieks, V.; Korosi, G.A.; Powell, K.S.; Robertson, E.; Potter, I.D. Early detection of grape phylloxera (daktulosphaira vitifoliae fitch) infestation through identification of chemical biomarkers. In V international Phylloxera Symposium; Griesser, M., Forneck, A., Eds.; International Society for Horticultural Science: Leuven, Belgium, 2011; Volume 904, pp. 17–24. [Google Scholar]
- Benheim, D.; Rochfort, S.; Korosi, G.A.; Powell, K.S.; Robertson, E.; Potter, I.D. Nuclear magnetic resonance metabolic profiling of leaves from vitis vinifera infested with root-feeding grape phylloxera (daktulosphaira vitifoliae fitch) under field conditions. In VI International Phylloxera Symposium; Ollat, N., Papura, D., Eds.; International Society for Horticultural Science: Leuven, Belgium, 2014; Volume 1045, pp. 59–66. [Google Scholar]
- Wishart, D.S. Advances in metabolite identification. Bioanalysis 2011, 3, 1769–1782. [Google Scholar] [CrossRef] [PubMed]
- Date, Y.; Kikuchi, J. Application of a deep neural network to metabolomics studies and its performance in determining important variables. Anal. Chem. 2018, 90, 1805–1810. [Google Scholar] [CrossRef] [PubMed]
- Trivedi, D.K.; Hollywood, K.A.; Goodacre, R. Metabolomics for the masses: The future of metabolomics in a personalized world. New Horiz. Transl. Med. 2017, 3, 294–305. [Google Scholar] [CrossRef] [PubMed]
- Scalbert, A.; Brennan, L.; Manach, C.; Andres-Lacueva, C.; Dragsted, L.O.; Draper, J.; Rappaport, S.M.; Van Der Hooft, J.J.J.; Wishart, D.S. The food metabolome: A window over dietary exposure. Am. J. Clin. Nutr. 2014, 99, 1286–1308. [Google Scholar] [CrossRef] [PubMed]
- Förster, J.; Famili, I.; Fu, P.; Palsson, B.Ø.; Nielsen, J. Genome-scale reconstruction of the saccharomyces cerevisiae metabolic network. Genome Res. 2003, 13, 244–253. [Google Scholar] [CrossRef] [PubMed]
- Zhang, C.; Hua, Q. Applications of genome-scale metabolic models in biotechnology and systems medicine. Front. Physiol. 2016, 6, 413. [Google Scholar] [CrossRef] [PubMed]
- Kerkhoven, E.J.; Lahtvee, P.-J.; Nielsen, J. Applications of computational modeling in metabolic engineering of yeast. FEMS Yeast Res. 2015, 15, 1–13. [Google Scholar] [CrossRef] [PubMed]
- Heavner, B.D.; Price, N.D. Comparative analysis of yeast metabolic network models highlights progress, opportunities for metabolic reconstruction. PLOS Comput. Biol. 2015, 11, e1004530. [Google Scholar] [CrossRef] [PubMed]
- Mendoza, S.N.; Cañón, P.M.; Contreras, Á.; Ribbeck, M.; Agosín, E. Genome-scale reconstruction of the metabolic network in Oenococcus oeni to assess wine malolactic fermentation. Front. Microbiol. 2017, 8, 534. [Google Scholar] [CrossRef] [PubMed]
- Saitua, F.; Torres, P.; Pérez-Correa, J.R.; Agosin, E. Dynamic genome-scale metabolic modeling of the yeast Pichia pastoris. BMC Syst. Biol. 2017, 11, 27. [Google Scholar] [CrossRef] [PubMed]
- Pizarro, F.; Varela, C.; Martabit, C.; Bruno, C.; Perez-Correa, J.R.; Agosin, E. Coupling kinetic expressions and metabolic networks for predicting wine fermentations. Biotechnol. Bioeng. 2007, 98, 986–998. [Google Scholar] [CrossRef] [PubMed]
- Vargas, F.A.; Pizarro, F.; Pérez-Correa, J.R.; Agosin, E. Expanding a dynamic flux balance model of yeast fermentation to genome-scale. BMC Syst. Biol. 2011, 5, 75. [Google Scholar] [CrossRef] [PubMed]
| Approach | Advantages | Disadvantages |
|---|---|---|
| Targeted metabolite analysis |
|
|
| Untargeted metabolite profiling |
|
|
| Analytical Technique | Advantages | Disadvantages | Use in Grape and Wine Research |
|---|---|---|---|
| GC-MS |
|
| [8,30,31,34,98,99,100,101,102,103,104,105,106,107,108,109,110] |
| LC-MS |
|
| [36,111,112,113,114,115,116,117,118,119] |
| NMR |
|
| [59,60,95,107,120,121,122,123] |
| CE-MS |
|
| [124,125] |
© 2018 by the author. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Pinu, F.R. Grape and Wine Metabolomics to Develop New Insights Using Untargeted and Targeted Approaches. Fermentation 2018, 4, 92. https://doi.org/10.3390/fermentation4040092
Pinu FR. Grape and Wine Metabolomics to Develop New Insights Using Untargeted and Targeted Approaches. Fermentation. 2018; 4(4):92. https://doi.org/10.3390/fermentation4040092
Chicago/Turabian StylePinu, Farhana R. 2018. "Grape and Wine Metabolomics to Develop New Insights Using Untargeted and Targeted Approaches" Fermentation 4, no. 4: 92. https://doi.org/10.3390/fermentation4040092
APA StylePinu, F. R. (2018). Grape and Wine Metabolomics to Develop New Insights Using Untargeted and Targeted Approaches. Fermentation, 4(4), 92. https://doi.org/10.3390/fermentation4040092





