Recent Progress in the Microbial Production of Pyruvic Acid
Abstract
:1. Introduction
2. Microbial Formation of Pyruvate
3. Medium Optimization
4. Metabolic Engineering of Pathways
5. Cofactor Engineering
6. Process Engineering
7. Other Biological Methods
8. Downstream Processes
9. Conclusions
Acknowledgments
Author Contributions
Conflicts of Interest
References
- Park, H.-S.; Lee, J.-Y.; Kim, H.-S. Production of L-DOPA (3,4,-dihydroxyphenyl-l-alanine) from benzene by using a hybrid pathway. Biotechnol. Bioeng. 1998, 58, 339–343. [Google Scholar] [CrossRef]
- Zhang, Y.; Tao, F.; Du, M.; Ma, C.; Qiu, J.; Gu, L.; He, X.; Xu, P. An efficient method for N-acetyl-d-neuraminic acid production using coupled bacterial cells with a safe temperature-induced system. Appl. Microbiol. Biotechnol. 2010, 86, 481–489. [Google Scholar] [CrossRef] [PubMed]
- Rosche, B.; Sandford, V.; Breuer, M.; Hauer, B.; Rogers, P. Biotransformation of benzaldehyde into (R)-phenylacetylcarbinol by filamentous fungi or their extracts. Appl. Microbiol. Biotechnol. 2001, 57, 309–315. [Google Scholar] [PubMed]
- Reisse, S.; Haack, M.; Garbe, D.; Sommer, B.; Steffler, F.; Carsten, J.; Bohnen, F.; Sieber, V.; Brück, T. In vitro bioconversion of pyruvate to n-butanol with minimized cofactor utilization. Front. Bioeng. Biotechnol. 2016, 4, 74. [Google Scholar] [CrossRef] [PubMed]
- Stine, A.; Zhang, M.; Ro, S.; Clendennen, S.; Shelton, M.C.; Tyo, K.E.J.; Broadbelt, L.J. Exploring De Novo metabolic pathways from pyruvate to propionic acid. Biotechnol. Prog. 2016, 32, 303–311. [Google Scholar] [CrossRef] [PubMed]
- Zhang, X.; Tervo, C.J.; Reed, J.L. Metabolic assessment of E. coli as a biofactory for commercial products. Metab. Eng. 2016, 35, 64–75. [Google Scholar] [CrossRef] [PubMed]
- Li, Y.; Chen, J.; Lun, S.-Y. Biotechnological production of pyruvic acid. Appl. Microbiol. Biotechnol. 2001, 57, 451–459. [Google Scholar] [PubMed]
- Xu, P.; Qiu, J.; Gao, C.; Ma, C. Biotechnological routes to pyruvate production. J. Biosci. Bioeng. 2008, 105, 169–175. [Google Scholar] [CrossRef] [PubMed]
- Mazzio, E.; Soliman, K.F. Cytoprotection of pyruvic acid and reduced beta-nicotinamide adenine dinucleotide against hydrogen peroxide toxicity in neuroblastoma cells. Neurochem. Res. 2003, 28, 733–741. [Google Scholar] [CrossRef] [PubMed]
- Wang, X.; Perez, E.; Liu, R.; Yan, L.-J.; Mallet, R.T.; Yang, S.-H. Pyruvate protects mitochondria from oxidative stress in human neuroblastoma SK-N-SH cells. Brain Res. 2007, 1132, 1–9. [Google Scholar] [CrossRef] [PubMed]
- Nakamichi, N.; Kambe, Y.; Oikawa, H.; Ogura, M.; Takano, K.; Tamaki, K.; Inoue, M.; Hinoi, E.; Yoneda, Y. Protection by exogenous pyruvate through a mechanism related to monocarboxylate transporters against cell death induced by hydrogen peroxide in cultured rat cortical neurons. J. Neurochem. 2005, 93, 84–93. [Google Scholar] [CrossRef] [PubMed]
- Yoo, M.H.; Lee, J.Y.; Lee, S.E.; Hok, J.Y.; Yoon, Y.H. Protection by pyruvate of rat retinal cells against zinc toxicity in vitro, and pressure-induced ischemia in vivo. Invest. Ophthalmol. Visual Sci. 2004, 45, 1523–1530. [Google Scholar] [CrossRef]
- Mongan, P.D.; Capacchione, J.; Fontana, J.L.; West, S.; Bünger, R. Pyruvate improves cerebral metabolism during hemorrhagic shock. Am. J. Physiol. Heart Circ. Physiol. 2001, 281, H854–H864. [Google Scholar] [PubMed]
- Ryou, M.-G.; Liu, R.; Ren, M.; Sun, J.; Mallet, R.T.; Yang, S.-H. Pyruvate protects the brain against ischemia-reperfusion injury by activating the erythropoietin signaling pathway. Stroke 2012, 43, 1101–1107. [Google Scholar] [CrossRef] [PubMed]
- Hasenfuss, G.; Maier, L.S.; Hermann, H.-P.; Lüers, C.; Hünlich, M.; Zeitz, O.; Janssen, P.M.L.; Pieske, B. Influence of pyruvate on contractile performance and Ca2+ cycling in isolated failing human myocardium. Circulation 2002, 105, 194–199. [Google Scholar] [CrossRef] [PubMed]
- Hermann, H.-P.; Arp, J.; Pieske, B.; Kögler, H.; Baron, S.; Hanssen, P.M. L.; Hasenfuss, F. Improved systolic and diastolic myocardial function with intracoronary pyruvate in patients with congestive heart failure. Eur. J. Heart Fail. 2004, 6, 213–218. [Google Scholar] [CrossRef] [PubMed]
- Kalman, D.; Colker, C.M.; Wilets, I.; Roufs, J.B.; Antonio, J. The effects of pyruvate supplementation on body composition in overweight individuals. Nutrition 1999, 15, 337–340. [Google Scholar] [CrossRef]
- Koh-Banerjee, P.K.; Ferreira, M.P.; Greenwood, M.; Bowden, R.G.; Cowan, P.N.; Almada, A.; Kreider, R.B. Effects of calcium pyruvate supplementation during training on body composition, exercise capacity, and metabolic responses to exercise. Nutrition 2005, 21, 312–319. [Google Scholar] [CrossRef] [PubMed]
- EFSA. Calcium acetate, calcium pyruvate, calcium succinate, magnesium pyruvate magnesium succinate and potassium malate added for nutritional purposes to food supplements. EFSA J. 2009, 1088, 1–25. [Google Scholar]
- Kim, J.-G.; Park, S.-J.; Damsté, J.S.S.; Schouten, S.; Rijpstra, W.I.C.; Jung, M.-Y.; Kim, S.-J.; Gwak, J.-H.; Hong, H.; Si, S.-J.; et al. Hydrogen peroxide detoxification is a key mechanism for growth of ammonia-oxidizing bacteria. Proc. Natl. Acad. Sci. USA 2016, 113, 7888–7893. [Google Scholar] [CrossRef] [PubMed]
- Kitahara, K.; Fukui, S. On the pyruvic acid fermentation (I): Selection of species from genus Mucor, and examination of production conditions. J. Ferment. Technol. 1951, 29, 227–233. [Google Scholar]
- Kitahara, K.; Fukui, S. On the pyruvic acid fermentation (II): Possibility of biochemical determination of vitamin B-1 by Mucor mandshuricus Saito. J. Ferment. Technol. 1951, 29, 287–290. [Google Scholar]
- Kitahara, K.; Fukui, S. On the pyruvic acid fermentation (III): Substances which inhibit the accumulation of pyruvic acid. J. Ferment. Technol. 1951, 29, 326–331. [Google Scholar]
- Kitahara, K.; Fukui, S. On the pyruvic acid fermentation (IV): The mechanism of pyruvate acid accumulation by Mucor mandshuricus. J. Ferment. Technol. 1951, 29, 378–384. [Google Scholar]
- Kitahara, K.; Fukui, S. Beriberi-like symptoms in microorganisms. J. Gen. Appl. Microbiol. 1955, 1, 61–76. [Google Scholar] [CrossRef]
- Yokota, A.; Takao, S. Pyruvic acid production by lipoic acid auxotrophs of Enterobacter aerogenes. Agric. Biol. Chem. 1989, 53, 705–711. [Google Scholar] [CrossRef]
- Izumi, Y.; Matsumura, Y.; Tani, Y.; Yamada, H. Pyruvic acid production from 1,2-propanediol by thiamin-requiring Acinetobacter sp. 80-M. Agri. Biol. Chem. 1982, 46, 2673–2679. [Google Scholar] [CrossRef]
- Takao, S.; Tanida, M. Pyruvic acid production by Schizophyllum commune: Pyruvic acid fermentation by Basidiomycetes (I). J. Ferm. Technol. 1982, 60, 277–280. [Google Scholar]
- Moriguchi, M. Fermentative production of pyruvic acid from citrus peel extract by Debaryomyces coudertii. Agric. Biol. Chem. 1982, 64, 955–961. [Google Scholar] [CrossRef]
- Morgunov, I.G.; Kamzolova, S.V.; Perevoznikova, O.A.; Shishkanova, N.V.; Finogenova, T.V. Pyruvic acid production by a thiamine auxotroph of Yarrowia lipolytica. Process Biochem. 2004, 39, 1469–1474. [Google Scholar] [CrossRef]
- Yokota, A.; Shimizu, H.; Terasawa, Y.; Takaoka, N.; Tomita, F. Pyruvic acid production by a lipoic acid auxotroph of Escherichia coli W1485. Appl. Microbiol. Biotechnol. 1994, 41, 638–643. [Google Scholar] [CrossRef]
- Wang, Q.; He, P.; Lu, D.; Shen, A.; Jiang, N. Screening of pyruvate-producing yeast and effect of nutritional conditions on pyruvate production. Lett. Appl. Microbiol. 2002, 35, 338–342. [Google Scholar] [CrossRef] [PubMed]
- Yonehara, T.; Miyata, R. Fermentative production of pyruvate from glucose by Torulopsis glabrata. J. Ferment. Bioeng. 1994, 78, 155–159. [Google Scholar] [CrossRef]
- Miyata, R.; Yonehara, T. Breeding of high-pyruvate-producing Torulopsis glabrata and amino acid auxotrophic mutants. J. Biosci. Bioeng. 2000, 90, 137–141. [Google Scholar] [CrossRef]
- Li, S.; Chen, X.; Liu, L.; Chen, J. Pyruvate production in Candida glabrata: Manipulation and optimization of physiological function. Crit. Rev. Biotechnol. 2016, 36, 1–10. [Google Scholar] [CrossRef] [PubMed]
- Kamzolova, S.V.; Morgunov, I.G. Biosynthesis of pyruvic acid from glucose by Blastobotrys adeninivorans. Appl. Microbiol. Biotechnol. 2016, 100, 7689–7697. [Google Scholar] [CrossRef] [PubMed]
- Miyata, R.; Yonehara, T. Improvement of fermentative production of pyruvate from glucose by Torulopsis glabrata IFO 0005. J. Ferment. Bioeng. 1996, 82, 475–479. [Google Scholar] [CrossRef]
- Li, Y.; Chen, J.; Liang, D.-F.; Lun, S.-Y. Effect of nitrogen source and nitrogen concentration on the production of pyruvate by Torulopsis glabrata. J. Biotechnol. 2000, 81, 27–34. [Google Scholar] [CrossRef]
- Li, Y.; Chen, J.; Lun, S.-Y.; Rui, X.-S. Efficient pyruvate production by a multi-vitamin auxotroph of Torulopsis glabrata: Key role and optimization of vitamin levels. Appl. Microbiol. Biotechnol. 2001, 55, 680–685. [Google Scholar] [CrossRef] [PubMed]
- Xu, N.; Liu, L.; Zou, W.; Liu, J.; Hua, Q.; Chen, J. Reconstruction and analysis of the genome-scale metabolic network of Candida glabrata. Mol. Biosyst. 2013, 9, 205–216. [Google Scholar] [CrossRef] [PubMed]
- Xu, N.; Ye, C.; Chen, X.; Liu, J.; Liu, L.; Chen, J. Genome sequencing of the pyruvate-producing strain Candida glabrata CCTCC M202019 and genomic comparison with strain CBS138. Sci. Rep. 2016, 6, 34893. [Google Scholar] [CrossRef] [PubMed]
- Yang, S.; Chen, X.; Xu, N.; Liu, L.; Chen, J. Urea enhances cell growth and pyruvate production in Torulopsis glabrata. Biotech. Prog. 2014, 30, 19–27. [Google Scholar] [CrossRef] [PubMed]
- Liu, L.; Xu, Q.; Li, Y.; Shi, Z.; Zhu, Y.; Du, G.; Chen, J. Enhancement of pyruvate production by osmotic-tolerant mutant of Torulopsis glabrata. Biotechnol. Bioeng. 2007, 97, 825–832. [Google Scholar] [CrossRef] [PubMed]
- Xu, S.; Zhou, J.; Liu, L.; Chen, J. Proline enhances Torulopsis glabrata growth during hyperosmotic stress. Biotechnol. Bioproc. Eng. 2010, 15, 285–292. [Google Scholar] [CrossRef]
- Calabia, B.P.; Tokiwa, Y.; Aiba, S. Fermentative production of l-(+)-lactic acid by an alkaliphilic marine microorganism. Biotechnol. Lett. 2011, 33, 1429–1433. [Google Scholar] [CrossRef] [PubMed]
- Yokaryo, H.; Tokiwa, Y. Isolation of alkaliphilic bacteria for production of high optically pure l-(+)-lactic acid. J. Gen. Appl. Microbiol. 2014, 60, 270–275. [Google Scholar] [CrossRef] [PubMed]
- Yin, J.; Chen, J.-C.; Wu, Q.; Chen, G.-Q. Halophiles, coming stars for industrial biotechnology. Biotechnol. Adv. 2015, 33, 1433–1442. [Google Scholar] [CrossRef] [PubMed]
- Kawata, Y.; Nishimura, T.; Matsushita, I.; Tsubota, J. Efficient production and secretion of pyruvate from Halomonas sp. KM-1 under aerobic conditions. AMB Express 2016, 6, 22. [Google Scholar] [CrossRef] [PubMed]
- Dietzler, D.N.; Leckie, M.P.; Magnani, J.L.; Sughrue, M.J.; Bergstein, P.E. Evidence for the coordinate control of glycogen synthesis, glucose utilization, and glycolysis in Escherichia coli. II. Quantitative correlation of the inhibition of glycogen synthesis and the stimulation of glucose utilization by 2,4-dinitrophenol with the effects on the cellular levels of glucose 6-phosphate, fructose, 1,6-diphosphate, and total adenylates. J. Biol. Chem. 1975, 250, 7195–7203. [Google Scholar] [PubMed]
- Jensen, P.R.; Michelsen, O. Carbon and energy metabolism of atp mutants of Escherichia coli. J. Bacteriol. 1992, 174, 7635–7641. [Google Scholar] [CrossRef] [PubMed]
- Yokota, A.; Terasawa, Y.; Takaoka, N.; Shimizu, H.; Fusao, T. Pyruvic acid production by an F1-ATPase-defective mutant of Escherichia coli W1485lip2. Biosci. Biotechnol. Biochem. 1994, 58, 2164–2167. [Google Scholar] [CrossRef] [PubMed]
- Yokota, A.; Henmi, M.; Takaoka, N.; Hayashi, C.; Takezawa, Y.; Fukumori, Y.; Tomita, F. Enhancement of glucose metabolism in a pyruvic acid-hyperproducing Escherichia coli mutant defective in F1-ATPase activity. J. Ferment. Bioeng. 1997, 83, 132–138. [Google Scholar] [CrossRef]
- Zhou, J.; Huang, L.; Liu, L.; Chen, J. Enhancement of pyruvate productivity by inducible expression of a F0F1-ATPase inhibitor INH1 in Torulopsis glabrata CCTCC M202019. J. Biotechnol. 2009, 144, 120–126. [Google Scholar] [CrossRef] [PubMed]
- Tomar, A.; Eiteman, M.A.; Altman, E. The effect of acetate pathway mutations on the production of pyruvate in Escherichia coli. Appl. Microbiol. Biotech. 2003, 62, 76–82. [Google Scholar] [CrossRef] [PubMed]
- Li, M.; Ho, P.Y.; Yao, S.; Shimizu, K. Effect of lpdA knockout on the metabolism in Escherichia coli based on enzyme activities, intracellular metabolite concentrations and metabolic flux analysis by 13C-labeling experiments. J. Biotechnol. 2006, 122, 254–266. [Google Scholar] [CrossRef] [PubMed]
- Zelić, B.; Gerharz, T.; Bott, M.; Vasić-Rački, Đ.; Wandrey, C.; Takors, R. Fed-batch process for pyruvate production by recombinant Escherichia coli YYC202 Strain. Eng. Life Sci. 2003, 3, 299–305. [Google Scholar] [CrossRef]
- Zelić, B.; Gostović, S.; Vuorilehto, K.; Vasić-Rački, Đ.; Takors, R. Process strategies to enhance pyruvate production with recombinant Escherichia coli: From repetitive fed-batch to in situ product recovery with fully integrated electrodialysis. Biotech. Bioeng. 2004, 85, 638–646. [Google Scholar] [CrossRef] [PubMed]
- Zelić, B.; Vasić-Rački, Đ.; Wandrey, C.; Takors, R. Modeling of the pyruvate production with Escherichia coli in a fed-batch bioreactor. Bioprocess Biosyst. Eng. 2004, 26, 249–258. [Google Scholar] [CrossRef] [PubMed]
- Zelić, B.; Bolf, N.; Vasić-Rački, Đ. Modeling of the pyruvate production with Escherichia coli: Comparison of mechanistic and neural networks-based models. Bioprocess Biosyst. Eng. 2006, 29, 39–47. [Google Scholar] [CrossRef] [PubMed]
- Causey, T.B.; Zhou, S.; Shanmugam, K.T.; Ingram, L.O. Engineering the metabolism of Escherichia coli W3110 for the conversion of sugar to redox-neutral and oxidized products: Homoacetate production. Proc. Natl. Acad. Sci. USA 2003, 100, 825–832. [Google Scholar] [CrossRef] [PubMed]
- Hansen, H.G.; Henning, U. Regulation of pyruvate dehydrogenase activity in Escherichia coli K12. Biochim. Biophys. Acta 1966, 122, 355–358. [Google Scholar] [CrossRef]
- Weitzman, P.D.J. Regulation of citrate synthase activity in Escherichia coli. Biochim. Biophys. Acta 1966, 128, 213–215. [Google Scholar] [CrossRef]
- Nakashima, N.; Ohno, S.; Yoshikawa, K.; Shimizu, H.; Tamura, T. A vector library for silencing central carbon metabolism genes with antisense RNAs in Escherichia coli. Appl. Environ. Microbiol. 2014, 80, 564–573. [Google Scholar] [CrossRef] [PubMed]
- Akita, H.; Nakashima, N.; Hoshino, T. Pyruvate production using engineered Escherichia coli. AMB Express 2016, 6, 94. [Google Scholar] [CrossRef] [PubMed]
- Wang, Q.; He, P.; Lu, D.; Shen, A.; Jiang, N. Metabolic engineering of Torulopsis glabrata for improved pyruvate production. Enzyme Microb. Technol. 2005, 36, 832–839. [Google Scholar] [CrossRef]
- Wang, D.; Wang, L.; Hou, L.; Deng, X.; Gao, Q.; Gao, N. Metabolic engineering of Saccharomyces cerevisiae for accumulating pyruvic acid. Ann. Microbiol. 2015, 65, 2323–2331. [Google Scholar] [CrossRef]
- Van Maris, A.J.A.; Geertman, J.-M.A.; Vermeulen, A.; Groothiuzen, M.K.; Winkler, A.A.; Piper, M.D.W.; van Dijken, J.P.; Pronk, J.T. Directed evolution of pyruvate decarboxylase-negative Saccharomyces cerevisiae, yielding a C2-independent, glucose-tolerant, and pyruvate-hyperproducing yeast. Appl. Environ. Microbiol. 2004, 70, 159–166. [Google Scholar] [CrossRef] [PubMed]
- Wang, Z.; Gao, C.; Wang, Q.; Liang, Q.; Qi, Q. Production of pyruvate in Saccharomyces cerevisiae through adaptive evolution and rational cofactor metabolic engineering. Biochem. Eng. J. 2012, 67, 126–131. [Google Scholar] [CrossRef]
- Zhang, B.; Zhu, Y.; Zhang, J.; Wang, D.; Sun, L.; Hong, J. Engineered Kluyveromyces marxianus for pyruvate production at elevated temperature with simultaneous consumption of xylose and glucose. Bioresour. Technol. 2017, 224, 553–562. [Google Scholar] [CrossRef] [PubMed]
- Kawai, S.; Ohashi, K.; Yoshida, S.; Fujii, M.; Mikami, S.; Sato, N.; Murata, K. Bacterial pyruvate production from alginate, a promising carbon source from marine brown macroalgae. J. Biosci. Bioeng. 2014, 3, 269–274. [Google Scholar] [CrossRef] [PubMed]
- Blombach, B.; Schreiner, M.E.; Holátko, J.; Bartek, T.; Oldiges, M.; Eikmanns, B.J. l-Valine production with pyruvate dehydrogenase complex-deficient Corynebacterium glutamicum. Appl. Environ. Microbiol. 2007, 73, 2079–2084. [Google Scholar] [CrossRef] [PubMed]
- Blombach, B.; Schreiner, M.E.; Bartek, T.; Oldiges, M.; Eikmanns, B.J. Corynebacterium glutamicum tailored for high-yield L-valine production. Appl. Microbiol. Biotechnol. 2008, 79, 471–479. [Google Scholar] [CrossRef] [PubMed]
- Wieschalka, S.; Blombach, B.; Eikmanns, B.J. Engineering Corynebacterium glutamicum for the production of pyruvate. Appl. Microbiol. Biotechnol. 2012, 94, 449–459. [Google Scholar] [CrossRef] [PubMed]
- Vemuri, G.N.; Altman, E.; Sangurdekar, D.P.; Khodursky, A.B.; Eiteman, M.A. Overflow metabolism in Escherichia coli during steady-state growth: Transcriptional regulation and effect of the redox ratio. Appl. Environ. Microbiol. 2006, 72, 3653–3661. [Google Scholar] [CrossRef] [PubMed]
- Zhu, Y.; Eiteman, M.A.; Altman, R.; Altman, E. High glycolytic flux improves pyruvate production by a metabolically engineered Escherichia coli strain. Appl. Environ. Microbiol. 2008, 74, 6649–6655. [Google Scholar] [CrossRef] [PubMed]
- Qin, Y.; Johnson, C.H.; Liu, L.; Chen, J. Introduction of heterogeneous NADH reoxidation pathways into Torulopsis glabrata significantly increases pyruvate production efficiency. Kor. J. Chem. Eng. 2011, 28, 1078–1084. [Google Scholar] [CrossRef]
- Ojima, Y.; Suryadarma, P.; Tsuchida, K.; Taya, M. Accumulation of pyruvate by changing the redox status in Escherichia coli. Biotech. Lett. 2012, 34, 889–893. [Google Scholar] [CrossRef] [PubMed]
- Hua, Q.; Araki, M.; Koide, Y.; Shimizu, K. Effects of glucose, vitamins, and DO concentrations on pyruvate fermentation using Torulopsis glabrata IFO 0005 with metabolic flux analysis. Biotechnol. Prog. 2001, 17, 62–68. [Google Scholar] [CrossRef] [PubMed]
- Li, Y.; Hugenholtz, J.; Chen, J.; Lun, S.-Y. Enhancement of pyruvate production by Torulopsis glabrata using a two-stage oxygen supply control strategy. Appl. Microbiol. Biotechnol. 2002, 60, 101–106. [Google Scholar] [PubMed]
- Causey, T.; Shanmugam, K.; Yomano, L.; Ingram, L. Engineering Escherichia coli for efficient conversion of glucose to pyruvate. Proc. Natl. Acad. Sci. USA 2004, 101, 2235–2240. [Google Scholar] [CrossRef] [PubMed]
- Zhu, Y.; Eiteman, M.A.; Lee, S.A. Conversion of glycerol to pyruvate by Escherichia coli using acetate- and acetate/glucose-limited fed-batch processes. J. Ind. Microbiol. Biotechnol. 2010, 37, 307–312. [Google Scholar] [CrossRef] [PubMed]
- Ma, C.Q.; Xu, P.; Dou, Y.M.; Qu, Y.B. Highly efficient conversion of lactate to pyruvate using whole cells of Acinetobacter sp. Biotechnol. Prog. 2003, 19, 1672–1676. [Google Scholar] [CrossRef] [PubMed]
- Eisenberg, A.; Seip, J.E.; Gavagan, J.E.; Payne, M.S.; Anton, D.L.; di Cosimo, R. Pyruvic acid production using methylotrophic yeast transformants as catalysts. J. Mol. Catal. B: Enzym. 1997, 2, 223–232. [Google Scholar] [CrossRef]
- Gough, S.; Dostal, L.; Howe, A.; Deshpande, M.; Scher, M.; Rosazza, J.N.P. Production of pyruvate from lactate using recombinant Pichia pastoris cells as catalyst. Proc. Biochem. 2005, 40, 2597–2601. [Google Scholar] [CrossRef]
- Hao, J.; Ma, C.; Gao, C.; Qiu, J.; Wang, M.; Zhang, Y.; Cui, X.; Xu, P. Pseudomonas stutzeri as a novel biocatalyst for pyruvate production from DL-lactate. Biotechnol. Lett. 2007, 29, 105–110. [Google Scholar] [CrossRef] [PubMed]
- Gao, C.; Qiu, J.; Ma, C.; Xu, P. Efficient production of pyruvate from DL-lactate by the lactate-utilizing strain Pseudomonas stutzeri SDM. PLoS ONE 2012, 7, e40755. [Google Scholar] [CrossRef] [PubMed]
- Hossain, G.S.; Shin, H.-D.; Li, J.; Du, G.; Chen, J.; Liu, L. Transporter engineering and enzyme evolution for pyruvate production from d/l-alanine with a whole-cell biocatalyst expressing l-amino acid deaminase from Proteus mirabilis. RSC Adv. 2016, 6, 82676–82684. [Google Scholar] [CrossRef]
- Johnson, C.W.; Beckham, G.T. Aromatic catabolic pathway selection for optimal production of pyruvate and lactate from lignin. Metab. Eng. 2015, 28, 240–247. [Google Scholar] [CrossRef] [PubMed]
- Hong, Y.K.; Hong, W.H.; Han, D.H. Application of reactive extraction to recovery of carboxylic acids. Biotechnol. Bioproc. Eng. 2001, 6, 386–394. [Google Scholar] [CrossRef]
- Wasewar, K.L.; Neesink, A.B.M.; Versteeg, G.F.; Pangarkar, V.G. Reactive extraction of lactic acid using Alamine 336 in MIBK: Equilibria and kinetics. J. Biotechnol. 2002, 97, 59–68. [Google Scholar] [CrossRef]
- Senol, A. Influence of diluent on amine extraction of pyruvic acid using Alamine system. Chem. Eng. Process. 2006, 45, 755–763. [Google Scholar] [CrossRef]
- Senol, A. Liquid-liquid equilibria for mixtures of (water + pyruvic acid + alcohol/Alamine). Modeling and optimization of extraction. J. Chem. Eng. Data 2013, 58, 528–536. [Google Scholar] [CrossRef]
- Pal, D.; Tripathi, A.; Shukla, A.; Gupta, K.R.; Keshav, A. Reactive extraction of pyruvic acid using tri-octylamine diluted in decanol/kerosene: Equilibrium and effect of temperature. J. Chem. Eng. Data 2015, 60, 860–869. [Google Scholar] [CrossRef]
- Pal, D.; Keshav, A. Kinetics of reactive extraction of pyruvic acid using tributylamine dissolved in n-butyl acetate. Int. J. Chem. React. Eng. 2015, 12, 63–69. [Google Scholar] [CrossRef]
- Marti, M.E.; Gurkan, T. Selective recovery of pyruvic acid from two and three acid aqueous solutions by reactive extraction. Separ. Purif. Technol. 2015, 156, 148–157. [Google Scholar] [CrossRef]
- Pal, D.; Thakre, N.; Kumar, A.; Keshav, A. Reactive extraction of pyruvic acid using mixed extractants. Separ. Sci. Technol. 2016, 51, 1141–1150. [Google Scholar] [CrossRef]
- Pal, D.; Keshav, A. Extraction equilibria of pyruvic acid using tri-n-butyl phosphate: Influence of diluents. J. Chem. Eng. Data. 2014, 59, 2709–2716. [Google Scholar] [CrossRef]
- Ma, C.Q.; Li, J.C.; Qiu, J.H.; Wang, M.; Xu, P. Recovery of pyruvic acid from biotransformation solutions. Appl. Microbiol. Biotechnol. 2006, 70, 308–314. [Google Scholar] [CrossRef] [PubMed]
- Marti, M.E.; Gurkan, T.; Doraiswamy, L.K. Equilibrium and kinetic studies on reactive extraction of pyruvic acid with trioctylamine in 1-octanol. Ind. Eng. Chem. Res. 2011, 50, 13518–13525. [Google Scholar] [CrossRef]
- Zhigang, T.; Rongqi, Z.; Zhanting, D. Separation of isopropanol from aqueous solution by salting-out extraction. J. Chem. Technol. Biotechnol. 2001, 76, 757–763. [Google Scholar] [CrossRef]
- Birajdar, S.D.; Padmanabhan, S.; Rajagopalan, S. Repulsive effect of salt on solvent extraction of 2,3-butanediol from aqueous fermentation solution. J. Chem. Technol. Biotechnol. 2015, 90, 1455–1462. [Google Scholar] [CrossRef]
- Ingle, U.; Rajagopalan, S.; Padmanabhan, S. Isolation and scale-up of sodium pyruvate from fermentation broth using a repulsive extraction and crystallization method. J. Chem. Technol. Biotechnol. 2015, 90, 2240–2248. [Google Scholar] [CrossRef]
- Zelić, B.; Vasić-Rački, Đ. Process development and modeling of pyruvate recovery from a model solution and fermentation broth. Desalination 2005, 174, 267–276. [Google Scholar] [CrossRef]
- Huang, S.; Qin, W.; Dai, Y. Recovery of pyruvic acid with weakly basic polymeric sorbents. J. Chem. Technol. Biotechnol. 2008, 83, 683–687. [Google Scholar] [CrossRef]
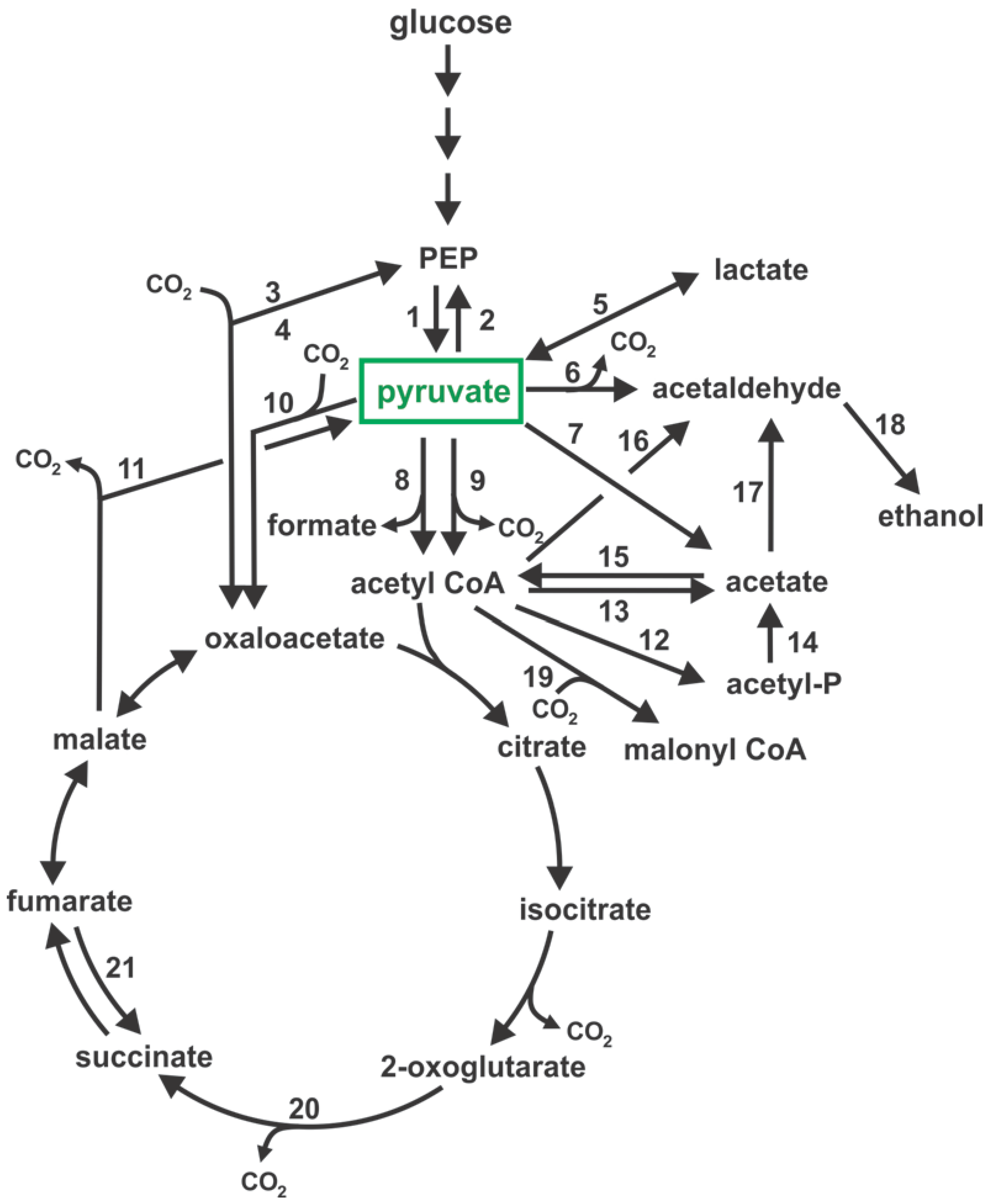
| Figure 1 (ref) | Enzyme Accepted Name | EC (1) Number | Reaction | E. coli (2) | C. glutamicum (3) | S. cerevisiae | C. glabrata |
|---|---|---|---|---|---|---|---|
| 1 | pyruvate kinase | 2.7.1.40 | PEP + ADP → pyruvate + ATP | pykA, pykF | pyk | PYK1 | CAGL0E05610g |
| PYK2 | CAGL0M12034g | ||||||
| 2 | PEP synthase | 2.7.9.2 | pyruvate + ATP + H2O → PEP + AMP + Pi | ppsA | cg0642, cg0644 | – | – |
| 3 | PEP carboxylase | 4.1.1.31 | PEP + HCO3− → oxaloacetate + Pi | ppc | ppc | – | – |
| 4 | PEP carboxykinase | 4.1.1.49 | PEP + CO2 + ADP → oxaloacetate + ATP | pck | – | PCK1 | CAGL0H06633g |
| 4.1.1.32 | PEP + CO2 + GDP → oxaloacetate + GTP | – | pck | – | – | ||
| 5 | d-lactate dehydrogenase | 1.1.1.28 | pyruvate + NADH + H+ → d-lactate + NAD+ | ldhA | dld | –(4) | –(4) |
| l-lactate dehydrogenase | 1.1.1.27 | pyruvate + NADH + H+ → l-lactate + NAD+ | – | ldh | –(4) | –(4) | |
| 6 | pyruvate decarboxylase | 4.1.1.1 | pyruvate → acetaldehyde + CO2 | – | – | THI3, PDC1, | CAGL0G02937g |
| PDC5, PDC6 | CAGL0L06842g | ||||||
| CAGL0M07920g | |||||||
| 7 | pyruvate oxidase | 1.2.5.1 | pyruvate + ubiquinone → acetate + CO2 + ubiquinol | poxB | poxB | – | – |
| 8 | pyruvate formate lyase | 2.3.1.54 | pyruvate + CoA → acetyl CoA + formate | pflB | – | – | – |
| 9 | pyruvate dehydrogenase complex: | pyruvate + CoA + NAD+ → acetyl CoA + NADH + H+ + CO2 | |||||
| pyruvate dehydrogenase (E1) | 1.2.4.1 | aceE | aceE | PDB1 | CAGL0K06831g | ||
| PDA1 | CAGL0L12078g | ||||||
| dihydrolipoamide acetyltransferase (E2) | 2.3.1.12 | aceF | – | PDA2 | CAGL0J10186g | ||
| dihydrolipoamide dehydrogenase (E3) | 1.8.1.4 | lpd | lpd | LPD1, IRC15 | CAGL0F01947g | ||
| 10 | pyruvate carboxylase | 6.4.1.1 | pyruvate + HCO3− + ATP → oxaloacetate + ADP | – | pyc | PYC1, PYC2 | CAGL0F06941g |
| 11 | malate dehydrogenase (NAD+, decarboxylating) | 1.1.1.38 | L-malate + NAD+ → pyruvate + CO2 + NADH | maeA | – | MAE1 | CAGL0L02035g |
| malate dehydrogenase (NADP+, decarboxylating) | 1.1.1.40 | L-malate + NADP+ → pyruvate + CO2 + NADPH | maeB | – | – | – | |
| 12 | phosphate acetyltransferase | 2.3.1.8 | acetyl CoA + Pi → acetyl-P + CoA | pta | pta | – | – |
| 13 | acetyl CoA hydrolase | 3.1.2.1 | acetyl CoA + H2O → acetate + CoA | – | – | ACH1 | CAGL0J04268g |
| 14 | acetate kinase | 2.7.2.1 | acetyl-P + ADP → acetate + ATP | ackA | ackA | – | – |
| 15 | acetyl CoA synthetase | 6.2.1.1 | acetate + ATP + CoA → acetyl CoA + AMP + PPi | acs | – | ACS1 | CAGL0B02717g |
| ACS2 | CAGL0L00649g | ||||||
| 16 | acetaldehyde dehydrogenase (acetylating) | 1.2.1.10 | acetyl CoA + NADH + H+ → acetaldehyde + NAD+ + CoA | adhE | – | – | – |
| 17 | acetaldehyde dehydrogenase (NAD) | 1.2.1.3 | acetate + NADH + H+ → acetaldehyde + NAD+ + H2O | – | xylC | ALD4-6 | CAGL0D06688g |
| HFD1 | CAGL0H05137g | ||||||
| CAGL0J03212g | |||||||
| CAGL0K03509g | |||||||
| acetaldehyde dehydrogenase (NADP) | 1.2.1.5 | acetate + NADPH + H+ → acetaldehyde + NADP+ + H2O | – | – | ALD2 ALD3 | CAGL0F07777g | |
| 18 | alcohol dehydrogenase (NAD) | 1.1.1.1 | acetaldehyde + NADH + H+ → ethanol + NAD+ | adhE | adhA | ADH1-5 | CAGL0I07843g |
| alcohol dehydrogenase (NADP) | 1.1.1.2 | acetaldehyde + NADPH + H+ → ethanol + NADP+ | – | – | ADH6-7 | CAGL0H06853g | |
| 19 | acetyl CoA carboxylase | 6.4.1.2 | acetyl CoA + HCO3− + ATP → malonyl CoA + ADP + Pi | accABCD | accABCD | HFA1, ACC1 | CAGL0L10780g |
| 20 | 2-oxoglutarate dehydrogenase complex: | 2-oxoglutarate + CoA + NAD+ → succinyl CoA + NADH + H+ + CO2 | |||||
| 2-oxoglutarate dehydrogenase (E1) | 1.2.4.2 | sucA | odhA | KGD1 | CAGL0G08712g | ||
| dihydrolipoamide succinyltransferase (E2) | 2.3.1.61 | sucB | sucB | KGD2 | CAGL0E01287g | ||
| dihydrolipoamide dehydrogenase (E3) | 1.8.1.4 | lpd | lpd | LPD1, IRC15 | CAGL0F01947g | ||
| 21 | fumarate reductase | 1.3.5.4 | fumarate + quinone → succinate + quinol | frdABCD | – | – | – |
© 2017 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license ( http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Maleki, N.; Eiteman, M.A. Recent Progress in the Microbial Production of Pyruvic Acid. Fermentation 2017, 3, 8. https://doi.org/10.3390/fermentation3010008
Maleki N, Eiteman MA. Recent Progress in the Microbial Production of Pyruvic Acid. Fermentation. 2017; 3(1):8. https://doi.org/10.3390/fermentation3010008
Chicago/Turabian StyleMaleki, Neda, and Mark A. Eiteman. 2017. "Recent Progress in the Microbial Production of Pyruvic Acid" Fermentation 3, no. 1: 8. https://doi.org/10.3390/fermentation3010008
APA StyleMaleki, N., & Eiteman, M. A. (2017). Recent Progress in the Microbial Production of Pyruvic Acid. Fermentation, 3(1), 8. https://doi.org/10.3390/fermentation3010008






