Deciphering Northeast–Northwest Differences in Steamed Bread Microbiota and Flavor via Metagenomics and Untargeted Metabolomics
Abstract
1. Introduction
2. Materials and Methods
2.1. Sourdough Materials and Experimental Grouping
2.2. Fermented Dough Preparation
2.3. Physicochemical Analysis
2.4. Metagenomics Analysis
2.4.1. Experimental Procedures
2.4.2. Bioinformatic Processing
2.5. Sample Preparation for Untargeted Metabolomics Analysis
2.5.1. Experimental Methods
2.5.2. Data Processing
3. Results and Discussion
3.1. Physical and Chemical Testing
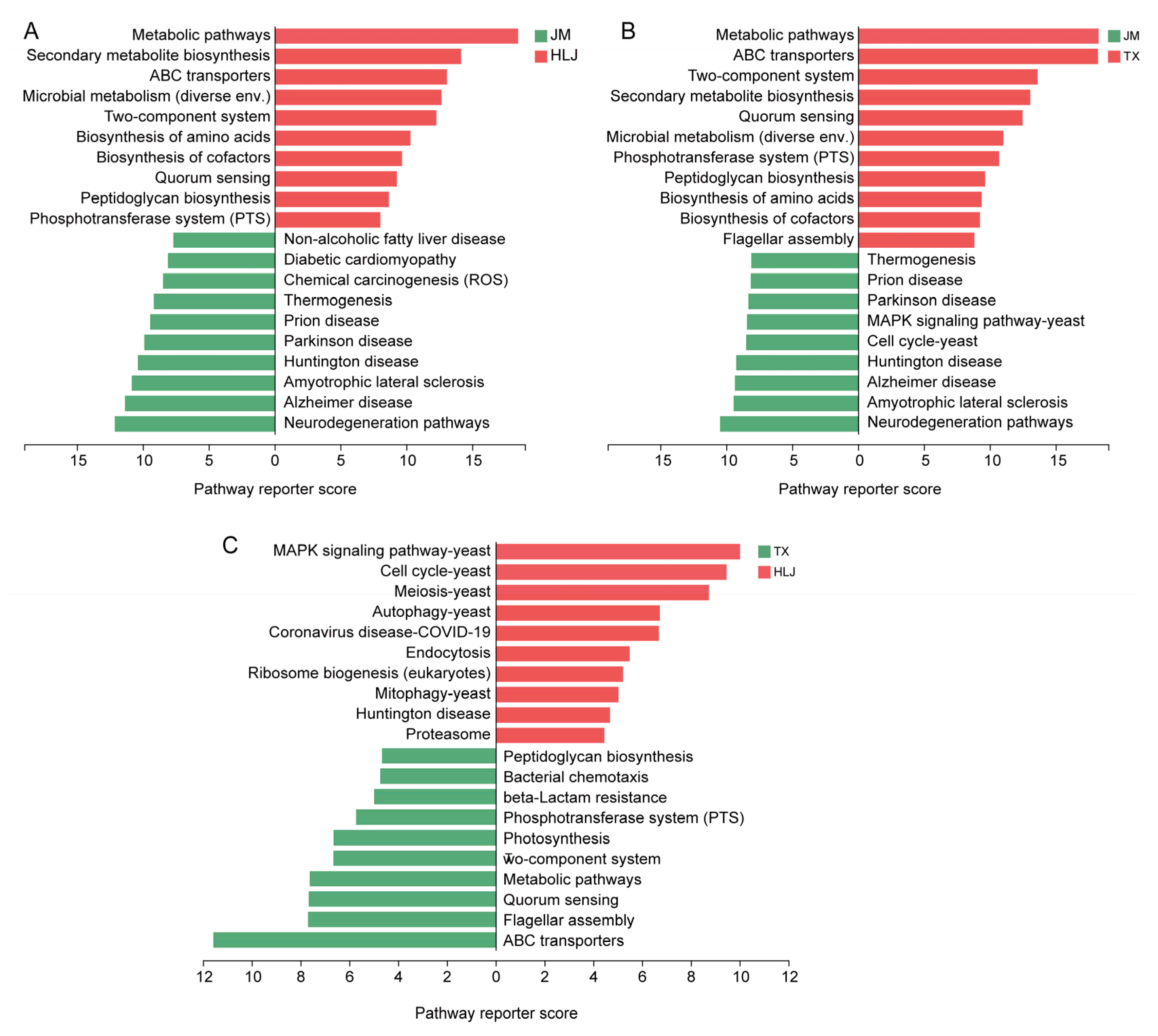
3.2. Metagenomic Analysis
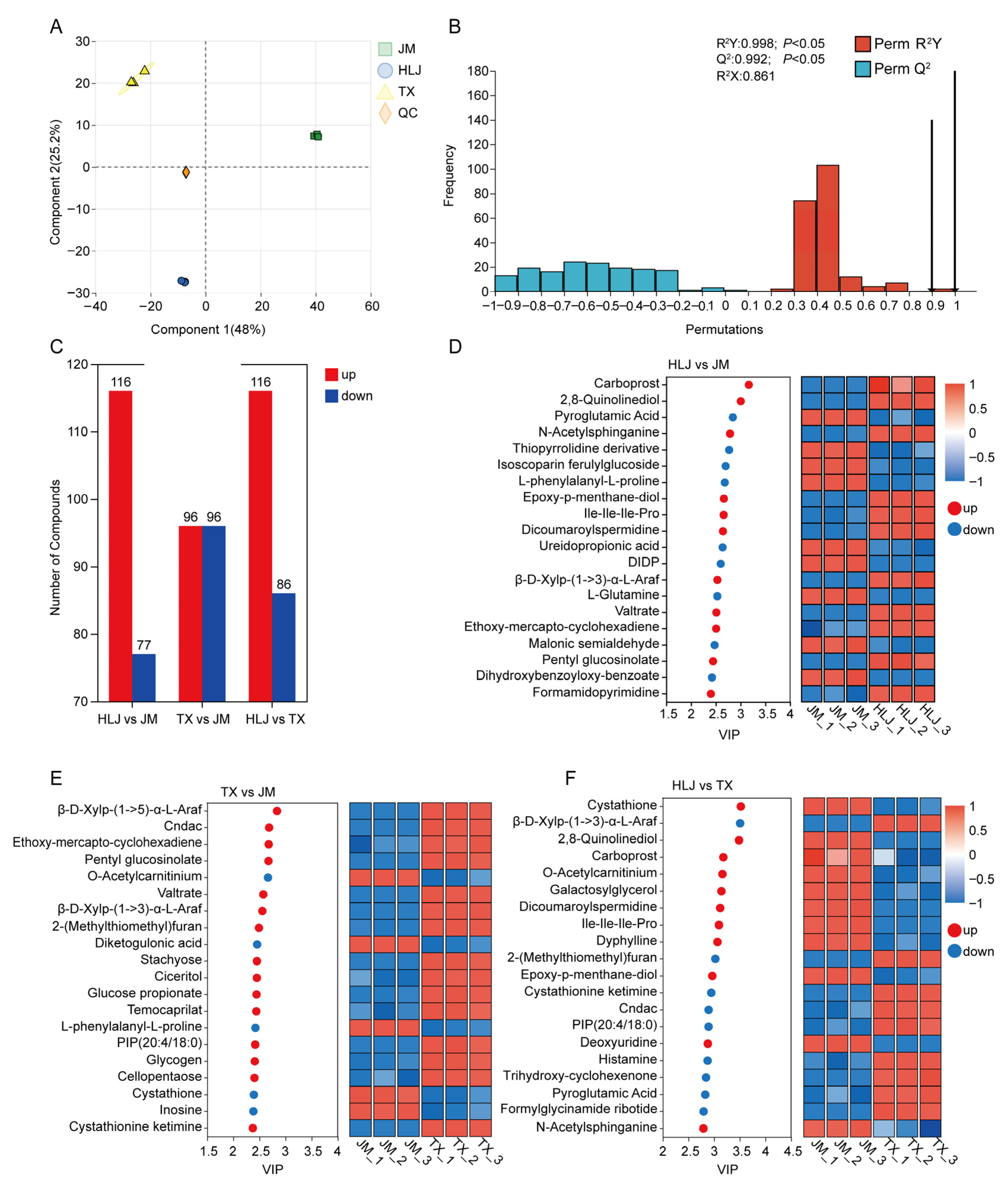
3.3. Untargeted Metabolomics Analysis
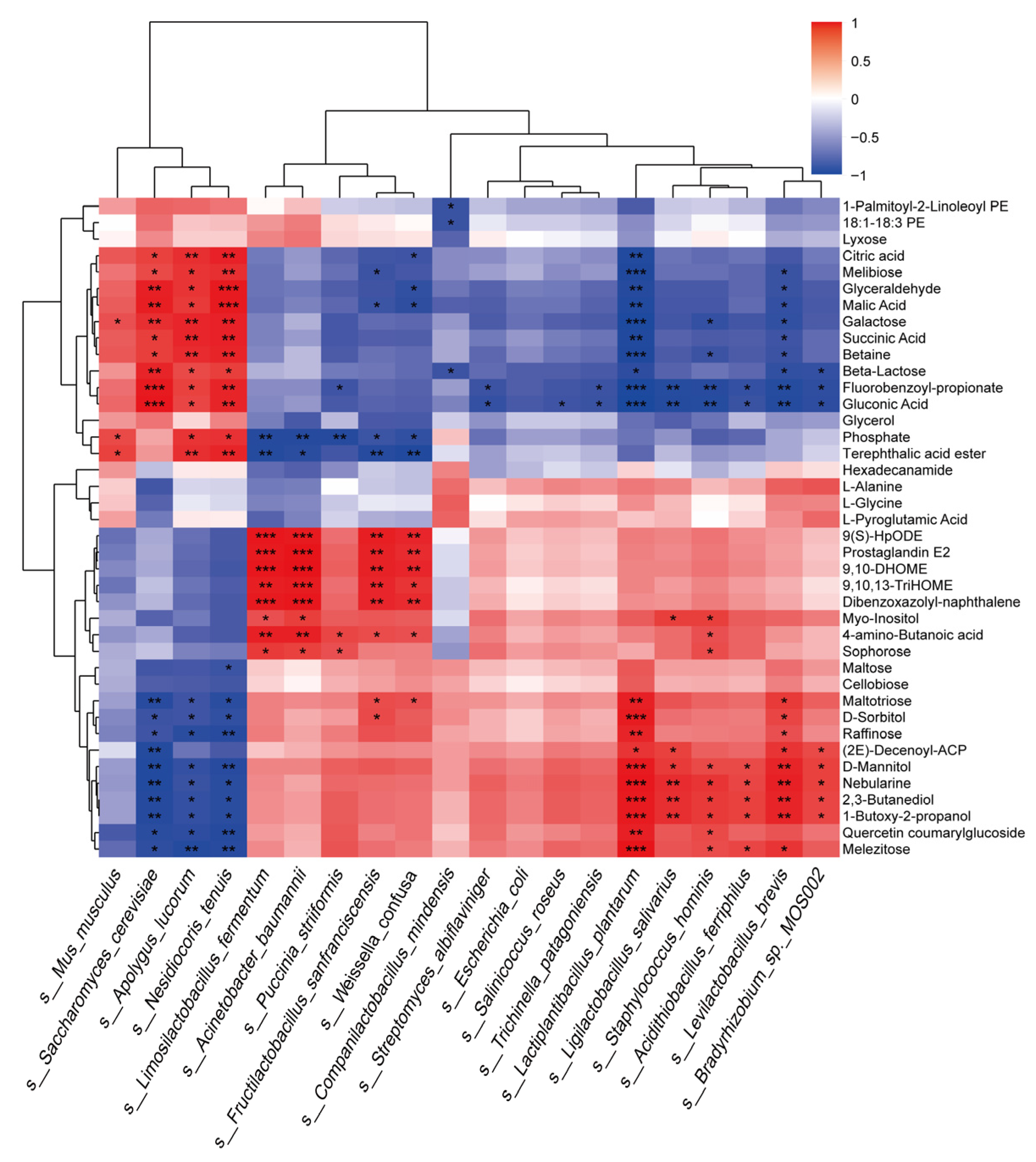
3.4. Analysis of Microbial Abundance and Metabolite Correlation
4. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Yang, Y.; Wang, X. Effects of coarse cereals on dough and Chinese steamed bread—A review. Front. Nutr. 2023, 10, 1186860. [Google Scholar] [CrossRef] [PubMed]
- Zhu, F. Influence of ingredients and chemical components on the quality of Chinese steamed bread. Food Chem. 2014, 163, 154–162. [Google Scholar] [CrossRef] [PubMed]
- Wei, G.; Zhang, Z.; Zhao, F.; Sang, Y.; Regenstein, J.M.; Zhou, P. Characteristic aroma compounds during the fermentation of Chinese steamed bread fermented with different starters. Food Chem. 2024, 457, 140151. [Google Scholar] [CrossRef]
- Fu, L.; Nowak, A.; Zhao, H.; Zhang, B. Relationship Between Microbial Composition of Sourdough and Texture, Volatile Compounds of Chinese Steamed Bread. Foods 2022, 11, 1908. [Google Scholar] [CrossRef] [PubMed]
- Ashaolu, T.J.; Reale, A. A Holistic Review on Euro-Asian Lactic Acid Bacteria Fermented Cereals and Vegetables. Microorganisms 2020, 8, 1176. [Google Scholar] [CrossRef]
- Yan, B.; Sadiq, F.A.; Cai, Y.; Fan, D.; Chen, W.; Zhang, H.; Zhao, J. Microbial diversity in traditional type I sourdough and jiaozi and its influence on volatiles in Chinese steamed bread. LWT 2019, 101, 764–773. [Google Scholar] [CrossRef]
- Gereková, P.; Petruláková, Z.; Šturdík, E. Importance of lactobacilli for bread-making industry. Acta Chim. Slovaca 2011, 4, 118–135. [Google Scholar]
- Hillmann, H.; Hofmann, T. Quantitation of Key Tastants and Re-engineering the Taste of Parmesan Cheese. J. Agric. Food Chem. 2016, 64, 1794–1805. [Google Scholar] [CrossRef]
- Pico, J.; Bernal, J.; Gómez, M. Wheat bread aroma compounds in crumb and crust: A review. Food Res. Int. 2015, 75, 200–215. [Google Scholar] [CrossRef]
- Ripari, V.; Cecchi, T.; Berardi, E. Microbiological characterisation and volatiles profile of model, ex-novo, and traditional Italian white wheat sourdoughs. Food Chem. 2016, 205, 297–307. [Google Scholar] [CrossRef]
- Pétel, C.; Onno, B.; Prost, C. Sourdough volatile compounds and their contribution to bread: A review. Trends Food Sci. Technol. 2017, 59, 105–123. [Google Scholar] [CrossRef]
- Tang, N.; Xing, X.; Li, H.; Suo, B.; Wang, Y.; Ai, Z.; Yang, Y. Co-culture fermentation by Saccharomycopsis fibuligera and lactic acid bacteria improves bioactivity and aroma profile of wheat bran and the bran-containing Chinese steamed bread. Food Res. Int. 2024, 182, 114179. [Google Scholar] [CrossRef] [PubMed]
- Wen, L.; Yang, L.; Chen, C.; Li, J.; Fu, J.; Liu, G.; Kan, Q.; Ho, C.-T.; Huang, Q.; Lan, Y.; et al. Applications of multi-omics techniques to unravel the fermentation process and the flavor formation mechanism in fermented foods. Crit. Rev. Food Sci. Nutr. 2024, 64, 8367–8383. [Google Scholar] [CrossRef] [PubMed]
- Sun, X.; Ye, Z.; Xiao, M.; Zhang, C.; Guo, M.; Zhao, J.; Chen, W.; Tian, F.; Yu, L.; Zhai, Q. The formation mechanism of volatile and nonvolatile flavor substances in sourdough based on genomics and metabolomics. Food Chem. 2025, 476, 143283. [Google Scholar] [CrossRef] [PubMed]
- Jiang, Z.J.; Sun, X.; Guo, X.-N.; Zhu, K.-X. Components and physicochemical properties of mill streams: Effects on freeze-thaw stability and quality of frozen steamed bread dough. Food Chem. 2025, 478, 143738. [Google Scholar] [CrossRef]
- Wang, H.; Wei, X.; Li, D.; Yan, J.; Wu, Y.; Zhou, Z. Impact of surfactin on the physicochemical properties of dough and quality of corresponding steamed bread. J. Sci. Food Agric. 2025, 105, 3122–3132. [Google Scholar] [CrossRef]
- Liu, J.; Zhao, W.; Zhang, A.; Li, P.; Liu, J. Effects of cereal flour types and sourdough on dough physicochemical properties and steamed bread quality. J. Food Sci. 2024, 89, 5434–5448. [Google Scholar] [CrossRef]
- Chen, S.; Zhou, Y.; Chen, Y.; Gu, J. fastp: An ultra-fast all-in-one FASTQ preprocessor. Bioinformatics 2018, 34, i884–i890. [Google Scholar] [CrossRef]
- Li, H.; Durbin, R. Fast and accurate short read alignment with Burrows-Wheeler transform. Bioinformatics 2009, 25, 1754–1760. [Google Scholar] [CrossRef]
- Li, D.; Liu, C.-M.; Luo, R.; Sadakane, K.; Lam, T.-W. MEGAHIT: An ultra-fast single-node solution for large and complex metagenomics assembly via succinct de Bruijn graph. Bioinformatics 2015, 31, 1674–1676. [Google Scholar] [CrossRef]
- Noguchi, H.; Park, J.; Takagi, T. MetaGene: Prokaryotic gene finding from environmental genome shotgun sequences. Nucleic Acids Res. 2006, 34, 5623–5630. [Google Scholar] [CrossRef]
- Fu, L.; Niu, B.; Zhu, Z.; Wu, S.; Li, W. CD-HIT: Accelerated for clustering the next-generation sequencing data. Bioinformatics 2012, 28, 3150–3152. [Google Scholar] [CrossRef]
- Li, R.; Li, Y.; Kristiansen, K.; Wang, J. SOAP: Short oligonucleotide alignment program. Bioinformatics 2008, 24, 713–714. [Google Scholar] [CrossRef]
- Buchfink, B.; Xie, C.; Huson, D.H. Fast and sensitive protein alignment using DIAMOND. Nat. Methods 2015, 12, 59–60. [Google Scholar] [CrossRef]
- Cooper, B.; Yang, R. An assessment of AcquireX and Compound Discoverer software 3.3 for non-targeted metabolomics. Sci. Rep. 2024, 14, 4841. [Google Scholar] [CrossRef]
- Wallace, W.E.; Moorthy, A.S. NIST Mass Spectrometry Data Center standard reference libraries and software tools: Application to seized drug analysis. J. Forensic Sci. 2023, 68, 1484–1493. [Google Scholar] [CrossRef] [PubMed]
- Misra, B.B.; Bassey, E.; Bishop, A.C.; Kusel, D.T.; Cox, L.A.; Olivier, M. High-resolution gas chromatography/mass spectrometry metabolomics of non-human primate serum. Rapid Commun. Mass. Spectrom. 2018, 32, 1497–1506. [Google Scholar] [CrossRef]
- Ren, Y.; Yu, G.; Shi, C.; Liu, L.; Guo, Q.; Han, C.; Zhang, D.; Zhang, L.; Liu, B.; Gao, H.; et al. Majorbio Cloud: A one-stop, comprehensive bioinformatic platform for multiomics analyses. Imeta 2022, 1, e12. [Google Scholar] [CrossRef] [PubMed]
- Parsell, G.; Bligh, J. The development of a questionnaire to assess the readiness of health care students for interprofessional learning (RIPLS). Med. Educ. 1999, 33, 95–100. [Google Scholar] [CrossRef] [PubMed]
- Hargreaves, J.; Popineau, Y.; Le Meste, M.; Hemminga, M.A. Molecular flexibility in wheat gluten proteins submitted to heating. FEBS Lett. 1995, 372, 103–107. [Google Scholar] [CrossRef]
- Patching, J.W.; Rose, A.H. Cold osmotic shock in Saccharomyces cerevisiae. J. Bacteriol. 1971, 108, 451–458. [Google Scholar] [CrossRef]
- Xu, D.; Hu, Y.; Wu, F.; Jin, Y.; Xu, X.; Gänzle, M.G. Comparison of the Functionality of Exopolysaccharides Produced by Sourdough Lactic Acid Bacteria in Bread and Steamed Bread. J. Agric. Food Chem. 2020, 68, 8907–8914. [Google Scholar] [CrossRef]
- Xie, J.; Zhao, Z.; Gänzle, M.G. Contribution of γ-Glutamyl-Cysteine Ligases of Limosilactobacillus reuteri to the Formation of Kokumi-Active γ-Glutamyl Dipeptides in Sourdough Bread. J. Agric. Food Chem. 2024, 72, 5935–5943. [Google Scholar] [CrossRef]
- Martínez-Anaya, M.A.; Benedito de Barber, C.; Collar Esteve, C. Effect of processing conditions on acidification properties of wheat sour doughs. Int. J. Food Microbiol. 1994, 22, 249–255. [Google Scholar] [CrossRef]
- Korakli, M.; Rossmann, A.; Gänzle, M.G.; Vogel, R.F. Sucrose metabolism and exopolysaccharide production in wheat and rye sourdoughs by Lactobacillus sanfranciscensis. J. Agric. Food Chem. 2001, 49, 5194–5200. [Google Scholar] [CrossRef]
- Menezes, L.A.A.; De Marco, I.; dos Santos, N.N.O.; Nunes, C.C.; Cartabiano, C.E.L.; Molognoni, L.; Pereira, G.V.d.M.; Daguer, H.; Lindner, J.D.D. Reducing FODMAPs and improving bread quality using type II sourdough with selected starter cultures. Int. J. Food Sci. Nutr. 2021, 72, 912–922. [Google Scholar] [CrossRef]
- Promdonkoy, P.; Watcharawipas, A.; Bubphasawan, S.; Sansatchanon, K.; Suwanakitti, N.; Kocharin, K.; Runguphan, W. Metabolic Engineering of Saccharomyces cerevisiae for Production of Canthaxanthin, Zeaxanthin, and Astaxanthin. J. Fungi 2024, 10, 433. [Google Scholar] [CrossRef]
- Martínez-Anaya, M.A.; Llin, M.L.; Macías, M.P.; Collar, C. Regulation of acetic acid production by homo- and heterofermentative lactobacilli in whole-wheat sour-doughs. Z. Lebensm. Unters. Forsch 1994, 199, 186–190. [Google Scholar] [CrossRef] [PubMed]
- Piper, P.; Mahé, Y.; Thompson, S.; Pandjaitan, R.; Holyoak, C.; Egner, R.; Mühlbauer, M.; Coote, P.; Kuchler, K. The pdr12 ABC transporter is required for the development of weak organic acid resistance in yeast. Embo J. 1998, 17, 4257–4265. [Google Scholar] [CrossRef] [PubMed]
- Fuqua, C.; Winans, S.C.; Greenberg, E.P. Census and consensus in bacterial ecosystems: The LuxR-LuxI family of quorum-sensing transcriptional regulators. Annu. Rev. Microbiol. 1996, 50, 727–751. [Google Scholar] [CrossRef] [PubMed]
- Halte, M.; Popp, P.F.; Hathcock, D.; Severn, J.; Fischer, S.; Goosmann, C.; Ducret, A.; Charpentier, E.; Tu, Y.; Lauga, E.; et al. Bacterial motility depends on a critical flagellum length and energy-optimized assembly. Proc. Natl. Acad. Sci. USA 2025, 122, e2413488122. [Google Scholar] [CrossRef]
- Luo, S.; Zhu, M.; Fu, C. The Obg-like ATPase Ola1 prevents excessive mitochondrial reactive oxygen species by inhibiting MAPK/Pmk1 signaling in fission yeast. Microbiol. Res. 2025, 299, 128259. [Google Scholar] [CrossRef]
- Yin, M.Q.; Xu, K.; Luan, T.; Kang, X.-L.; Yang, X.-Y.; Li, H.-X.; Hou, Y.-H.; Zhao, J.-Z.; Bao, X.-M. Metabolic engineering for compartmentalized biosynthesis of the valuable compounds in Saccharomyces cerevisiae. Microbiol. Res. 2024, 286, 127815. [Google Scholar] [CrossRef]
- Wu, J.; Wang, Z.; Hu, J.; Liu, J.; Han, X.; Chen, H.; Zhu, S.; Deng, J. Bacillus subtilis-Derived Postbiotics as a Multifunctional Bio-Catalyst for Enhancing Lactic Acid Bacteria Viability and Yogurt Quality. Foods 2025, 14, 1806. [Google Scholar] [CrossRef]
- Zheng, C.; Yang, Y.; Wei, F.; Lv, X.; Xia, Z.; Qi, M.; Zhou, Q. Widely targeted metabolomics reveal the glucosinolate profile and odor-active compounds in flowering Chinese cabbage powder. Food Res. Int. 2023, 172, 113121. [Google Scholar] [CrossRef]
- Elattar, M.M.; Hammoda, H.M.; Ghareeb, D.A.; Abdulmalek, S.A.; Abdelrahim, F.A.; Seif, I.A.K.; Dawood, H.M.; Darwish, R.S. Insights into bioactive constituents of onion (Allium cepa L.) waste: A comparative metabolomics study enhanced by chemometric tools. BMC Complement. Med. Ther. 2024, 24, 271. [Google Scholar] [CrossRef]
- Huang, Y.; He, X.; Han, Y.; Chen, L.; Yu, X.; Li, J.; Yun, X.; Sha, R.; Borjigin, G. Age-Related Meat Flavor Precursors of Naturally Grazed Sunit Sheep: Metabolomics and Transcriptomics Approaches. Foods 2025, 14, 1616. [Google Scholar] [CrossRef] [PubMed]
- Shulami, S.; Zehavi, A.; Belakhov, V.; Salama, R.; Lansky, S.; Baasov, T.; Shoham, G.; Shoham, Y. Cross-utilization of β-galactosides and cellobiose in Geobacillus stearothermophilus. J. Biol. Chem. 2020, 295, 10766–10780. [Google Scholar] [CrossRef] [PubMed]
- Xiao, M.; Bi, J.; Yi, J.; Zhao, Y.; Peng, J.; Zhou, L.; Chen, Q. Osmotic pretreatment for instant controlled pressure drop dried apple chips: Impact of the type of saccharides and treatment conditions. Dry. Technol. 2019, 37, 896–905. [Google Scholar] [CrossRef]
- Zhang, L.; Chen, J.; Guo, X.; Cao, Y.; Qu, G.; Li, Q.; Gao, Y.; Yu, X. Microwave pretreatment effects on the aroma precursors, sensory characteristics and flavor profiles of fragrant rapeseed oil. Food Chem. X 2024, 22, 101381. [Google Scholar] [CrossRef]
- Weerawatanakorn, M.; Wu, J.-C.; Pan, M.-H.; Ho, C.-T. Reactivity and stability of selected flavor compounds. J. Food Drug Anal. 2015, 23, 176–190. [Google Scholar] [CrossRef]
- Thomas, D.; Surdin-Kerjan, Y. Metabolism of sulfur amino acids in Saccharomyces cerevisiae. Microbiol. Mol. Biol. Rev. 1997, 61, 503–532. [Google Scholar]
- Zhu, Q.; Zhang, L.; Sun, X.; Sun, B.; Zhang, Y. Analyzing the Effect of Dried Shrimp on the Flavor of Sheep Bone Soup Through Sensory Evaluation Combined with Untargeted Approaches. Foods 2025, 14, 1425. [Google Scholar] [CrossRef]
- Gaquerel, E.; Kotkar, H.; Onkokesung, N.; Galis, I.; Baldwin, I.T. Silencing an N-acyltransferase-like involved in lignin biosynthesis in Nicotiana attenuata dramatically alters herbivory-induced phenolamide metabolism. PLoS ONE 2013, 8, e62336. [Google Scholar] [CrossRef]
- Wu, J.; Lin, X.; Cui, C.; Li, J. Comparative analysis of kokumi effects in Ile-Leu, Ile-Ile, Leu-Leu, and Leu-Ile: Sensory, simulation, and structural insights. Food Chem. 2025, 474, 143004. [Google Scholar] [CrossRef]
- Zhang, X.; Li, H.; Nie, J.; Wu, D.; Huang, Q. Unveiling flavor formation and variation in fermented vinasse grass carp based on the dynamic correlation of microbiota with metabolites by multi-omics and bioinformatics approaches. Food Chem. 2025, 487, 144730. [Google Scholar] [CrossRef]
- Eguchi, K.; Ohtubo, Y.; Yoshii, K. Functional expression of M3, a muscarinic acetylcholine receptor subtype, in taste bud cells of mouse fungiform papillae. Chem. Senses 2008, 33, 47–55. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Nimker, V.; Patel, A.K.; Chen, C.-W.; Giri, B.S.; Dong, C.-D.; Singhania, R.R. Molecular engineering and biotechnological advancements in β-glucosidase for industrial impact: A review. Int. J. Biol. Macromol. 2025, 318, 145133. [Google Scholar] [CrossRef]
- Lin, P.; Shi, M.; Wang, F.; Lin, Y.; Zheng, Y. The α-Glucosidase Inhibition Activities of Phaeochromycins D and E Isolated from Marine Streptomyces sp. FJ0218. Molecules 2025, 30, 1993. [Google Scholar] [CrossRef] [PubMed]
- Mosharov, E.; Cranford, M.R.; Banerjee, R. The quantitatively important relationship between homocysteine metabolism and glutathione synthesis by the transsulfuration pathway and its regulation by redox changes. Biochemistry 2000, 39, 13005–13011. [Google Scholar] [CrossRef]


| Group | TX (N = 4) | HLJ (N = 4) | JM (N = 4) | Total (N = 12) |
|---|---|---|---|---|
| Hardness | ||||
| Mean ± SD | 8369.65 ± 1300.18 | 9685.97 ± 1433.13 | 6523.82 ± 1406.14 | 8193.15 ± 1842.67 |
| Median [min–max] | 8120.23 [7155.98, 10,082.16] | 9832.19 [7912.17, 11,167.32] | 7098.89 [4441.36, 7456.14] | 7756.09 [4441.36, 11,167.32] |
| Springiness | ||||
| Mean ± SD | 0.62 ± 0.10 | 0.59 ± 0.05 | 0.69 ± 0.08 | 0.63 ± 0.08 |
| Median [min–max] | 0.62 [0.53, 0.72] | 0.60 [0.52, 0.64] | 0.69 [0.60, 0.77] | 0.62 [0.52, 0.77] |
| Cohesiveness | ||||
| Mean ± SD | 0.53 ± 0.06 | 0.62 ± 0.02 | 0.52 ± 0.03 | 0.56 ± 0.06 |
| Median [min–max] | 0.54 [0.45, 0.58] | 0.63 [0.59, 0.64] | 0.52 [0.50, 0.56] | 0.56 [0.45, 0.64] |
| Gumminess | ||||
| Mean ± SD | 4363.10 ± 490.53 | 5989.08 ± 771.53 | 3396.43 ± 620.81 | 4582.87 ± 1257.61 |
| Median [min–max] | 4331.43 [3830.61, 4958.93] | 6017.58 [5030.37, 6890.80] | 3702.94 [2465.28, 3714.55] | 4331.43 [2465.28, 6890.80] |
| Chewiness | ||||
| Mean ± SD | 2685.98 ± 323.74 | 3517.26 ± 330.94 | 2339.58 ± 562.29 | 2847.60 ± 641.17 |
| Median [min–max] | 2690.10 [2294.56, 3069.16] | 3644.23 [3029.38, 3751.21] | 2458.84 [1594.48, 2846.14] | 2810.49 [1594.48, 3751.21] |
| Resilience | ||||
| Mean ± SD | 0.19 ± 0.04 | 0.27 ± 0.01 | 0.20 ± 0.02 | 0.22 ± 0.04 |
| Median [min–max] | 0.20 [0.15, 0.23] | 0.27 [0.26, 0.28] | 0.20 [0.18, 0.22] | 0.22 [0.15, 0.28] |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license.
Share and Cite
Wu, Q.; Zhang, H.; Xin, S.; Guo, J.; Yang, X.; Wang, Q.; Fang, H. Deciphering Northeast–Northwest Differences in Steamed Bread Microbiota and Flavor via Metagenomics and Untargeted Metabolomics. Fermentation 2026, 12, 153. https://doi.org/10.3390/fermentation12030153
Wu Q, Zhang H, Xin S, Guo J, Yang X, Wang Q, Fang H. Deciphering Northeast–Northwest Differences in Steamed Bread Microbiota and Flavor via Metagenomics and Untargeted Metabolomics. Fermentation. 2026; 12(3):153. https://doi.org/10.3390/fermentation12030153
Chicago/Turabian StyleWu, Qing, Heyu Zhang, Shihua Xin, Jianhong Guo, Xiaoping Yang, Qi Wang, and Haitian Fang. 2026. "Deciphering Northeast–Northwest Differences in Steamed Bread Microbiota and Flavor via Metagenomics and Untargeted Metabolomics" Fermentation 12, no. 3: 153. https://doi.org/10.3390/fermentation12030153
APA StyleWu, Q., Zhang, H., Xin, S., Guo, J., Yang, X., Wang, Q., & Fang, H. (2026). Deciphering Northeast–Northwest Differences in Steamed Bread Microbiota and Flavor via Metagenomics and Untargeted Metabolomics. Fermentation, 12(3), 153. https://doi.org/10.3390/fermentation12030153






