Functional Relevance of the Long Intergenic Non-Coding RNA Regulator of Reprogramming (Linc-ROR) in Cancer Proliferation, Metastasis, and Drug Resistance
Abstract
1. Introduction
2. Overview of Long Non-Coding RNAs
3. Linc-ROR in Disease
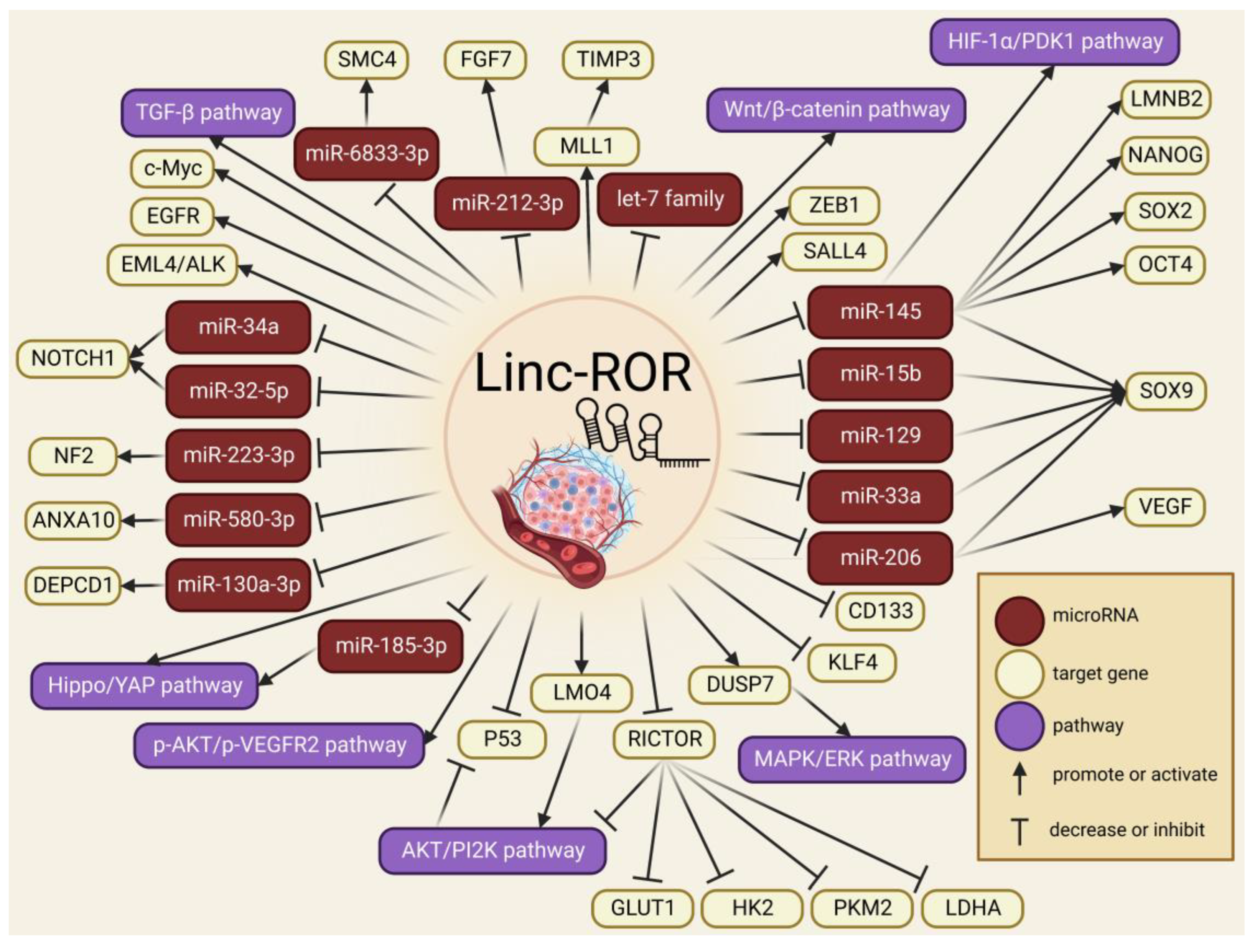
4. Linc-ROR in Cancer Proliferation and Progression
5. Linc-ROR in EMT, Cancer Invasion and Metastasis
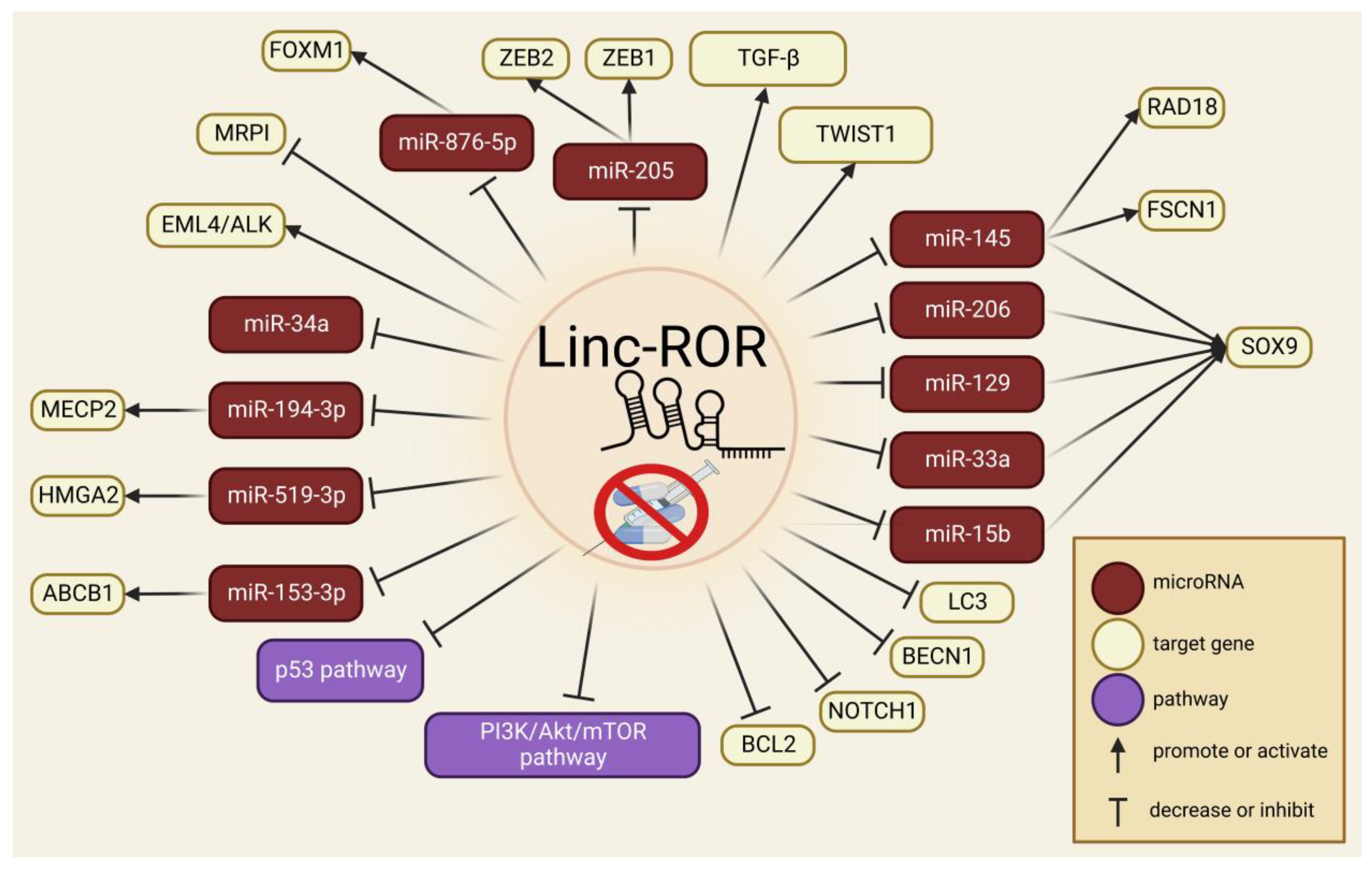
6. Linc-ROR in Drug Resistance
7. Clinical Relevance of Linc-ROR in Cancer
8. Conclusions
Author Contributions
Funding
Conflicts of Interest
References
- Sung, H.; Ferlay, J.; Siegel, R.L.; Laversanne, M.; Soerjomataram, I.; Jemal, A.; Bray, F. Global Cancer Statistics 2020: GLOBOCAN Estimates of Incidence and Mortality Worldwide for 36 Cancers in 185 Countries. CA Cancer J. Clin. 2021, 71, 209–249. [Google Scholar] [CrossRef] [PubMed]
- Lambert, A.W.; Pattabiraman, D.R.; Weinberg, R.A. Emerging Biological Principles of Metastasis. Cell 2017, 168, 670–691. [Google Scholar] [CrossRef] [PubMed]
- Eggert, J.; Society, O.N. Cancer Basics; Oncology Nursing Society: Pittsburgh Metropolitan Area, PA, USA, 2010. [Google Scholar]
- Massagué, J.; Batlle, E.; Gomis, R.R. Understanding the molecular mechanisms driving metastasis. Mol. Oncol. 2017, 11, 3–4. [Google Scholar] [CrossRef]
- Bukowski, K.; Kciuk, M.; Kontek, R. Mechanisms of Multidrug Resistance in Cancer Chemotherapy. Int. J. Mol. Sci. 2020, 21, 3233. [Google Scholar] [CrossRef] [PubMed]
- Zhou, Y.; Sun, W.; Qin, Z.; Guo, S.; Kang, Y.; Zeng, S.; Yu, L. LncRNA regulation: New frontiers in epigenetic solutions to drug chemoresistance. Biochem. Pharmacol. 2021, 189, 114228. [Google Scholar] [CrossRef]
- Zhang, P.; Wu, W.; Chen, Q.; Chen, M. Non-Coding RNAs and their Integrated Networks. J. Integr. Bioinform. 2019, 16, 20190027. [Google Scholar] [CrossRef]
- Saw, P.E.; Xu, X.; Chen, J.; Song, E.W. Non-coding RNAs: The new central dogma of cancer biology. Sci. China Life Sci. 2021, 64, 22–50. [Google Scholar] [CrossRef]
- Peña-Flores, J.A.; Bermúdez, M.; Ramos-Payán, R.; Villegas-Mercado, C.E.; Soto-Barreras, U.; Muela-Campos, D.; Álvarez-Ramírez, A.; Pérez-Aguirre, B.; Larrinua-Pacheco, A.D.; López-Camarillo, C.; et al. Emerging role of lncRNAs in drug resistance mechanisms in head and neck squamous cell carcinoma. Front. Oncol. 2022, 12, 965628. [Google Scholar] [CrossRef]
- Hombach, S.; Kretz, M. Non-coding RNAs: Classification, Biology and Functioning. Adv. Exp. Med. Biol. 2016, 937, 3–17. [Google Scholar] [CrossRef]
- Yan, H.; Bu, P. Non-coding RNA in cancer. Essays Biochem. 2021, 65, 625–639. [Google Scholar] [CrossRef]
- Ali Syeda, Z.; Langden, S.S.S.; Munkhzul, C.; Lee, M.; Song, S.J. Regulatory Mechanism of MicroRNA Expression in Cancer. Int. J. Mol. Sci. 2020, 21, 1723. [Google Scholar] [CrossRef]
- Fabian, M.R.; Sonenberg, N.; Filipowicz, W. Regulation of mRNA translation and stability by microRNAs. Annu. Rev. Biochem. 2010, 79, 351–379. [Google Scholar] [CrossRef]
- Engreitz, J.M.; Haines, J.E.; Perez, E.M.; Munson, G.; Chen, J.; Kane, M.; McDonel, P.E.; Guttman, M.; Lander, E.S. Local regulation of gene expression by lncRNA promoters, transcription and splicing. Nature 2016, 539, 452–455. [Google Scholar] [CrossRef] [PubMed]
- Laham-Karam, N.; Laitinen, P.; Turunen, T.A.; Ylä-Herttuala, S. Activating the Chromatin by Noncoding RNAs. Antioxid. Redox Signal. 2018, 29, 813–831. [Google Scholar] [CrossRef] [PubMed]
- Liu, Y.; Liu, X.; Lin, C.; Jia, X.; Zhu, H.; Song, J.; Zhang, Y. Noncoding RNAs regulate alternative splicing in Cancer. J. Exp. Clin. Cancer Res. 2021, 40, 11. [Google Scholar] [CrossRef] [PubMed]
- Teng, L.; Feng, Y.C.; Guo, S.T.; Wang, P.L.; Qi, T.F.; Yue, Y.M.; Wang, S.X.; Zhang, S.N.; Tang, C.X.; La, T.; et al. The pan-cancer lncRNA PLANE regulates an alternative splicing program to promote cancer pathogenesis. Nat. Commun. 2021, 12, 3734. [Google Scholar] [CrossRef] [PubMed]
- Cao, H.L.; Liu, Z.J.; Huang, P.L.; Yue, Y.L.; Xi, J.N. lncRNA-RMRP promotes proliferation, migration and invasion of bladder cancer via miR-206. Eur. Rev. Med. Pharmacol. Sci. 2019, 23, 1012–1021. [Google Scholar] [CrossRef]
- Li, D.; Zhang, J.; Li, J. Role of miRNA sponges in hepatocellular carcinoma. Clin. Chim. Acta 2020, 500, 10–19. [Google Scholar] [CrossRef]
- Zhao, Y.; Yuan, D.; Zhu, D.; Xu, T.; Huang, A.; Jiang, L.; Liu, C.; Qian, H.; Bu, X. LncRNA-MSC-AS1 inhibits the ovarian cancer progression by targeting miR-425-5p. J. Ovarian Res. 2021, 14, 109. [Google Scholar] [CrossRef]
- Ding, X.; Jia, X.; Wang, C.; Xu, J.; Gao, S.J.; Lu, C. A DHX9-lncRNA-MDM2 interaction regulates cell invasion and angiogenesis of cervical cancer. Cell Death Differ. 2019, 26, 1750–1765. [Google Scholar] [CrossRef]
- Dykes, I.M.; Emanueli, C. Transcriptional and Post-transcriptional Gene Regulation by Long Non-coding RNA. Genom. Proteom. Bioinform. 2017, 15, 177–186. [Google Scholar] [CrossRef]
- Sanchez Calle, A.; Kawamura, Y.; Yamamoto, Y.; Takeshita, F.; Ochiya, T. Emerging roles of long non-coding RNA in cancer. Cancer Sci. 2018, 109, 2093–2100. [Google Scholar] [CrossRef] [PubMed]
- Chen, W.; Wang, H.; Liu, Y.; Xu, W.; Ling, C.; Li, Y.; Liu, J.; Chen, M.; Zhang, Y.; Chen, B.; et al. Linc-RoR promotes proliferation, migration, and invasion via the Hippo/YAP pathway in pancreatic cancer cells. J Cell. Biochem. 2020, 121, 632–641. [Google Scholar] [CrossRef]
- Li, C.; Lu, L.; Feng, B.; Zhang, K.; Han, S.; Hou, D.; Chen, L.; Chu, X.; Wang, R. The lincRNA-ROR/miR-145 axis promotes invasion and metastasis in hepatocellular carcinoma via induction of epithelial-mesenchymal transition by targeting ZEB2. Sci. Rep. 2017, 7, 4637. [Google Scholar] [CrossRef] [PubMed]
- Yue, B.; Liu, C.; Sun, H.; Liu, M.; Song, C.; Cui, R.; Qiu, S.; Zhong, M. A Positive Feed-Forward Loop between LncRNA-CYTOR and Wnt/β-Catenin Signaling Promotes Metastasis of Colon Cancer. Mol. Ther. 2018, 26, 1287–1298. [Google Scholar] [CrossRef] [PubMed]
- Wang, Y.; Gao, L.; Zhu, B.; Zhu, H.; Luo, Y.; Wang, Q.; Zuo, J. Integrative analysis of long non-coding RNA acting as ceRNAs involved in chilling injury in tomato fruit. Gene 2018, 667, 25–33. [Google Scholar] [CrossRef] [PubMed]
- Bhan, A.; Soleimani, M.; Mandal, S.S. Long Noncoding RNA and Cancer: A New Paradigm. Cancer Res. 2017, 77, 3965–3981. [Google Scholar] [CrossRef]
- Jarroux, J.; Morillon, A.; Pinskaya, M. History, Discovery, and Classification of lncRNAs. Adv. Exp. Med. Biol. 2017, 1008, 1–46. [Google Scholar] [CrossRef]
- Institute, E.E.B. Statistics about the Current GENCODE Release (Version 42). Available online: https://www.gencodegenes.org/human/stats.html (accessed on 16 December 2022).
- Liu, Y.; Yang, Y.; Li, L.; Liu, Y.; Geng, P.; Li, G.; Song, H. LncRNA SNHG1 enhances cell proliferation, migration, and invasion in cervical cancer. Biochem. Cell Biol. 2018, 96, 38–43. [Google Scholar] [CrossRef]
- Luo, H.; Xu, C.; Le, W.; Ge, B.; Wang, T. lncRNA CASC11 promotes cancer cell proliferation in bladder cancer through miRNA-150. J. Cell. Biochem. 2019, 120, 13487–13493. [Google Scholar] [CrossRef]
- Luo, J.; Wang, K.; Yeh, S.; Sun, Y.; Liang, L.; Xiao, Y.; Xu, W.; Niu, Y.; Cheng, L.; Maity, S.N.; et al. LncRNA-p21 alters the antiandrogen enzalutamide-induced prostate cancer neuroendocrine differentiation via modulating the EZH2/STAT3 signaling. Nat. Commun. 2019, 10, 2571. [Google Scholar] [CrossRef] [PubMed]
- Sun, Z.; Xue, S.; Zhang, M.; Xu, H.; Hu, X.; Chen, S.; Liu, Y.; Guo, M.; Cui, H. Aberrant NSUN2-mediated m(5)C modification of H19 lncRNA is associated with poor differentiation of hepatocellular carcinoma. Oncogene 2020, 39, 6906–6919. [Google Scholar] [CrossRef]
- Li, X.; Jin, F.; Li, Y. A novel autophagy-related lncRNA prognostic risk model for breast cancer. J. Cell. Mol. Med. 2021, 25, 4–14. [Google Scholar] [CrossRef] [PubMed]
- Luo, Y.; Zheng, S.; Wu, Q.; Wu, J.; Zhou, R.; Wang, C.; Wu, Z.; Rong, X.; Huang, N.; Sun, L.; et al. Long noncoding RNA (lncRNA) EIF3J-DT induces chemoresistance of gastric cancer via autophagy activation. Autophagy 2021, 17, 4083–4101. [Google Scholar] [CrossRef] [PubMed]
- Wang, L.; Cho, K.B.; Li, Y.; Tao, G.; Xie, Z.; Guo, B. Long Noncoding RNA (lncRNA)-Mediated Competing Endogenous RNA Networks Provide Novel Potential Biomarkers and Therapeutic Targets for Colorectal Cancer. Int. J. Mol. Sci. 2019, 20, 5758. [Google Scholar] [CrossRef]
- Cui, P.H.; Li, Z.Y.; Li, D.H.; Han, S.Y.; Zhang, Y.J. SP1-induced lncRNA DANCR contributes to proliferation and invasion of ovarian cancer. Kaohsiung J. Med. Sci. 2021, 37, 371–378. [Google Scholar] [CrossRef]
- Zhang, Y.X.; Yuan, J.; Gao, Z.M.; Zhang, Z.G. LncRNA TUC338 promotes invasion of lung cancer by activating MAPK pathway. Eur. Rev. Med. Pharmacol. Sci. 2018, 22, 443–449. [Google Scholar] [CrossRef]
- Zhao, W.; Geng, D.; Li, S.; Chen, Z.; Sun, M. LncRNA HOTAIR influences cell growth, migration, invasion, and apoptosis via the miR-20a-5p/HMGA2 axis in breast cancer. Cancer Med. 2018, 7, 842–855. [Google Scholar] [CrossRef]
- Li, J.; Meng, H.; Bai, Y.; Wang, K. Regulation of lncRNA and Its Role in Cancer Metastasis. Oncol. Res. 2016, 23, 205–217. [Google Scholar] [CrossRef]
- Luan, X.; Wang, Y. LncRNA XLOC_006390 facilitates cervical cancer tumorigenesis and metastasis as a ceRNA against miR-331-3p and miR-338-3p. J. Gynecol. Oncol. 2018, 29, e95. [Google Scholar] [CrossRef]
- Park, E.G.; Pyo, S.J.; Cui, Y.; Yoon, S.H.; Nam, J.W. Tumor immune microenvironment lncRNAs. Brief. Bioinform. 2022, 23, bbab504. [Google Scholar] [CrossRef] [PubMed]
- Lin, W.; Zhou, Q.; Wang, C.Q.; Zhu, L.; Bi, C.; Zhang, S.; Wang, X.; Jin, H. LncRNAs regulate metabolism in cancer. Int. J. Biol. Sci. 2020, 16, 1194–1206. [Google Scholar] [CrossRef]
- Schmitz, S.U.; Grote, P.; Herrmann, B.G. Mechanisms of long noncoding RNA function in development and disease. Cell. Mol. Life Sci. 2016, 73, 2491–2509. [Google Scholar] [CrossRef] [PubMed]
- Chan, J.J.; Tay, Y. Noncoding RNA:RNA Regulatory Networks in Cancer. Int. J. Mol. Sci. 2018, 19, 1310. [Google Scholar] [CrossRef] [PubMed]
- Li, X.; Yang, L.; Chen, L.L. The Biogenesis, Functions, and Challenges of Circular RNAs. Mol. Cell 2018, 71, 428–442. [Google Scholar] [CrossRef]
- Charles Richard, J.L.; Eichhorn, P.J.A. Platforms for Investigating LncRNA Functions. SLAS Technol. 2018, 23, 493–506. [Google Scholar] [CrossRef] [PubMed]
- Zibitt, M.S.; Hartford, C.C.R.; Lal, A. Interrogating lncRNA functions via CRISPR/Cas systems. RNA Biol. 2021, 18, 2097–2106. [Google Scholar] [CrossRef]
- Dahariya, S.; Paddibhatla, I.; Kumar, S.; Raghuwanshi, S.; Pallepati, A.; Gutti, R.K. Long non-coding RNA: Classification, biogenesis and functions in blood cells. Mol. Immunol. 2019, 112, 82–92. [Google Scholar] [CrossRef]
- Long, Y.; Wang, X.; Youmans, D.T.; Cech, T.R. How do lncRNAs regulate transcription? Sci. Adv. 2017, 3, eaao2110. [Google Scholar] [CrossRef]
- Paraskevopoulou, M.D.; Hatzigeorgiou, A.G. Analyzing MiRNA-LncRNA Interactions. Methods Mol. Biol. 2016, 1402, 271–286. [Google Scholar] [CrossRef]
- He, Y.; Dan, Y.; Gao, X.; Huang, L.; Lv, H.; Chen, J. DNMT1-mediated lncRNA MEG3 methylation accelerates endothelial-mesenchymal transition in diabetic retinopathy through the PI3K/Akt/mTOR signaling pathway. Am. J. Physiol. Endocrinol. Metab. 2021, 320, E598–E608. [Google Scholar] [CrossRef]
- Thomas, A.A.; Biswas, S.; Feng, B.; Chen, S.; Gonder, J.; Chakrabarti, S. lncRNA H19 prevents endothelial-mesenchymal transition in diabetic retinopathy. Diabetologia 2019, 62, 517–530. [Google Scholar] [CrossRef] [PubMed]
- Fasolo, F.; Jin, H.; Winski, G.; Chernogubova, E.; Pauli, J.; Winter, H.; Li, D.Y.; Glukha, N.; Bauer, S.; Metschl, S.; et al. Long Noncoding RNA MIAT Controls Advanced Atherosclerotic Lesion Formation and Plaque Destabilization. Circulation 2021, 144, 1567–1583. [Google Scholar] [CrossRef] [PubMed]
- Ye, Z.M.; Yang, S.; Xia, Y.P.; Hu, R.T.; Chen, S.; Li, B.W.; Chen, S.L.; Luo, X.Y.; Mao, L.; Li, Y.; et al. LncRNA MIAT sponges miR-149-5p to inhibit efferocytosis in advanced atherosclerosis through CD47 upregulation. Cell Death. Dis. 2019, 10, 138. [Google Scholar] [CrossRef] [PubMed]
- Cai, L.J.; Tu, L.; Huang, X.M.; Huang, J.; Qiu, N.; Xie, G.H.; Liao, J.X.; Du, W.; Zhang, Y.Y.; Tian, J.Y. LncRNA MALAT1 facilitates inflammasome activation via epigenetic suppression of Nrf2 in Parkinson′s disease. Mol. Brain. 2020, 13, 130. [Google Scholar] [CrossRef] [PubMed]
- Cheng, J.; Duan, Y.; Zhang, F.; Shi, J.; Li, H.; Wang, F.; Li, H. The Role of lncRNA TUG1 in the Parkinson Disease and Its Effect on Microglial Inflammatory Response. Neuromol. Med. 2021, 23, 327–334. [Google Scholar] [CrossRef]
- Zhang, M.; Lu, N.; Guo, X.Y.; Li, H.J.; Guo, Y.; Lu, L. Influences of the lncRNA TUG1-miRNA-34a-5p network on fibroblast-like synoviocytes (FLSs) dysfunction in rheumatoid arthritis through targeting the lactate dehydrogenase A (LDHA). J. Clin. Lab. Anal. 2021, 35, e23969. [Google Scholar] [CrossRef]
- Najafi, S.; Khatami, S.H.; Khorsand, M.; Jamali, Z.; Shabaninejad, Z.; Moazamfard, M.; Majidpoor, J.; Aghaei Zarch, S.M.; Movahedpour, A. Long non-coding RNAs (lncRNAs); roles in tumorigenesis and potentials as biomarkers in cancer diagnosis. Exp. Cell Res. 2022, 418, 113294. [Google Scholar] [CrossRef]
- Zhang, E.; He, X.; Zhang, C.; Su, J.; Lu, X.; Si, X.; Chen, J.; Yin, D.; Han, L.; De, W. A novel long noncoding RNA HOXC-AS3 mediates tumorigenesis of gastric cancer by binding to YBX1. Genome Biol. 2018, 19, 154. [Google Scholar] [CrossRef]
- Su, H.; Fan, G.; Huang, J.; Qiu, X. LncRNA HOXC-AS3 promotes non-small-cell lung cancer growth and metastasis through upregulation of YBX1. Cell Death Dis. 2022, 13, 307. [Google Scholar] [CrossRef]
- Han, X.; Jiang, H.; Qi, J.; Li, J.; Yang, J.; Tian, Y.; Li, W.; Jing, Q.; Wang, C. Novel lncRNA UPLA1 mediates tumorigenesis and prognosis in lung adenocarcinoma. Cell Death Dis. 2020, 11, 999. [Google Scholar] [CrossRef] [PubMed]
- Ni, W.; Zhang, Y.; Zhan, Z.; Ye, F.; Liang, Y.; Huang, J.; Chen, K.; Chen, L.; Ding, Y. A novel lncRNA uc.134 represses hepatocellular carcinoma progression by inhibiting CUL4A-mediated ubiquitination of LATS1. J. Hematol. Oncol. 2017, 10, 91. [Google Scholar] [CrossRef] [PubMed]
- Ransohoff, J.D.; Wei, Y.; Khavari, P.A. The functions and unique features of long intergenic non-coding RNA. Nat. Rev. Mol. Cell Biol. 2018, 19, 143–157. [Google Scholar] [CrossRef]
- Chen, Y.M.; Liu, Y.; Wei, H.Y.; Lv, K.Z.; Fu, P.F. Large intergenic non-coding RNA-ROR reverses gemcitabine-induced autophagy and apoptosis in breast cancer cells. Oncotarget 2016, 7, 59604–59617. [Google Scholar] [CrossRef] [PubMed]
- Quek, X.C.; Thomson, D.W.; Maag, J.L.; Bartonicek, N.; Signal, B.; Clark, M.B.; Gloss, B.S.; Dinger, M.E. lncRNAdb v2.0: Expanding the reference database for functional long noncoding RNAs. Nucleic Acids Res. 2015, 43, D168–D173. [Google Scholar] [CrossRef] [PubMed]
- Gao, S.; Wang, P.; Hua, Y.; Xi, H.; Meng, Z.; Liu, T.; Chen, Z.; Liu, L. ROR functions as a ceRNA to regulate Nanog expression by sponging miR-145 and predicts poor prognosis in pancreatic cancer. Oncotarget 2016, 7, 1608–1618. [Google Scholar] [CrossRef] [PubMed]
- Zhang, H.Y.; Liang, F.; Zhang, J.W.; Wang, F.; Wang, L.; Kang, X.G. Effects of long noncoding RNA-ROR on tamoxifen resistance of breast cancer cells by regulating microRNA-205. Cancer Chemother. Pharmacol. 2017, 79, 327–337. [Google Scholar] [CrossRef]
- Zhou, Q.; Guo, J.; Huang, W.; Yu, X.; Xu, C.; Long, X. Linc-ROR promotes the progression of breast cancer and decreases the sensitivity to rapamycin through miR-194-3p targeting MECP2. Mol. Oncol. 2020, 14, 2231–2250. [Google Scholar] [CrossRef]
- Wang, Y.; Xu, Z.; Jiang, J.; Xu, C.; Kang, J.; Xiao, L.; Wu, M.; Xiong, J.; Guo, X.; Liu, H. Endogenous miRNA sponge lincRNA-RoR regulates Oct4, Nanog, and Sox2 in human embryonic stem cell self-renewal. Dev. Cell 2013, 25, 69–80. [Google Scholar] [CrossRef]
- Zou, G.; Liu, T.; Guo, L.; Huang, Y.; Feng, Y.; Huang, Q.; Duan, T. miR-145 modulates lncRNA-ROR and Sox2 expression to maintain human amniotic epithelial stem cell pluripotency and β islet-like cell differentiation efficiency. Gene 2016, 591, 48–57. [Google Scholar] [CrossRef]
- Fu, Y.; Hu, X.; Gao, Y.; Li, K.; Fu, Q.; Liu, Q.; Liu, D.; Zhang, Z.; Qiao, J. LncRNA ROR/miR-145-5p axis modulates the osteoblasts proliferation and apoptosis in osteoporosis. Bioengineered 2021, 12, 7714–7723. [Google Scholar] [CrossRef] [PubMed]
- Feng, L.; Yang, Z.M.; Li, Y.C.; Wang, H.X.; Lo, J.H.T.; Zhang, X.T.; Li, G. Linc-ROR promotes mesenchymal stem cells chondrogenesis and cartilage formation via regulating SOX9 expression. Osteoarthr. Cartil. 2021, 29, 568–578. [Google Scholar] [CrossRef] [PubMed]
- Qin, W.W.; Xin, Z.L.; Wang, H.Q.; Wang, K.P.; Li, X.Y.; Wang, X. Inhibiting lncRNA ROR suppresses growth, migration and angiogenesis in microvascular endothelial cells by up-regulating miR-26. Eur. Rev. Med. Pharm. Sci. 2018, 22, 7985–7993. [Google Scholar] [CrossRef]
- Chen, G.; Xu, C.; Gillette, T.G.; Huang, T.; Huang, P.; Li, Q.; Li, X.; Li, Q.; Ning, Y.; Tang, R.; et al. Cardiomyocyte-derived small extracellular vesicles can signal eNOS activation in cardiac microvascular endothelial cells to protect against Ischemia/Reperfusion injury. Theranostics 2020, 10, 11754–11774. [Google Scholar] [CrossRef] [PubMed]
- Hu, Y.H.; Sun, J.; Zhang, J.; Hua, F.Z.; Liu, Q.; Liang, Y.P. Long non-coding RNA ROR sponges miR-138 to aggravate hypoxia/reoxygenation-induced cardiomyocyte apoptosis via upregulating Mst1. Exp. Mol. Pathol. 2020, 114, 104430. [Google Scholar] [CrossRef]
- Zeng, M.; Yi, S.; Xiao, Y.; Chen, Z. LncRNA ROR promotes NLRP3-mediated cardiomyocyte pyroptosis by upregulating FOXP1 via interactions with PTBP1. Cytokine 2022, 152, 155812. [Google Scholar] [CrossRef]
- Honarmand Tamizkar, K.; Gorji, P.; Gholipour, M.; Hussen, B.M.; Mazdeh, M.; Eslami, S.; Taheri, M.; Ghafouri-Fard, S. Parkinson’s Disease Is Associated With Dysregulation of Circulatory Levels of lncRNAs. Front. Immunol. 2021, 12, 763323. [Google Scholar] [CrossRef]
- Fallah, H.; Azari, I.; Neishabouri, S.M.; Oskooei, V.K.; Taheri, M.; Ghafouri-Fard, S. Sex-specific up-regulation of lncRNAs in peripheral blood of patients with schizophrenia. Sci. Rep. 2019, 9, 12737. [Google Scholar] [CrossRef]
- Ghafouri-Fard, S.; Eghtedarian, R.; Taheri, M.; Beatrix Brühl, A.; Sadeghi-Bahmani, D.; Brand, S. A Review on the Expression Pattern of Non-coding RNAs in Patients With Schizophrenia: With a Special Focus on Peripheral Blood as a Source of Expression Analysis. Front. Psychiatry 2021, 12, 640463. [Google Scholar] [CrossRef]
- Hashemian, F.; Ghafouri-Fard, S.; Arsang-Jang, S.; Mirzajani, S.; Fallah, H.; Mehvari Habibabadi, J.; Sayad, A.; Taheri, M. Epilepsy Is Associated With Dysregulation of Long Non-coding RNAs in the Peripheral Blood. Front. Mol. Biosci. 2019, 6, 113. [Google Scholar] [CrossRef]
- Liu, X.; Wang, L.; Wang, Q.; Zhao, J.; Chang, H.; Zhu, R. Association Between Genetic Variants in the lncRNA-p53 Regulatory Network and Ischemic Stroke Prognosis. Neurotox Res. 2021, 39, 1171–1180. [Google Scholar] [CrossRef] [PubMed]
- Zhou, Q.; An, Y.; Tang, Y. Long noncoding RNA-regulator of reprogramming alleviates hypoxia-induced cerebral injury in mice model and human via modulating apoptosis complexes. J. Integr. Neurosci. 2019, 18, 431–437. [Google Scholar] [CrossRef]
- Chen, W.; Yang, J.; Fang, H.; Li, L.; Sun, J. Relevance Function of Linc-ROR in the Pathogenesis of Cancer. Front. Cell Dev. Biol. 2020, 8, 696. [Google Scholar] [CrossRef]
- Yu, X.; Ding, H.; Shi, Y.; Yang, L.; Zhou, J.; Yan, Z.; Xiao, B. Downregulated Expression of linc-ROR in Gastric Cancer and Its Potential Diagnostic and Prognosis Value. Dis. Markers 2020, 2020, 7347298. [Google Scholar] [CrossRef]
- Feng, S.; Yao, J.; Chen, Y.; Geng, P.; Zhang, H.; Ma, X.; Zhao, J.; Yu, X. Expression and Functional Role of Reprogramming-Related Long Noncoding RNA (lincRNA-ROR) in Glioma. J. Mol. Neurosci. 2015, 56, 623–630. [Google Scholar] [CrossRef] [PubMed]
- Shi, H.; Pu, J.; Zhou, X.L.; Ning, Y.Y.; Bai, C. Silencing long non-coding RNA ROR improves sensitivity of non-small-cell lung cancer to cisplatin resistance by inhibiting PI3K/Akt/mTOR signaling pathway. Tumour Biol. 2017, 39, 1010428317697568. [Google Scholar] [CrossRef]
- Peng, W.X.; Huang, J.G.; Yang, L.; Gong, A.H.; Mo, Y.Y. Linc-RoR promotes MAPK/ERK signaling and confers estrogen-independent growth of breast cancer. Mol. Cancer 2017, 16, 161. [Google Scholar] [CrossRef]
- Chen, Y.M.; Liu, Y.; Wei, H.Y.; Lv, K.Z.; Fu, P. Linc-ROR induces epithelial-mesenchymal transition and contributes to drug resistance and invasion of breast cancer cells. Tumour Biol. 2016, 37, 10861–10870. [Google Scholar] [CrossRef]
- Li, Y.; Jiang, B.; Zhu, H.; Qu, X.; Zhao, L.; Tan, Y.; Jiang, Y.; Liao, M.; Wu, X. Inhibition of long non-coding RNA ROR reverses resistance to Tamoxifen by inducing autophagy in breast cancer. Tumour Biol. 2017, 39, 1010428317705790. [Google Scholar] [CrossRef] [PubMed]
- Hou, P.; Zhao, Y.; Li, Z.; Yao, R.; Ma, M.; Gao, Y.; Zhao, L.; Zhang, Y.; Huang, B.; Lu, J. LincRNA-ROR induces epithelial-to-mesenchymal transition and contributes to breast cancer tumorigenesis and metastasis. Cell Death. Dis. 2014, 5, e1287. [Google Scholar] [CrossRef]
- Zhao, T.; Wu, L.; Li, X.; Dai, H.; Zhang, Z. Large intergenic non-coding RNA-ROR as a potential biomarker for the diagnosis and dynamic monitoring of breast cancer. Cancer Biomark. 2017, 20, 165–173. [Google Scholar] [CrossRef]
- Zhang, K.; Luo, Z.; Zhang, Y.; Wang, Y.; Cui, M.; Liu, L.; Zhang, L.; Liu, J. Detection and analysis of circulating large intergenic non-coding RNA regulator of reprogramming in plasma for breast cancer. Thorac. Cancer 2018, 9, 66–74. [Google Scholar] [CrossRef]
- Luo, C.; Cao, J.; Peng, R.; Guo, Q.; Ye, H.; Wang, P.; Wang, K.; Song, C. Functional Variants in Linc-ROR are Associated with mRNA Expression of Linc-ROR and Breast Cancer Susceptibility. Sci. Rep. 2018, 8, 4680. [Google Scholar] [CrossRef]
- Huang, J.; Zhang, A.; Ho, T.T.; Zhang, Z.; Zhou, N.; Ding, X.; Zhang, X.; Xu, M.; Mo, Y.Y. Linc-RoR promotes c-Myc expression through hnRNP I and AUF1. Nucleic Acids Res. 2016, 44, 3059–3069. [Google Scholar] [CrossRef] [PubMed]
- Hu, A.; Hong, F.; Li, D.; Jin, Y.; Kon, L.; Xu, Z.; He, H.; Xie, Q. Long non-coding RNA ROR recruits histone transmethylase MLL1 to up-regulate TIMP3 expression and promote breast cancer progression. J. Transl. Med. 2021, 19, 95. [Google Scholar] [CrossRef] [PubMed]
- Mohebi, M.; Sattari, A.; Ghafouri-Fard, S.; Modarressi, M.H.; Kholghi-Oskooei, V.; Taheri, M. Expression profiling revealed up-regulation of three lncRNAs in breast cancer samples. Exp. Mol. Pathol. 2020, 117, 104544. [Google Scholar] [CrossRef] [PubMed]
- Eades, G.; Wolfson, B.; Zhang, Y.; Li, Q.; Yao, Y.; Zhou, Q. lincRNA-RoR and miR-145 regulate invasion in triple-negative breast cancer via targeting ARF6. Mol. Cancer Res. 2015, 13, 330–338. [Google Scholar] [CrossRef]
- Ma, J.; Yang, Y.; Huo, D.; Wang, Z.; Zhai, X.; Chen, J.; Sun, H.; An, W.; Jie, J.; Yang, P. LincRNA-RoR/miR-145 promote invasion and metastasis in triple-negative breast cancer via targeting MUC1. Biochem. Biophys. Res. Commun. 2018, 500, 614–620. [Google Scholar] [CrossRef] [PubMed]
- Hou, L.; Tu, J.; Cheng, F.; Yang, H.; Yu, F.; Wang, M.; Liu, J.; Fan, J.; Zhou, G. Long noncoding RNA ROR promotes breast cancer by regulating the TGF-β pathway. Cancer Cell Int. 2018, 18, 142. [Google Scholar] [CrossRef] [PubMed]
- Jiang, B.; Zhu, H.; Tang, L.; Gao, T.; Zhou, Y.; Gong, F.; Tan, Y.; Xie, L.; Wu, X.; Li, Y. Apatinib Inhibits Stem Properties and Malignant Biological Behaviors of Breast Cancer Stem Cells by Blocking Wnt/β-catenin Signal Pathway through Downregulating LncRNA ROR. Anti-Cancer Agents Med. Chem. 2022, 22, 1723–1734. [Google Scholar] [CrossRef]
- Toraih, E.A.; El-Wazir, A.; Ageeli, E.A.; Hussein, M.H.; Eltoukhy, M.M.; Killackey, M.T.; Kandil, E.; Fawzy, M.S. Unleash multifunctional role of long noncoding RNAs biomarker panel in breast cancer: A predictor classification model. Epigenomics 2020, 12, 1215–1237. [Google Scholar] [CrossRef]
- Lou, Y.; Jiang, H.; Cui, Z.; Wang, L.; Wang, X.; Tian, T. Linc-ROR induces epithelial-to-mesenchymal transition in ovarian cancer by increasing Wnt/β-catenin signaling. Oncotarget 2017, 8, 69983–69994. [Google Scholar] [CrossRef]
- Jiang, H.H.; Lou, Y.H.; Wang, X.Y.; Han, Y.; Cui, Z.M. Expression and function of long intergenic non-protein coding RNA-regulator of reprogramming in high-grade ovarian serous cancer. Zhonghua Fu Chan Ke Za Zhi 2016, 51, 921–927. [Google Scholar] [CrossRef] [PubMed]
- Shen, W.; Xie, X.; Liu, M.; Wang, L. Diagnostic Value of lncRNA ROR in Differentiating Ovarian Cancer Patients. Clin. Lab. 2020, 66. [Google Scholar] [CrossRef]
- Li, J.; Zhang, S.; Wu, L.; Pei, M. Interaction between LncRNA-ROR and miR-145 contributes to epithelial-mesenchymal transition of ovarian cancer cells. Gen. Physiol. Biophys. 2019, 38, 461–471. [Google Scholar] [CrossRef] [PubMed]
- Zhou, X.; Gao, Q.; Wang, J.; Zhang, X.; Liu, K.; Duan, Z. Linc-RNA-RoR acts as a “sponge” against mediation of the differentiation of endometrial cancer stem cells by microRNA-145. Gynecol. Oncol. 2014, 133, 333–339. [Google Scholar] [CrossRef] [PubMed]
- Zeng, S.Y.; Liu, C.Q.; Zhuang, Y.; Chen, Y.; Gu, L.L.; Shi, S.Q. LncRNA ROR promotes proliferation of endometrial cancer cells via regulating Notch1 pathway. Eur. Rev. Med. Pharmacol. Sci. 2020, 24, 5970–5978. [Google Scholar] [CrossRef]
- Alsadat Mahmoudian, R.; Lotfi Gharaie, M.; Abbaszadegan, R.; Forghanifard, M.M.; Abbaszadegan, M.R. Interaction between LINC-ROR and Stemness State in Gastric Cancer Cells with Helicobacter pylori Infection. Iran. Biomed. J. 2021, 25, 157–168. [Google Scholar] [CrossRef]
- Jin, W.; Zhang, H.; Li, M.; Lin, S. Long Noncoding RNA Regulator of Reprogramming Regulates Cell Growth, Metastasis, and Cisplatin Resistance in Gastric Cancer via miR-519d-3p/HMGA2 Axis. Cancer Biother. Radiopharm. 2020. [Google Scholar] [CrossRef]
- Mi, Y.; Li, Y.; He, Z.; Chen, D.; Hong, Q.; You, J. Upregulation of Linc-ROR Promotes the Proliferation, Migration, and Invasion of Gastric Cancer Cells Through miR-212-3p/FGF7 Axis. Cancer Manag. Res. 2021, 13, 899–912. [Google Scholar] [CrossRef]
- Liu, M.; Zhang, M.; Yin, H. Linc-ROR promotes invasion and metastasis of gastric cancer by activating epithelial-mesenchymal transition. Indian J. Pathol. Microbiol. 2022, 65, 545–550. [Google Scholar] [CrossRef] [PubMed]
- Wang, S.; Liu, F.; Deng, J.; Cai, X.; Han, J.; Liu, Q. Long Noncoding RNA ROR Regulates Proliferation, Invasion, and Stemness of Gastric Cancer Stem Cell. Cell. Reprogram. 2016, 18, 319–326. [Google Scholar] [CrossRef] [PubMed]
- Wang, S.; Chen, W.; Yu, H.; Song, Z.; Li, Q.; Shen, X.; Wu, Y.; Zhu, L.; Ma, Q.; Xing, D. lncRNA ROR Promotes Gastric Cancer Drug Resistance. Cancer Control 2020, 27, 1073274820904694. [Google Scholar] [CrossRef] [PubMed]
- Soghala, S.; Harsiny, K.; Momeni, P.; Hatami, M.; Kholghi Oskooei, V.; Hussen, B.M.; Taheri, M.; Ghafouri-Fard, S. Down-regulation of LINC-ROR, HOXA-AS2 and MEG3 in gastric cancer. Heliyon 2022, 8, e11155. [Google Scholar] [CrossRef]
- Bai, L.; Zhuang, Y.; Xie, J.; Liu, K.; Yin, S.; Yan, F. SOX2-Induced Linc-ROR Upregulation Inhibits Gastric Carcinoma Cell Proliferation and Metastasis Via the miR-580-3p/ANXA10 Pathway. Biochem. Genet. 2022. [Google Scholar] [CrossRef]
- Pan, Y.; Chen, J.; Tao, L.; Zhang, K.; Wang, R.; Chu, X.; Chen, L. Long noncoding RNA ROR regulates chemoresistance in docetaxel-resistant lung adenocarcinoma cells via epithelial mesenchymal transition pathway. Oncotarget 2017, 8, 33144–33158. [Google Scholar] [CrossRef]
- Qu, C.H.; Sun, Q.Y.; Zhang, F.M.; Jia, Y.M. Long non-coding RNA ROR is a novel prognosis factor associated with non-small-cell lung cancer progression. Eur. Rev. Med. Pharmacol. Sci. 2017, 21, 4087–4091. [Google Scholar]
- Yang, Y.; Huang, J.; Xie, N.; Huang, H.; Xu, S.; Cai, J.; Qi, S. lincROR influences the stemness and crizotinib resistance in EML-ALK(+) non-small-cell lung cancer cells. OncoTargets Ther. 2018, 11, 3649–3657. [Google Scholar] [CrossRef]
- Xia, F.; Xiong, Y.; Li, Q. Interaction of lincRNA ROR and p53/miR-145 correlates with lung cancer stem cell signatures. J. Cell. Biochem. 2017. [Google Scholar] [CrossRef]
- Chen, Y.; Shen, Z.; Zhi, Y.; Zhou, H.; Zhang, K.; Wang, T.; Feng, B.; Chen, Y.; Song, H.; Wang, R.; et al. Long non-coding RNA ROR promotes radioresistance in hepatocelluar carcinoma cells by acting as a ceRNA for microRNA-145 to regulate RAD18 expression. Arch. Biochem. Biophys. 2018, 645, 117–125. [Google Scholar] [CrossRef]
- Li, X.; Li, N. LINC ROR from Hepatocarcinoma Cell-derived Exosomes Modulates Inflammation in Human Macrophages. Sichuan Da Xue Xue Bao Yi Xue Ban 2019, 50, 177–181. [Google Scholar] [PubMed]
- Zhang, Y.; Wu, W.; Sun, Q.; Ye, L.; Zhou, D.; Wang, W. Linc-ROR facilitates hepatocellular carcinoma resistance to doxorubicin by regulating TWIST1-mediated epithelial-mesenchymal transition. Mol. Med. Rep. 2021, 23, 340. [Google Scholar] [CrossRef] [PubMed]
- Tian, C.; Abudoureyimu, M.; Lin, X.; Chu, X.; Wang, R. Linc-ROR facilitates progression and angiogenesis of hepatocellular carcinoma by modulating DEPDC1 expression. Cell Death Dis. 2021, 12, 1047. [Google Scholar] [CrossRef] [PubMed]
- Zhi, Y.; Abudoureyimu, M.; Zhou, H.; Wang, T.; Feng, B.; Wang, R.; Chu, X. FOXM1-Mediated LINC-ROR Regulates the Proliferation and Sensitivity to Sorafenib in Hepatocellular Carcinoma. Mol. Ther. Nucleic Acids 2019, 16, 576–588. [Google Scholar] [CrossRef] [PubMed]
- Takahashi, K.; Yan, I.K.; Kogure, T.; Haga, H.; Patel, T. Extracellular vesicle-mediated transfer of long non-coding RNA ROR modulates chemosensitivity in human hepatocellular cancer. FEBS Open Bio 2014, 4, 458–467. [Google Scholar] [CrossRef]
- Takahashi, K.; Yan, I.K.; Haga, H.; Patel, T. Modulation of hypoxia-signaling pathways by extracellular linc-RoR. J. Cell Sci. 2014, 127, 1585–1594. [Google Scholar] [CrossRef]
- Li, X.; Sun, D.; Zhao, T.; Zhang, Z. Long non-coding RNA ROR confers arsenic trioxide resistance to HepG2 cells by inhibiting p53 expression. Eur. J. Pharmacol. 2020, 872, 172982. [Google Scholar] [CrossRef]
- Ma, Y.L.; Wang, C.Y.; Guan, Y.J.; Gao, F.M. Long noncoding RNA ROR promotes proliferation and invasion of colorectal cancer by inhibiting tumor suppressor gene NF2 through interacting with miR-223-3p. Eur. Rev. Med. Pharmacol. Sci. 2020, 24, 2401–2411. [Google Scholar] [CrossRef]
- He, X.; Yu, J.; Xiong, L.; Liu, Y.; Fan, L.; Li, Y.; Chen, B.; Chen, J.; Xu, X. Exosomes derived from liver cancer cells reprogram biological behaviors of LO2 cells by transferring Linc-ROR. Gene 2019, 719, 144044. [Google Scholar] [CrossRef]
- Zhan, H.X.; Wang, Y.; Li, C.; Xu, J.W.; Zhou, B.; Zhu, J.K.; Han, H.F.; Wang, L.; Wang, Y.S.; Hu, S.Y. LincRNA-ROR promotes invasion, metastasis and tumor growth in pancreatic cancer through activating ZEB1 pathway. Cancer Lett. 2016, 374, 261–271. [Google Scholar] [CrossRef]
- Sun, Z.; Sun, D.; Feng, Y.; Zhang, B.; Sun, P.; Zhou, B.; Du, L.; Wang, Y.; Fan, Z.; Yang, J.; et al. Exosomal linc-ROR mediates crosstalk between cancer cells and adipocytes to promote tumor growth in pancreatic cancer. Mol. Ther. Nucleic Acids 2021, 26, 253–268. [Google Scholar] [CrossRef]
- Fu, Z.; Li, G.; Li, Z.; Wang, Y.; Zhao, Y.; Zheng, S.; Ye, H.; Luo, Y.; Zhao, X.; Wei, L.; et al. Endogenous miRNA Sponge LincRNA-ROR promotes proliferation, invasion and stem cell-like phenotype of pancreatic cancer cells. Cell Death Discov. 2017, 3, 17004. [Google Scholar] [CrossRef]
- Arunkumar, G.; Deva Magendhra Rao, A.K.; Manikandan, M.; Arun, K.; Vinothkumar, V.; Revathidevi, S.; Rajkumar, K.S.; Rajaraman, R.; Munirajan, A.K. Expression profiling of long non-coding RNA identifies linc-RoR as a prognostic biomarker in oral cancer. Tumour Biol. 2017, 39, 1010428317698366. [Google Scholar] [CrossRef] [PubMed]
- Rose, M.M.; Dhamodharan, S.; Bharath, G.; Murali, K.; Subbiah, S.; Bhaskar, L.V.; Murugan, A.K.; Munirajan, A.K. Linc-ROR genetic variants are associated with the advanced disease in oral squamous cell carcinoma. Arch. Oral Biol. 2022, 139, 105428. [Google Scholar] [CrossRef] [PubMed]
- Ma, X.; Zhang, H.; Li, Q.; Schiferle, E.; Qin, Y.; Xiao, S.; Li, T. FOXM1 Promotes Head and Neck Squamous Cell Carcinoma via Activation of the Linc-ROR/LMO4/AKT/PI3K Axis. Front. Oncol. 2021, 11, 658712. [Google Scholar] [CrossRef] [PubMed]
- Zhang, S.; Cai, J.; Ji, Y.; Zhou, S.; Miao, M.; Zhu, R.; Li, K.; Xue, Z.; Hu, S. Tumor-derived exosomal lincRNA ROR promotes angiogenesis in nasopharyngeal carcinoma. Mol. Cell. Probes 2022, 66, 101868. [Google Scholar] [CrossRef]
- Hong, F.; Gu, W.; Jiang, J.; Liu, X.; Jiang, H. Anticancer activity of polyphyllin I in nasopharyngeal carcinoma by modulation of lncRNA ROR and P53 signalling. J. Drug Target. 2019, 27, 806–811. [Google Scholar] [CrossRef] [PubMed]
- Li, L.; Gu, M.; You, B.; Shi, S.; Shan, Y.; Bao, L.; You, Y. Long non-coding RNA ROR promotes proliferation, migration and chemoresistance of nasopharyngeal carcinoma. Cancer Sci. 2016, 107, 1215–1222. [Google Scholar] [CrossRef] [PubMed]
- Wei, J.; Meng, X.; Wei, X.; Zhu, K.; Du, L.; Wang, H. Down-regulated lncRNA ROR in tumor-educated platelets as a liquid-biopsy biomarker for nasopharyngeal carcinoma. J. Cancer Res. Clin. Oncol. 2022. [Google Scholar] [CrossRef]
- Wang, L.; Yu, X.; Zhang, Z.; Pang, L.; Xu, J.; Jiang, J.; Liang, W.; Chai, Y.; Hou, J.; Li, F. Linc-ROR promotes esophageal squamous cell carcinoma progression through the derepression of SOX9. J. Exp. Clin. Cancer Res. 2017, 36, 182. [Google Scholar] [CrossRef] [PubMed]
- Sahebi, R.; Malakootian, M.; Balalaee, B.; Shahryari, A.; Khoshnia, M.; Abbaszadegan, M.R.; Moradi, A.; Javad Mowla, S. Linc-ROR and its spliced variants 2 and 4 are significantly up-regulated in esophageal squamous cell carcinoma. Iran. J. Basic Med. Sci. 2016, 19, 1131–1135. [Google Scholar] [PubMed]
- Gao, H.; Wang, T.; Zhang, P.; Shang, M.; Gao, Z.; Yang, F.; Liu, R. Linc-ROR regulates apoptosis in esophageal squamous cell carcinoma via modulation of p53 ubiquitination by targeting miR-204-5p/MDM2. J. Cell. Physiol. 2020, 235, 2325–2335. [Google Scholar] [CrossRef] [PubMed]
- Shang, M.; Wang, X.; Zhang, Y.; Gao, Z.; Wang, T.; Liu, R. LincRNA-ROR promotes metastasis and invasion of esophageal squamous cell carcinoma by regulating miR-145/FSCN1. OncoTargets Ther. 2018, 11, 639–649. [Google Scholar] [CrossRef] [PubMed]
- Su, X.; Feng, X.; Gao, C.; Wang, G.; Liu, L. ROR promotes the proliferation and migration of esophageal cancer through regulating miR-145/LMNB2 signal axis. Am. J. Transl. Res. 2020, 12, 7223–7235. [Google Scholar] [PubMed]
- Yan, Z.Y.; Sun, X.C. LincRNA-ROR functions as a ceRNA to regulate Oct4, Sox2, and Nanog expression by sponging miR-145 and its effect on biologic characteristics of colonic cancer stem cells. Zhonghua Bing Li Xue Za Zhi 2018, 47, 284–290. [Google Scholar] [CrossRef]
- Shaalan, A.A.M.; Mokhtar, S.H.; Ahmedah, H.T.; Almars, A.I.; Toraih, E.A.; Ibrahiem, A.T.; Fawzy, M.S.; Salem, M.A. Prognostic Value of LINC-ROR (rs1942347) Variant in Patients with Colon Cancer Harboring BRAF Mutation: A Propensity Score-Matched Analysis. Biomolecules 2022, 12, 569. [Google Scholar] [CrossRef]
- Yang, P.; Yang, Y.; An, W.; Xu, J.; Zhang, G.; Jie, J.; Zhang, Q. The long noncoding RNA-ROR promotes the resistance of radiotherapy for human colorectal cancer cells by targeting the p53/miR-145 pathway. J. Gastroenterol. Hepatol. 2017, 32, 837–845. [Google Scholar] [CrossRef]
- Zhou, P.; Sun, L.; Liu, D.; Liu, C.; Sun, L. Long Non-Coding RNA lincRNA-ROR Promotes the Progression of Colon Cancer and Holds Prognostic Value by Associating with miR-145. Pathol. Oncol. Res. 2016, 22, 733–740. [Google Scholar] [CrossRef]
- Chen, Y.; Yang, L.; Yin, D.; Feng, X.; Jie, J.; Yao, D.; Chen, J. Role of Long Noncoding RNA Regulator of Reprogramming in Colon Cancer Progression via Epidermal Growth Factor Receptor Signaling. Technol. Cancer Res. Treat. 2022, 21, 15330338221114707. [Google Scholar] [CrossRef]
- Li, X.; Chen, W.; Jia, J.; You, Z.; Hu, C.; Zhuang, Y.; Lin, Z.; Liu, Y.; Yang, C.; Xu, R. The Long Non-Coding RNA-RoR Promotes the Tumorigenesis of Human Colorectal Cancer by Targeting miR-6833-3p Through SMC4. OncoTargets Ther. 2020, 13, 2573–2581. [Google Scholar] [CrossRef]
- Li, H.; Jiang, X.; Niu, X. Long Non-Coding RNA Reprogramming (ROR) Promotes Cell Proliferation in Colorectal Cancer via Affecting P53. Med. Sci. Monit. 2017, 23, 919–928. [Google Scholar] [CrossRef]
- Chaleshi, V.; Irani, S.; Alebouyeh, M.; Mirfakhraie, R.; Aghdaei, H.A. Association of lncRNA-p53 regulatory network (lincRNA-p21, lincRNA-ROR and MALAT1) and p53 with the clinicopathological features of colorectal primary lesions and tumors. Oncol. Lett. 2020, 19, 3937–3949. [Google Scholar] [CrossRef] [PubMed]
- Thiele, J.A.; Hosek, P.; Kralovcova, E.; Ostasov, P.; Liska, V.; Bruha, J.; Vycital, O.; Rosendorf, J.; Opattova, A.; Horak, J.; et al. lncRNAs in Non-Malignant Tissue Have Prognostic Value in Colorectal Cancer. Int. J. Mol. Sci. 2018, 19, 2672. [Google Scholar] [CrossRef] [PubMed]
- Fawzy, M.S.; Toraih, E.A.; El-Wazir, A.; Hosny, M.M.; Badran, D.I.; El Kelish, A. Long intergenic non-coding RNA, regulator of reprogramming (LINC-ROR) over-expression predicts poor prognosis in renal cell carcinoma. Arch. Med. Sci. 2021, 17, 1016–1027. [Google Scholar] [CrossRef] [PubMed]
- Chen, Y.; Peng, Y.; Xu, Z.; Ge, B.; Xiang, X.; Zhang, T.; Gao, L.; Shi, H.; Wang, C.; Huang, J. LncROR Promotes Bladder Cancer Cell Proliferation, Migration, and Epithelial-Mesenchymal Transition. Cell. Physiol. Biochem. 2017, 41, 2399–2410. [Google Scholar] [CrossRef]
- Shi, J.; Zhang, W.; Tian, H.; Zhang, Q.; Men, T. lncRNA ROR promotes the proliferation of renal cancer and is negatively associated with favorable prognosis. Mol. Med. Rep. 2017, 16, 9561–9566. [Google Scholar] [CrossRef] [PubMed]
- Shi, J.; Zhang, D.; Zhong, Z.; Zhang, W. lncRNA ROR promotes the progression of renal cell carcinoma through the miR-206/VEGF axis. Mol. Med. Rep. 2019, 20, 3782–3792. [Google Scholar] [CrossRef] [PubMed]
- Fan, J.; Xing, Y.; Wen, X.; Jia, R.; Ni, H.; He, J.; Ding, X.; Pan, H.; Qian, G.; Ge, S.; et al. Long non-coding RNA ROR decoys gene-specific histone methylation to promote tumorigenesis. Genome Biol. 2015, 16, 139. [Google Scholar] [CrossRef]
- Fan, Y.; Fan, X.; Yan, H.; Liu, Z.; Wang, X.; Yuan, Q.; Xie, J.; Lu, X.; Yang, Y. Long non-coding ROR promotes the progression of papillary thyroid carcinoma through regulation of the TESC/ALDH1A1/TUBB3/PTEN axis. Cell Death Dis. 2022, 13, 157. [Google Scholar] [CrossRef]
- Zhang, R.; Hardin, H.; Huang, W.; Buehler, D.; Lloyd, R.V. Long Non-coding RNA Linc-ROR Is Upregulated in Papillary Thyroid Carcinoma. Endocr. Pathol. 2018, 29, 1–8. [Google Scholar] [CrossRef]
- Hardin, H.; Helein, H.; Meyer, K.; Robertson, S.; Zhang, R.; Zhong, W.; Lloyd, R.V. Thyroid cancer stem-like cell exosomes: Regulation of EMT via transfer of lncRNAs. Lab. Investig. 2018, 98, 1133–1142. [Google Scholar] [CrossRef] [PubMed]
- Yu, Q.; Hardin, H.; Chu, Y.H.; Rehrauer, W.; Lloyd, R.V. Parathyroid Neoplasms: Immunohistochemical Characterization and Long Noncoding RNA (lncRNA) Expression. Endocr. Pathol. 2019, 30, 96–105. [Google Scholar] [CrossRef] [PubMed]
- Toraih, E.A.; El-Wazir, A.; Hussein, M.H.; Khashana, M.S.; Matter, A.; Fawzy, M.S.; Hosny, S. Expression of long intergenic non-coding RNA, regulator of reprogramming, and its prognostic value in patients with glioblastoma. Int. J. Biol. Markers 2019, 34, 69–79. [Google Scholar] [CrossRef] [PubMed]
- Kovalenko, T.F.; Yadav, B.; Anufrieva, K.S.; Rubtsov, Y.P.; Zatsepin, T.S.; Shcherbinina, E.Y.; Solyus, E.M.; Staroverov, D.B.; Larionova, T.D.; Latyshev, Y.A.; et al. Functions of long non-coding RNA ROR in patient-derived glioblastoma cells. Biochimie 2022, 200, 131–139. [Google Scholar] [CrossRef] [PubMed]
- Ruiz Esparza-Garrido, R.; Rodríguez-Corona, J.M.; López-Aguilar, J.E.; Rodríguez-Florido, M.A.; Velázquez-Wong, A.C.; Viedma-Rodríguez, R.; Salamanca-Gómez, F.; Velázquez-Flores, M. Differentially Expressed Long Non-Coding RNAs Were Predicted to Be Involved in the Control of Signaling Pathways in Pediatric Astrocytoma. Mol. Neurobiol. 2017, 54, 6598–6608. [Google Scholar] [CrossRef]
- Li, Y.; He, Z.C.; Liu, Q.; Zhou, K.; Shi, Y.; Yao, X.H.; Zhang, X.; Kung, H.F.; Ping, Y.F.; Bian, X.W. Large Intergenic Non-coding RNA-RoR Inhibits Aerobic Glycolysis of Glioblastoma Cells via Akt Pathway. J. Cancer 2018, 9, 880–889. [Google Scholar] [CrossRef]
- Gao, Y.; Luo, X.; Zhang, J. LincRNA-ROR is activated by H3K27 acetylation and induces EMT in retinoblastoma by acting as a sponge of miR-32 to activate the Notch signaling pathway. Cancer Gene Ther. 2021, 28, 42–54. [Google Scholar] [CrossRef]
- Cheng, F.H.; Zhao, Z.S.; Liu, W.D. Long non-coding RNA ROR regulated ABCB1 to induce cisplatin resistance in osteosarcoma by sponging miR-153-3p. Eur. Rev. Med. Pharmacol. Sci. 2019, 23, 7256–7265. [Google Scholar] [CrossRef]
- Wang, W.; Li, Y.; Zhi, S.; Li, J.; Miao, J.; Ding, Z.; Peng, Y.; Huang, Y.; Zheng, R.; Yu, H.; et al. LncRNA-ROR/microRNA-185-3p/YAP1 axis exerts function in biological characteristics of osteosarcoma cells. Genomics 2021, 113, 450–461. [Google Scholar] [CrossRef]
- Fei, D.; Sui, G.; Lu, Y.; Tan, L.; Dongxu, Z.; Zhang, K. The long non-coding RNA-ROR promotes osteosarcoma progression by targeting miR-206. J. Cell. Mol. Med. 2019, 23, 1865–1872. [Google Scholar] [CrossRef]
- Li, X.; Zuo, C.; Wu, M.; Zhang, Z. Linc-ROR promotes arsenite-transformed keratinocyte proliferation by inhibiting P53 activity. Metallomics 2020, 12, 963–973. [Google Scholar] [CrossRef] [PubMed]
- Liu, T.; Chi, H.; Chen, J.; Chen, C.; Huang, Y.; Xi, H.; Xue, J.; Si, Y. Curcumin suppresses proliferation and in vitro invasion of human prostate cancer stem cells by ceRNA effect of miR-145 and lncRNA-ROR. Gene 2017, 631, 29–38. [Google Scholar] [CrossRef] [PubMed]
- Institute, N.C. What is Cancer? Available online: https://www.cancer.gov/about-cancer/understanding/what-is-cancer (accessed on 13 November 2022).
- Mathur, G.; Nain, S.; Sharma, P.K. Cancer: An overview. Acad. J. Cancer Res. 2015, 8, 1–9. [Google Scholar]
- Compton, C. Cancer: The Enemy from Within: A Comprehensive Textbook of Cancer’s Causes, Complexities and Consequences; Springer International Publishing: Berlin/Heidelberg, Germany, 2020. [Google Scholar]
- Hejmadi, M. Introduction to Cancer Biology; BoonBooks: Haywards Heath, UK, 2014. [Google Scholar]
- Kontomanolis, E.N.; Koutras, A.; Syllaios, A.; Schizas, D.; Mastoraki, A.; Garmpis, N.; Diakosavvas, M.; Angelou, K.; Tsatsaris, G.; Pagkalos, A.; et al. Role of Oncogenes and Tumor-suppressor Genes in Carcinogenesis: A Review. Anticancer Res. 2020, 40, 6009–6015. [Google Scholar] [CrossRef] [PubMed]
- Tao, F.; Xu, Y.; Yang, D.; Tian, B.; Jia, Y.; Hou, J.; Dong, M. LncRNA NKILA correlates with the malignant status and serves as a tumor-suppressive role in rectal cancer. J. Cell. Biochem. 2018, 119, 9809–9816. [Google Scholar] [CrossRef] [PubMed]
- Wei, G.H.; Wang, X. lncRNA MEG3 inhibit proliferation and metastasis of gastric cancer via p53 signaling pathway. Eur. Rev. Med. Pharmacol. Sci. 2017, 21, 3850–3856. [Google Scholar]
- Yang, X.; Xie, Z.; Lei, X.; Gan, R. Long non-coding RNA GAS5 in human cancer. Oncol. Lett. 2020, 20, 2587–2594. [Google Scholar] [CrossRef]
- Kadkhoda, S.; Ghafouri-Fard, S. Function of miRNA-145-5p in the pathogenesis of human disorders. Pathol. Res. Pract. 2022, 231, 153780. [Google Scholar] [CrossRef]
- Dongre, A.; Weinberg, R.A. New insights into the mechanisms of epithelial-mesenchymal transition and implications for cancer. Nat. Rev. Mol. Cell Biol. 2019, 20, 69–84. [Google Scholar] [CrossRef]
- Jayachandran, J.; Srinivasan, H.; Mani, K.P. Molecular mechanism involved in epithelial to mesenchymal transition. Arch. Biochem. Biophys. 2021, 710, 108984. [Google Scholar] [CrossRef]
- Lu, W.; Kang, Y. Epithelial-Mesenchymal Plasticity in Cancer Progression and Metastasis. Dev. Cell 2019, 49, 361–374. [Google Scholar] [CrossRef]
- Lee, J.; You, J.H.; Kim, M.S.; Roh, J.L. Epigenetic reprogramming of epithelial-mesenchymal transition promotes ferroptosis of head and neck cancer. Redox Biol. 2020, 37, 101697. [Google Scholar] [CrossRef] [PubMed]
- Zhang, N.; Ng, A.S.; Cai, S.; Li, Q.; Yang, L.; Kerr, D. Novel therapeutic strategies: Targeting epithelial-mesenchymal transition in colorectal cancer. Lancet Oncol. 2021, 22, e358–e368. [Google Scholar] [CrossRef] [PubMed]
- Gregory, P.A.; Bert, A.G.; Paterson, E.L.; Barry, S.C.; Tsykin, A.; Farshid, G.; Vadas, M.A.; Khew-Goodall, Y.; Goodall, G.J. The miR-200 family and miR-205 regulate epithelial to mesenchymal transition by targeting ZEB1 and SIP1. Nat. Cell Biol. 2008, 10, 593–601. [Google Scholar] [CrossRef] [PubMed]
- Zhang, Y.; Wang, X. Targeting the Wnt/β-catenin signaling pathway in cancer. J. Hematol. Oncol. 2020, 13, 165. [Google Scholar] [CrossRef] [PubMed]
- Wang, S.H.; Zhang, M.D.; Wu, X.C.; Weng, M.Z.; Zhou, D.; Quan, Z.W. Overexpression of LncRNA-ROR predicts a poor outcome in gallbladder cancer patients and promotes the tumor cells proliferation, migration, and invasion. Tumour Biol. 2016, 37, 12867–12875. [Google Scholar] [CrossRef]
- Roden, D.M.; McLeod, H.L.; Relling, M.V.; Williams, M.S.; Mensah, G.A.; Peterson, J.F.; Van Driest, S.L. Pharmacogenomics. Lancet 2019, 394, 521–532. [Google Scholar] [CrossRef]
- Wang, C.; Ding, T.; Yang, D.; Zhang, P.; Hu, X.; Qin, W.; Zheng, J. The lncRNA OGFRP1/miR-149-5p/IL-6 axis regulates prostate cancer chemoresistance. Pathol. Res. Pract. 2021, 224, 153535. [Google Scholar] [CrossRef]
- Xia, C.; Sun, Y.; Li, Y.; Ma, J.; Shi, J. LncRNA CCAT1 enhances chemoresistance in hepatocellular carcinoma by targeting QKI-5. Sci. Rep. 2022, 12, 7826. [Google Scholar] [CrossRef]
- Yang, J.; Yang, M.; Lv, H.; Zhou, M.; Mao, X.; Qin, X.; Xu, Y.; Li, L.; Xing, H. lncRNA SNHG15 Induced by SOX12 Promotes the Tumorigenic Properties and Chemoresistance in Cervical Cancer via the miR-4735-3p/HIF1a Pathway. Oxid. Med. Cell. Longev. 2022, 2022, 8548461. [Google Scholar] [CrossRef]
- Zhang, F.; Wang, H.; Yu, J.; Yao, X.; Yang, S.; Li, W.; Xu, L.; Zhao, L. LncRNA CRNDE attenuates chemoresistance in gastric cancer via SRSF6-regulated alternative splicing of PICALM. Mol. Cancer 2021, 20, 6. [Google Scholar] [CrossRef] [PubMed]
- Wang, H.; Hang, C.; Ou, X.L.; Nie, J.S.; Ding, Y.T.; Xue, S.G.; Gao, H.; Zhu, J.X. MiR-145 functions as a tumor suppressor via regulating angiopoietin-2 in pancreatic cancer cells. Cancer Cell Int. 2016, 16, 65. [Google Scholar] [CrossRef] [PubMed]
- Zhang, W.; Ji, W.; Li, T.; Liu, T.; Zhao, X. MiR-145 functions as a tumor suppressor in Papillary Thyroid Cancer by inhibiting RAB5C. Int. J. Med. Sci. 2020, 17, 1992–2001. [Google Scholar] [CrossRef]
- Li, S.; Yue, X.C.; Sun, C.Y.; Qin, H.Y.; Zhang, X.Y. Prognostic value of long noncoding RNA ROR in patients with cancer in China: A systematic review and meta-analysis. Medicine 2019, 98, e15758. [Google Scholar] [CrossRef] [PubMed]
- Lu, R.; Chen, J.; Kong, L.; Zhu, H. Prognostic value of lncRNA ROR expression in various cancers: A meta-analysis. Biosci. Rep. 2018, 38, BSR20181095. [Google Scholar] [CrossRef] [PubMed]
- Arantes, L.; De Carvalho, A.C.; Melendez, M.E.; Lopes Carvalho, A. Serum, plasma and saliva biomarkers for head and neck cancer. Expert Rev. Mol. Diagn. 2018, 18, 85–112. [Google Scholar] [CrossRef]
- Müller, V.; Oliveira-Ferrer, L.; Steinbach, B.; Pantel, K.; Schwarzenbach, H. Interplay of lncRNA H19/miR-675 and lncRNA NEAT1/miR-204 in breast cancer. Mol. Oncol. 2019, 13, 1137–1149. [Google Scholar] [CrossRef]
- Nagasaka, M.; Uddin, M.H.; Al-Hallak, M.N.; Rahman, S.; Balasubramanian, S.; Sukari, A.; Azmi, A.S. Liquid biopsy for therapy monitoring in early-stage non-small cell lung cancer. Mol. Cancer 2021, 20, 82. [Google Scholar] [CrossRef]
- Wang, Y.; Guo, Z.; Zhao, Y.; Jin, Y.; An, L.; Wu, B.; Liu, Z.; Chen, X.; Chen, X.; Zhou, H.; et al. Genetic polymorphisms of lncRNA-p53 regulatory network genes are associated with concurrent chemoradiotherapy toxicities and efficacy in nasopharyngeal carcinoma patients. Sci. Rep. 2017, 7, 8320. [Google Scholar] [CrossRef]

| Type of Cancer | Target/Relation | Effect | Reference |
|---|---|---|---|
| Breast | MAPK/ERK pathway | Promotes estrogen-independent proliferation | [89] |
| miR-194-3p | Promotes rapamycin resistance | [70] | |
| N- and E-cadherin, vimentin | Promotes 5-FU and paclitaxel resistance and EMT | [90] | |
| miR-205, ZEB1, ZEB2 | Promotes tamoxifen resistance and EMT | [69] | |
| LC2, Beclin 1 | Promotes tamoxifen resistance by autophagy | [91] | |
| miR-205, ZEB2 | Promotes EMT | [92] | |
| Estrogen and progesterone receptors | Promotes lymph node metastasis | [93,94] | |
| Reproductive factors | Higher risk | [95] | |
| hnRNPI, AUF1 | Promotes proliferation and tumorigenesis | [96] | |
| MLL1/H3K4/TIMP3 | Promotes progression | [97] | |
| CTBP1-AS2, SPRY4-IT1 | Promotes pathogenesis | [98] | |
| miR-145/ARF6 | Promotes metastasis and invasion | [99] | |
| miR-145/MUC1/E-cadherin | Promotes metastasis and invasion | [100] | |
| TGF-β pathway | Promotes proliferation and invasion | [101] | |
| miR-34a | Promotes autophagy and gemcitabine resistance | [66] | |
| Wnt/β-catenin pathway | Promotes viability, migration, and invasion | [102] | |
| ND | Promotes metastasis | [103] | |
| Ovarian and Endometrial | Wnt/β-catenin pathway | Promotes EMT and metastasis | [104] |
| ND | Promotes proliferation, invasion, and metastasis | [105] | |
| CA125 | Promotes lymph node metastasis | [106] | |
| miR-145/FLNB | Promotes EMT and invasion | [107] | |
| miR-145 | Promotes stemness | [108] | |
| miR-34a, Notch | Promotes proliferation and suppresses apoptosis | [109] | |
| Gastric | SALL4 | Promotes maintenance and aggressiveness | [110] |
| miR-519d-3p/HMGA2 | Promotes proliferation, EMT and cisplatin resistance | [111] | |
| miR-212-3p/FGF7 | Promotes proliferation, migration, and invasion | [112] | |
| Vimentin, E-cadherin, β-catenin, c-Myc | Promotes EMT and lymph node metastasis | [113] | |
| OCT4, SOX2, NANOG, CD133 | Promotes proliferation and invasion | [114] | |
| MRPI | Promotes Adriamycin and vincristine resistance | [115] | |
| ADAR, FUS | Increased survival rate | [86] | |
| HOXA-AS1 | Downregulated | [116] | |
| miR-580-3p/ANXA10 | Suppresses proliferation, migration, and invasion | [117] | |
| Lung | miR-145/FSCN1 | Promotes docetaxel resistance | [118] |
| NA | Promotes distant and lymph node metastasis | [119] | |
| EML4-ALK | Promotes stemness and crizotinib resistance | [120] | |
| P53/miR-145 | Promotes proliferation, migration, and invasion | [121] | |
| PI3K/Akt/mTOR | Suppresses cisplatin resistance | [88] | |
| Liver | miR-145/ZEB2 | Promotes EMT and metastasis | [25] |
| miR-145/RAD18 | Promotes radioresistance | [122] | |
| IL-1β | Promotes release of pro-inflammatory cytokines | [123] | |
| E-cadherin, vimentin, TWIST1 | Promotes EMT and Adriamycin resistance | [124] | |
| DEPCD1 | Promotes progression and angiogenesis | [125] | |
| miR-876-5p/FOXM1 | Promotes sorafenib resistance | [126] | |
| TGF-β | Promotes sorafenib resistance | [127] | |
| miR-145/HIF-1α | Promotes survival during hypoxic stress | [128] | |
| P53 | Promotes arsenic trioxide resistance | [129] | |
| miR-223-3p/NF2 | Promotes proliferation and invasion | [130] | |
| OCT4, NANOG, SOX2, p53, CD133 | Promotes proliferation | [131] | |
| Pancreatic | ZEB1 | Promotes EMT and aggressiveness | [132] |
| Hippo/YAP pathway | Promotes EMT, proliferation, and invasion | [24] | |
| HIF1-α/ZEB1 | Promotes EMT | [133] | |
| miR-145, NANOG | Promotes proliferation and decreases migration | [68] | |
| Let-7 family | Promotes migration, invasion, and EMT | [134] | |
| Head and Neck | miR-145-5p | Promotes stemness | [135] |
| ND | Promotes progression and metastasis | [136] | |
| LMO4/AKT/PI3K | Promotes proliferation and invasion | [137] | |
| p-AKT/p-VEGFR2 | Promotes proliferation, migration, and angiogenesis | [138] | |
| P53 | Promotes proliferation, metastasis and inhibits apoptosis | [139,140] | |
| ND | Downregulated in plasma | [141] | |
| P53 | |||
| Esophageal | miR-15b, miR33a, miR-129, miR-145, miR-206 | Promotes proliferation, motility, chemoresistance, and renewal capacity | [142] |
| ND | Promotes initiation and progression | [143] | |
| miR-204-5p/MDM2/p53 | Suppresses apoptosis | [144] | |
| miR-145/FSCNI | Promotes metastasis | [145] | |
| miR-145/LMNB2 | Promotes proliferation and migration | [146] | |
| Colorectal | miR-145 | Promotes stemness and metastasis | [147] |
| hnRNPI, AUF1 | Promotes proliferation and tumorigenesis | [96] | |
| NA | Related with larger tumor size, metastasis, and mortality | [148] | |
| P53/miR-145 | Promotes radioresistance and suppresses apoptosis | [149] | |
| miR-145 | Promotes lower survival rate | [150] | |
| EGFR | Promotes proliferation invasion, and migration | [151] | |
| miR-6833-3p/SMC4 | Promotes proliferation and lower survival rate | [152] | |
| P53 | Promotes proliferation and viability | [153,154] | |
| CCAT1 | Promotes metastasis | [155] | |
| Kidney and Bladder | SOX2, Nanog, POU5F1 | Promotes stemness, infiltration and shorter survival | [156] |
| ZEB1 | Promotes proliferation, metastasis, EMT, and inhibits apoptosis | [157] | |
| P53, c-Myc | Promotes shorter survival rate | [158] | |
| miR-206/VEGF | Promotes proliferation and metastasis | [159] | |
| TESC | Promotes tumorigenesis | [160] | |
| Thyroid and Parathyroid | TESC/ALDH1A1/ TUBB3/PTEN | Promotes progression | [161] |
| miR-145 | Promotes EMT | [162] | |
| ND | Promotes EMT and metastasis | [163] | |
| ND | Suppresses progression | [164] | |
| Brain and Retina | ND | Promotes poor overall survival | [165] |
| EGFR | Promotes proliferation and stemness | [166] | |
| KLF4/CD133 | Suppresses proliferation | [87] | |
| ND | Promotes proliferation and angiogenesis | [167] | |
| Akt pathway | Suppresses proliferation | [168] | |
| miR-32-5p/Notch | Promotes EMT, invasion and metastasis | [169] | |
| Bone | miR-153-3p/ABCB1 | Promotes cisplatin resistance | [170] |
| miR-185-3p/YAP1 | Promotes growth and metastasis | [171] | |
| miR-206 | Relates to advanced TNM, metastasis and poor survival | [172] | |
| Skin | P53, PI3K/Akt | Promotes proliferation | [173] |
| Prostate | miR-145/Oct4 | Promotes proliferation, invasion, and tumorigenicity | [174] |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2023 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Peña-Flores, J.A.; Enríquez-Espinoza, D.; Muela-Campos, D.; Álvarez-Ramírez, A.; Sáenz, A.; Barraza-Gómez, A.A.; Bravo, K.; Estrada-Macías, M.E.; González-Alvarado, K. Functional Relevance of the Long Intergenic Non-Coding RNA Regulator of Reprogramming (Linc-ROR) in Cancer Proliferation, Metastasis, and Drug Resistance. Non-Coding RNA 2023, 9, 12. https://doi.org/10.3390/ncrna9010012
Peña-Flores JA, Enríquez-Espinoza D, Muela-Campos D, Álvarez-Ramírez A, Sáenz A, Barraza-Gómez AA, Bravo K, Estrada-Macías ME, González-Alvarado K. Functional Relevance of the Long Intergenic Non-Coding RNA Regulator of Reprogramming (Linc-ROR) in Cancer Proliferation, Metastasis, and Drug Resistance. Non-Coding RNA. 2023; 9(1):12. https://doi.org/10.3390/ncrna9010012
Chicago/Turabian StylePeña-Flores, José A., Diego Enríquez-Espinoza, Daniela Muela-Campos, Alexis Álvarez-Ramírez, Angel Sáenz, Andrés A. Barraza-Gómez, Kenia Bravo, Marvin E. Estrada-Macías, and Karla González-Alvarado. 2023. "Functional Relevance of the Long Intergenic Non-Coding RNA Regulator of Reprogramming (Linc-ROR) in Cancer Proliferation, Metastasis, and Drug Resistance" Non-Coding RNA 9, no. 1: 12. https://doi.org/10.3390/ncrna9010012
APA StylePeña-Flores, J. A., Enríquez-Espinoza, D., Muela-Campos, D., Álvarez-Ramírez, A., Sáenz, A., Barraza-Gómez, A. A., Bravo, K., Estrada-Macías, M. E., & González-Alvarado, K. (2023). Functional Relevance of the Long Intergenic Non-Coding RNA Regulator of Reprogramming (Linc-ROR) in Cancer Proliferation, Metastasis, and Drug Resistance. Non-Coding RNA, 9(1), 12. https://doi.org/10.3390/ncrna9010012







