FuncPEP: A Database of Functional Peptides Encoded by Non-Coding RNAs
Abstract
1. Introduction
2. Data Collection and Computational Methods
2.1. Data Collection and Database Construction
- (i)
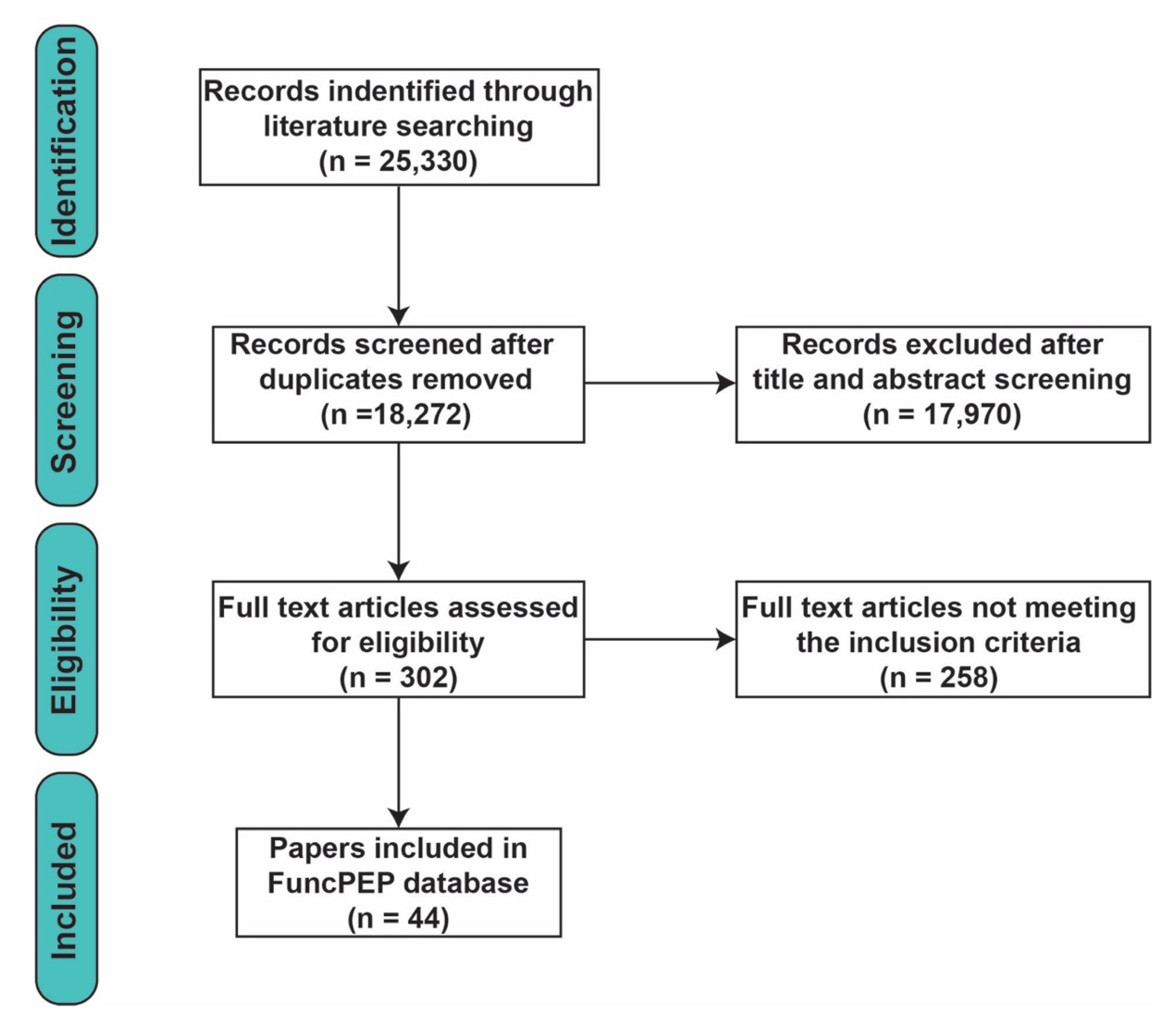
- We included only ncPEPs that were directly (MS, Western blotting, immunohistochemistry, immunofluorescence) or indirectly (ribosome profiling, loss/gain of function studies) experimentally confirmed;
- (ii)
- All included ncPEPs were not merely experimentally confirmed but also functionally characterized (linked to a physiological or a pathological process); and
- (iii)
- For concordance with the definitions discussed above, we considered ncPEPs of only ≤ 100 aa in length.
2.2. Sequence and Molecular Weight Prediction
2.3. Comparison of ncPEPs with Known Peptides Encoded by Messenger RNAs of Coding Genes
3. Implementation and Results
3.1. Systematic Review
3.2. Database Interface
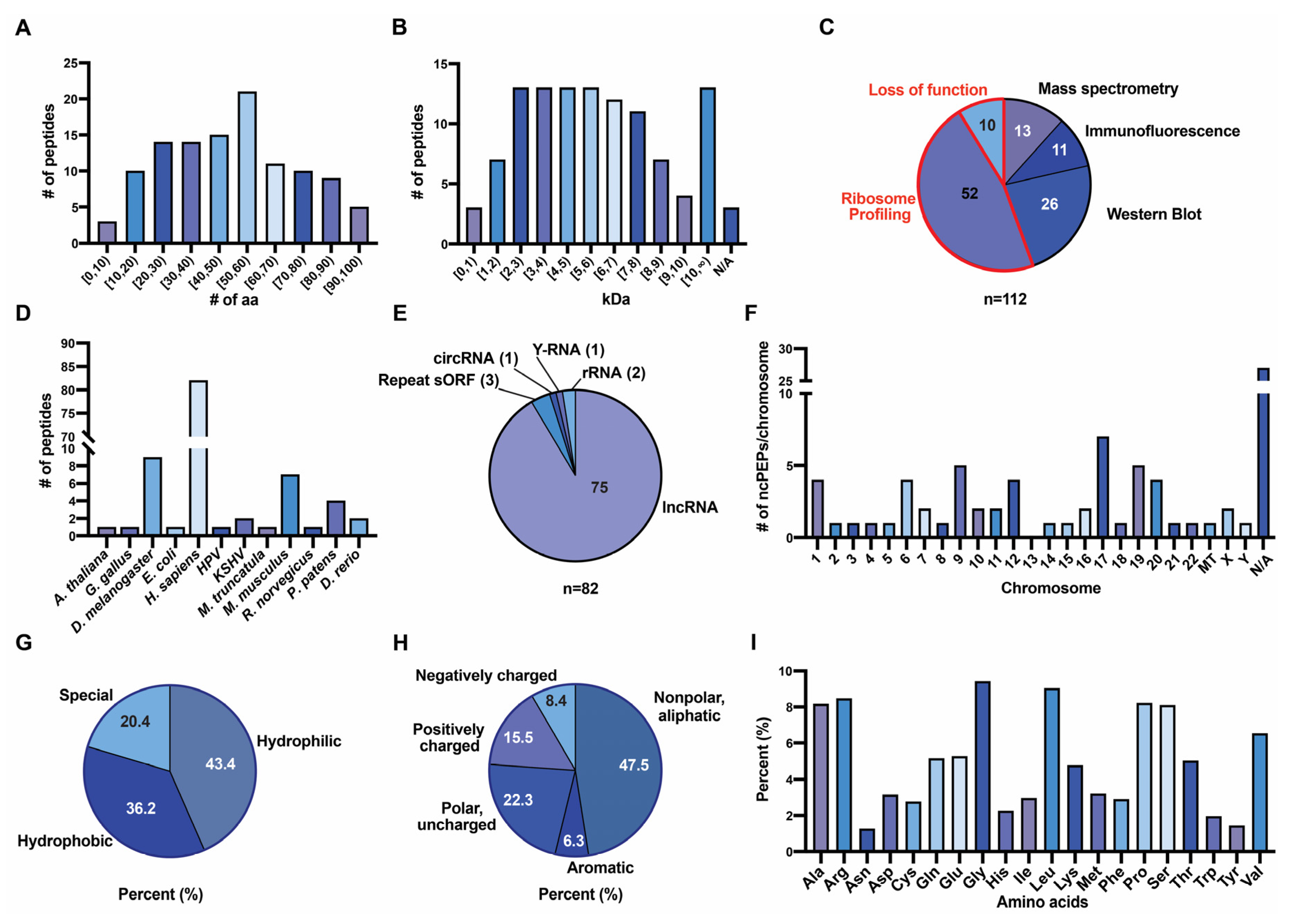
3.3. Characteristics of ncPEPs from the FuncPEP Database
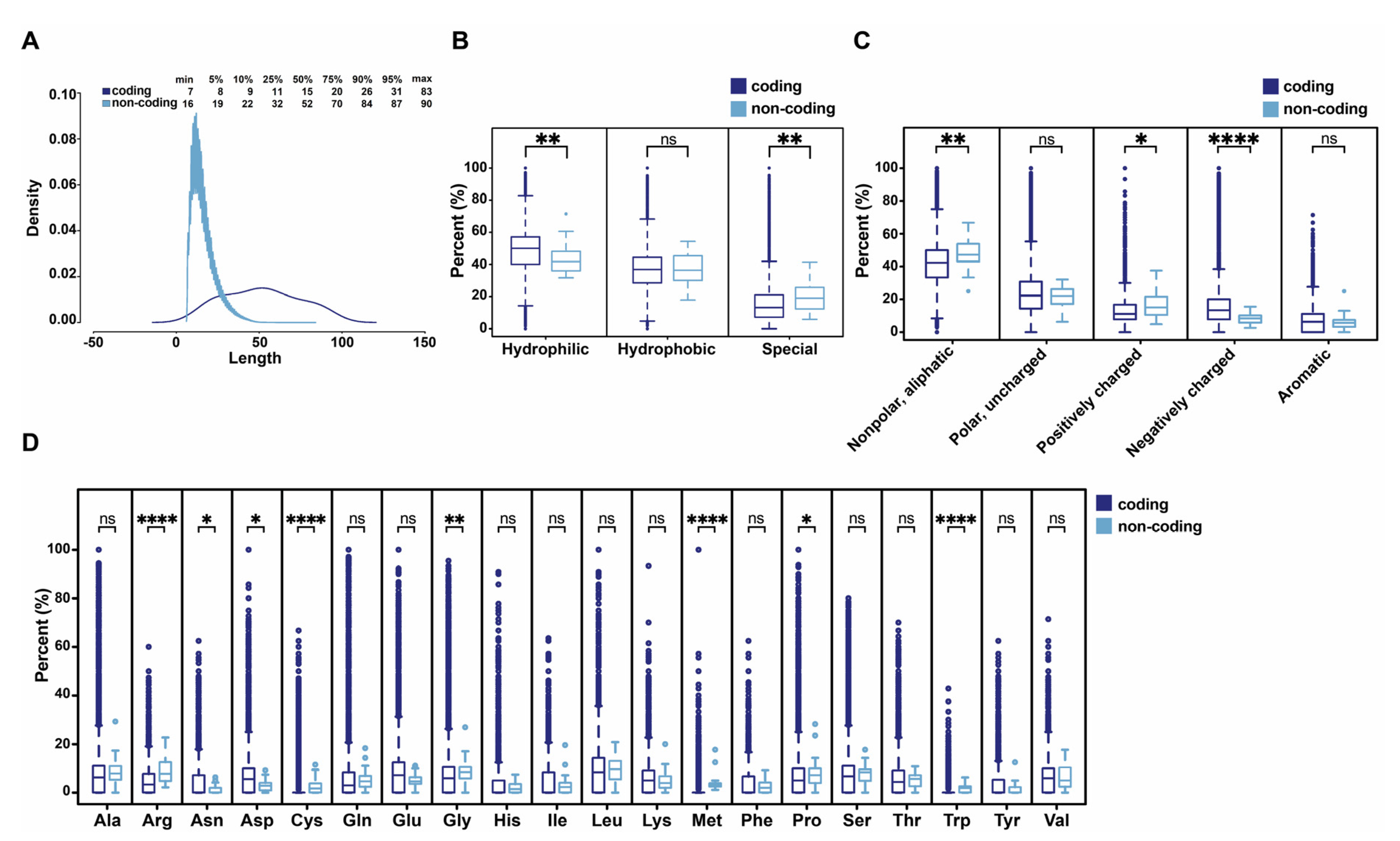
3.4. Comparison of Human Functional ncPEPs with Human Peptides from Coding Regions
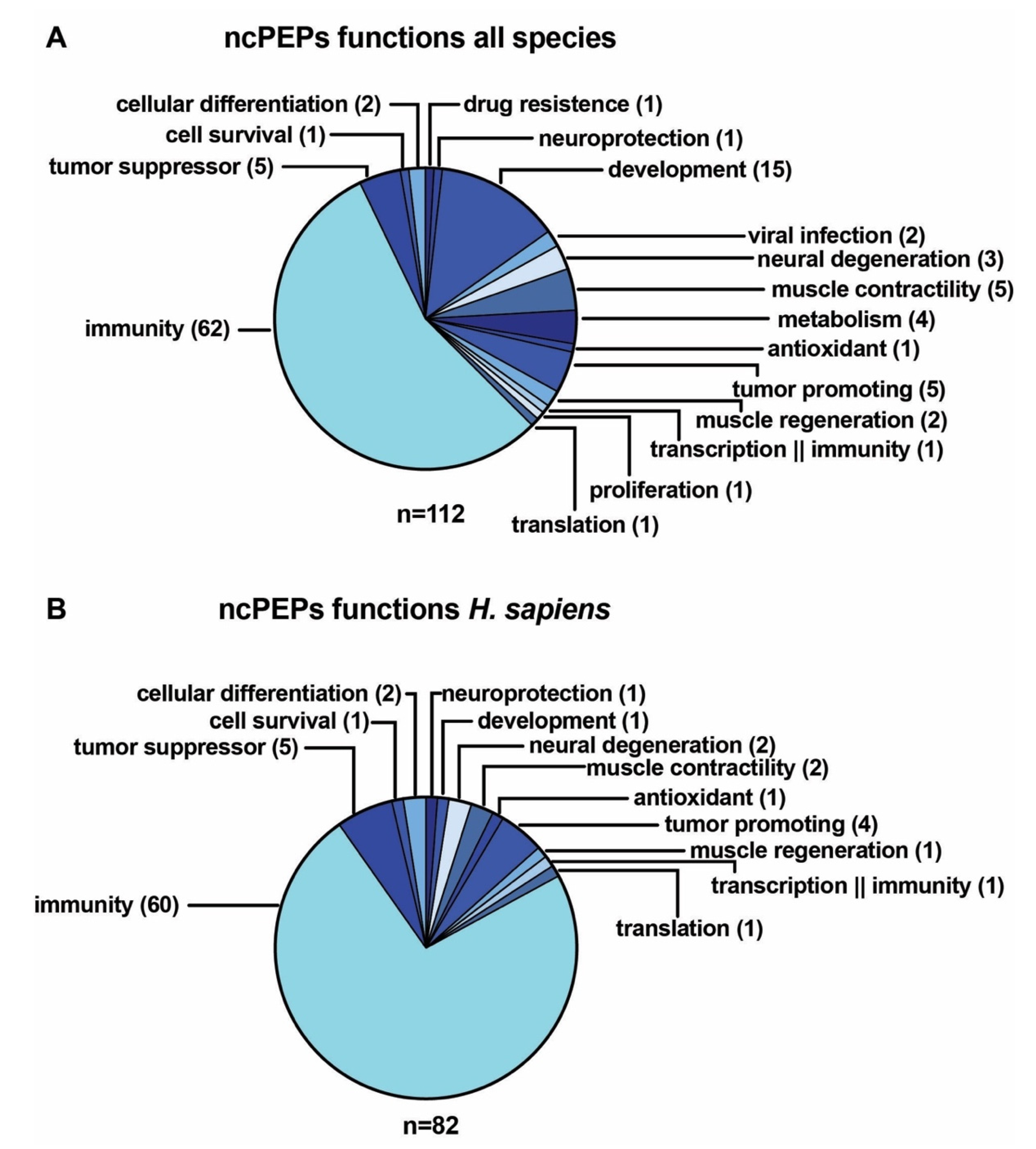
3.5. Examples of ncPEPs Functions
4. Discussion
5. Conclusions
Author Contributions
Funding
Conflicts of Interest
Availability
References
- The FANTOM consortium; Carninci, P.; Kasukawa, T.; Katayama, S.; Gough, J.; Frith, M.; Maeda, N.; Oyama, R.; Ravasi, T.; Lenhard, B.; et al. The Transcriptional Landscape of the Mammalian Genome. Science 2005, 309, 1559–1563. [Google Scholar] [CrossRef]
- Mudge, J.; Frankish, A.; Harrow, J. Functional Transcriptomics in the Post-ENCODE era. Genome Res. 2013, 23, 1961–1973. [Google Scholar] [CrossRef]
- Derrien, T.; Johnson, R.; Bussotti, G.; Tanzer, A.; Djebali, S.; Tilgner, H.; Guernec, G.; Martin, D.; Merkel, A.; Knowles, D.G.; et al. The GENCODE v7 Catalog of Human Long Noncoding RNAs: Analysis of Their Gene Structure, Evolution, and Expression. Genome Res. 2012, 22, 1775–1789. [Google Scholar] [CrossRef]
- Frankish, A.; Diekhans, M.; Ferreira, A.-M.; Johnson, R.; Jungreis, I.; Loveland, J.E.; Mudge, J.; Sisu, C.; Wright, J.C.; Armstrong, J.; et al. GENCODE Reference Annotation for the Human and Mouse Genomes. Nucleic Acids Res. 2018, 47, D766–D773. [Google Scholar] [CrossRef]
- Dinger, M.E.; Pang, K.C.; Mercer, T.R.; Mattick, J.S. Differentiating Protein-Coding and Noncoding RNA: Challenges and Ambiguities. PLoS Comput. Boil. 2008, 4, e1000176. [Google Scholar] [CrossRef] [PubMed]
- Slavoff, S.A.; Mitchell, A.J.; Schwaid, A.G.; Cabili, M.N.; Ma, J.; Levin, J.Z.; Karger, A.; Budnik, B.A.; Rinn, J.L.; Saghatelian, A. Peptidomic Discovery of Short Open Reading Frame–Encoded Peptides in Human Cells. Nat. Methods 2012, 9, 59–64. [Google Scholar] [CrossRef]
- Banfai, B.; Jia, H.; Khatun, J.; Wood, E.; Risk, B.; Gundling, W.E.; Kundaje, A.; Gunawardena, H.P.; Yu, Y.; Xie, L.; et al. Long Noncoding RNAs are Rarely Translated in Two Human Cell Lines. Genome Res. 2012, 22, 1646–1657. [Google Scholar] [CrossRef] [PubMed]
- Ruiz-Orera, J.; Messeguer, X.; Subirana, J.A.; Albà, M.M. Long Non-coding RNAs as a Source of New Peptides. eLife 2014, 3, e03523. [Google Scholar] [CrossRef] [PubMed]
- Makarewich, C.A.; Olson, E.N. Mining for Micropeptides. Trends Cell Boil. 2017, 27, 685–696. [Google Scholar] [CrossRef] [PubMed]
- Lin, M.F.; Jungreis, I.; Kellis, M. PhyloCSF: A Comparative Genomics Method to Distinguish Protein Coding and Non-coding Regions. Bioinformatics 2011, 27, i275–i282. [Google Scholar] [CrossRef] [PubMed]
- Aspden, J.; Eyre-Walker, Y.C.; Phillips, R.J.; Amin, U.; Mumtaz, M.A.S.; Brocard, M.; Couso, J.P. Extensive Translation of Small Open Reading Frames Revealed by Poly-Ribo-Seq. eLife 2014, 3, e03528. [Google Scholar] [CrossRef] [PubMed]
- Anderson, U.M.; Anderson, K.M.; Chang, C.-L.; Makarewich, C.A.; Nelson, B.R.; McAnally, J.R.; Kasaragod, P.; Shelton, J.M.; Liou, J.; Bassel-Duby, R.; et al. A Micropeptide Encoded by a Putative Long Noncoding RNA Regulates Muscle Performance. Cell 2015, 160, 595–606. [Google Scholar] [CrossRef] [PubMed]
- Nelson, B.R.; Makarewich, C.A.; Anderson, D.M.; Winders, B.R.; Troupes, C.D.; Wu, F.; Reese, A.L.; McAnally, J.R.; Chen, X.; Kavalali, E.T.; et al. A Peptide Encoded by a Transcript Annotated as Long Noncoding RNA Enhances SERCA Activity in Muscle. Science 2016, 351, 271–275. [Google Scholar] [CrossRef] [PubMed]
- Li, L.-J.; Leng, R.-X.; Fan, Y.-G.; Pan, H.-F.; Ye, D.-Q. Translation of Noncoding RNAs: Focus on lncRNAs, pri-miRNAs, and circRNAs. Exp. Cell Res. 2017, 361, 1–8. [Google Scholar] [CrossRef] [PubMed]
- Ramakrishnan, A.; Janga, S.C. Human Protein-RNA Interaction Network is Highly Stable Across Mammals. BMC Genom. 2019, 20, 1004–1014. [Google Scholar] [CrossRef]
- López-Bigas, N.; De, S.; Teichmann, S.A. Functional Protein Divergence in the Evolution of Homo Sapiens. Genome Boil. 2008, 9, R33. [Google Scholar] [CrossRef]
- NCBI. Open Reading Frame Finder. Available online: https://www.ncbi.nlm.nih.gov/orffinder/ (accessed on 1 April 2020).
- Jia, H.; Osak, M.; Bogu, G.K.; Stanton, L.W.; Johnson, R.; Lipovich, L. Genome-Wide Computational Identification and Manual Annotation of Human Long Noncoding RNA Genes. RNA 2010, 16, 1478–1487. [Google Scholar] [CrossRef]
- Esteller, M. Non-coding RNAs in Human Disease. Nat. Rev. Genet. 2011, 12, 861–874. [Google Scholar] [CrossRef]
- Hombach, S.; Kretz, M. Non-coding RNAs: Classification, Biology and Functioning. Adv. Exp. Med. Biol. 2016, 937, 3–17. [Google Scholar] [CrossRef]
- Dragomir, M.P.; Knutsen, E.; Calin, G.A. SnapShot: Unconventional miRNA Functions. Cell. 2018, 174, 1038. [Google Scholar] [CrossRef]
- Dragomir, M.; Calin, G.A. Circular RNAs in Cancer-Lessons Learned From microRNAs. Front. Oncol. 2018, 8, 179. [Google Scholar] [CrossRef] [PubMed]
- Geisler, S.; Coller, J. RNA in Unexpected Places: Long Non-coding RNA Functions in Diverse Cellular Contexts. Nat. Rev. Mol. Cell Boil. 2013, 14, 699–712. [Google Scholar] [CrossRef] [PubMed]
- Koerner, M.V.; Pauler, F.M.; Huang, R.; Barlow, D.P. The Function of Non-coding RNAs in Genomic Imprinting. Development 2009, 136, 1771–1783. [Google Scholar] [CrossRef] [PubMed]
- Chooniedass-Kothari, S.; Emberley, E.; Hamedani, M.; Troup, S.; Wang, X.; Czosnek, A.; Hube, F.; Mutawe, M.; Watson, P.; Leygue, E. The Steroid Receptor RNA Activator is the First Functional RNA Encoding a Protein. FEBS Lett. 2004, 566, 43–47. [Google Scholar] [CrossRef] [PubMed]
- Tenson, T.; DeBlasio, A.; Mankin, A. A Functional Peptide Encoded in the Escherichia coli 23S rRNA. Proc. Natl. Acad. Sci. USA 1996, 93, 5641–5646. [Google Scholar] [CrossRef]
- Bileschi, M.; Belanger, D.; Bryant, D.H.; Sanderson, T.; Carter, B.; Sculley, D.; Depristo, M.A.; Colwell, L. Using Deep Learning to Annotate the Protein Universe. bioRxiv 2019, 626507. [Google Scholar]
- Mattick, J.S.; Dinger, M.E. The Extent of Functionality in the Human Genome. HUGO J. 2013, 7, 1–4. [Google Scholar] [CrossRef]
- Lauressergues, M.; Couzigou, J.-M.; Clemente, H.S.; Martinez, Y.; Dunand, C.; Becard, G.; Combier, J.-P. Primary Transcripts of MicroRNAs Encode Regulatory Peptides. Nature 2015, 520, 90–93. [Google Scholar] [CrossRef]
- Cai, X.; Hagedorn, C.H.; Cullen, B.R. Human microRNAs are Processed from Capped, Polyadenylated Transcripts That Can Also Function as mRNAs. RNA 2004, 10, 1957–1966. [Google Scholar] [CrossRef]
- Jeck, W.R.; Sorrentino, J.A.; Wang, K.; Slevin, M.K.; Burd, C.E.; Liu, J.; Marzluff, W.F.; Sharpless, N.E. Circular RNAs Are Abundant, Conserved, and Associated with ALU Repeats. RNA 2012, 19, 141–157. [Google Scholar] [CrossRef]
- Lodish, H.; Berk, A.; Kaiser, C.A.; Krieger, M.; Bretscher, A.; Ploegh, H.; Amon, A.; Martin, K.; Scott, M.P. Molecular Cell Biology, 8th ed.; W.H.Freeman & Co Ltd.: New York, NY, USA, 2016; p. 1280. [Google Scholar]
- Nelson, D.L.; Cox, M.M.; Lehninger, A.L. Lehninger Principles of Biochemistry, 7th ed.; W.H. Freeman and Company: New York, NY, USA; Macmillan Higher Education: Houndmills, Basingstoke, 2017. [Google Scholar]
- Desiere, F. The Peptide Atlas Project. Nucleic Acids Res. 2006, 34, D655–D658. [Google Scholar] [CrossRef] [PubMed]
- Huang, H.-H.; Chen, F.-Y.; Chou, W.-C.; Hou, H.-A.; Ko, B.-S.; Lin, C.-T.; Tang, J.-L.; Li, C.-C.; Yao, M.; Tsay, W.; et al. Long Non-coding RNA HOXB-AS3 Promotes Myeloid Cell Proliferation and Its Higher Expression Is an Adverse Prognostic Marker in Patients with Acute Myeloid Leukemia and Myelodysplastic Syndrome. BMC Cancer 2019, 19, 1–14. [Google Scholar] [CrossRef] [PubMed]
- Huang, J.-Z.; Chen, M.; Chen, D.; Gao, X.-C.; Zhu, S.; Huang, H.; Hu, M.; Zhu, H.; Yan, G.-R. A Peptide Encoded by a Putative lncRNA HOXB-AS3 Suppresses Colon Cancer Growth. Mol. Cell. 2017, 68, 171–184.e6. [Google Scholar] [CrossRef]
- Pauli, A.; Norris, M.L.; Valen, E.; Chew, G.-L.; Gagnon, J.A.; Zimmerman, S.; Mitchell, A.; Ma, J.; Dubrulle, J.; Reyon, D.; et al. Toddler: An Embryonic Signal That Promotes Cell Movement via Apelin Receptors. Science 2014, 343, 1248636. [Google Scholar] [CrossRef]
- D’Lima, N.G.; Ma, J.; Winkler, L.; Chu, Q.; Loh, K.H.; Corpuz, E.O.; Budnik, B.A.; Lykke-Andersen, J.; Saghatelian, A.; Slavoff, S.A. A Human Microprotein That Interacts with the mRNA Decapping Complex. Nat. Methods 2016, 13, 174–180. [Google Scholar] [CrossRef] [PubMed]
- Galindo, M.I.; Pueyo, J.I.; Fouix, S.; Bishop, S.A.; Couso, J.P. Peptides Encoded by Short ORFs Control Development and Define a New Eukaryotic Eene Family. PLoS Boil. 2007, 5, e106. [Google Scholar] [CrossRef]
- Cai, B.; Li, Z.; Ma, M.; Wang, Z.; Han, P.; Abdalla, B.A.; Nie, Q.; Zhang, X. LncRNA-Six1 Encodes a Micropeptide to Activate Six1 in Cis and Is Involved in Cell Proliferation and Muscle Growth. Front. Physiol. 2017, 8. [Google Scholar] [CrossRef]
- Razooky, B.S.; Obermayer, B.; O’May, J.B.; Tarakhovsky, A. Viral Infection Identifies Micropeptides Differentially Regulated in smORF-Containing lncRNAs. Genes 2017, 8, 206. [Google Scholar] [CrossRef]
- Makarewich, C.A.; Baskin, K.K.; Munir, A.Z.; Bezprozvannaya, S.; Sharma, G.; Khemtong, C.; Shah, A.M.; McAnally, J.R.; Malloy, C.R.; Szweda, L.I.; et al. MOXI Is a Mitochondrial Micropeptide That Enhances Fatty Acid β-Oxidation. Cell Rep. 2018, 23, 3701–3709. [Google Scholar] [CrossRef]
- Stein, C.S.; Jadiya, P.; Zhang, X.; McLendon, J.M.; Abouassaly, G.M.; Witmer, N.; Anderson, E.J.; Elrod, J.W.; Boudreau, R.L. Mitoregulin: A lncRNA-Encoded Microprotein that Supports Mitochondrial Supercomplexes and Respiratory Efficiency. Cell Rep. 2018, 23, 3710–3720.e8. [Google Scholar] [CrossRef]
- Zheng, X.; Chen, L.; Zhou, Y.; Wang, Q.; Zheng, Z.; Xu, B.; Wu, C.; Zhou, Q.; Hu, W.; Jiang, J.; et al. A Novel Protein Encoded by a Circular RNA circPPP1R12A Promotes Tumor Pathogenesis and Metastasis of Colon Cancer via Hippo-YAP Signaling. Mol. Cancer 2019, 18, 47. [Google Scholar] [CrossRef] [PubMed]
- Polycarpou-Schwarz, M.; Groß, M.; Mestdagh, P.; Schott, J.; Grund, S.E.; Hildenbrand, C.; Rom, J.; Aulmann, S.; Sinn, P.; Vandesompele, J.; et al. The Cancer-Associated Microprotein CASIMO1 Controls Cell Proliferation and Interacts with Squalene Epoxidase Modulating Lipid Droplet Formation. Oncogene 2018, 37, 4750–4768. [Google Scholar] [CrossRef] [PubMed]
- Zhao, J.; Lee, E.E.; Kim, J.; Yang, R.; Chamseddin, B.; Ni, C.; Gusho, E.; Xie, Y.; Chiang, C.-M.; Buszczak, M.; et al. Transforming Activity of an Oncoprotein-Encoding Circular RNA From Human Papillomavirus. Nat. Commun. 2019, 10, 2300. [Google Scholar] [CrossRef] [PubMed]
- Charpentier, M.; Croyal, M.; Carbonnelle, D.; Fortun, A.; Florenceau, L.; Rabu, C.; Krempf, M.; Labarrière, N.; Lang, F. IRES-Dependent Translation of the Long Non coding RNA Meloe in Melanoma Cells Produces the Most Immunogenic MELOE Antigens. Oncotarget 2016, 7, 59704–59713. [Google Scholar] [CrossRef] [PubMed]
- Wu, S.; Zhang, L.; Deng, J.; Guo, B.; Li, F.; Wang, Y.; Wu, R.; Zhang, S.; Lu, J.; Zhou, Y. A Novel Micropeptide Encoded by Y-Linked LINC00278 Links Cigarette Smoking and AR Signaling in Male Esophageal Squamous Cell Carcinoma. Cancer Res. 2020, 80, 2790–2803. [Google Scholar] [CrossRef] [PubMed]
- Guo, B.; Wu, S.; Zhu, X.; Zhang, L.; Deng, J.; Li, F.; Wang, Y.; Zhang, S.; Wu, R.; Lu, J.; et al. Micropeptide CIP 2A- BP Encoded by LINC 00665 Inhibits Triple-Negative Breast Cancer Progression. EMBO J. 2019, 39. [Google Scholar] [CrossRef] [PubMed]
- Wang, Y.; Wu, S.; Zhu, X.; Zhang, L.; Deng, J.; Li, F.; Guo, B.; Zhang, S.; Wu, R.; Zhang, Z.; et al. LncRNA-Encoded Polypeptide ASRPS Inhibits Triple-Negative Breast Cancer Angiogenesis. J. Exp. Med. 2019, 217. [Google Scholar] [CrossRef]
- Li, M.; Li, X.; Zhang, Y.; Wu, H.; Zhou, H.; Ding, X.; Zhang, X.; Jin, X.; Wang, Y.; Yin, X.; et al. Micropeptide MIAC Inhibits HNSCC Progression by Interacting with Aquaporin 2. J. Am. Chem. Soc. 2020, 142, 6708–6716. [Google Scholar] [CrossRef]
- Zhang, M.; Zhao, K.; Xu, X.; Yang, Y.; Yan, S.; Wei, P.; Liu, H.; Xu, J.; Xiao, F.; Zhou, H.; et al. A Peptide Encoded by Circular Form of LINC-PINT Suppresses Oncogenic Transcriptional Elongation in Glioblastoma. Nat. Commun. 2018, 9, 4475. [Google Scholar] [CrossRef]
- Illing, P.T.; Vivian, J.P.; Dudek, N.L.; Kostenko, L.; Chen, Z.; Bharadwaj, M.; Miles, J.J.; Kjer-Nielsen, L.; Gras, S.; Williamson, N.A.; et al. Immune Self-Reactivity Triggered by Drug-Modified HLA-peptide Repertoire. Nature 2012, 486, 554–558. [Google Scholar] [CrossRef]
- Saini, S.K.; Ostermeir, K.; Ramnarayan, V.R.; Schuster, H.; Zacharias, M.; Springer, S. Dipeptides Promote Folding and Peptide Binding of MHC Class I Molecules. Proc. Natl. Acad. Sci. USA 2013, 110, 15383–15388. [Google Scholar] [CrossRef]
- Pollard, K.S.; Hubisz, M.J.; Rosenbloom, K.R.; Siepel, A. Detection of Nonneutral Substitution Rates on Mammalian Phylogenies. Genome Res. 2009, 20, 110–121. [Google Scholar] [CrossRef] [PubMed]
- Wang, J.; Zhu, S.; Meng, N.; He, Y.; Lu, R.; Yan, G.-R. ncRNA-Encoded Peptides or Proteins and Cancer. Mol. Ther. 2019, 27, 1718–1725. [Google Scholar] [CrossRef] [PubMed]
- Washietl, S.; Kellis, M.; Garber, M. Evolutionary Dynamics and Tissue Specificity of Human Long Noncoding RNAs in Six Mammals. Genome Res. 2014, 24, 616–628. [Google Scholar] [CrossRef]
- Necsulea, A.; Soumillon, M.; Warnefors, M.; Liechti, A.; Daish, T.; Zeller, U.; Baker, J.C.; Grützner, F.; Kaessmann, H. The Evolution of lncRNA Repertoires and Expression Patterns in Tetrapods. Nature 2014, 505, 635–640. [Google Scholar] [CrossRef]
- Ingolia, N.T.; Ghaemmaghami, S.; Newman, J.R.S.; Weissman, J.S. Genome-Wide Analysis In Vivo of Translation with Nucleotide Resolution Using Ribosome Profiling. Science 2009, 324, 218–223. [Google Scholar] [CrossRef]
- Gebetsberger, J.; Żywicki, M.; Künzi, A.; Polacek, N. tRNA-Derived Fragments Target the Ribosome and Function as Regulatory Non-Coding RNA in Haloferax Volcanii. Archaea 2012, 2012, 1–11. [Google Scholar] [CrossRef] [PubMed]
- Pircher, A.; Bakowska-Zywicka, K.; Schneider, L.; Żywicki, M.; Polacek, N. An mRNA-Derived Noncoding RNA Targets and Regulates the Ribosome. Mol. Cell. 2014, 54, 147–155. [Google Scholar] [CrossRef]
- Van Heesch, S.; Van Iterson, M.; Jacobi, J.; Boymans, S.; Essers, P.B.; De Bruijn, E.; Hao, W.; MacInnes, A.W.; Cuppen, E.; Simonis, M. Extensive Localization of Long Noncoding RNAs to the Cytosol and Mono- and Polyribosomal Complexes. Genome Boil. 2014, 15, R6. [Google Scholar] [CrossRef]
- Calviello, L.; Mukherjee, N.; Wyler, E.; Zauber, H.; Hirsekorn, A.; Selbach, M.; Landthaler, M.; Obermayer, B.; Ohler, U. Detecting Actively Translated Ppen Reading Frames in Ribosome Profiling Data. Nat. Methods 2015, 13, 165–170. [Google Scholar] [CrossRef]
- Fournaise, E.; Chaurand, P. Increasing Specificity in Imaging Mass Spectrometry: High Spatial Fidelity Transfer of Proteins from Tissue Sections to Functionalized Surfaces. Anal. Bioanal. Chem. 2014, 407, 2159–2166. [Google Scholar] [CrossRef] [PubMed]
- Natsume, T.; Yamauchi, Y.; Nakayama, H.; Shinkawa, T.; Yanagida, M.; Takahashi, N.; Isobe, T. A Direct Nanoflow Liquid Chromatography−Tandem Mass Spectrometry System for Interaction Proteomics. Anal. Chem. 2002, 74, 4725–4733. [Google Scholar] [CrossRef] [PubMed]
- Chen, K.-Y.; Knoepfler, P. To CRISPR and Beyond: The Evolution of Genome Editing in Stem Cells. Regen. Med. 2016, 11, 801–816. [Google Scholar] [CrossRef] [PubMed]

© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Dragomir, M.P.; Manyam, G.C.; Ott, L.F.; Berland, L.; Knutsen, E.; Ivan, C.; Lipovich, L.; Broom, B.M.; Calin, G.A. FuncPEP: A Database of Functional Peptides Encoded by Non-Coding RNAs. Non-Coding RNA 2020, 6, 41. https://doi.org/10.3390/ncrna6040041
Dragomir MP, Manyam GC, Ott LF, Berland L, Knutsen E, Ivan C, Lipovich L, Broom BM, Calin GA. FuncPEP: A Database of Functional Peptides Encoded by Non-Coding RNAs. Non-Coding RNA. 2020; 6(4):41. https://doi.org/10.3390/ncrna6040041
Chicago/Turabian StyleDragomir, Mihnea P., Ganiraju C. Manyam, Leonie Florence Ott, Léa Berland, Erik Knutsen, Cristina Ivan, Leonard Lipovich, Bradley M. Broom, and George A. Calin. 2020. "FuncPEP: A Database of Functional Peptides Encoded by Non-Coding RNAs" Non-Coding RNA 6, no. 4: 41. https://doi.org/10.3390/ncrna6040041
APA StyleDragomir, M. P., Manyam, G. C., Ott, L. F., Berland, L., Knutsen, E., Ivan, C., Lipovich, L., Broom, B. M., & Calin, G. A. (2020). FuncPEP: A Database of Functional Peptides Encoded by Non-Coding RNAs. Non-Coding RNA, 6(4), 41. https://doi.org/10.3390/ncrna6040041








