Crosstalk Between Mammalian Antiviral Pathways
Abstract
1. Introduction
1.1. The Mammalian Type-I IFN Response
1.2. Viral-Derived Nucleic Acids Sensing Mechanisms
1.2.1. TLR-Detection of Viral Nucleic Acids
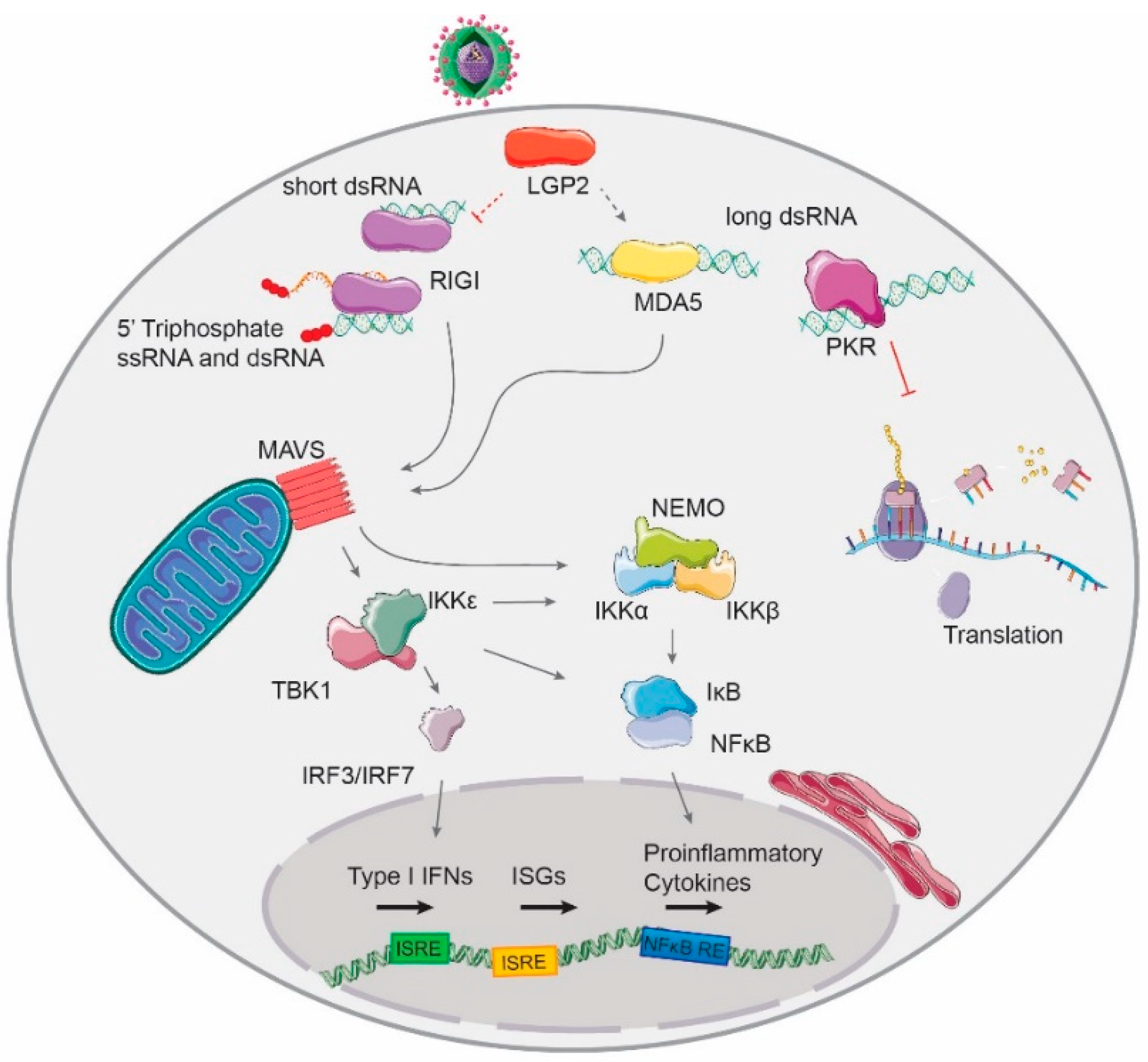
1.2.2. RLR-Detection of Virus-Derived RNA
1.2.3. Other Antiviral dsRNA-Activated Pathways
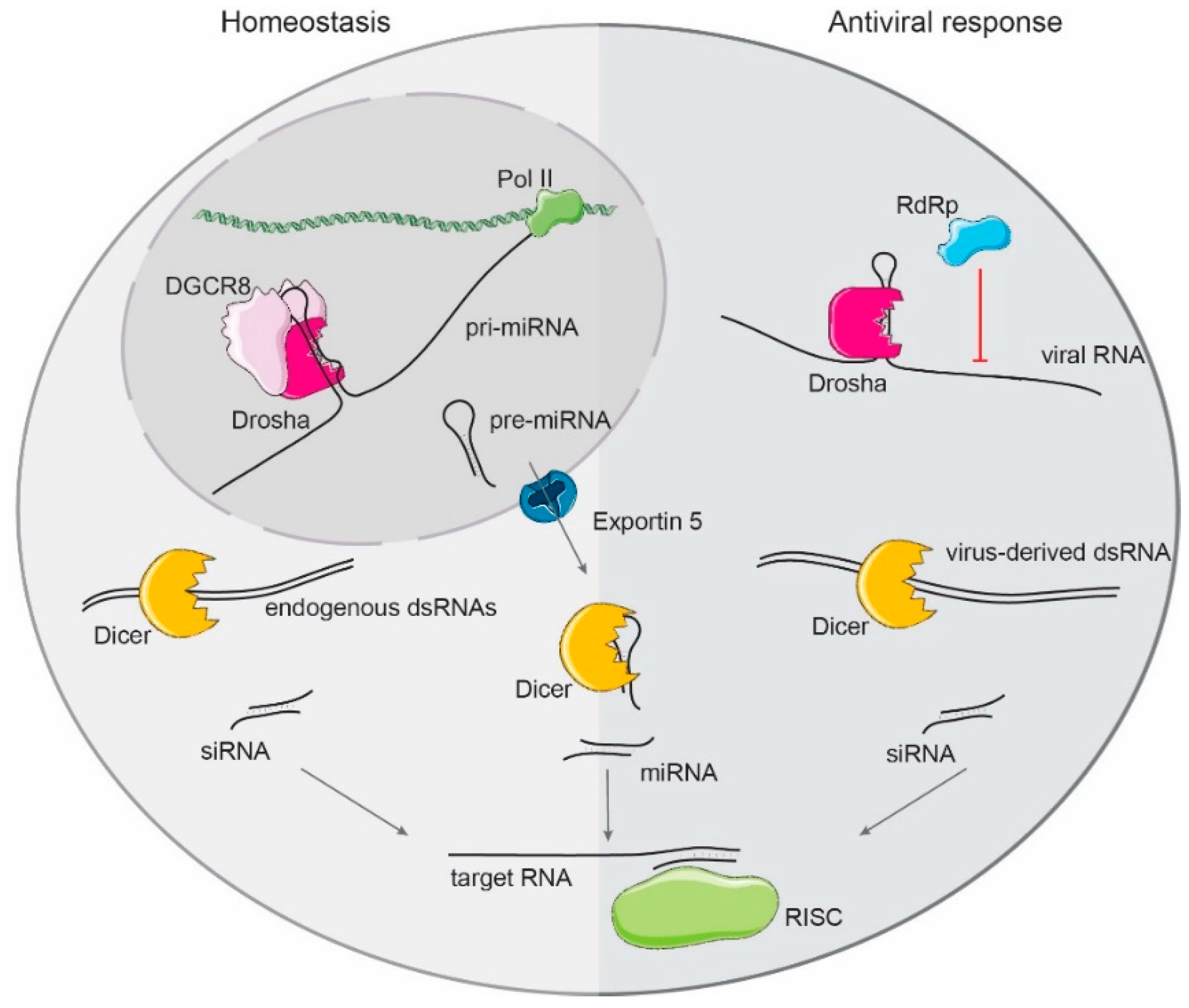
1.3. Mammalian Small RNA Biogenesis
2. Drosha and Dicer as Direct Antiviral Factors
2.1. Drosha
2.2. Dicer
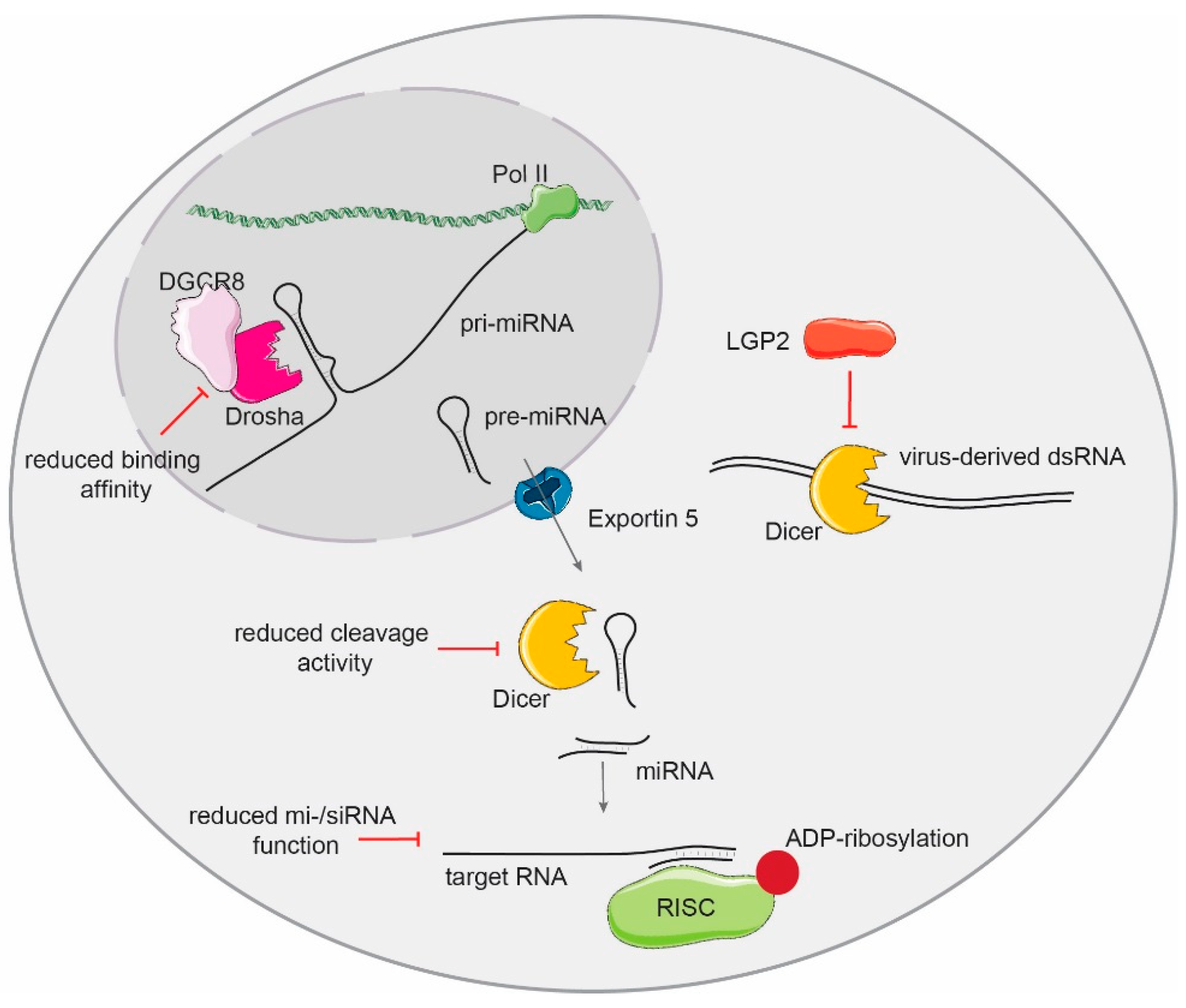
3. Type I IFNs Modulate the Activity of The Small RNA Biogenesis Pathway
4. MiRNA-Mediated Regulation of Viruses
5. Consequences of Small RNA Biogenesis Manipulation on the IFN and Antiviral Response
5.1. Knock-Downs of Small RNA Biogenesis in Somatic Cells
5.2. Knock-Out of Small RNA Biogenesis in Somatic Cells
5.3. Small RNAs and IFN Response in Embryonic Stem Cells
6. Conclusions
Author Contributions
Funding
Conflicts of Interest
References
- González-Navajas, J.M.; Lee, J.; David, M.; Raz, E. Immunomodulatory functions of type i interferons. Nat. Rev. Immunol. 2012, 12, 125–135. [Google Scholar] [CrossRef] [PubMed]
- Di Franco, S.; Turdo, A.; Todaro, M.; Stassi, G. Role of type i and ii interferons in colorectal cancer and melanoma. Front. Immunol. 2017, 8, 878. [Google Scholar] [CrossRef]
- Hall, J.C.; Rosen, A. Type i interferons: Crucial participants in disease amplification in autoimmunity. Nat. Rev. Rheumatol. 2010, 6, 40–49. [Google Scholar] [CrossRef]
- Venkatesh, B.; Lee, A.P.; Ravi, V.; Maurya, A.K.; Lian, M.M.; Swann, J.B.; Ohta, Y.; Flajnik, M.F.; Sutoh, Y.; Kasahara, M.; et al. Elephant shark genome provides unique insights into gnathostome evolution. Nature 2014, 505, 174–179. [Google Scholar] [CrossRef]
- Secombes, C.J.; Zou, J. Evolution of interferons and interferon receptors. Front. Immunol. 2017, 8, 209. [Google Scholar] [CrossRef]
- Swiecki, M.; Colonna, M. Type i interferons: Diversity of sources, production pathways and effects on immune responses. Curr. Opin. Virol. 2011, 1, 463–475. [Google Scholar] [CrossRef] [PubMed]
- Pestka, S.; Krause, C.D.; Walter, M.R. Interferons, interferon-like cytokines, and their receptors. Immunol. Rev. 2004, 202, 8–32. [Google Scholar] [CrossRef]
- Havell, E.A.; Berman, B.; Ogburn, C.A.; Berg, K.; Paucker, K.; Vilcek, J. Two antigenically distinct species of human interferon. Proc. Natl. Acad. Sci. USA 1975, 72, 2185–2187. [Google Scholar] [CrossRef] [PubMed]
- Klein, J.R.; Raulet, D.H.; Pasternack, M.S.; Bevan, M.J. Cytotoxic t lymphocytes produce immune interferon in response to antigen or mitogen. J. Exp. Med. 1982, 155, 1198–1203. [Google Scholar] [CrossRef] [PubMed]
- Scharton, T.M.; Scott, P. Natural killer cells are a source of interferon gamma that drives differentiation of cd4+ t cell subsets and induces early resistance to leishmania major in mice. J. Exp. Med. 1993, 178, 567–577. [Google Scholar] [CrossRef] [PubMed]
- Coccia, E.M.; Severa, M.; Giacomini, E.; Monneron, D.; Remoli, M.E.; Julkunen, I.; Cella, M.; Lande, R.; Uzé, G. Viral infection and toll-like receptor agonists induce a differential expression of type i and λ interferons in human plasmacytoid and monocyte-derived dendritic cells. Eur. J. Immunol. 2004, 34, 796–805. [Google Scholar] [CrossRef]
- Karpala, A.J.; Morris, K.R.; Broadway, M.M.; McWaters, P.G.D.; O’Neil, T.E.; Goossens, K.E.; Lowenthal, J.W.; Bean, A.G.D. Molecular cloning, expression, and characterization of chicken ifn -λ. J. Interferon Cytokine Res. 2008, 28, 341–350. [Google Scholar] [CrossRef] [PubMed]
- Kotenko, S.V.; Gallagher, G.; Baurin, V.V.; Lewis-Antes, A.; Shen, M.; Shah, N.K.; Langer, J.A.; Sheikh, F.; Dickensheets, H.; Donnelly, R.P. Ifn-λs mediate antiviral protection through a distinct class ii cytokine receptor complex. Nat. Immunol. 2003, 4, 69–77. [Google Scholar] [CrossRef]
- Spann, K.M.; Tran, K.-C.; Chi, B.; Rabin, R.L.; Collins, P.L. Suppression of the induction of alpha, beta, and lambda interferons by the ns1 and ns2 proteins of human respiratory syncytial virus in human epithelial cells and macrophages. J. Virol. 2004, 78, 4363–4369. [Google Scholar] [CrossRef]
- Gürtler, C.; Bowie, A.G. Innate immune detection of microbial nucleic acids. Trends Microbiol. 2013, 21, 413–420. [Google Scholar] [CrossRef] [PubMed]
- Roers, A.; Hiller, B.; Hornung, V. Recognition of endogenous nucleic acids by the innate immune system. Immunity 2016, 44, 739–754. [Google Scholar] [CrossRef] [PubMed]
- Novick, D.; Cohen, B.; Rubinstein, M. The human interferon alpha/beta receptor: Characterization and molecular cloning. Cell 1994, 77, 391–400. [Google Scholar] [CrossRef]
- Schneider, W.M.; Chevillotte, M.D.; Rice, C.M. Interferon-stimulated genes: A complex web of host defenses. Annu. Rev. Immunol. 2014, 32, 513–545. [Google Scholar] [CrossRef]
- Lester, S.N.; Li, K. Toll-like receptors in antiviral innate immunity. J. Mol. Biol. 2014, 426, 1246–1264. [Google Scholar] [CrossRef]
- Takeda, K.; Akira, S. Toll-like receptors in innate immunity. Int. Immunol. 2004, 17, 1–14. [Google Scholar] [CrossRef]
- Alexopoulou, L.; Holt, A.C.; Medzhitov, R.; Flavell, R.A. Recognition of double-stranded RNA and activation of nf-κb by toll-like receptor 3. Nature 2001, 413, 732–738. [Google Scholar] [CrossRef]
- Deguine, J.; Barton, G.M. Myd88: A central player in innate immune signaling. F1000Prime Rep. 2014, 6, 97. [Google Scholar] [CrossRef]
- Honda, K.; Yanai, H.; Mizutani, T.; Negishi, H.; Shimada, N.; Suzuki, N.; Ohba, Y.; Takaoka, A.; Yeh, W.-C.; Taniguchi, T. Role of a transductional-transcriptional processor complex involving myd88 and irf-7 in toll-like receptor signaling. Proc. Natl. Acad. Sci. USA 2004, 101, 15416–15421. [Google Scholar] [CrossRef]
- Vercammen, E.; Staal, J.; Beyaert, R. Sensing of viral infection and activation of innate immunity by toll-like receptor 3. Clin. Microbiol. Rev. 2008, 21, 13–25. [Google Scholar] [CrossRef]
- Yoneyama, M.; Kikuchi, M.; Matsumoto, K.; Imaizumi, T.; Miyagishi, M.; Taira, K.; Foy, E.; Loo, Y.M.; Gale, M.; Akira, S.; et al. Shared and unique functions of the dexd/h-box helicases rig-i, mda5, and lgp2 in antiviral innate immunity. J. Immunol. 2005, 175, 2851–2858. [Google Scholar] [CrossRef]
- Yoneyama, M.; Kikuchi, M.; Natsukawa, T.; Shinobu, N.; Imaizumi, T.; Miyagishi, M.; Taira, K.; Akira, S.; Fujita, T. The RNA helicase rig-i has an essential function in double-stranded RNA-induced innate antiviral responses. Nat. Immunol. 2004, 5, 730–737. [Google Scholar] [CrossRef]
- Hornung, V.; Ellegast, J.; Kim, S.; Brzózka, K.; Jung, A.; Kato, H.; Poeck, H.; Akira, S.; Conzelmann, K.-K.; Schlee, M.; et al. 5’-triphosphate RNA is the ligand for rig-i. Science 2006, 314, 994–997. [Google Scholar] [CrossRef]
- Kato, H.; Takeuchi, O.; Mikamo-Satoh, E.; Hirai, R.; Kawai, T.; Matsushita, K.; Hiiragi, A.; Dermody, T.S.; Fujita, T.; Akira, S. Length-dependent recognition of double-stranded ribonucleic acids by retinoic acid-inducible gene-i and melanoma differentiation-associated gene 5. J. Exp. Med. 2008, 205, 1601–1610. [Google Scholar] [CrossRef]
- Kato, H.; Takeuchi, O.; Sato, S.; Yoneyama, M.; Yamamoto, M.; Matsui, K.; Uematsu, S.; Jung, A.; Kawai, T.; Ishii, K.J.; et al. Differential roles of mda5 and rig-i helicases in the recognition of Rna viruses. Nature 2006, 441, 101–105. [Google Scholar] [CrossRef]
- Jiang, F.; Ramanathan, A.; Miller, M.T.; Tang, G.-Q.Q.; Gale, M.; Patel, S.S.; Marcotrigiano, J. Structural basis of RNA recognition and activation by innate immune receptor rig-i. Nature 2011, 479, 423–427. [Google Scholar] [CrossRef]
- Takahasi, K.; Yoneyama, M.; Nishihori, T.; Hirai, R.; Kumeta, H.; Narita, R.; Gale, M.; Inagaki, F.; Fujita, T. Nonself RNA-sensing mechanism of rig-i helicase and activation of antiviral immune responses. Mol. Cell 2008, 29, 428–440. [Google Scholar] [CrossRef]
- Peisley, A.; Lin, C.; Wu, B.; Orme-Johnson, M.; Liu, M.; Walz, T.; Hur, S. Cooperative assembly and dynamic disassembly of mda5 filaments for viral dsRNA recognition. Proc. Natl. Acad. Sci. USA 2011, 108, 21010–21015. [Google Scholar] [CrossRef]
- Peisley, A.; Wu, B.; Xu, H.; Chen, Z.J.; Hur, S. Structural basis for ubiquitin-mediated antiviral signal activation by rig-i. Nature 2014, 509, 110–114. [Google Scholar] [CrossRef]
- Hou, F.; Sun, L.; Zheng, H.; Skaug, B.; Jiang, Q.-X.; Chen, Z.J. Mavs forms functional prion-like aggregates to activate and propagate antiviral innate immune response. Cell 2011, 146, 448–461. [Google Scholar] [CrossRef]
- Kawai, T.; Takahashi, K.; Sato, S.; Coban, C.; Kumar, H.; Kato, H.; Ishii, K.J.; Takeuchi, O.; Akira, S. Ips-1, an adaptor triggering rig-i- and mda5-mediated type i interferon induction. Nat. Immunol. 2005, 6, 981–988. [Google Scholar] [CrossRef]
- Hiscott, J. Triggering the innate antiviral response through irf-3 activation. J. Biol. Chem. 2007, 282, 15325–15329. [Google Scholar] [CrossRef]
- Reikine, S.; Nguyen, J.B.; Modis, Y. Pattern recognition and signaling mechanisms of rig-i and mda5. Front. Immunol. 2014, 5, 342. [Google Scholar] [CrossRef]
- Bruns, A.M.; Leser, G.P.; Lamb, R.A.; Horvath, C.M. The innate immune sensor lgp2 activates antiviral signaling by regulating mda5-RNA interaction and filament assembly. Mol. Cell 2014, 55, 771–781. [Google Scholar] [CrossRef]
- Satoh, T.; Kato, H.; Kumagai, Y.; Yoneyama, M.; Sato, S.; Matsushita, K.; Tsujimura, T.; Fujita, T.; Akira, S.; Takeuchi, O. Lgp2 is a positive regulator of rig-i- and mda5-mediated antiviral responses. Proc. Natl. Acad. Sci. USA 2010, 107, 1512–1517. [Google Scholar] [CrossRef]
- Rothenfusser, S.; Goutagny, N.; DiPerna, G.; Gong, M.; Monks, B.G.; Schoenemeyer, A.; Yamamoto, M.; Akira, S.; Fitzgerald, K.A. The RNA helicase lgp2 inhibits tlr-independent sensing of viral replication by retinoic acid-inducible gene-i. J. Immunol. 2005, 175, 5260–5268. [Google Scholar] [CrossRef]
- Bruns, A.M.; Horvath, C.M. Lgp2 synergy with mda5 in rlr-mediated RNA recognition and antiviral signaling. Cytokine 2015, 74, 198–206. [Google Scholar] [CrossRef]
- Komuro, A.; Horvath, C.M. Rna- and virus-independent inhibition of antiviral signaling by RNA helicase lgp2. J. Virol. 2006, 80, 12332–12342. [Google Scholar] [CrossRef]
- Vitour, D.; Meurs, E.F. Regulation of interferon production by rig-i and lgp2: A lesson in self-control. Sci. Signal. 2007, 2007, pe20. [Google Scholar] [CrossRef]
- Meurs, E.; Chong, K.; Galabru, J.; Thomas, N.S.B.; Kerr, I.M.; Williams, B.R.G.; Hovanessian, A.G. Molecular cloning and characterization of the human double-stranded RNA-activated protein kinase induced by interferon. Cell 1990, 62, 379–390. [Google Scholar] [CrossRef]
- Gross, M.; Wing, M.; Rundquist, C.; Rubino, S. Evidence that phosphorylation of eif-2 (4 prevents the eif-2b-mediated dissociation of eif-z * gdp from the 60 s subunit of complete initiation complexes. J. Biol. Chem. 1987, 262, 6899–6907. [Google Scholar]
- Meurs, E.F.; Watanabe, Y.; Kadereit, S.; Barber, G.N.; Katze, M.G.; Chong, K.; Williams, B.R.; Hovanessian, A.G. Constitutive expression of human double-stranded RNA-activated p68 kinase in murine cells mediates phosphorylation of eukaryotic initiation factor 2 and partial resistance to encephalomyocarditis virus growth. J. Virol. 1992, 66, 5805–5814. [Google Scholar]
- Sudhakar, A.; Ramachandran, A.; Ghosh, S.; Hasnain, S.E.; Kaufman, R.J.; Ramaiah, K.V.A. Phosphorylation of serine 51 in initiation factor 2α (eif2α) promotes complex formation between eif2α(p) and eif2b and causes inhibition in the guanine nucleotide exchange activity of eif2b. Biochemistry 2000, 39, 12929–12938. [Google Scholar] [CrossRef]
- Hovanessian, A.G.; Justesen, J. The human 2′-5′oligoadenylate synthetase family: Unique interferon-inducible enzymes catalyzing 2′-5′ instead of 3′-5′ phosphodiester bond formation. Biochimie 2007, 89, 779–788. [Google Scholar] [CrossRef]
- Floyd-Smith, G.; Slattery, E.; Lengyel, P. Interferon action: Rna cleavage pattern of a (2’-5’) oligoadenylate--dependent endonuclease. Science 1981, 212, 1030–1032. [Google Scholar] [CrossRef]
- Nakanishi, M.; Goto, Y.; Kitade, Y. 2-5a induces a conformational change in the ankyrin-repeat domain of RNAse L. Proteins: Struct. Funct. Bioinform. 2005, 60, 131–138. [Google Scholar] [CrossRef]
- Wreschner, D.H.; McCauley, J.W.; Skehel, J.J.; Kerr, I.M. Interferon action—sequence specificity of the ppp(a2′p)na-dependent ribonuclease. Nature 1981, 289, 414–417. [Google Scholar] [CrossRef] [PubMed]
- Malathi, K.; Dong, B.; Gale, M.; Silverman, R.H. Small self-RNA generated by RNAse l amplifies antiviral innate immunity. Nature 2007, 448, 816–819. [Google Scholar] [CrossRef] [PubMed]
- Vitali, P.; Scadden, A.D. Double-stranded RNAs containing multiple iu pairs are sufficient to suppress interferon induction and apoptosis. Nat. Struct Mol. Biol. 2010, 17, 1043–1050. [Google Scholar] [CrossRef] [PubMed]
- Mannion, N.M.; Greenwood, S.M.; Young, R.; Cox, S.; Brindle, J.; Read, D.; Nellaker, C.; Vesely, C.; Ponting, C.P.; McLaughlin, P.J. , et al. The RNA-editing enzyme adar1 controls innate immune responses to RNA. Cell Rep. 2014, 9, 1482–1494. [Google Scholar] [CrossRef] [PubMed]
- Samuel, C.E. Adenosine deaminases acting on RNA (adars) are both antiviral and proviral. Virology 2011, 411, 180–193. [Google Scholar] [CrossRef]
- Farazi, T.A.; Juranek, S.A.; Tuschl, T. Development. Development 2008, 132, 4645–4652. [Google Scholar]
- Siomi, M.C.; Sato, K.; Pezic, D.; Aravin, A.A. Piwi-interacting small RNAs: The vanguard of genome defence. Nat. Rev. Mol. Cell Biol. 2011, 12, 246–258. [Google Scholar] [CrossRef]
- Lee, Y.; Jeon, K.; Lee, J.-T.; Kim, S.; Kim, V.N.; Kim, V.N. MicroRNA maturation: Stepwise processing and subcellular localization. EMBO J. 2002, 21, 4663–4670. [Google Scholar] [CrossRef]
- Lee, Y.; Kim, M.; Han, J.; Yeom, K.-H.; Lee, S.; Baek, S.H.; Kim, V.N. MicroRNA genes are transcribed by RNA polymerase ii. EMBO J. 2004, 23, 4051–4060. [Google Scholar] [CrossRef]
- Denli, A.M.; Tops, B.B.J.; Plasterk, R.H.A.; Ketting, R.F.; Hannon, G.J. Processing of primary microRNAs by the microprocessor complex. Nature 2004, 432, 231–235. [Google Scholar] [CrossRef]
- Gregory, R.I.; Yan, K.-P.; Amuthan, G.; Chendrimada, T.; Doratotaj, B.; Cooch, N.; Shiekhattar, R. The microprocessor complex mediates the genesis of microRNAs. Nature 2004, 432, 235–240. [Google Scholar] [CrossRef]
- Han, J.; Lee, Y.; Yeom, K.-H.; Kim, Y.-K.; Jin, H.; Kim, V.N. The drosha-dgcr8 complex in primary microRNA processing. Genes Dev. 2004, 18, 3016–3027. [Google Scholar] [CrossRef]
- Landthaler, M.; Yalcin, A.; Tuschl, T. The human digeorge syndrome critical region gene 8 and its d. Melanogaster homolog are required for miRNA biogenesis. Curr. Biol. 2004, 14, 2162–2167. [Google Scholar] [CrossRef]
- Lee, Y.; Ahn, C.; Han, J.; Choi, H.; Kim, J.; Yim, J.; Lee, J.; Provost, P.; Rådmark, O.; Kim, S.; et al. The nuclear RNAse iii drosha initiates microRNA processing. Nature 2003, 425, 415–419. [Google Scholar] [CrossRef] [PubMed]
- Nguyen, T.A.; Jo, M.H.; Choi, Y.-G.; Park, J.; Kwon, S.C.; Hohng, S.; Kim, V.N.; Woo, J.-S. Functional anatomy of the human microprocessor. Cell 2015, 161, 1374–1387. [Google Scholar] [CrossRef] [PubMed]
- Kwon, S.C.; Baek, S.C.; Choi, Y.-G.; Yang, J.; Lee, Y.-S.; Woo, J.-S.; Kim, V.N. Molecular basis for the single-nucleotide precision of primary microRNA processing. Mol. Cell 2018, 73, 505–518. [Google Scholar] [CrossRef] [PubMed]
- Zeng, Y.; Yi, R.; Cullen, B.R. Recognition and cleavage of primary microRNA precursors by the nuclear processing enzyme drosha. EMBO J. 2005, 24, 138–148. [Google Scholar] [CrossRef]
- Auyeung, V.C.; Ulitsky, I.; McGeary, S.E.; Bartel, D.P. Beyond secondary structure: Primary-sequence determinants license pri-miRNA hairpins for processing. Cell 2013, 152, 844–858. [Google Scholar] [CrossRef]
- Fang, W.; Bartel, D.P. The menu of features that define primary microRNAs and enable de novo design of microRNA genes. Mol. Cell 2015, 60, 131–145. [Google Scholar] [CrossRef]
- Michlewski, G.; Cáceres, J.F. Post-transcriptional control of miRNA biogenesis. RNA 2019, 25, 1–16. [Google Scholar] [CrossRef]
- Bohnsack, M.T.; Czaplinski, K.; Gorlich, D. Exportin 5 is a rangtp-dependent dsRNA-binding protein that mediates nuclear export of pre-miRNAs. RNA 2004, 10, 185–191. [Google Scholar] [CrossRef] [PubMed]
- Yi, R.; Qin, Y.; Macara, I.G.; Cullen, B.R. Exportin-5 mediates the nuclear export of pre-microRNAs and short hairpin RNAs. Genes Dev. 2003, 17, 3011–3016. [Google Scholar] [CrossRef] [PubMed]
- Bernstein, E.; Caudy, A.A.; Hammond, S.M.; Hannon, G.J. Role for a bidentate ribonuclease in the initiation step of RNA interference. Nature 2001, 409, 363–366. [Google Scholar] [CrossRef] [PubMed]
- Hutvágner, G.; McLachlan, J.; Pasquinelli, A.E.; Bálint, E.; Tuschl, T.; Zamore, P.D. A cellular function for the RNA-interference enzyme dicer in the maturation of the let-7 small temporal RNA. Science 2001, 293, 834–838. [Google Scholar] [CrossRef]
- Ketting, R.F.; Fischer, S.E.; Bernstein, E.; Sijen, T.; Hannon, G.J.; Plasterk, R.H. Dicer functions in RNA interference and in synthesis of small RNA involved in developmental timing in c. Elegans. Genes Dev. 2001, 15, 2654–2659. [Google Scholar] [CrossRef] [PubMed]
- Zhang, H.; Kolb, F.A.; Jaskiewicz, L.; Westhof, E.; Filipowicz, W. Single processing center models for human dicer and bacterial RNAse iii. Cell 2004, 118, 57–68. [Google Scholar] [CrossRef]
- Iwasaki, S.; Kobayashi, M.; Yoda, M.; Sakaguchi, Y.; Katsuma, S.; Suzuki, T.; Tomari, Y. Hsc70/hsp90 chaperone machinery mediates atp-dependent risc loading of small RNA duplexes. Mol. Cell 2010, 39, 292–299. [Google Scholar] [CrossRef] [PubMed]
- Khvorova, A.; Reynolds, A.; Jayasena, S.D. Functional siRNAs and miRNAs exhibit strand bias. Cell 2003, 115, 209–216. [Google Scholar] [CrossRef]
- Schwarz, D.S.; Hutvágner, G.; Du, T.; Xu, Z.; Aronin, N.; Zamore, P.D. Asymmetry in the assembly of the RNAi enzyme complex. Cell 2003, 115, 199–208. [Google Scholar] [CrossRef]
- Dueck, A.; Ziegler, C.; Eichner, A.; Berezikov, E.; Meister, G. MicroRNAs associated with the different human argonaute proteins. Nucleic Acids Res. 2012, 40, 9850–9862. [Google Scholar] [CrossRef]
- Su, H.; Trombly, M.I.; Chen, J.; Wang, X. Essential and overlapping functions for mammalian argonautes in microRNA silencing. Genes Dev. 2009, 23, 304–317. [Google Scholar] [CrossRef]
- Liu, J.; Carmell, M.A.; Rivas, F.V.; Marsden, C.G.; Thomson, J.M.; Song, J.-J.; Hammond, S.M.; Joshua-Tor, L.; Hannon, G.J. Argonaute2 is the catalytic engine of mammalian RNAi. Science 2004, 305, 1437–1441. [Google Scholar] [CrossRef]
- Meister, G.; Landthaler, M.; Patkaniowska, A.; Dorsett, Y.; Teng, G.; Tuschl, T. Human argonaute2 mediates RNA cleavage targeted by miRNAs and siRNAs. Mol. Cell 2004, 15, 185–197. [Google Scholar] [CrossRef]
- Chendrimada, T.P.; Gregory, R.I.; Kumaraswamy, E.; Norman, J.; Cooch, N.; Nishikura, K.; Shiekhattar, R. Trbp recruits the dicer complex to ago2 for microRNA processing and gene silencing. Nature 2005, 436, 740–744. [Google Scholar] [CrossRef]
- Fukunaga, R.; Han, B.W.; Hung, J.H.; Xu, J.; Weng, Z.; Zamore, P.D. Dicer partner proteins tune the length of mature miRNAs in flies and mammals. Cell 2012, 151, 533–546. [Google Scholar] [CrossRef]
- Gregory, R.I.; Chendrimada, T.P.; Cooch, N.; Shiekhattar, R. Human risc couples microRNA biogenesis and posttranscriptional gene silencing. Cell 2005, 123, 631–640. [Google Scholar] [CrossRef]
- Haase, A.D.; Jaskiewicz, L.; Zhang, H.; Lainé, S.; Sack, R.; Gatignol, A.; Filipowicz, W. Trbp, a regulator of cellular pkr and hiv-1 virus expression, interacts with dicer and functions in RNA silencing. EMBO Rep. 2005, 6, 961–967. [Google Scholar] [CrossRef]
- MacRae, I.J.; Ma, E.; Zhou, M.; Robinson, C.V.; Doudna, J.A. In vitro reconstitution of the human risc-loading complex. Proc. Natl. Acad. Sci. USA 2008, 105, 512–517. [Google Scholar] [CrossRef]
- Maniataki, E.; Mourelatos, Z. A human, atp-independent, risc assembly machine fueled by pre-miRNA. Genes Dev. 2005, 19, 2979–2990. [Google Scholar] [CrossRef]
- Bartel, D.P. MicroRNAs: Target recognition and regulatory functions. Cell 2009, 136, 215–233. [Google Scholar] [CrossRef]
- Huntzinger, E.; Izaurralde, E. Gene silencing by microRNAs: Contributions of translational repression and mRNA decay. Nat. Rev. Genet. 2011, 12, 99–110. [Google Scholar] [CrossRef]
- Jonas, S.; Izaurralde, E. Towards a molecular understanding of microRNA-mediated gene silencing. Nat. Rev. Genet. 2015, 16, 421–433. [Google Scholar] [CrossRef]
- Lewis, B.P.; Shih, I.-h.; Jones-Rhoades, M.W.; Bartel, D.P.; Burge, C.B. Prediction of mammalian microRNA targets. Cell 2003, 115, 787–798. [Google Scholar] [CrossRef]
- Okamura, K.; Hagen, J.W.; Duan, H.; Tyler, D.M.; Lai, E.C. The mirtron pathway generates microRNA-class regulatory RNAs in drosophila. Cell 2007, 130, 89–100. [Google Scholar] [CrossRef]
- Ruby, J.G.; Jan, C.H.; Bartel, D.P. Intronic microRNA precursors that bypass drosha processing. Nature 2007, 448, 83–86. [Google Scholar] [CrossRef]
- Westholm, J.O.; Lai, E.C. Mirtrons: MicroRNA biogenesis via splicing. Biochimie 2011, 93, 1897–1904. [Google Scholar] [CrossRef]
- Cheloufi, S.; Dos Santos, C.O.; Chong, M.M.W.; Hannon, G.J. A dicer-independent miRNA biogenesis pathway that requires ago catalysis. Nature 2010, 465, 584–589. [Google Scholar] [CrossRef]
- Cifuentes, D.; Xue, H.; Taylor, D.W.; Patnode, H.; Mishima, Y.; Cheloufi, S.; Ma, E.; Mane, S.; Hannon, G.J.; Lawson, N.D.; et al. A novel miRNA processing pathway independent of dicer requires argonaute2 catalytic activity. Science 2010, 328, 1694–1698. [Google Scholar] [CrossRef]
- Yang, J.-S.; Maurin, T.; Robine, N.; Rasmussen, K.D.; Jeffrey, K.L.; Chandwani, R.; Papapetrou, E.P.; Sadelain, M.; O’Carroll, D.; Lai, E.C. Conserved vertebrate mir-451 provides a platform for dicer-independent, ago2-mediated microRNA biogenesis. Proc. Natl. Acad. Sci. USA 2010, 107, 15163–15168. [Google Scholar] [CrossRef]
- Daugaard, I.; Hansen, T.B. Biogenesis and function of ago-associated RNAs. Trends Genet. 2017, 33. [Google Scholar] [CrossRef]
- Hansen, T.B.; Venø, M.T.; Jensen, T.I.; Schaefer, A.; Damgaard, C.K.; Kjems, J. Argonaute-associated short introns are a novel class of gene regulators. Nat. Commun. 2016, 7, 11538. [Google Scholar] [CrossRef]
- Babiarz, J.E.; Ruby, J.G.; Wang, Y.; Bartel, D.P.; Blelloch, R. Mouse es cells express endogenous shRNAs, siRNAs, and other microprocessor-independent, dicer-dependent small RNAs. Genes Dev. 2008, 22, 2773–2785. [Google Scholar] [CrossRef]
- Chung, W.-J.; Okamura, K.; Martin, R.; Lai, E.C. Endogenous RNA interference provides a somatic defense against drosophila transposons. Curr. Biol. 2008, 18, 795–802. [Google Scholar] [CrossRef]
- Czech, B.; Malone, C.D.; Zhou, R.; Stark, A.; Schlingeheyde, C.; Dus, M.; Perrimon, N.; Kellis, M.; Wohlschlegel, J.A.; Sachidanandam, R.; et al. An endogenous small interfering RNA pathway in drosophila. Nature 2008, 453, 798–802. [Google Scholar] [CrossRef]
- Song, J.-J.; Smith, S.K.; Hannon, G.J.; Joshua-Tor, L. Crystal structure of argonaute and its implications for risc slicer activity. Science 2004, 305, 1434–1437. [Google Scholar] [CrossRef]
- Tam, O.H.; Aravin, A.A.; Stein, P.; Girard, A.; Murchison, E.P.; Cheloufi, S.; Hodges, E.; Anger, M.; Sachidanandam, R.; Schultz, R.M.; et al. Pseudogene-derived small interfering RNAs regulate gene expression in mouse oocytes. Nature 2008, 453, 534–538. [Google Scholar] [CrossRef]
- Watanabe, T.; Totoki, Y.; Toyoda, A.; Kaneda, M.; Kuramochi-Miyagawa, S.; Obata, Y.; Chiba, H.; Kohara, Y.; Kono, T.; Nakano, T.; et al. Endogenous siRNAs from naturally formed dsRNAs regulate transcripts in mouse oocytes. Nature 2008, 453, 539–543. [Google Scholar] [CrossRef]
- Gu, W.; Shirayama, M.; Conte, D.; Vasale, J.; Batista, P.J.; Claycomb, J.M.; Moresco, J.J.; Youngman, E.M.; Keys, J.; Stoltz, M.J.; et al. Distinct argonaute-mediated 22g-RNA pathways direct genome surveillance in the c. Elegans germline. Mol. Cell 2009, 36, 231–244. [Google Scholar] [CrossRef]
- Pak, J.; Fire, A. Distinct populations of primary and secondary effectors during RNAi in c. Elegans. Science 2007, 315, 241–244. [Google Scholar] [CrossRef]
- Sjien, T.; Steiner, F.A.; Thijssen, K.L.; Plasterk, R.H.A.; Sijen, T.; Steiner, F.A.; Thijssen, K.L.; Plasterk, R.H.A. Secondary siRNAs result from form a distinct class. Science 2007, 315, 2005–2008. [Google Scholar]
- Ghildiyal, M.; Seitz, H.; Horwich, M.D.; Li, C.; Du, T.; Lee, S.; Xu, J.; Kittler, E.L.W.; Zapp, M.L.; Weng, Z.; et al. Endogenous siRNAs derived from transposons and mRNAs in drosophila somatic cells. Science 2008, 320, 1077–1081. [Google Scholar] [CrossRef] [PubMed]
- Chen, L.-L.; Yang, L.; Carmichael, G.G. Molecular basis for an attenuated cytoplasmic dsRNA response in human embryonic stem cells. Cell Cycle 2010, 9, 3552–3564. [Google Scholar] [CrossRef]
- Nejepinska, J.; Malik, R.; Filkowski, J.; Flemr, M.; Filipowicz, W.; Svoboda, P. DsRNA expression in the mouse elicits RNAi in oocytes and low adenosine deamination in somatic cells. Nucleic Acids Res. 2012, 40, 399–413. [Google Scholar] [CrossRef] [PubMed]
- Stein, P.; Zeng, F.; Pan, H.; Schultz, R.M. Absence of non-specific effects of RNA interference triggered by long double-stranded RNA in mouse oocytes. Dev. Biol. 2005, 286, 464–471. [Google Scholar] [CrossRef]
- Wang, R.; Wang, J.; Paul, A.M.; Acharya, D.; Bai, F.; Huang, F.; Guo, Y.-L. Mouse embryonic stem cells are deficient in type i interferon expression in response to viral infections and double-stranded RNA. J. Biol. Chem. 2013, 288, 15926–15936. [Google Scholar] [CrossRef] [PubMed]
- Lee, D.; Shin, C. Emerging roles of drosha beyond primary microRNA processing. RNA Biol. 2018, 15, 186–193. [Google Scholar] [CrossRef]
- Macias, S.; Cordiner, R.A.; Cáceres, J.F. Cellular functions of the microprocessor. Biochem. Soc. Trans. 2013, 41, 838–843. [Google Scholar] [CrossRef]
- Macias, S.; Plass, M.; Stajuda, A.; Michlewski, G.; Eyras, E.; Cáceres, J.F. Dgcr8 hits-clip reveals novel functions for the microprocessor. Nat. Struct. Mol. Biol. 2012, 19, 760–766. [Google Scholar] [CrossRef]
- Han, J.; Pedersen, J.S.; Kwon, S.C.; Belair, C.D.; Kim, Y.-K.; Yeom, K.-H.; Yang, W.-Y.; Haussler, D.; Blelloch, R.; Kim, V.N. Posttranscriptional crossregulation between drosha and dgcr8. Cell 2009, 136, 75–84. [Google Scholar] [CrossRef]
- Kadener, S.; Rodriguez, J.; Abruzzi, K.C.; Khodor, Y.L.; Sugino, K.; Marr, M.T.; Nelson, S.; Rosbash, M. Genome-wide identification of targets of the drosha-pasha/dgcr8 complex. RNA 2009, 15, 537–545. [Google Scholar] [CrossRef]
- Triboulet, R.; Chang, H.-M.; LaPierre, R.J.; Gregory, R.I. Post-transcriptional control of dgcr8 expression by the microprocessor. RNA 2009, 15, 1005–1011. [Google Scholar] [CrossRef]
- Heras, S.R.; Macias, S.; Plass, M.; Fernandez, N.; Cano, D.; Eyras, E.; Garcia-Perez, J.L.; Cáceres, J.F. The microprocessor controls the activity of mammalian retrotransposons. Nat. Struct. Mol. Biol. 2013, 20, 1173–1181. [Google Scholar] [CrossRef] [PubMed]
- Rouha, H.; Thurner, C.; Mandl, C.W. Functional microRNA generated from a cytoplasmic RNA virus. Nucleic Acids Res. 2010, 38, 8328–8337. [Google Scholar] [CrossRef] [PubMed]
- Shapiro, J.S.; Langlois, R.A.; Pham, A.M.; Tenoever, B.R. Evidence for a cytoplasmic microprocessor of pri-miRNAs. RNA 2012, 18, 1338–1346. [Google Scholar] [CrossRef]
- Shapiro, J.S.; Schmid, S.; Aguado, L.C.; Sabin, L.R.; Yasunaga, A.; Shim, J.V.; Sachs, D.; Cherry, S.; TenOever, B.R. Drosha as an interferon-independent antiviral factor. Proc. Natl. Acad. Sci. USA 2014, 111, 7108–7113. [Google Scholar] [CrossRef]
- Shapiro, J.S.; Varble, A.; Pham, A.M.; Tenoever, B.R. Noncanonical cytoplasmic processing of viral microRNAs. RNA 2010, 16, 2068–2074. [Google Scholar] [CrossRef] [PubMed]
- Varble, A.; Chua, M.A.; Perez, J.T.; Manicassamy, B.; Garcia-Sastre, A.; tenOever, B.R. Engineered RNA viral synthesis of microRNAs. Proc. Natl. Acad. Sci. USA 2010, 107, 11519–11524. [Google Scholar] [CrossRef] [PubMed]
- Park, H.H.; Triboulet, R.; Bentler, M.; Guda, S.; Du, P.; Xu, H.; Gregory, R.I.; Brendel, C.; Williams, D.A. Drosha knockout leads to enhancement of viral titers for vectors encoding miRNA-adapted shRNAs. Mol. Ther. Nucleic Acids 2018, 12, 591–599. [Google Scholar] [CrossRef] [PubMed]
- Aguado, L.C.; Schmid, S.; May, J.; Sabin, L.R.; Panis, M.; Blanco-Melo, D.; Shim, J.V.; Sachs, D.; Cherry, S.; Simon, A.E.; et al. Rnase iii nucleases from diverse kingdoms serve as antiviral effectors. Nature 2017, 547, 114–117. [Google Scholar] [CrossRef]
- Feldman, E.R.; Kara, M.; Coleman, C.B.; Grau, K.R.; Oko, L.M.; Krueger, B.J.; Renne, R.; van Dyk, L.F.; Tibbetts, S.A. Virus-encoded microRNAs facilitate gammaherpesvirus latency and pathogenesis in vivo. MBio 2014, 5, e00981-14. [Google Scholar] [CrossRef]
- Guo, Y.E.; Oei, T.; Steitz, J.A. Herpesvirus saimiri microRNAs preferentially target host cell cycle regulators. J. Virol. 2015, 89, 10901–10911. [Google Scholar] [CrossRef] [PubMed]
- Lo, A.K.; To, K.F.; Lo, K.W.; Lung, R.W.; Hui, J.W.; Liao, G.; Hayward, S.D. Modulation of lmp1 protein expression by ebv-encoded microRNAs. Proc. Natl. Acad. Sci. USA 2007, 104, 16164–16169. [Google Scholar] [CrossRef] [PubMed]
- Pfeffer, S.; Zavolan, M.; Grasser, F.A.; Chien, M.; Russo, J.J.; Ju, J.; John, B.; Enright, A.J.; Marks, D.; Sander, C.; et al. Identification of virus-encoded microRNAs. Science 2004, 304, 734–736. [Google Scholar] [CrossRef] [PubMed]
- Riley, K.J.; Rabinowitz, G.S.; Steitz, J.A. Comprehensive analysis of rhesus lymphocryptovirus microRNA expression. J. Virol. 2010, 84, 5148–5157. [Google Scholar] [CrossRef] [PubMed]
- Umbach, J.L.; Kramer, M.F.; Jurak, I.; Karnowski, H.W.; Coen, D.M.; Cullen, B.R. MicroRNAs expressed by herpes simplex virus 1 during latent infection regulate viral mRNAs. Nature 2008, 454, 780–783. [Google Scholar] [CrossRef] [PubMed]
- Bogerd, H.P.; Karnowski, H.W.; Cai, X.; Shin, J.; Pohlers, M.; Cullen, B.R. A mammalian herpesvirus uses noncanonical expression and processing mechanisms to generate viral microRNAs. Mol. Cell 2010, 37, 135–142. [Google Scholar] [CrossRef]
- Cazalla, D.; Xie, M.; Steitz, J.A. A primate herpesvirus uses the integrator complex to generate viral microRNAs. Mol. Cell 2011, 43, 982–992. [Google Scholar] [CrossRef]
- Lin, Y.T.; Sullivan, C.S. Expanding the role of drosha to the regulation of viral gene expression. Proc. Natl. Acad. Sci. USA 2011, 108, 11229–11234. [Google Scholar] [CrossRef]
- Xing, L.; Kieff, E. Epstein-barr virus bhrf1 micro- and stable RNAs during latency iii and after induction of replication. J. Virol. 2007, 81, 9967–9975. [Google Scholar] [CrossRef]
- Dang, Y.; Yang, Q.; Xue, Z.; Liu, Y. Rna interference in fungi: Pathways, functions, and applications. Eukaryot. Cell 2011, 10, 1148–1155. [Google Scholar] [CrossRef]
- Lee, Y.S.; Nakahara, K.; Pham, J.W.; Kim, K.; He, Z.; Sontheimer, E.J.; Carthew, R.W. Distinct roles for drosophila dicer-1 and dicer-2 in the siRNA/miRNA silencing pathways. Cell 2004, 117, 69–81. [Google Scholar] [CrossRef]
- Margis, R.; Fusaro, A.F.; Smith, N.A.; Curtin, S.J.; Watson, J.M.; Finnegan, E.J.; Waterhouse, P.M. The evolution and diversification of dicers in plants. FEBS Lett. 2006, 580, 2442–2450. [Google Scholar] [CrossRef]
- Backes, S.; Langlois, R.A.; Schmid, S.; Varble, A.; Shim, J.V.; Sachs, D.; tenOever, B.R. The mammalian response to virus infection is independent of small RNA silencing. Cell Rep. 2014, 8, 114–125. [Google Scholar] [CrossRef]
- Bogerd, H.P.; Skalsky, R.L.; Kennedy, E.M.; Furuse, Y.; Whisnant, A.W.; Flores, O.; Schultz, K.L.; Putnam, N.; Barrows, N.J.; Sherry, B.; et al. Replication of many human viruses is refractory to inhibition by endogenous cellular microRNAs. J. Virol. 2014, 88, 8065–8076. [Google Scholar] [CrossRef]
- Girardi, E.; Chane-Woon-Ming, B.; Messmer, M.; Kaukinen, P.; Pfeffer, S. Identification of RNAse l-dependent, 3’-end-modified, viral small RNAs in sindbis virus-infected mammalian cells. MBio 2013, 4, e00698-13. [Google Scholar] [CrossRef]
- Kennedy, E.M.; Whisnant, A.W.; Kornepati, A.V.; Marshall, J.B.; Bogerd, H.P.; Cullen, B.R. Production of functional small interfering RNAs by an amino-terminal deletion mutant of human dicer. Proc. Natl. Acad. Sci. USA 2015, 112, E6945–E6954. [Google Scholar] [CrossRef]
- Parameswaran, P.; Sklan, E.; Wilkins, C.; Burgon, T.; Samuel, M.A.; Lu, R.; Ansel, K.M.; Heissmeyer, V.; Einav, S.; Jackson, W.; et al. Six RNA viruses and forty-one hosts: Viral small RNAs and modulation of small RNA repertoires in vertebrate and invertebrate systems. PLoS Pathog 2010, 6, e1000764. [Google Scholar] [CrossRef]
- Qiu, Y.; Xu, Y.; Zhang, Y.; Zhou, H.; Deng, Y.Q.; Li, X.F.; Miao, M.; Zhang, Q.; Zhong, B.; Hu, Y.; et al. Human virus-derived small RNAs can confer antiviral immunity in mammals. Immunity 2017, 46, 992–1004. [Google Scholar] [CrossRef]
- Li, Y.; Lu, J.; Han, Y.; Fan, X.; Ding, S.-W.W. Rna interference functions as an antiviral immunity mechanism in mammals. Science 2013, 342, 231–234. [Google Scholar] [CrossRef]
- Maillard, P.V.; Ciaudo, C.; Marchais, A.; Li, Y.; Jay, F.; Ding, S.W.; Voinnet, O. Antiviral RNA interference in mammalian cells. Science 2013, 342, 235–238. [Google Scholar] [CrossRef]
- Rossi, J.J. Mammalian dicer finds a partner. EMBO Rep. 2005, 6, 927–929. [Google Scholar] [CrossRef] [PubMed]
- Provost, P.; Dishart, D.; Doucet, J.; Frendewey, D.; Samuelsson, B.; Rådmark, O. Ribonuclease activity and RNA binding of recombinant human dicer. EMBO J. 2002, 21, 5864–5874. [Google Scholar] [CrossRef] [PubMed]
- Girardi, E.; Lefèvre, M.; Chane-Woon-Ming, B.; Paro, S.; Claydon, B.; Imler, J.-L.; Meignin, C.; Pfeffer, S. Cross-species comparative analysis of dicer proteins during sindbis virus infection. Sci. Rep. 2015, 5, 10693. [Google Scholar] [CrossRef] [PubMed]
- Ma, E.; MacRae, I.J.; Kirsch, J.F.; Doudna, J.A. Autoinhibition of human dicer by its internal helicase domain. J. Mol. Biol 2008, 380, 237–243. [Google Scholar] [CrossRef] [PubMed]
- Flemr, M.; Malik, R.; Franke, V.; Nejepinska, J.; Sedlacek, R.; Vlahovicek, K.; Svoboda, P. A retrotransposon-driven dicer isoform directs endogenous small interfering RNA production in mouse oocytes. Cell 2013, 155, 807–816. [Google Scholar] [CrossRef] [PubMed]
- Cui, L.; Wang, H.; Ji, Y.; Yang, J.; Xu, S.; Huang, X.; Wang, Z.; Qin, L.; Tien, P.; Zhou, X.; et al. The nucleocapsid protein of coronaviruses acts as a viral suppressor of RNA silencing in mammalian cells. J. Virol 2015, 89, 9029–9043. [Google Scholar] [CrossRef] [PubMed]
- Haasnoot, J.; de Vries, W.; Geutjes, E.J.; Prins, M.; de Haan, P.; Berkhout, B. The ebola virus vp35 protein is a suppressor of RNA silencing. PLoS Pathog 2007, 3, e86. [Google Scholar] [CrossRef]
- Li, Y.; Basavappa, M.; Lu, J.; Dong, S.; Cronkite, D.A.; Prior, J.T.; Reinecker, H.C.; Hertzog, P.; Han, Y.; Li, W.X.; et al. Induction and suppression of antiviral RNA interference by influenza a virus in mammalian cells. Nat. Microbiol 2016, 2, 16250. [Google Scholar] [CrossRef]
- Andersson, M.G.; Haasnoot, P.C.J.; Xu, N.; Berenjian, S.; Berkhout, B.; Akusjärvi, G. Suppression of RNA interference by adenovirus virus-associated RNA. J. Virol. 2005, 79, 9556–9565. [Google Scholar] [CrossRef]
- Lu, S.; Cullen, B.R. Adenovirus va1 noncoding RNA can inhibit small interfering RNA and microRNA biogenesis. J. Virol. 2004, 78, 12868–12876. [Google Scholar] [CrossRef]
- Kitajewski, J.; Schneider, R.J.; Safer, B.; Munemitsu, S.M.; Samuel, C.E.; Thimmappaya, B.; Shenk, T. Adenovirus vai RNA antagonizes the antiviral action of interferon by preventing activation of the interferon-induced eif-2 alpha kinase. Cell 1986, 45, 195–200. [Google Scholar] [CrossRef]
- O’Malley, R.P.; Mariano, T.M.; Siekierka, J.; Mathews, M.B. A mechanism for the control of protein synthesis by adenovirus va RNAi. Cell 1986, 44, 391–400. [Google Scholar] [CrossRef]
- Machitani, M.; Sakurai, F.; Wakabayashi, K.; Tomita, K.; Tachibana, M.; Mizuguchi, H. Dicer functions as an antiviral system against human adenoviruses via cleavage of adenovirus-encoded noncoding RNA. Sci Rep. 2016, 6, 27598. [Google Scholar] [CrossRef]
- Ablasser, A.; Gulen, M.F. The role of cgas in innate immunity and beyond. J. Mol. Med. 2016, 94, 1085–1093. [Google Scholar] [CrossRef]
- Lee-Kirsch, M.A. The type i interferonopathies. Annu Rev. Med. 2017, 68, 297–315. [Google Scholar] [CrossRef]
- Forster, S.C.; Tate, M.D.; Hertzog, P.J. MicroRNA as type i interferon-regulated transcripts and modulators of the innate immune response. Front. Immunol 2015, 6, 334. [Google Scholar] [CrossRef]
- Li, Y.; Shi, X. MicroRNAs in the regulation of tlr and rig-i pathways. Cell Mol. Immunol 2013, 10, 65–71. [Google Scholar] [CrossRef]
- Momen-Heravi, F.; Bala, S. MiRNA regulation of innate immunity. J. Leukoc Biol 2018. [Google Scholar] [CrossRef]
- Chiappinelli, K.B.; Haynes, B.C.; Brent, M.R.; Goodfellow, P.J. Reduced dicer1 elicits an interferon response in endometrial cancer cells. Mol. Cancer Res. 2012, 10, 316–325. [Google Scholar] [CrossRef]
- Papadopoulou, A.S.; Dooley, J.; Linterman, M.A.; Pierson, W.; Ucar, O.; Kyewski, B.; Zuklys, S.; Hollander, G.A.; Matthys, P.; Gray, D.H.D.; et al. The thymic epithelial microRNA network elevates the threshold for infection-associated thymic involution via mir-29a mediated suppression of the ifn-α receptor. Nat. Immunol. 2012, 13, 181–187. [Google Scholar] [CrossRef]
- Varol, D.; Mildner, A.; Blank, T.; Maggio, N.; Prinz, M.; Jung, S. Dicer deficiency differentially impacts microglia of the developing and adult brain. Immunity 2017, 46, 1030–1044. [Google Scholar] [CrossRef]
- Seo, G.J.; Kincaid, R.P.; Phanaksri, T.; Burke, J.M.; Pare, J.M.; Cox, J.E.; Hsiang, T.Y.; Krug, R.M.; Sullivan, C.S. Reciprocal inhibition between intracellular antiviral signaling and the RNAi machinery in mammalian cells. Cell Host Microbe 2013, 14, 435–445. [Google Scholar] [CrossRef]
- O’Connell, R.M.; Taganov, K.D.; Boldin, M.P.; Cheng, G.; Baltimore, D. MicroRNA-155 is induced during the macrophage inflammatory response. Proc. Natl. Acad. Sci. USA 2007, 104, 1604–1609. [Google Scholar] [CrossRef]
- Taganov, K.D.; Boldin, M.P.; Chang, K.J.; Baltimore, D. Nf-kappab-dependent induction of microRNA mir-146, an inhibitor targeted to signaling proteins of innate immune responses. Proc. Natl. Acad. Sci. USA 2006, 103, 12481–12486. [Google Scholar] [CrossRef]
- Chen, Y.; Chen, J.; Wang, H.; Shi, J.; Wu, K.; Liu, S.; Liu, Y.; Wu, J. Hcv-induced mir-21 contributes to evasion of host immune system by targeting myd88 and irak1. PLoS Pathog 2013, 9, e1003248. [Google Scholar] [CrossRef]
- Hou, J.; Wang, P.; Lin, L.; Liu, X.; Ma, F.; An, H.; Wang, Z.; Cao, X. MicroRNA-146a feedback inhibits rig-i-dependent type i ifn production in macrophages by targeting traf6, irak1, and irak2. J. Immunol 2009, 183, 2150–2158. [Google Scholar] [CrossRef]
- Tili, E.; Michaille, J.J.; Cimino, A.; Costinean, S.; Dumitru, C.D.; Adair, B.; Fabbri, M.; Alder, H.; Liu, C.G.; Calin, G.A.; et al. Modulation of mir-155 and mir-125b levels following lipopolysaccharide/tnf-alpha stimulation and their possible roles in regulating the response to endotoxin shock. J. Immunol 2007, 179, 5082–5089. [Google Scholar] [CrossRef]
- Witwer, K.W.; Sisk, J.M.; Gama, L.; Clements, J.E. MicroRNA regulation of ifn-beta protein expression: Rapid and sensitive modulation of the innate immune response. J. Immunol. 2010, 184, 2369–2376. [Google Scholar] [CrossRef]
- Witteveldt, J.; Ivens, A.; Macias, S. Inhibition of microprocessor function during the activation of the type i interferon response. Cell Rep. 2018, 23, 3275–3285. [Google Scholar] [CrossRef]
- Wiesen, J.L.; Tomasi, T.B. Dicer is regulated by cellular stresses and interferons. Mol. Immunol. 2009, 46, 1222–1228. [Google Scholar] [CrossRef]
- Machitani, M.; Sakurai, F.; Wakabayashi, K.; Takayama, K.; Tachibana, M.; Mizuguchi, H. Type i interferons impede short hairpin RNA-mediated RNAi via inhibition of dicer-mediated processing to small interfering RNA. Mol. Ther. Nucleic Acids 2017, 6, 173–182. [Google Scholar] [CrossRef]
- van der Veen, A.G.; Maillard, P.V.; Schmidt, J.M.; Lee, S.A.; Deddouche-Grass, S.; Borg, A.; Kjaer, S.; Snijders, A.P.; Reis e Sousa, C. The rig-i-like receptor lgp2 inhibits dicer-dependent processing of long double-stranded RNA and blocks RNA interference in mammalian cells. EMBO J. 2018, 37. [Google Scholar] [CrossRef]
- Ota, H.; Sakurai, M.; Gupta, R.; Valente, L.; Wulff, B.E.; Ariyoshi, K.; Iizasa, H.; Davuluri, R.V.; Nishikura, K. Adar1 forms a complex with dicer to promote microRNA processing and RNA-induced gene silencing. Cell 2013, 153, 575–589. [Google Scholar] [CrossRef]
- Kawahara, Y.; Zinshteyn, B.; Chendrimada, T.P.; Shiekhattar, R.; Nishikura, K. Rna editing of the microRNA-151 precursor blocks cleavage by the dicer-trbp complex. EMBO Rep. 2007, 8, 763–769. [Google Scholar] [CrossRef]
- Yang, W.; Chendrimada, T.P.; Wang, Q.; Higuchi, M.; Seeburg, P.H.; Shiekhattar, R.; Nishikura, K. Modulation of microRNA processing and expression through RNA editing by adar deaminases. Nat. Struct Mol. Biol 2006, 13, 13–21. [Google Scholar] [CrossRef]
- Patel, R.C.; Sen, G.C. Pact, a protein activator of the interferon-induced protein kinase, pkr. EMBO J. 1998, 17, 4379–4390. [Google Scholar] [CrossRef]
- Park, H.; Davies, M.V.; Langland, J.O.; Chang, H.W.; Nam, Y.S.; Tartaglia, J.; Paoletti, E.; Jacobs, B.L.; Kaufman, R.J.; Venkatesan, S. Tar RNA-binding protein is an inhibitor of the interferon-induced protein kinase pkr. Proc. Natl. Acad. Sci. USA 1994, 91, 4713–4717. [Google Scholar] [CrossRef]
- Sanghvi, V.R.; Steel, L.F. The cellular tar RNA binding protein, trbp, promotes hiv-1 replication primarily by inhibiting the activation of double-stranded RNA-dependent kinase pkr. J. Virol. 2011, 85, 12614–12621. [Google Scholar] [CrossRef]
- Chakravarthy, S.; Sternberg, S.H.; Kellenberger, C.A.; Doudna, J.A. Substrate-specific kinetics of dicer-catalyzed RNA processing. J. Mol. Biol. 2010, 404, 392–402. [Google Scholar] [CrossRef]
- Kok, K.H.; Ng, M.-H.J.; Ching, Y.-P.; Jin, D.-Y. Human trbp and pact directly interact with each other and associate with dicer to facilitate the production of small interfering RNA. J. Biol. Chem. 2007, 282, 17649–17657. [Google Scholar] [CrossRef]
- Lee, H.Y.; Zhou, K.; Smith, A.M.; Noland, C.L.; Doudna, J.A. Differential roles of human dicer-binding proteins trbp and pact in small RNA processing. Nucleic Acids Res. 2013, 41, 6568–6576. [Google Scholar] [CrossRef]
- Kok, K.-H.; Lui, P.-Y.; Ng, M.-H.J.; Siu, K.-L.; Au, S.W.N.; Jin, D.-Y. The double-stranded RNA-binding protein pact functions as a cellular activator of rig-i to facilitate innate antiviral response. Cell Host Microbe 2011, 9, 299–309. [Google Scholar] [CrossRef]
- Saunders, L.R.; Barber, G.N. The dsRNA binding protein family: Critical roles, diverse cellular functions. FASEB J. 2003, 17, 961–983. [Google Scholar] [CrossRef]
- Kolakofsky, D.; Kowalinski, E.; Cusack, S. A structure-based model of rig-i activation. RNA 2012, 18, 2118–2127. [Google Scholar] [CrossRef]
- Rawling, D.C.; Pyle, A.M. Parts, assembly and operation of the rig-i family of motors. Curr. Opin. Struct. Biol. 2014, 25, 25–33. [Google Scholar] [CrossRef]
- Burke, D.C.; Graham, C.F.; Lehman, J.M. Appearance of interferon inducibility and sensitivity during differentiation of murine teratocarcinoma cells in vitro. Cell 1978, 13, 243–248. [Google Scholar] [CrossRef]
- Billy, E.; Brondani, V.; Zhang, H.; Muller, U.; Filipowicz, W. Specific interference with gene expression induced by long, double-stranded RNA in mouse embryonal teratocarcinoma cell lines. Proc. Natl. Acad. Sci. USA 2001, 98, 14428–14433. [Google Scholar] [CrossRef]
- Paddison, P.J.; Caudy, A.A.; Hannon, G.J. Stable suppression of gene expression by RNAi in mammalian cells. Proc. Natl. Acad. Sci. USA 2002, 99, 1443–1448. [Google Scholar] [CrossRef]
- Maillard, P.V.; Van der Veen, A.G.; Deddouche-Grass, S.; Rogers, N.C.; Merits, A.; e Sousa, C.R. Inactivation of the type i interferon pathway reveals long double-stranded RNA-mediated RNA interference in mammalian cells. EMBO J. 2016, 35, 2505–2518. [Google Scholar] [CrossRef]
- Hou, W.; Tian, Q.; Zheng, J.; Bonkovsky, H.L. MicroRNA-196 represses bach1 protein and hepatitis c virus gene expression in human hepatoma cells expressing hepatitis c viral proteins. Hepatology 2010, 51, 1494–1504. [Google Scholar] [CrossRef]
- Murakami, Y.; Aly, H.H.; Tajima, A.; Inoue, I.; Shimotohno, K. Regulation of the hepatitis c virus genome replication by mir-199a. J. Hepatol 2009, 50, 453–460. [Google Scholar] [CrossRef]
- Pedersen, I.M.; Cheng, G.; Wieland, S.; Volinia, S.; Croce, C.M.; Chisari, F.V.; David, M. Interferon modulation of cellular microRNAs as an antiviral mechanism. Nature 2007, 449, 919–922. [Google Scholar] [CrossRef]
- Russo, A.; Potenza, N. Antiviral effects of human microRNAs and conservation of their target sites. FEBS Lett 2011, 585, 2551–2555. [Google Scholar] [CrossRef]
- Swaminathan, G.; Martin-Garcia, J.; Navas-Martin, S. Rna viruses and microRNAs: Challenging discoveries for the 21st century. Physiol Genom. 2013, 45, 1035–1048. [Google Scholar] [CrossRef]
- Lecellier, C.H.; Dunoyer, P.; Arar, K.; Lehmann-Che, J.; Eyquem, S.; Himber, C.; Saib, A.; Voinnet, O. A cellular microRNA mediates antiviral defense in human cells. Science 2005, 308, 557–560. [Google Scholar] [CrossRef]
- Bennasser, Y.; Le, S.-Y.; Benkirane, M.; Jeang, K.-T. Evidence that hiv-1 encodes an siRNA and a suppressor of RNA silencing. Immunity 2005, 22, 607–619. [Google Scholar] [CrossRef]
- Bennasser, Y.; Yeung, M.L.; Jeang, K.T. Hiv-1 tar RNA subverts RNA interference in transfected cells through sequestration of tar RNA-binding protein, trbp. J. Biol Chem 2006, 281, 27674–27678. [Google Scholar] [CrossRef]
- de Vries, W.; Haasnoot, J.; Fouchier, R.; de Haan, P.; Berkhout, B. Differential RNA silencing suppression activity of ns1 proteins from different influenza a virus strains. J. Gen. Virol 2009, 90, 1916–1922. [Google Scholar] [CrossRef]
- Backes, S.; Shapiro, J.S.; Sabin, L.R.; Pham, A.M.; Reyes, I.; Moss, B.; Cherry, S.; tenOever, B.R. Degradation of host microRNAs by poxvirus poly(a) polymerase reveals terminal RNA methylation as a protective antiviral mechanism. Cell Host Microbe 2012, 12, 200–210. [Google Scholar] [CrossRef]
- Jopling, C.L.; Yi, M.; Lancaster, A.M.; Lemon, S.M.; Sarnow, P. Modulation of hepatitis c virus RNA abundance by a liver-specific microRNA. Science 2005, 309, 1577–1581. [Google Scholar] [CrossRef]
- Sedano, C.D.; Sarnow, P. Hepatitis c virus subverts liver-specific mir-122 to protect the viral genome from exoribonuclease xrn2. Cell Host Microbe 2014, 16, 257–264. [Google Scholar] [CrossRef]
- Jopling, C.L. Regulation of hepatitis c virus by microRNA-122. Biochem Soc. Trans. 2008, 36, 1220–1223. [Google Scholar] [CrossRef]
- Wilson, J.A.; Zhang, C.; Huys, A.; Richardson, C.D. Human ago2 is required for efficient microRNA 122 regulation of hepatitis c virus RNA accumulation and translation. J. Virol 2011, 85, 2342–2350. [Google Scholar] [CrossRef]
- Ziv, O.; Gabryelska, M.M.; Lun, A.T.L.; Gebert, L.F.R.; Sheu-Gruttadauria, J.; Meredith, L.W.; Liu, Z.Y.; Kwok, C.K.; Qin, C.F.; MacRae, I.J.; et al. Comrades determines in vivo RNA structures and interactions. Nat. Methods 2018, 15, 785–788. [Google Scholar] [CrossRef]
- Chiang, K.; Liu, H.; Rice, A.P. Mir-132 enhances hiv-1 replication. Virology 2013, 438, 1–4. [Google Scholar] [CrossRef]
- O’Connor, C.M.; Vanicek, J.; Murphy, E.A. Host microRNA regulation of human cytomegalovirus immediate early protein translation promotes viral latency. J. Virol 2014, 88, 5524–5532. [Google Scholar] [CrossRef]
- White, E.; Schlackow, M.; Kamieniarz-Gdula, K.; Proudfoot, N.J.; Gullerova, M. Human nuclear dicer restricts the deleterious accumulation of endogenous double-stranded RNA. Nat. Struct Mol. Biol 2014, 21, 552–559. [Google Scholar] [CrossRef]
- Matskevich, A.A.; Moelling, K. Dicer is involved in protection against influenza a virus infection. J. Gen. Virol. 2007, 88, 2627–2635. [Google Scholar] [CrossRef]
- Bucher, E.; Hemmes, H.; de Haan, P.; Goldbach, R.; Prins, M. The influenza a virus ns1 protein binds small interfering RNAs and suppresses RNA silencing in plants. J. Gen. Virol 2004, 85, 983–991. [Google Scholar] [CrossRef]
- Ma, Y.; Ouyang, J.; Wei, J.; Maarouf, M.; Chen, J.L. Involvement of host non-coding RNAs in the pathogenesis of the influenza virus. Int. J. Mol. Sci 2016, 18, 39. [Google Scholar] [CrossRef]
- Chen, J.S.; Li, H.C.; Lin, S.I.; Yang, C.H.; Chien, W.Y.; Syu, C.L.; Lo, S.Y. Cleavage of dicer protein by i7 protease during vaccinia virus infection. PLoS ONE 2015, 10, e0120390. [Google Scholar] [CrossRef]
- Bernstein, E.; Kim, S.Y.; Carmell, M.A.; Murchison, E.P.; Alcorn, H.; Li, M.Z.; Mills, A.A.; Elledge, S.J.; Anderson, K.V.; Hannon, G.J. Dicer is essential for mouse development. Nat. Genet. 2003, 35, 215–217. [Google Scholar] [CrossRef]
- Otsuka, M.; Jing, Q.; Georgel, P.; New, L.; Chen, J.; Mols, J.; Kang, Y.J.; Jiang, Z.; Du, X.; Cook, R.; et al. Hypersusceptibility to vesicular stomatitis virus infection in dicer1-deficient mice is due to impaired mir24 and mir93 expression. Immunity 2007, 27, 123–134. [Google Scholar] [CrossRef]
- Ostermann, E.; Tuddenham, L.; Macquin, C.; Alsaleh, G.; Schreiber-Becker, J.; Tanguy, M.; Bahram, S.; Pfeffer, S.; Georgel, P. Deregulation of type i ifn-dependent genes correlates with increased susceptibility to cytomegalovirus acute infection of dicer mutant mice. PLoS ONE 2012, 7, e43744. [Google Scholar] [CrossRef]
- Aguado, L.C.; Schmid, S.; Sachs, D.; Shim, J.V.; Lim, J.K.; tenOever, B.R. MicroRNA function is limited to cytokine control in the acute response to virus infection. Cell Host Microbe 2015, 18, 714–722. [Google Scholar] [CrossRef]
- Bogerd, H.P.; Whisnant, A.W.; Kennedy, E.M.; Flores, O.; Cullen, B.R. Derivation and characterization of dicer- and microRNA-deficient human cells. RNA 2014, 20, 923–937. [Google Scholar] [CrossRef]
- Barlow, D.P.; Randle, B.J.; Burke, D.C. Interferon synthesis in the early post-implantation mouse embryo. Differentiation 1984, 27, 229–235. [Google Scholar] [CrossRef]
- Murchison, E.P.; Partridge, J.F.; Tam, O.H.; Cheloufi, S.; Hannon, G.J. Characterization of dicer-deficient murine embryonic stem cells. Proc. Natl. Acad. Sci. USA 2005, 102, 12135–12140. [Google Scholar] [CrossRef]
- Bodak, M.; Cirera-Salinas, D.; Yu, J.; Ngondo, R.P.; Ciaudo, C. Dicer, a new regulator of pluripotency exit and line-1 elements in mouse embryonic stem cells. FEBS Open Bio 2017, 7, 204–220. [Google Scholar] [CrossRef]
- Calabrese, J.M.; Seila, A.C.; Yeo, G.W.; Sharp, P.A. Rna sequence analysis defines dicer’s role in mouse embryonic stem cells. Proc. Natl. Acad. Sci. USA 2007, 104, 18097–18102. [Google Scholar] [CrossRef]
- Witteveldt, J.; Knol, L.I.; Macias, S. MicroRNA-deficient embryonic stem cells acquire a functional interferon response. bioRxiv 2018, 501254. [Google Scholar]
| Pattern Recognition Receptors (PRRs) | Location | Nucleic Acid Ligand | Signaling Adaptor | |
|---|---|---|---|---|
| Toll like receptors | TLR3 | Endosome | dsRNA | TRIF |
| TLR7/8 | Endosome | GU-rich ssRNA | MyD88 | |
| TLR9 | Endosome | Unmethylated CpG DNA | MyD88 | |
| RIG-I like receptors | RIG-I | Cytoplasm | 5′ppp RNA, short dsRNA | MAVS |
| MDA5 | Cytoplasm | Long dsRNA | MAVS | |
| LGP2 | Cytoplasm | Termini of dsRNA | ||
| Others | cGAS | Cytoplasm | dsDNA | STING |
| PKR | Cytoplasm | dsRNA | eIF2α | |
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Watson, S.F.; Knol, L.I.; Witteveldt, J.; Macias, S. Crosstalk Between Mammalian Antiviral Pathways. Non-Coding RNA 2019, 5, 29. https://doi.org/10.3390/ncrna5010029
Watson SF, Knol LI, Witteveldt J, Macias S. Crosstalk Between Mammalian Antiviral Pathways. Non-Coding RNA. 2019; 5(1):29. https://doi.org/10.3390/ncrna5010029
Chicago/Turabian StyleWatson, Samir F., Lisanne I. Knol, Jeroen Witteveldt, and Sara Macias. 2019. "Crosstalk Between Mammalian Antiviral Pathways" Non-Coding RNA 5, no. 1: 29. https://doi.org/10.3390/ncrna5010029
APA StyleWatson, S. F., Knol, L. I., Witteveldt, J., & Macias, S. (2019). Crosstalk Between Mammalian Antiviral Pathways. Non-Coding RNA, 5(1), 29. https://doi.org/10.3390/ncrna5010029





