The Novel Effector Ue943 Is Essential for Host Plant Colonization by Ustilago esculenta
Abstract
1. Introduction
2. Materials and Methods
2.1. Strains and Growth Conditions
2.2. Construction of Deletion and Overexpression Strains
2.3. Quantitative PCR
2.4. Plant Inoculation
2.5. Subcellular Localization
2.6. Western Blot
2.7. Microscopy Observation
2.8. Transcriptome Analysis
2.9. Bioinformatic Analysis
2.10. Statistical Analysis
3. Results
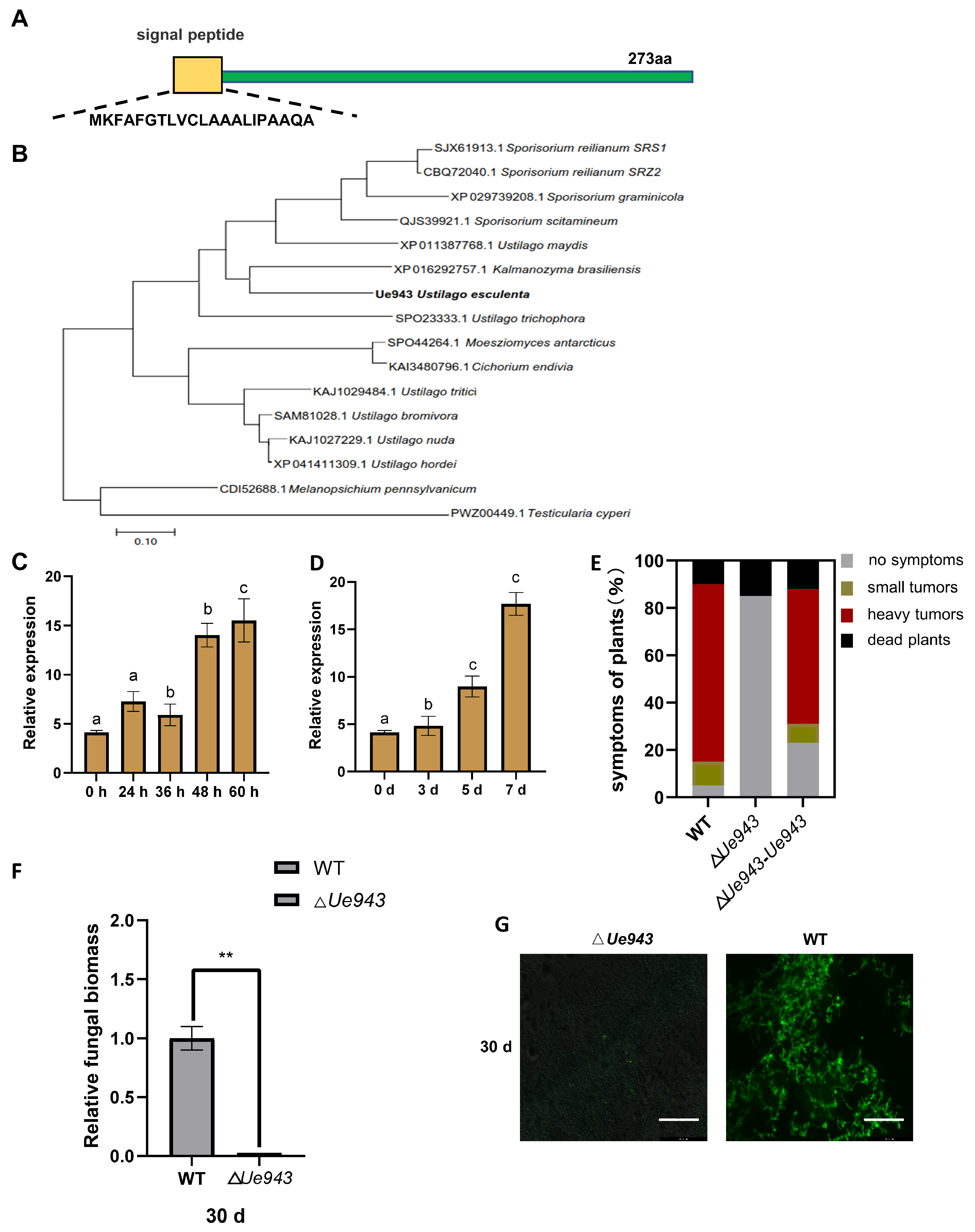
3.1. Ue943 Encodes an Important Virulence Factor in U. esculenta
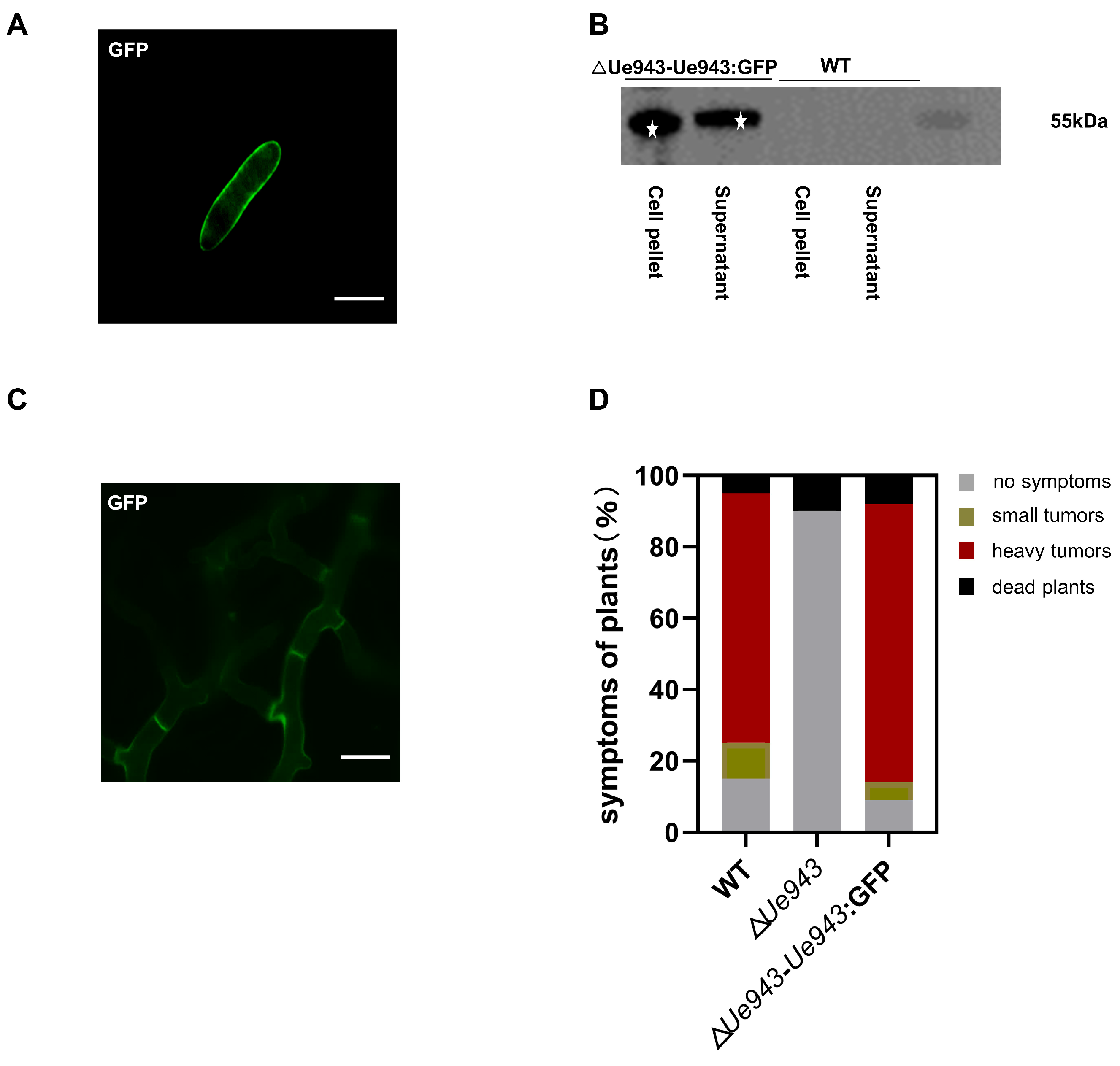
3.2. Ue943 Is Secreted by U. esculenta
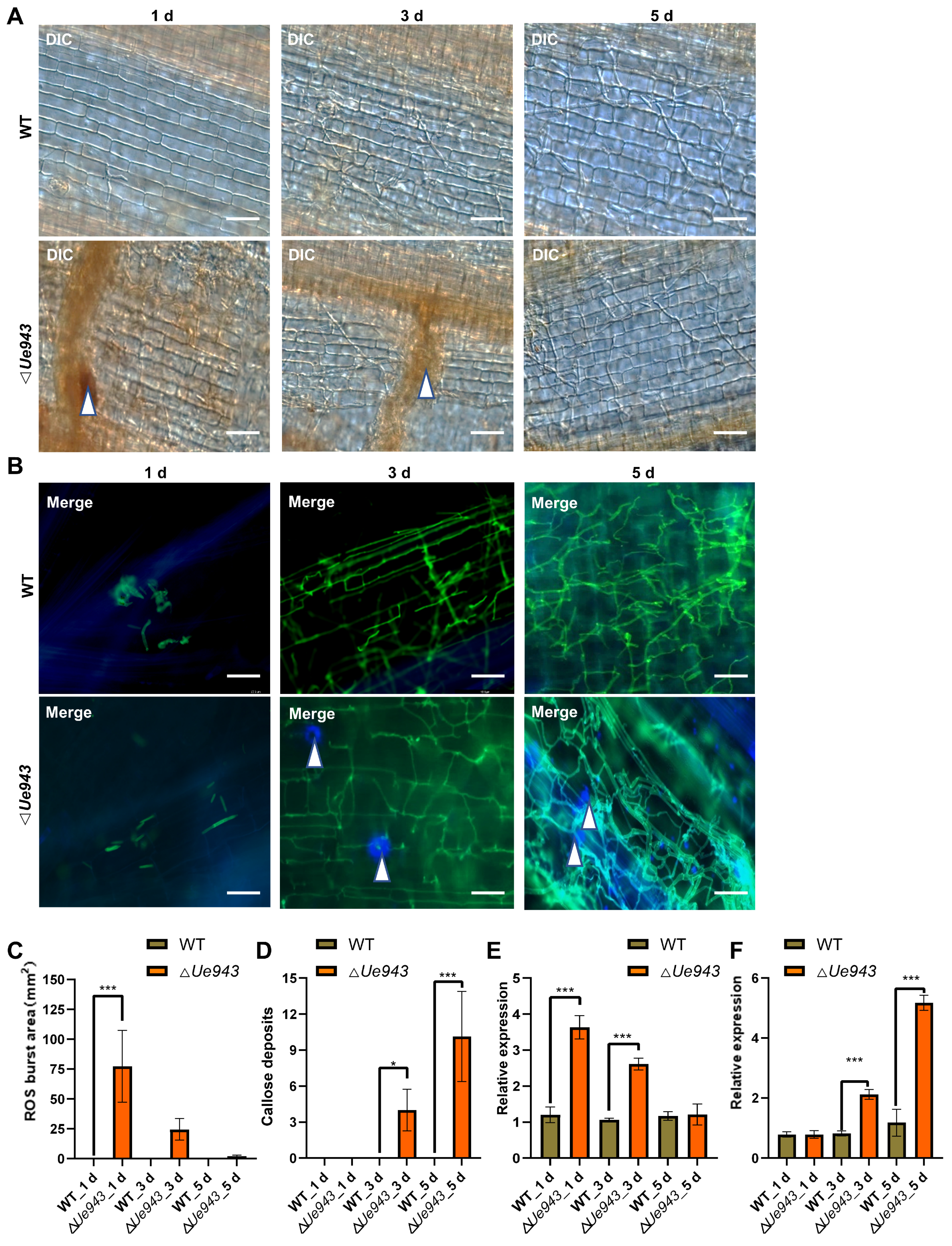
3.3. ΔUe943 Mutant Induces a Hypersensitive Response
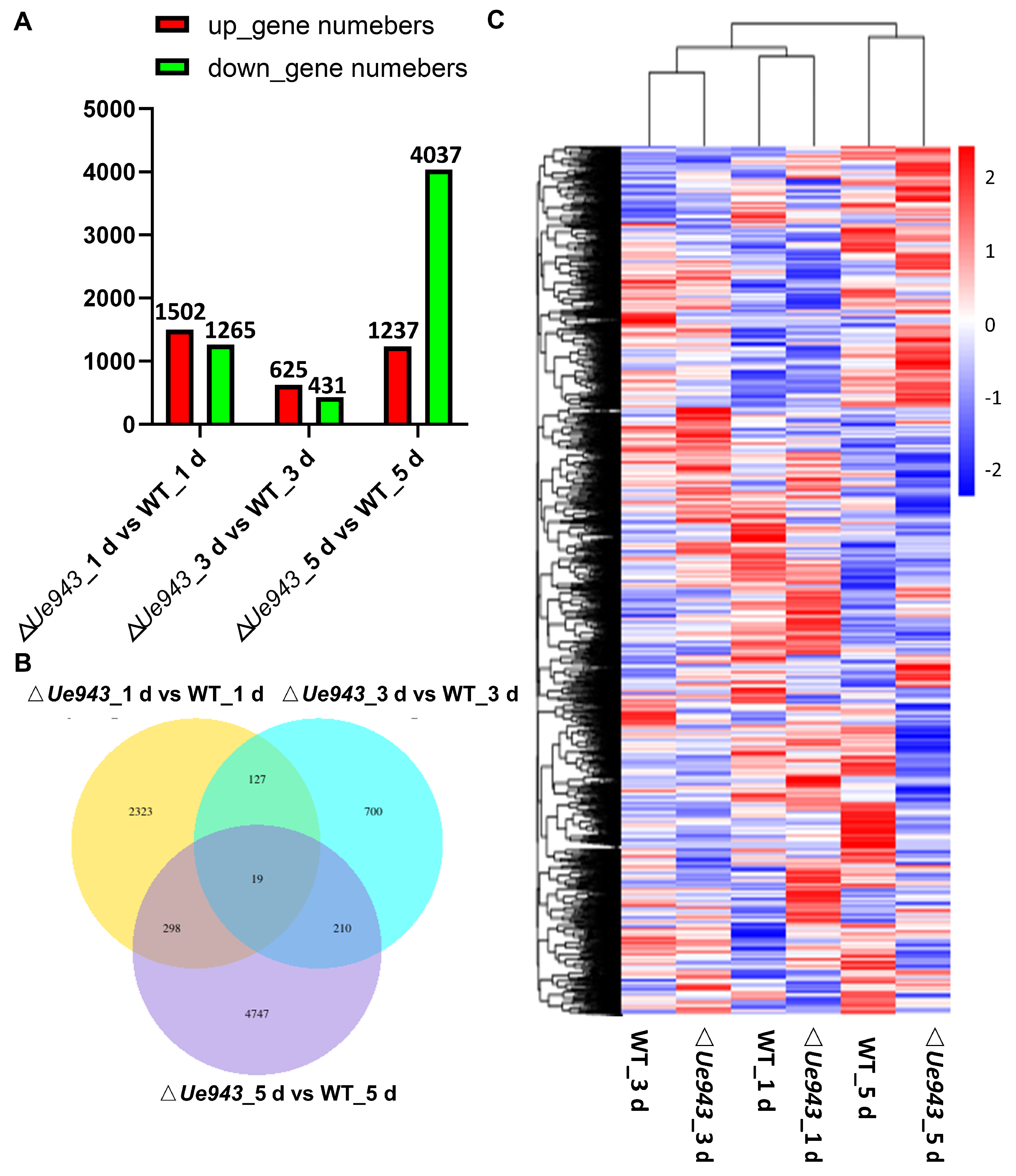
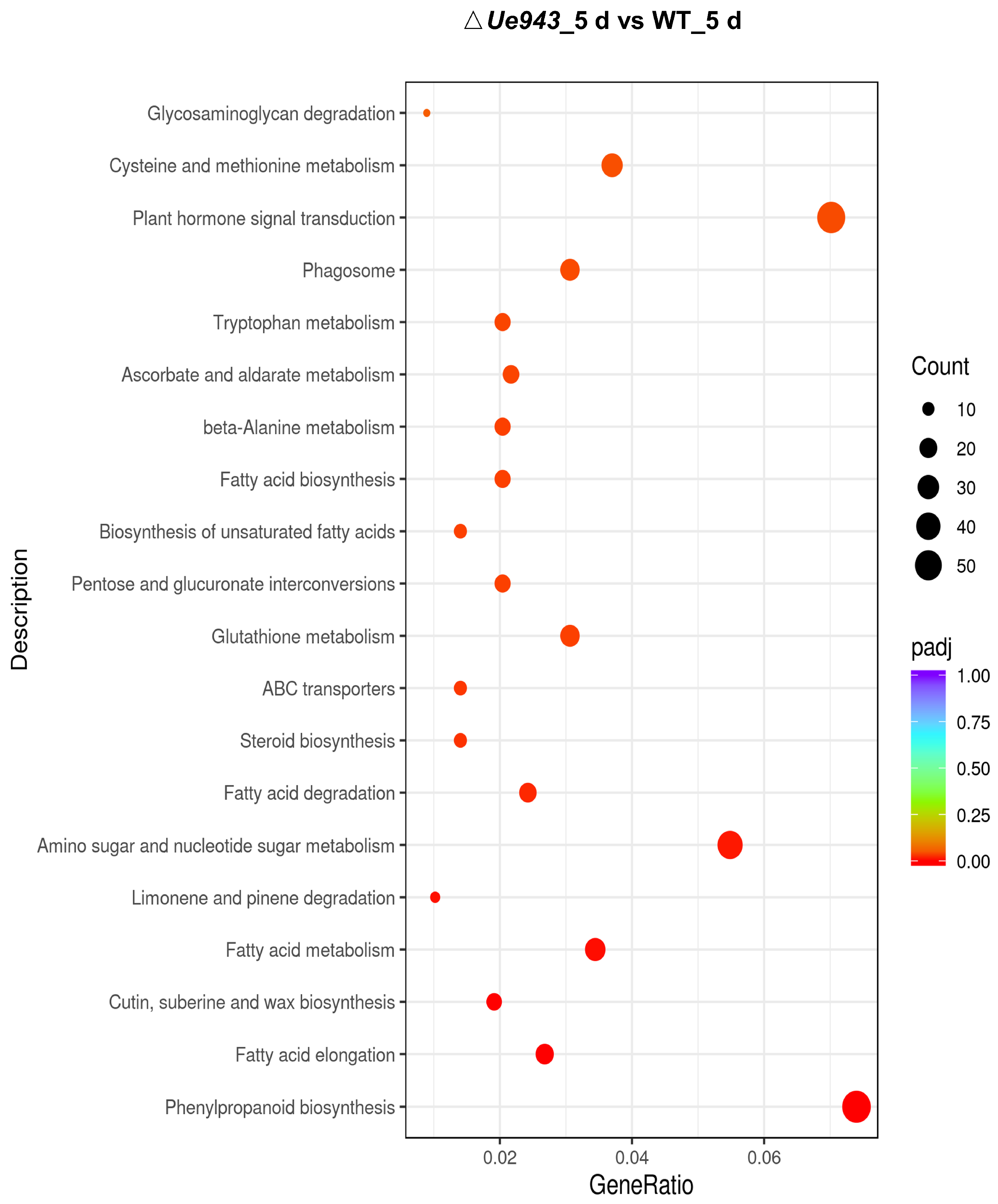
3.4. Transcriptome Analysis of WT Strains and ΔUe943 Strains
3.5. Gene Expression Changes Related to Plant Hormones
4. Discussion
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Tanaka, S.; Brefort, T.; Neidig, N.; Djamei, A.; Kahnt, J.; Vermerris, W.; Koenig, S.; Feussner, K.; Feussner, I.; Kahmann, R. A secreted Ustilago maydis effector promotes virulence by targeting anthocyanin biosynthesis in maize. eLife 2014, 3, e01355. [Google Scholar] [CrossRef] [PubMed]
- Mueller, A.N.; Ziemann, S.; Treitschke, S.; Aßmann, D.; Doehlemann, G. Compatibility in the Ustilago maydis-maize interaction requires inhibition of host cysteine proteases by the fungal effector Pit2. PLoS Pathog. 2013, 9, e1003177. [Google Scholar] [CrossRef] [PubMed]
- Doehlemann, G.; Linde, K.; Assmann, D. Pep1, a secreted effector protein of Ustilago maydis, is required for successful invasion of plant cells. PLoS Pathog. 2009, 5, e1000290. [Google Scholar] [CrossRef]
- Redkar, A.; Villajuana-Bonequi, M.; Doehlemann, G. Conservation of the Ustilago maydis effector See1 in related smuts. Plant Signal. Behav. 2015, 10, e1086855. [Google Scholar] [CrossRef]
- Storfie, E.; Saville, B.J. Fungal Pathogen Emergence: Investigations with an Ustilago maydis × Sporisorium reilianum Hybrid. J. Fungi 2021, 7, 672. [Google Scholar] [CrossRef] [PubMed]
- Tanaka, S.; Schweizer, G.; Rössel, N.; Fukada, F.; Thines, M.; Kahmann, R. Neofunctionalization of the secreted Tin2 effector in the fungal pathogen Ustilago maydis. Nat. Microbiol. 2019, 4, 251–257. [Google Scholar] [CrossRef] [PubMed]
- Rahul, S.; Bilal, Ö.; Gunther, D. Saprotrophic yeasts formerly classified as Pseudozyma have retained a large effector arsenal, including functional Pep1 orthologs. Mycol. Prog. 2019, 18, 763–768. [Google Scholar]
- Wang, S.; Gao, L.; Yin, Y.; Zhang, Y.; Tang, J.; Cui, H.; Li, S.; Zhang, Z.; Yu, X.; Ye, Z.; et al. Transcriptome Comparison between Two Strains of Ustilago esculenta during the Mating. J. Fungi 2022, 9, 32. [Google Scholar] [CrossRef]
- Zhang, Z.; Bian, J.; Zhang, Y.; Xia, W.; Li, S.; Ye, Z. An Endoglucanase Secreted by Ustilago esculenta Promotes Fungal Proliferation. J. Fungi 2022, 8, 1050. [Google Scholar] [CrossRef]
- Yu, J.; Zhang, Y.; Cui, H.; Hu, P.; Yu, X.; Ye, Z. An efficient genetic manipulation protocol for Ustilago esculenta. FEMS Microbiol. Lett. 2015, 362, fnv087. [Google Scholar] [CrossRef]
- Terfrüchte, M.; Joehnk, B.; Fajardo-Somera, R.; Braus, G.H.; Riquelme, M.; Schipper, K.; Feldbrügge, M. Establishing a versatile Golden Gate cloning system for genetic engineering in fungi. Fungal Genet. Biol. 2014, 62, 1–10. [Google Scholar] [CrossRef] [PubMed]
- Zhang, Y.; Wu, M.; Ge, Q.; Yang, M.; Xia, W.; Cui, H.; Yu, X.; Zhang, S.; Ye, Z. Cloning and disruption of the UeArginase in Ustilago esculenta: Evidence for a role of arginine in its dimorphic transition. BMC Microbiol. 2019, 19, 208. [Google Scholar] [CrossRef]
- Wang, Z.-D.; Yan, N.; Wang, Z.-H.; Zhang, X.-H.; Zhang, J.-Z.; Xue, H.-M.; Wang, L.-X.; Zhan, Q.; Xu, Y.-P.; Guo, D.-P. RNA-seq analysis provides insight into reprogramming of culm development in Zizania latifolia induced by Ustilago esculenta. Plant Mol. Biol. 2017, 95, 533–547. [Google Scholar] [CrossRef] [PubMed]
- Zhang, Y.; Cao, Q.; Hu, P.; Cui, H.; Yu, X.; Ye, Z. Investigation on the differentiation of two Ustilago esculenta strains—Implications of a relationship with the host phenotypes appearing in the fields. BMC Microbiol. 2017, 17, 228. [Google Scholar] [CrossRef]
- Herburger, K.; Holzinger, A. Aniline blue and Calcofluor white staining of callose and cellulose in the streptophyte green algae Zygnema and Klebsormidium. Bio-Protocol 2016, 6, e1969. [Google Scholar] [CrossRef] [PubMed]
- Jambunathan, N. Determination and detection of reactive oxygen species (ROS), lipid peroxidation, and electrolyte leakage in plants. In Plant Stress Tolerance; Methods in Molecular Biology; Humana Press: Totowa, NJ, USA, 2010; Volume 639, pp. 292–298. [Google Scholar]
- Kitin, P.; Nakaba, S.; Hunt, C.G.; Lim, S.; Funada, R. Direct fluorescence imaging of lignocellulosic and suberized cell walls in roots and stems. AoB Plants 2020, 12, plaa032. [Google Scholar] [CrossRef]
- Matei, A.; Ernst, C.; Gunl, M.; Thiele, B.; Altmuller, J.; Walbot, V.; Usadel, B.; Doehlemann, G. How to make a tumour: Cell type specific dissection of Ustilago maydis-induced tumour development in maize leaves. New Phytol. 2018, 217, 1681–1695. [Google Scholar] [CrossRef]
- Husin, N.A.; Rahman, S.; Karunakaran, R.; Bhore, S.J. Transcriptome analysis during fruit developmental stages in durian (Durio zibethinus Murr.) var. D24. Genet. Mol. Biol. 2023, 45, e20210379. [Google Scholar] [CrossRef]
- Shahriyari, L. Effect of normalization methods on the performance of supervised learning algorithms applied to HTSeq-FPKM-UQ data sets: 7SK RNA expression as a predictor of survival in patients with colon adenocarcinoma. Brief. Bioinform. 2019, 20, 985–994. [Google Scholar] [CrossRef]
- Lu, C.; Chi, H.; Wang, Y. Transcriptome analysis of PCOS arrested 2-cell embryos. Cell Cycle 2018, 17, 1007–1013. [Google Scholar] [CrossRef] [PubMed]
- Palombo, V.; Milanesi, M.; Sferra, G.; Capomaccio, S.; Sgorlon, S.; D’Andrea, M. PANEV: An R package for a pathway-based network visualization. BMC Bioinform. 2020, 21, 46. [Google Scholar] [CrossRef] [PubMed]
- Gao, C.; Ren, X.; Mason, A.S. Horizontal gene transfer in plants. Funct. Integr. Genom. 2018, 14, 23–29. [Google Scholar] [CrossRef] [PubMed]
- Yaeno, T.; Matsuda, O.; Iba, K. Role of chloroplast trienoic fatty acids in plant disease defense responses. Plant J. 2004, 40, 931–941. [Google Scholar] [CrossRef] [PubMed]
- Pretorius, C.J.; Zeiss, D.R.; Dubery, I.A. The presence of oxygenated lipids in plant defense in response to biotic stress: A metabolomics appraisal. Plant Signal. Behav. 2021, 16, 1989215. [Google Scholar] [CrossRef] [PubMed]
- Molinari, S.; Fanelli, E.; Leonetti, P. Expression of tomato salicylic acid (SA)-responsive pathogenesis-related genes in Mi-1-mediated and SA-induced resistance to root-knot nematodes. Mol. Plant Pathol. 2014, 15, 255–264. [Google Scholar] [CrossRef]
- Lanver, D.; Tollot, M.; Schweizer, G.; Lo Presti, L.; Reissmann, S.; Ma, L.S.; Schuster, M.; Tanaka, S.; Liang, L.; Ludwig, N.; et al. Ustilago maydis effectors and their impact on virulence. Nat. Rev. Microbiol. 2017, 15, 409–421. [Google Scholar] [CrossRef]
- Sharma, R.; Mishra, B.; Runge, F.; Thines, M. Gene loss rather than gene gain is associated with a host jump from monocots to dicots in the Smut Fungus Melanopsichium pennsylvanicum. Genome Biol. Evol. 2014, 6, 2034–2049. [Google Scholar] [CrossRef]
- Pazzagli, L.; Seidl-Seiboth, V.; Barsottini, M. Cerato-platanins: Elicitors and effectors. Plant Sci. 2014, 228, 79–87. [Google Scholar] [CrossRef]
- Weiland, P.; Dempwolff, F.; Steinchen, W.; Freibert, S.A.; Tian, H.; Glatter, T.; Martin, R.; Thomma, B.P.; Bange, G.; Altegoer, F. Structural and functional analysis of the cerato-platanin-like protein Cpl1 suggests diverging functions in smut fungi. Mol. Plant Pathol. 2023, 10, 1111. [Google Scholar] [CrossRef]
- Ökmen, B.; Mathow, D.; Hof, A.; Lahrmann, U.; Aßmann, D.; Doehlemann, G. Mining the effector repertoire of the biotrophic fungal pathogen Ustilago hordei during host and non-host infection. Mol. Plant Pathol. 2018, 19, 2603–2622. [Google Scholar] [CrossRef]
- Müller, O.; Schreier, P.H.; Uhrig, J.F. Identification and characterization of secreted and pathogenesis-related proteins in Ustilago maydis. Mol. Genet. Genom. 2008, 279, 27–39. [Google Scholar] [CrossRef] [PubMed]
- Ma, L.S.; Wang, L.; Trippel, C.; Mendoza-Mendoza, A.; Ullmann, S.; Moretti, M.; Carsten, A.; Kahnt, J.; Reissmann, S.; Zechmann, B.; et al. The Ustilago maydis repetitive effector Rsp3 blocks the antifungal activity of mannose-binding maize proteins. Nat. Commun. 2018, 9, 1711. [Google Scholar] [CrossRef] [PubMed]
- Ökmen, B.; Kemmerich, B.; Hilbig, D.; Wemhöner, R.; Aschenbroich, J.; Perrar, A.; Huesgen, P.F.; Schipper, K.; Doehlemann, G. Dual function of a secreted fungalysin metalloprotease in Ustilago maydis. New Phytol. 2018, 220, 249–261. [Google Scholar] [CrossRef] [PubMed]
- Torres, M.-A. ROS in biotic interactions. Physiol. Plant 2010, 138, 414–429. [Google Scholar] [CrossRef]
- Zhang, W.; Chen, W. Role of salicylic acid in alleviating photochemical damage and autophagic cell death induction of cadmium stress in Arabidopsis thaliana. Photochem. Photobiol. Sci. 2011, 10, 947–955. [Google Scholar] [CrossRef]
- Li, S.; Liu, S.; Zhang, Q.; Cui, M.; Zhao, M.; Li, N.; Wang, S.; Wu, R.; Zhang, L.; Cao, Y.; et al. The interaction of ABA and ROS in plant growth and stress resistances. Front. Plant Sci. 2022, 13, 1050132. [Google Scholar] [CrossRef]
- Seo, P.J.; Park, C.M. MYB96-mediated abscisic acid signals induce pathogen resistance response by promoting salicylic acid biosynthesis in Arabidopsis. New Phytol. 2010, 186, 471–483. [Google Scholar] [CrossRef]
- Wang, D.; Pajerowska-Mukhtar, K.; Culler, A.H. Salicylic acid inhibits pathogen growth in plants through repression of the auxin signaling pathway. Curr. Biol. 2007, 17, 1784–1790. [Google Scholar] [CrossRef]

Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2023 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Wang, S.; Xia, W.; Li, Y.; Peng, Y.; Zhang, Y.; Tang, J.; Cui, H.; Qu, L.; Yao, T.; Yu, Z.; et al. The Novel Effector Ue943 Is Essential for Host Plant Colonization by Ustilago esculenta. J. Fungi 2023, 9, 593. https://doi.org/10.3390/jof9050593
Wang S, Xia W, Li Y, Peng Y, Zhang Y, Tang J, Cui H, Qu L, Yao T, Yu Z, et al. The Novel Effector Ue943 Is Essential for Host Plant Colonization by Ustilago esculenta. Journal of Fungi. 2023; 9(5):593. https://doi.org/10.3390/jof9050593
Chicago/Turabian StyleWang, Shuqing, Wenqiang Xia, Yani Li, Yuyan Peng, Yafen Zhang, Jintian Tang, Haifeng Cui, Lisi Qu, Tongfu Yao, Zetao Yu, and et al. 2023. "The Novel Effector Ue943 Is Essential for Host Plant Colonization by Ustilago esculenta" Journal of Fungi 9, no. 5: 593. https://doi.org/10.3390/jof9050593
APA StyleWang, S., Xia, W., Li, Y., Peng, Y., Zhang, Y., Tang, J., Cui, H., Qu, L., Yao, T., Yu, Z., & Ye, Z. (2023). The Novel Effector Ue943 Is Essential for Host Plant Colonization by Ustilago esculenta. Journal of Fungi, 9(5), 593. https://doi.org/10.3390/jof9050593




