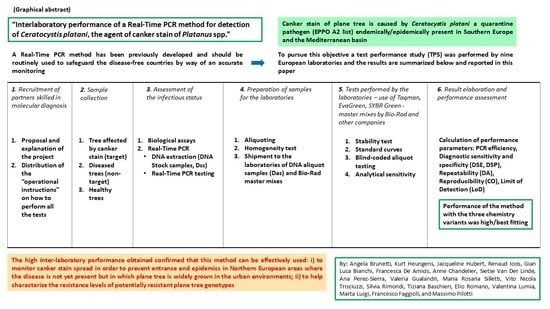
Interlaboratory Performance of a Real-Time PCR Method for Detection of Ceratocystis platani, the Agent of Canker Stain of Platanus spp.
Abstract
:1. Introduction
2. Materials and Methods
2.1. Wood Sample Collection and Assessment of Their Infectious Status
2.2. DNA Extraction
2.3. The Participating Laboratories (PL), the Real-Time PCR Methods, Master Mixes, and Cycling Protocols
2.4. Diagnostic Confirmation of DNA-Stock-Samples (Dss), Aliquoting, Homogeneity Testing, and the Shipping Material
- -
- 16 blind test samples—15 DNA extracts from wood, and one from an axenic C. platani culture.
- -
- 2 controls: PAC and NAC obtained from NI.3 and H.1 (also supplied as blind Das).
- -
- 1 positive wood extract sample for the standard curve experiments (obtained from NI.2, also supplied as a blind Das), labelled DNA-aliquot-St.Cu. (Da.St.Cu.)
- -
- The gDNA from an axenic culture of C. platani for use in the analytical sensitivity test (also supplied as a blind Das)
2.5. Stability Test
2.6. The Test Performance Study with Real-Time PCR: (i) Generating Standard Curves, (ii) Testing Blind-Coded DNA-Aliquot-Samples (iii) Testing Analytical Sensitivity
2.7. Performance Criteria, Nomenclature, and Statistical Analysis
2.8. Outliers
2.9. Disclosure Policy by the Organiser
3. Results
3.1. Sample Preparation
3.2. Thermal Cycling Adjustments
3.3. Test Performance Study: Generating Standard Curves
3.4. Test Performance Study: Blind Testing of DNA-Aliquot-Samples
3.5. Test Performance Study: Testing Analytical Sensitivity
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
Appendix A

References
- Engelbrecht, C.J.B.; Harrington, T.C. Intersterility, morphology and taxonomy of Ceratocystis fimbriata on sweet potato, cacao and sycamore. Mycologia 2005, 97, 57–69. [Google Scholar] [CrossRef]
- De Beer, Z.W.; Duong, T.A.; Barnes, I.; Wingfield, B.D.; Wingfield, M.J. Redefining Ceratocystis and allied genera. Stud. Mycol. 2014, 79, 187–219. [Google Scholar] [CrossRef] [Green Version]
- Tsopelas, P.; Santini, A.; Wingfield, M.J.; De Beer, Z.W. Canker stain: A lethal disease destroying iconic plane trees. Plant Dis. 2017, 101, 645–658. [Google Scholar] [CrossRef] [Green Version]
- Pilotti, M.; Brunetti, A.; Tizzani, L.; Marani, O. Platanus × acerifolia genotypes surviving to inoculation with Ceratocystis platani (the agent of canker stain): First screening and molecular characterization. Euphytica 2009, 169, 1–17. [Google Scholar] [CrossRef]
- Harrington, T.C. Ceratocystis diseases. In Infectious Forest Diseases; Gonthier, P., Nicolotti, G., Eds.; CAB International: Wallingford, UK, 2013; pp. 230–255. [Google Scholar] [CrossRef]
- Panconesi, A.; Moricca, S.; Della Valle, I.; Torraca, C. The epidemiology of canker stain of plane tree and its spread from urban plantings to spontaneous groves and natural forests. In Proceedings of the II International Symposium on Plant Health in Urban Horticulture, Mitt. Biol. Bundesanst Land-Forstwirtsch, Berlin, Germany, 27–29 August 2003; Volume 394, pp. 84–91. [Google Scholar]
- Luchi, N.; Ghelardini, L.; Belbahri, L.; Quartier, M.; Santini, A. Rapid detection of Ceratocystis platani inoculum by quantitative Real-Time PCR assay. Appl. Environ. Microb. 2013, 79, 5394–5404. [Google Scholar] [CrossRef] [Green Version]
- Soulioti, N.; Tsopelas, P.; Woodward, S. Platypus cylindrus, a vector of Ceratocystis platani in Platanus orientalis stands in Greece. Forest Pathol. 2015, 45, 367–372. [Google Scholar] [CrossRef]
- Panconesi, A. Canker stain of plane trees: A serious danger to urban plantings. J. Plant Pathol. 1999, 81, 3–15. [Google Scholar]
- Lehtijärvi, A.; Oskay, F.; Doğmuş Lehtijärvi, H.T.; Aday Kaya, A.G.; Pecori, F.; Santini, A.; Woodward, S. Ceratocystis platani is killing plane trees in Istanbul (Turkey). Forest Pathol. 2018, 48, e12375. [Google Scholar] [CrossRef]
- Commission Implementing Regulation (EU) 2019/2072. Annex II part B: Pests known to occur in the Union territory. Off. J. Eur. Union 2019, 319, 1–279. [Google Scholar]
- OEPP/EPPO. Ceratocystis fimbriata (Ell. Halsted) f.sp. platani (Walter). OEPP/EPPO Bull. 1986, 16, 21–24. [Google Scholar] [CrossRef]
- OEPP/EPPO. PM 7/014 (1): Ceratocystis fimbriata f. sp. Platani. OEPP/EPPO Bull. 2003, 33, 249–255. [Google Scholar] [CrossRef]
- OEPP/EPPO. PM 7/014 (2): Ceratocystis platani. OEPP/EPPO Bull. 2014, 44, 338–349. [Google Scholar] [CrossRef] [Green Version]
- Ministerial Decree—MIPAAF (Ministero delle Politiche Agricole Alimentari e Forestali). Misure di emergenza per la prevenzione, il controllo e l’eradicazione del cancro colorato del platano causato da Ceratocystis fimbriata [Emergency measures for the prevention, control and eradication of canker stain of plane tree caused by Ceratocystis fimbriata]. In Gazzetta Ufficiale della Repubblica Italiana; Serie Generale n. 100; MIPAAF: Rome, Italy, 2012; pp. 4–8. (In Italian) [Google Scholar]
- Ministerial Decree—MIPAAF (Ministero delle Politiche Agricole Alimentari e Forestali). Modifica del decreto 29 Febbraio 2012 recante misure di emergenza per la prevenzione, il controllo e l’eradicazione del cancro colorato del platano causato da Ceratocystis fimbriata [Modification of decree 29 February 2012 dealing with emergency measures for the prevention, control and eradication of canker stain of plane tree caused by Ceratocystis fimbriata]. In Gazzetta Ufficiale della Repubblica Italiana; Serie Generale n. 222; MIPAAF: Rome, Italy, 2015; pp. 10–11. (In Italian) [Google Scholar]
- Pilotti, M.; Lumia, V.; Di Lernia, G.; Brunetti, A. Development of Real-Time PCR for in wood-detection of Ceratocystis platani, the agent of canker stain of Platanus spp. Eur. J. Plant Pathol. 2012, 134, 61–79. [Google Scholar] [CrossRef]
- Lumia, V.; Modesti, V.; Brunetti, A.; Wilkinson, C.L.; Di Lernia, G.; Harrington, T.C.; Pilotti, M. Real-Time PCR for Ceratocystis platani detection: In-depth validation to assess the diagnostic potential and include additional technical options. iForest 2018, 11, 499–509. [Google Scholar] [CrossRef] [Green Version]
- OEPP/EPPO PM 7/122 (1). Guidelines for the organization of interlaboratory comparisons by plant pest diagnostic laboratories. OEPP/EPPO Bull. 2014, 44, 390–399. [Google Scholar] [CrossRef]
- OEPP/EPPO PM 7/98 (2). Specific requirements for laboratories preparing accreditation for a plant pest diagnostic activity. OEPP/EPPO Bull. 2014, 44, 117–127. [Google Scholar] [CrossRef] [Green Version]
- OEPP/EPPO PM 7/98 (5). Specific requirements for laboratories preparing accreditation for a plant pest diagnostic activity. OEPP/EPPO Bull. 2021, 51, 468–498. [Google Scholar] [CrossRef]
- Pilotti, M.; Gervasi, F.; Brunetti, A. Molecular identification of Fomitiporia mediterranea and Eutypa lata/Libertella blepharis in Platanus × acerifolia. J. Phytopathol 2005, 153, 193–202. [Google Scholar] [CrossRef]
- ISO 16140; Microbiology of Foods and Animal Feeding Stuffs—Protocol for the Validation of Alternative Methods. International Organization for Standardization: Geneva, Switzerland, 2003.
- Chabirand, A.; Anthoine, G.; Pierson, O.; Hostachy, B. The organization of proficiency testing in plant pathology (qualitative methods of analysis) according to the ISO/IEC 17043: Example of the French national reference laboratory. Accred. Qual. Assur. 2014, 19, 111–125. [Google Scholar] [CrossRef] [Green Version]
- Chabirand, A.; Loiseau, M.; Renaudin, I.; Poliakoff, F. Data processing of qualitative results from an interlaboratory comparison for the detection of “Flavescence dorée” phytoplasma: How the use of statistics can improve the reliability of the method validation process in plant pathology. PLoS ONE 2017, 12, e0175247. [Google Scholar] [CrossRef] [Green Version]
- Langton, S.D.; Chevennement, R.; Nagelkerke, N.; Lombard, B. Analysis collaborative trials for qualitative microbiological methods. Int. J. Food Microbiol. 2002, 79, 175–181. [Google Scholar] [CrossRef]
- R Core Team. R: A Language and Environment for Statistical Computing. R Foundation for Statistical Computing, Vienna, Austria. 2021. Available online: https://www.R-project.org/ (accessed on 15 February 2022).
- Wilson, E.B. Probable inference, the law of succession, and statistical inference. J. Am. Stat. Assoc. 1927, 22, 209–212. [Google Scholar] [CrossRef]
- Luigi, M.; Manglli, A.; Tiberini, A.; Bertin, S.; Ferretti, L.; Taglienti, A.; Faggioli, F.; Tomassoli, L. Inter-laboratory Comparison of RT-PCR-Based Methods for the Detection of Tomato Brown Rugose Fruit Virus on Tomato. Pathogens 2022, 11, 207. [Google Scholar] [CrossRef]
- Ioos, R.; Aloi, F.; Piškur, B.; Guinet, C.; Mullett, M.; Berbegal, M.; Bragança, H.; Cacciola, S.O.; Oskay, F.; Cornejo, C.; et al. Transferability of PCR-based diagnostic protocols: An international collaborative case study assessing protocols targeting the quarantine pine pathogen Fusarium circinatum. Sci. Rep. 2019, 9, 8195. [Google Scholar] [CrossRef] [Green Version]
- Ahmed, Y.; Hubert, J.; Fourrier-Jeandel, C.; Dewdney, M.M.; Aguayo, J.; Ioos, R. A Set of Conventional and Multiplex Real-Time PCR Assays for Direct Detection of Elsinoë fawcettii, E. australis, and Pseudocercospora angolensis in Citrus Fruits. Plant Dis. 2019, 103, 345–356. [Google Scholar] [CrossRef] [Green Version]
- Ahmed, Y.; Hussein, A.; Hubert, J.; Fourrier-Jeandel, C.; Aguayo, J.; Ioos, R. New multiplex conventional PCR and quadruplex real-time PCR assays for one-tube detection of Phyllosticta citricarpa, Elsinoë fawcettii, Elsinoë australis, and Pseudocercospora angolensis in Citrus: Development and validation. Appl. Microbiol. Biot. 2020, 104, 9363–9385. [Google Scholar] [CrossRef]
- Benzigar, M.R.; Bhattacharjee, R.; Baharfar, M.; Liu, G. Current methods for diagnosis of human coronaviruses: Pros and cons. Anal. Bioanal. Chem. 2021, 413, 2311–2330. [Google Scholar] [CrossRef]
- Petersen, L.M.; Martin, I.W.; Moschetti, W.E.; Kershaw, C.M.; Tsongalis, G.J. Third-Generation sequencing in the clinical laboratory: Exploring the advantages and challenges of nanopore sequencing. J. Clin. Microbiol. 2020, 58, e01315-19. [Google Scholar] [CrossRef]
- Hoang, M.T.V.; Irinyi, L.; Hu, Y.; Schwessinger, B.; Meyer, W. Long-reads-based metagenomics in clinical diagnosis with a special focus on fungal infections. Front. Microbiol. 2022, 12, 708550. [Google Scholar] [CrossRef]
- Mao, F.; Leung, W.Y.; Xin, X. Characterization of EvaGreen and the implication of its physicochemical properties for qPCR applications. BMC Biotechnol. 2007, 7, 76. [Google Scholar] [CrossRef] [Green Version]
- Sjöback, R.; Nygren, J.; Kubista, M. Absorption and Fluorescence Properties of Fluorescein. Spectrochim. Acta A-M 1995, 51, L7–L21. [Google Scholar] [CrossRef]





| Sample ID | Nature of the Sample | Number of Samples | Sample Type | Expected Detection |
|---|---|---|---|---|
| NI.1→ NI.3 | C. platani Naturally-Infected tree | 3 | Blind samples | Positive |
| AI.1→AI.6 | C. platani Artificially-Infected tree | 6 | Blind samples | Positive |
| H.1→H.3 | Healthy tree | 3 | Blind samples | Negative |
| D.1→D.3 | Diseased tree infected with non-target species | 3 | Blind samples | Negative |
| gDNA C.P. 32 | Pure C. platani colony | 1 | Blind sample | Positive |
| NAC | Healthy tree (=H.1) | 1 | Negative amplification control | Negative |
| PAC | C. platani Naturally-Infected tree (=NI.3) | 1 | Positive amplification control | Positive |
| DNA-aliquot-St.Cu | C. platani Naturally-Infected tree (=NI.2) | 1 | Standard curve validation | Positive |
| 1 | 2 | 3 | 4 | 5 | 6 | 7 | 8 | 9 | 10 | 11 | 12 | 13 | 14 | 15 | 16 | 17 | 18 | 19 | 20 | |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Dss 1 | NI.1 | NI.2 | NI3 | AI.1 | AI2 | AI3 | AI4 | AI5 | AI6 | H.1 | H.2 | H.3 | D.1 | D.2 | D.3 | gDNA C.P. 32 | PAC (=NI.3) | NAC (=H.1) | Da.St.Cu. (=NI.2) | gDNA C.P. 32 |
| Z (OL) | 6 | 3 | 11 | 12 | 14 | 15 | 10 | 2 | 1 | 4 | 7 | 5 | 8 | 16 | 9 | 13 | + | + | + | + |
| A | 3 | 14 | 13 | 10 | 7 | 9 | 11 | 15 | 12 | 1 | 8 | 16 | 4 | 6 | 5 | 2 | + | + | + | + |
| B | 12 | 7 | 14 | 16 | 15 | 11 | 8 | 6 | 10 | 2 | 13 | 3 | 1 | 9 | 4 | 5 | + | + | + | + |
| C | 11 | 12 | 15 | 14 | 6 | 4 | 3 | 10 | 7 | 9 | 1 | 2 | 16 | 5 | 13 | 8 | + | + | + | + |
| D | 9 | 5 | 7 | 8 | 13 | 10 | 16 | 11 | 15 | 12 | 4 | 1 | 6 | 2 | 3 | 14 | + | + | + | + |
| E | 7 | 9 | 2 | 5 | 12 | 1 | 4 | 13 | 3 | 11 | 6 | 8 | 15 | 14 | 16 | 10 | + | + | + | + |
| F | 4 | 16 | 10 | 11 | 9 | 5 | 13 | 1 | 6 | 7 | 3 | 15 | 2 | 8 | 14 | 12 | + | + | − | + |
| G | 2 | 13 | 8 | 7 | 10 | 14 | 12 | 5 | 16 | 15 | 9 | 11 | 3 | 4 | 6 | 1 | + | + | + | + |
| H | 10 | 15 | 6 | 9 | 2 | 13 | 5 | 12 | 14 | 3 | 16 | 4 | 7 | 1 | 8 | 11 | + | + | − | + |
| OL, PL | Taqman | EvaGreen | SYBR Green |
|---|---|---|---|
| Z |
|
|
|
| A1 |
| ||
| A2 |
| ||
| A3 |
| ||
| A4 |
| ||
| B |
| ||
| B1 |
| ||
| B2 |
| ||
| C1 |
| ||
| C2 |
| ||
| D |
|
| |
| E |
|
| |
| F |
|
| |
| G |
|
| |
| H1 |
| ||
| H2 |
|
| Performance Criteria | Acronyms and Calculation | Legenda | Best Performance Level (%) |
|---|---|---|---|
| Accuracy | AC = (NTP + NTN)/N | NTP NTN = number of true positives and true negatives N = total number of tested sample | 100 |
| Diagnostic sensitivity | DSE = NTP/N+ | N+ = number of samples for which the assigned value is positive (i.e., Ceratocystis platani-positive) | 100 |
| Diagnostic specificity | DSP = NTN/N− | N− = number of samples for which the assigned value is negative (i.e., Ceratocystis platani-negative) | 100 |
| Repeatability | DA = (NTP/N)2 + (NTN/N)2 | See above | 1 |
| Reproducibility | CO—Calculate the interlaboratory pairs sharing the same (and conforming) results and infer the percentage compared to the total number of the interlaboratory pairs | 100 | |
| Laboratory Code 1 | Assay | E 2 (%) | R˄2 3 | Slope 4 | Intercept |
|---|---|---|---|---|---|
| Z | Taqman | 96.3 | 0.999 | −3.413 | 34.041 |
| A1 | Taqman | 94.9 | 0.999 | −3.451 | 34.426 |
| A3 | Taqman | 95.3 | 0.999 | −3.439 | 33.987 |
| A4 | Taqman | 101.4 | 0.999 | −3.288 | 31.492 |
| B1 | Taqman | 99.9 | 0.999 | −3.323 | 34.102 |
| B2 | Taqman | 102.3 | 0.999 | −3.269 | 34.129 |
| C1 | Taqman | 97.9 | 0.999 | −3.373 | 29.367 |
| C2 | Taqman | 101.4 | 0.999 | −3.288 | 29.236 |
| D | Taqman | 79.7 | 0.998 | −3.930 | 35.751 |
| E | Taqman | 96.3 | 0.999 | −3.413 | 33.749 |
| F | Taqman | 94.1 | 0.999 | −3.472 | 34.460 |
| Z | EvaGreen | 98.3 | 1.000 | −3.362 | 33.720 |
| D | EvaGreen | 74.3 | 0.998 | −4.143 | 35.968 |
| E | EvaGreen | 99.5 | 0.999 | −3.335 | 33.484 |
| F | EvaGreen | 98.4 | 0.999 | −3.360 | 33.967 |
| G | EvaGreen | 100.7 | 0.999 | −3.304 | 33.512 |
| Z | SYBR Green | 98.8 | 0.999 | −3.352 | 30.738 |
| B | SYBR Green | 99.1 | 0.992 | −3.343 | 30.451 |
| H1 | SYBR Green | 96.7 | 0.999 | −3.404 | 32.052 |
| H2 | SYBR Green | 95.8 | 0.999 | −3.427 | 32.092 |
| Performance Parameters | TM.Z | TM.A1 | TM.A2 | TM.B1 | TM.B2 | TM.C1 | TM.C2 | TM.D | TM.E | TM.F | TM.G | TM Global |
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| DSE | 100 | 100 | 100 | 100 | 100 | 100 | 100 | 100 | 100 | 100 | 100 | 100 |
| DSP | 100 | 100 | 100 | 100 | 88.9 * (81.2–93.7) | 100 | 100 | 100 | 100 | 100 | 100 | 99.0 (94.5–99.9) |
| AC | 100 | 100 | 100 | 100 | 95.8 * (89.9–98.3) | 100 | 100 | 100 | 100 | 100 | 100 | 99.6 (95.6–99.9) |
| CO | 99.2 (94.8–99.9) | |||||||||||
| Performance parameters | EG.Z | EG.D | EG.E | EG.F | EG.G | EG global | SG.Z | SG.B | SG.H1 | SG.H2 | SG global | |
| DSE | 100 | 100 | 100 | 100 | 100 | 100 | 100 | 100 | 100 | 100 | 100 | |
| DSP | 100 | 100 | 100 | 100 | 100 | 100 | 100 | 100 | 100 | 66.7 * (57.0–75.2) | 91.7 (84.6–95.7) | |
| AC | 100 | 100 | 100 | 100 | 100 | 100 | 100 | 100 | 100 | 87.5 * (79.6–92.6) | 96.9 (91.4–98.9) | |
| CO | 100 | 93.2 (86.5–96.7) |
| Performance Parameters at 15fg 1 | TM.Z | TM.A1 | TM.A2 | TM.B | TM.C | TM.E | TM.F | TM.G | TM. Global |
|---|---|---|---|---|---|---|---|---|---|
| DSE | 100 | 100 | 100 | 100 | 100 | 100 | 83.3 * (74.8–89.3) | 83.3 * (74.8–89.3) | 95.7 (89.8–98.3) |
| DA | 1 | 1 | 1 | 1 | 1 | 1 | 0.7 * (0.10–0.98) | 0.7 * (0.10–0.98) | N.E.2 |
| AC | 100 | 100 | 100 | 100 | 100 | 100 | 83.3 * (74.8–89.3) | 83.3 * (74.8–89.3) | 95.7 (89.8–98.3) |
| CO | 92.9 (86.1–96.5) | ||||||||
| Performance parameters at 15fg 1 | EG.Z | EG.E | EG. global | SG.Z | SG.H1 | SG.H2 | SG. global | ||
| DSE | 100 | 100 | 100 | 100 | 100 | 100 | 100 | ||
| DA | 1 | 1 | N.E. 2 | 1 | 1 | 1 | N.E. 2 | ||
| AC | 100 | 100 | 100 | 100 | 100 | 100 | 100 | ||
| CO | 100 | 100 | |||||||
| Performance parameters at 3 fg 1 | TM.Z | TM.A1 | TM.A2 | TM.B | TM.C | TM.E | TM.F | TM.G | TM. global |
| DSE | 100 | 50 * (40.3–59.6) | 83.3 * (74.8–89.3) | 100 | 100 | 83.3 * (74.8–89.3) | 83.3 * (74.8–89.3) | 100 | 87.2 (79.2–92.4) |
| DA | 1 | 0.2 * (0.01–0.86) | 0.7 * (0.1–0.98) | 1 | 1 | 0.7 * (0.1–0.98) | 0.7 * (0.1–0.98) | 1 | N.E. 2 |
| AC | 100 | 50 * (40.3–59.6) | 83.3 * (74.8–89.3) | 100 | 100 | 83.3 * (74.8–89.3) | 83.3 * (74.8–89.3) | 100 | 87.2 (79.2–92.4) |
| CO | 76.2 (67.0–83.4) | ||||||||
| Performance parameters at 3 fg 1 | EG.Z | EG.E | EG. global | SG.Z | SG.H1 | SG.H2 | SG. global | ||
| DSE | 100 | 100 | 100 | 100 | 100 | 100 | 100 | ||
| DA | 1 | 1 | N.E. 2 | 1 | 1 | 1 | N.E. 2 | ||
| AC | 100 | 100 | 100 | 100 | 100 | 100 | 100 | ||
| CO | 100 | 100 |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Brunetti, A.; Heungens, K.; Hubert, J.; Ioos, R.; Bianchi, G.L.; De Amicis, F.; Chandelier, A.; Van Der Linde, S.; Perez-Sierra, A.; Gualandri, V.; et al. Interlaboratory Performance of a Real-Time PCR Method for Detection of Ceratocystis platani, the Agent of Canker Stain of Platanus spp. J. Fungi 2022, 8, 778. https://doi.org/10.3390/jof8080778
Brunetti A, Heungens K, Hubert J, Ioos R, Bianchi GL, De Amicis F, Chandelier A, Van Der Linde S, Perez-Sierra A, Gualandri V, et al. Interlaboratory Performance of a Real-Time PCR Method for Detection of Ceratocystis platani, the Agent of Canker Stain of Platanus spp. Journal of Fungi. 2022; 8(8):778. https://doi.org/10.3390/jof8080778
Chicago/Turabian StyleBrunetti, Angela, Kurt Heungens, Jacqueline Hubert, Renaud Ioos, Gian Luca Bianchi, Francesca De Amicis, Anne Chandelier, Sietse Van Der Linde, Ana Perez-Sierra, Valeria Gualandri, and et al. 2022. "Interlaboratory Performance of a Real-Time PCR Method for Detection of Ceratocystis platani, the Agent of Canker Stain of Platanus spp." Journal of Fungi 8, no. 8: 778. https://doi.org/10.3390/jof8080778