Seventeen Ustilaginaceae High-Quality Genome Sequences Allow Phylogenomic Analysis and Provide Insights into Secondary Metabolite Synthesis
Abstract
1. Introduction
2. Materials and Methods
2.1. Selection of Fungal Strains
2.2. Genomic DNA Preparation
2.3. Nanopore Library Preparation and GridION® Sequencing
2.4. Illumina Library Preparation and MiSeq Sequencing
2.5. Base Calling, Reads Processing, and Assembly
2.6. Gene Prediction and Genome Annotation
2.7. Comparative Genome Analyses and Phylogenetic Analysis
3. Results and Discussion
3.1. Genomic Data
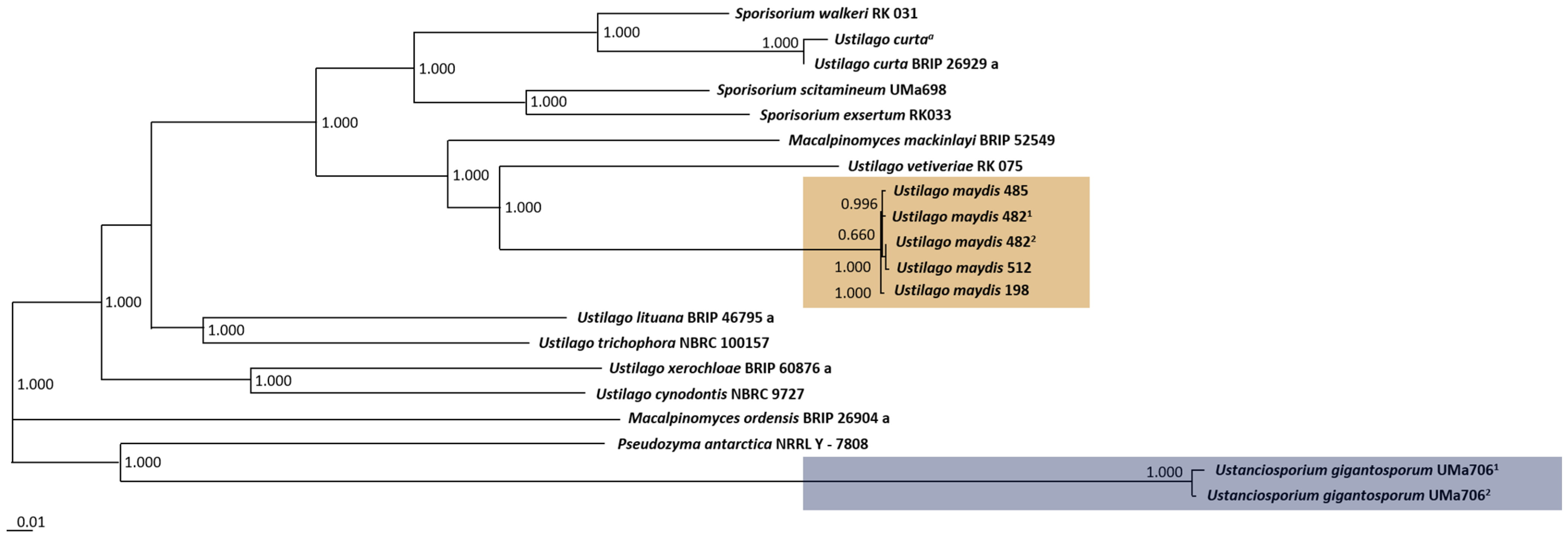
3.2. Phylogenomic Analysis
3.3. Comparative Genomics
3.3.1. Pairwise Percentage of Conserved Proteins
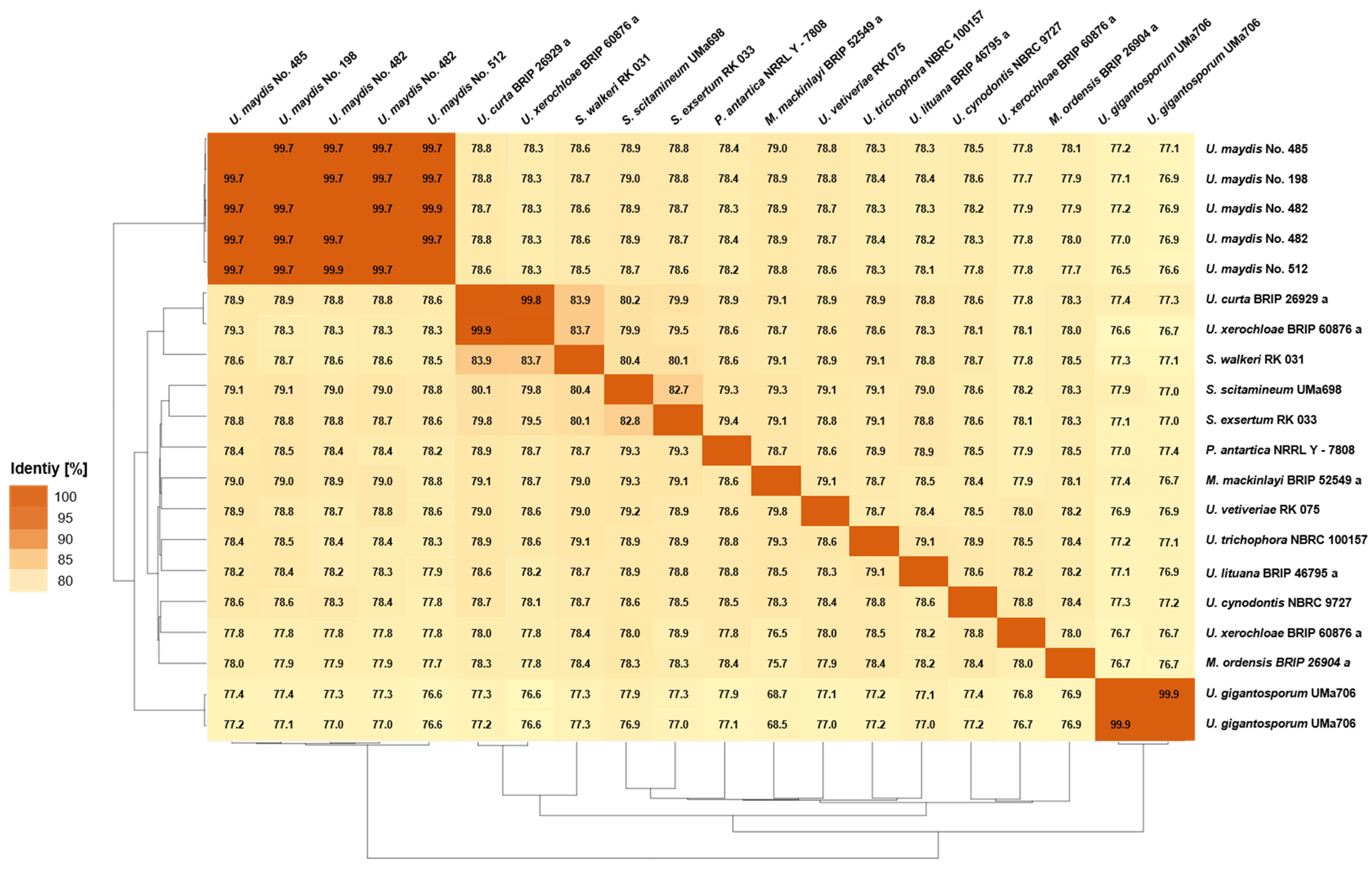
3.3.2. Average Nucleotide Identity
3.3.3. Average Amino Acid Identity
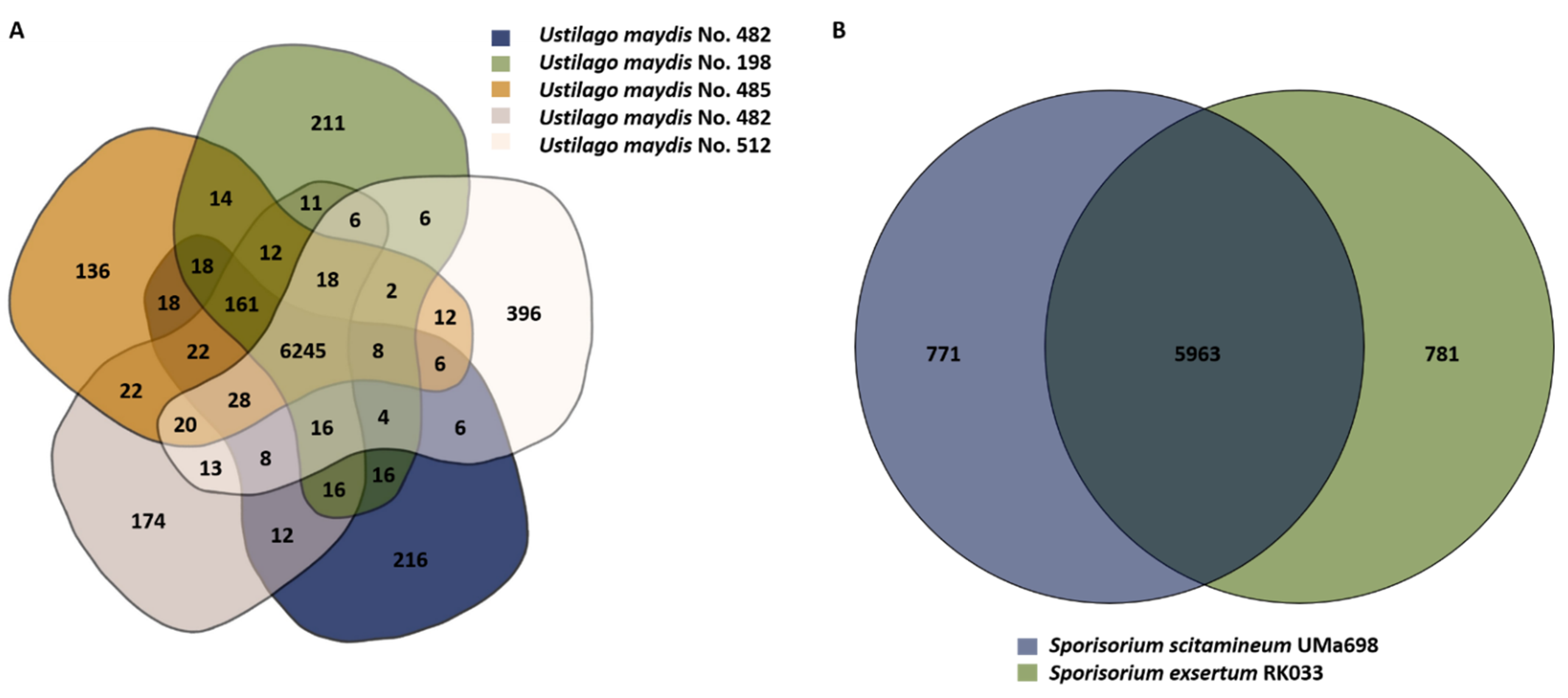
3.4. Gene-Based Comparison
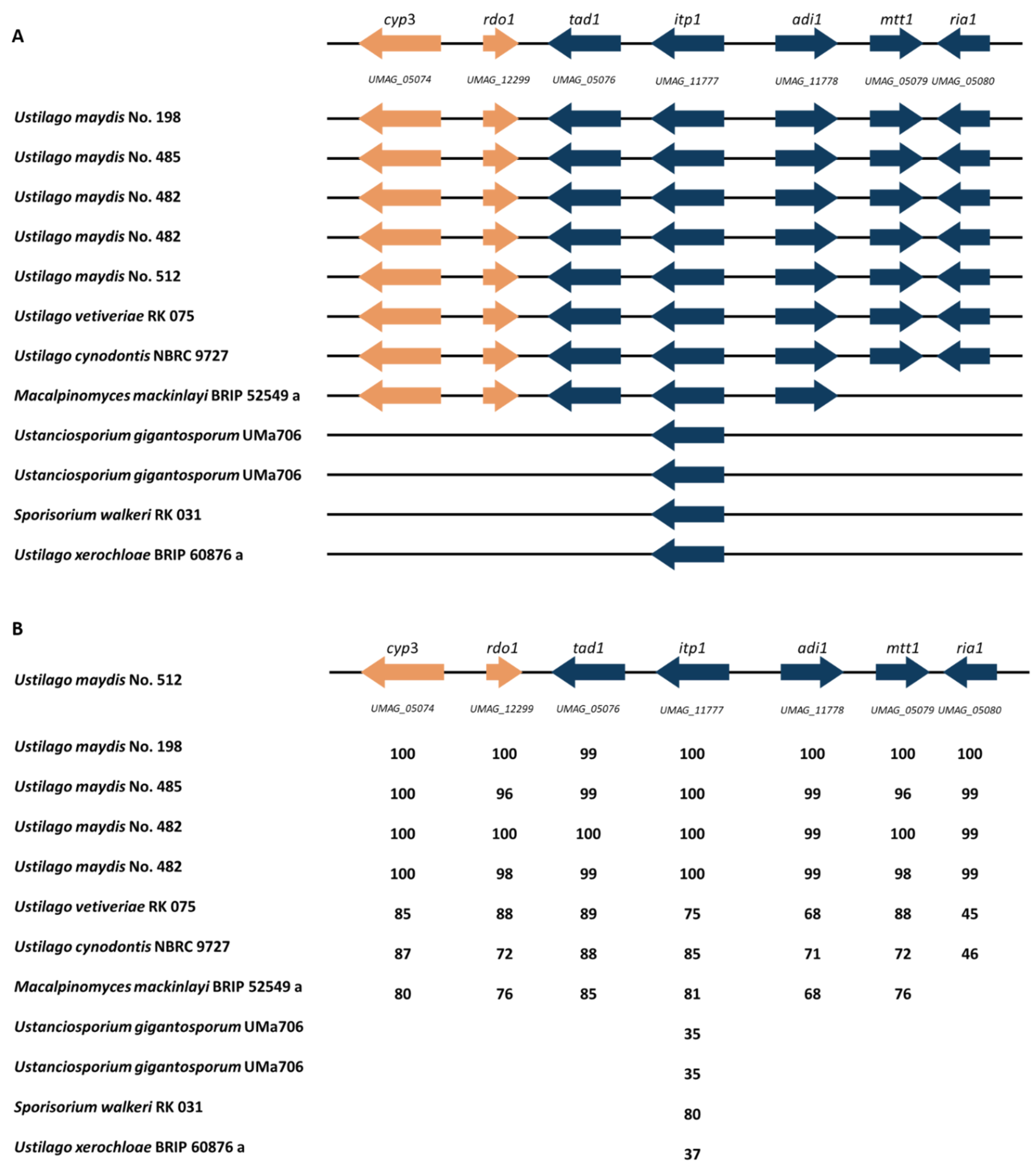
3.5. Itaconate Cluster Identification
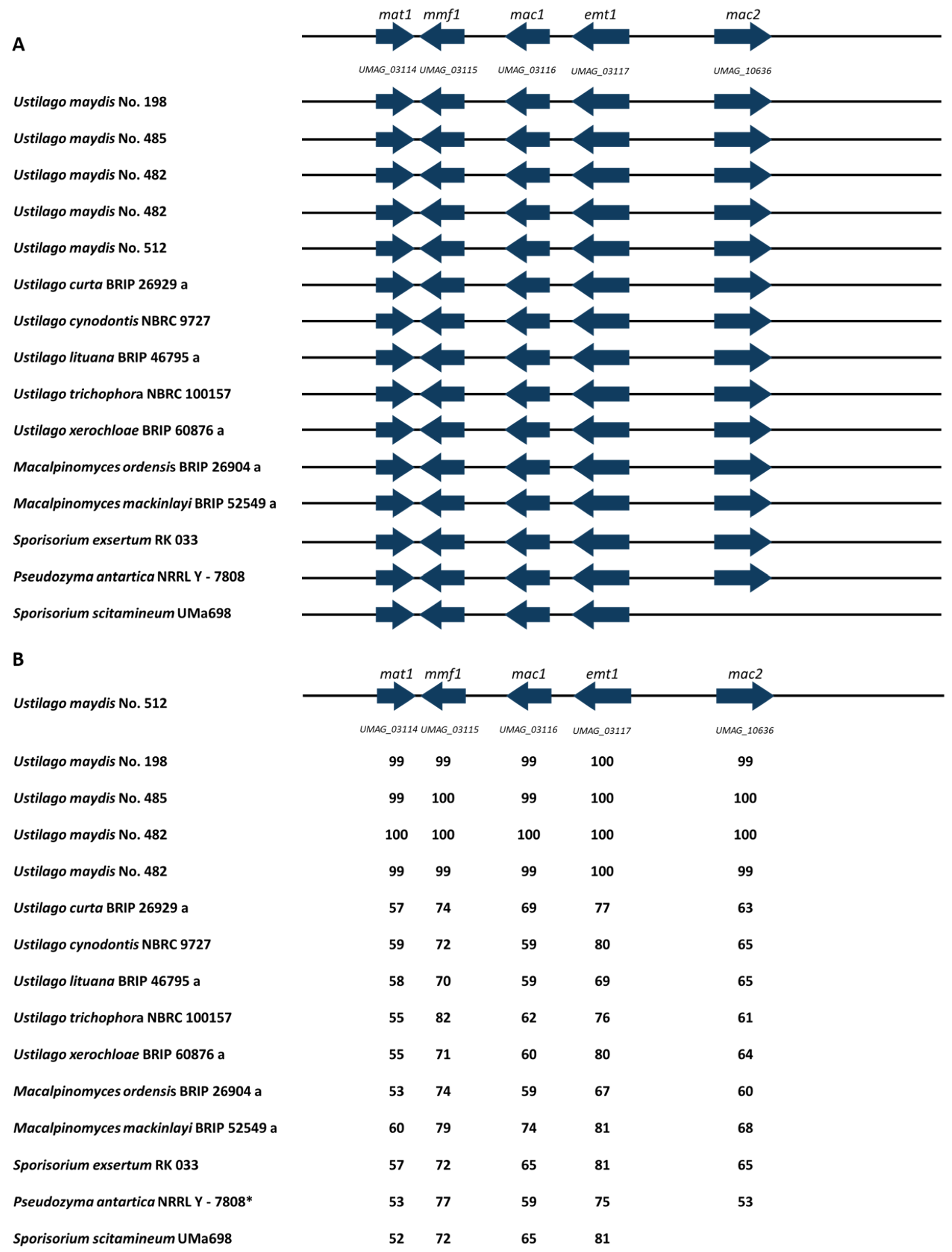
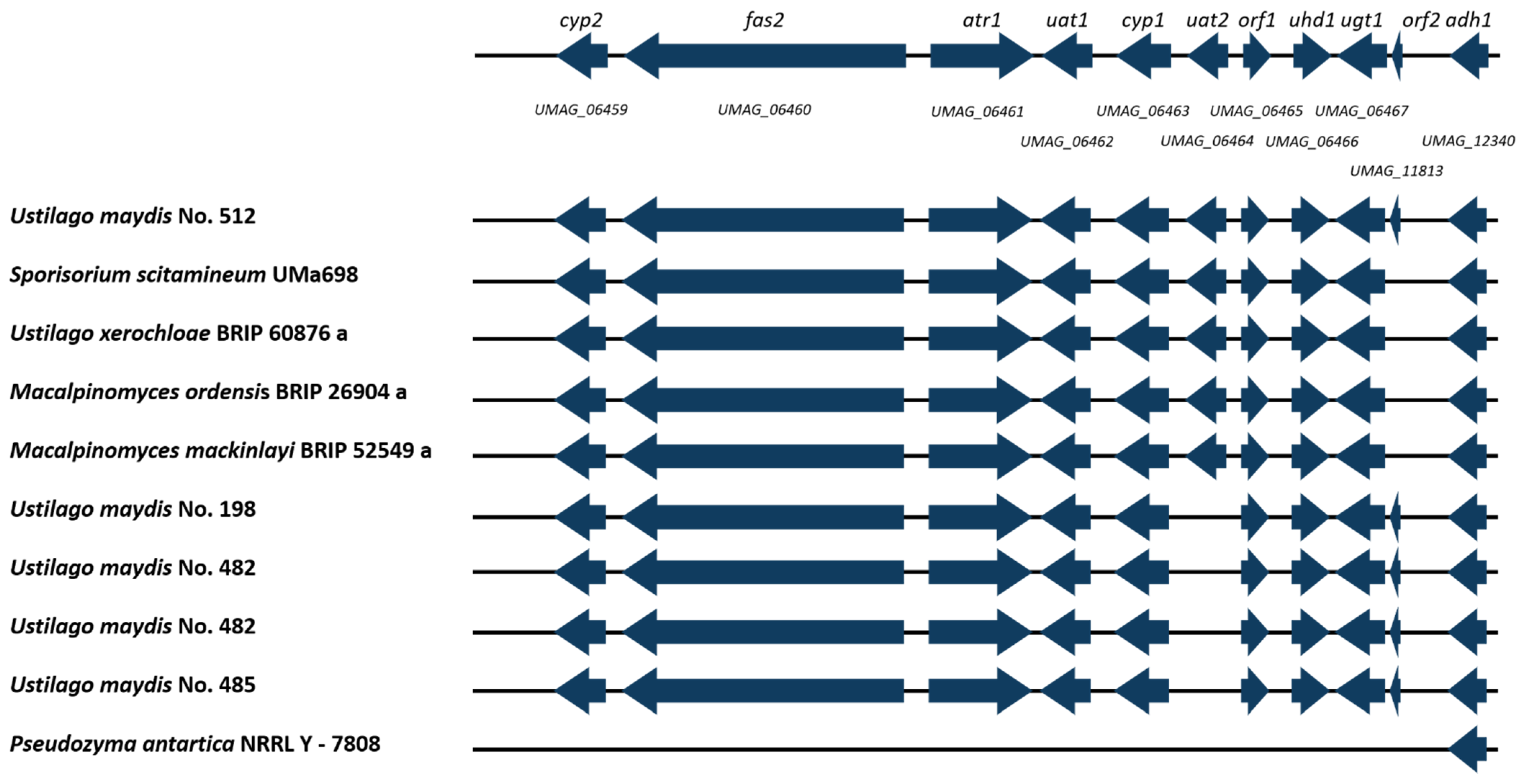
3.6. MEL Cluster Identification
3.7. Ustilagic Acid Cluster Identification
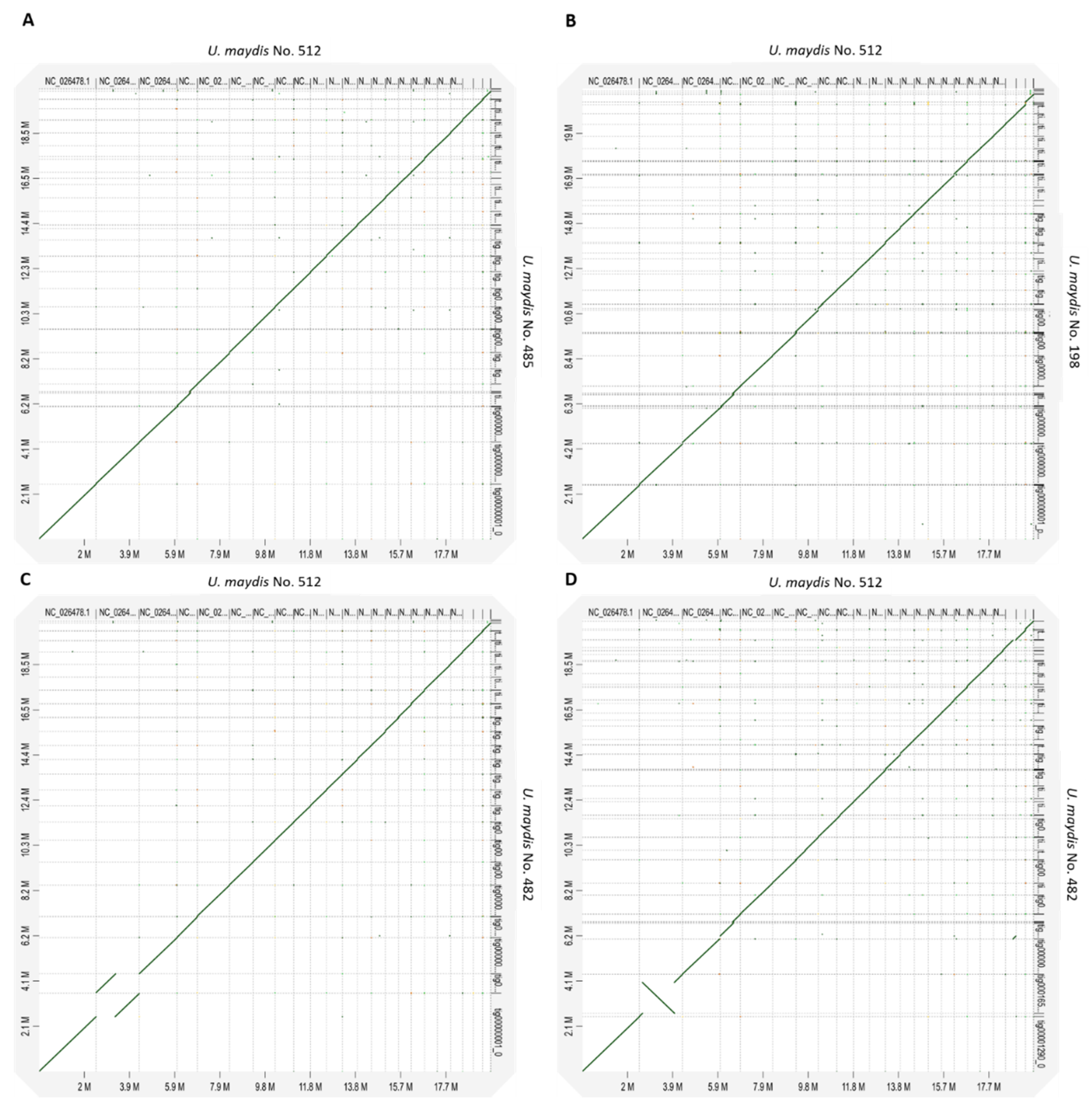
3.8. Macro Synteny Plot of U. maydis Isolates
4. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- McTaggart, A.; Shivas, R.G.; Boekhout, T.; Oberwinkler, F.; Vánky, K.; Pennycook, S.R.; Begerow, D. Mycosarcoma (Ustilaginaceae), a resurrected generic name for corn smut (Ustilago maydis) and its close relatives with hypertrophied, tubular sori. IMA Fungus 2016, 7, 309–315. [Google Scholar] [CrossRef] [PubMed]
- Vánky, K.; Lutz, M. Tubisorus, a new genus of smut fungi (Ustilaginomycetes) for Sorosporium pachycarpum. Mycol. Balc. 2011, 8, 129–135. [Google Scholar]
- Bakkeren, G.; Kämper, J.; Schirawski, J. Sex in smut fungi: Structure, function and evolution of mating-type complexes. Fungal Genet. Biol. 2008, 45 (Suppl. 1), S15–S21. [Google Scholar] [CrossRef] [PubMed]
- Saha, B.C. Emerging biotechnologies for production of itaconic acid and its applications as a platform chemical. J. Ind. Microbiol. Biotechnol. 2017, 44, 303–315. [Google Scholar] [CrossRef] [PubMed]
- Geiser, E.; Wiebach, V.; Wierckx, N.; Blank, L.M. Prospecting the biodiversity of the fungal family Ustilaginaceae for the production of value-added chemicals. Fungal Biol. Biotechnol. 2014, 1, 2. [Google Scholar] [CrossRef]
- Saika, A.; Koike, H.; Fukuoka, T.; Morita, T. Tailor-made mannosylerythritol lipids: Current state and perspectives. Appl. Microbiol. Biotechnol. 2018, 102, 6877–6884. [Google Scholar] [CrossRef] [PubMed]
- Guevarra, E.D.; Tabuchi, T. Accumulation of Itaconic, 2-Hydroxyparaconic, Itatartaric, and Malic Acids by Strains of the Genus Ustilago. Agric. Biol. Chem. 1990, 54, 2353–2358. [Google Scholar] [CrossRef][Green Version]
- Liu, Y.; Koh, C.M.J.; Ji, L. Bioconversion of crude glycerol to glycolipids in Ustilago maydis. Bioresour. Technol. 2011, 102, 3927–3933. [Google Scholar] [CrossRef]
- Maassen, N.; Panakova, M.; Wierckx, N.; Geiser, E.; Zimmermann, M.; Bölker, M.; Klinner, U.; Blank, L.M. Influence of carbon and nitrogen concentration on itaconic acid production by the smut fungus Ustilago maydis. Eng. Life Sci. 2013, 14, 129–134. [Google Scholar] [CrossRef]
- Ullmann, L.; Phan, A.N.T.; Kaplan, D.K.P.; Blank, L.M. Ustilaginaceae Biocatalyst for Co-Metabolism of CO2-Derived Substrates toward Carbon-Neutral Itaconate Production. J. Fungi 2021, 7, 98. [Google Scholar] [CrossRef] [PubMed]
- Zambanini, T.; Hosseinpour Tehrani, H.; Geiser, E.; Merker, D.; Schleese, S.; Krabbe, J.; Buescher, J.M.; Meurer, G.; Wierckx, N.; Blank, L.M. Efficient itaconic acid production from glycerol with Ustilago vetiveriae TZ1. Biotechnol. Biofuels 2017, 10, 131. [Google Scholar] [CrossRef]
- Steinberg, G.; Perez-Martin, J. Ustilago maydis, a new fungal model system for cell biology Introduction: Fungal model systems in cell biology. Trends Cell Biol. 2008, 18, 61–67. [Google Scholar] [CrossRef]
- Geiser, E.; Hosseinpour Tehrani, H.; Meyer, S.; Blank, L.M.; Wierckx, N. Evolutionary freedom in the regulation of the conserved itaconate cluster by Ria1 in related Ustilaginaceae. Fungal Biol. Biotechnol. 2018, 5, 14. [Google Scholar] [CrossRef]
- Geiser, E.; Przybilla, S.K.; Friedrich, A.; Buckel, W.; Wierckx, N.; Blank, L.M.; Bölker, M. Ustilago maydis produces itaconic acid via the unusual intermediate trans -aconitate. Microb. Biotechnol. 2015, 9, 116–126. [Google Scholar] [CrossRef]
- Vollmeister, E.; Schipper, K.; Feldbrügge, M. Microtubule-dependent mRNA transport in the model microorganism Ustilago maydis. RNA Biol. 2012, 9, 261–268. [Google Scholar] [CrossRef]
- Kämper, J.; Kahmann, R.; Bölker, M.; Ma, L.-J.; Brefort, T.; Saville, B.J.; Banuett, F.; Kronstad, J.W.; Gold, S.E.; Müller, O.; et al. Insights from the genome of the biotrophic fungal plant pathogen Ustilago maydis. Nature 2006, 444, 97–101. [Google Scholar] [CrossRef] [PubMed]
- Geiser, E.; Ludwig, F.; Zambanini, T.; Wierckx, N.; Blank, L.M. Draft Genome Sequences of Itaconate-Producing Ustilaginaceae. Genome Announc. 2016, 4, e01291-16. [Google Scholar] [CrossRef]
- Goodwin, S.; McPherson, J.D.; McCombie, W.R. Coming of age: Ten years of next-generation sequencing technologies. Nat. Rev. Genet. 2016, 17, 333–351. [Google Scholar] [CrossRef] [PubMed]
- Wibberg, D.; Stadler, M.; Lambert, C.; Bunk, B.; Spröer, C.; Rückert, C.; Kalinowski, J.; Cox, R.J.; Kuhnert, E. High quality genome sequences of thirteen Hypoxylaceae (Ascomycota) strengthen the phylogenetic family backbone and enable the discovery of new taxa. Fungal Divers. 2021, 106, 7–28. [Google Scholar] [CrossRef]
- Geiser, E.; Przybilla, S.K.; Engel, M.; Kleineberg, W.; Büttner, L.; Sarikaya, E.; Hartog, T.D.; Klankermayer, J.; Leitner, W.; Bölker, M.; et al. Genetic and biochemical insights into the itaconate pathway of Ustilago maydis enable enhanced production. Metab. Eng. 2016, 38, 427–435. [Google Scholar] [CrossRef]
- Wibberg, D.; Andersson, L.; Tzelepis, G.; Rupp, O.; Blom, J.; Jelonek, L.; Pühler, A.; Fogelqvist, J.; Varrelmann, M.; Schlüter, A.; et al. Genome analysis of the sugar beet pathogen Rhizoctonia solani AG2-2IIIB revealed high numbers in secreted proteins and cell wall degrading enzymes. BMC Genom. 2016, 17, 245. [Google Scholar] [CrossRef]
- Koren, S.; Walenz, B.P.; Berlin, K.; Miller, J.R.; Bergmann, N.H.; Phillippy, A.M. Canu: scalable and accurate long-read assembly via adaptive k-mer weighting and repeat separation. bioRxiv 2016, 25, 1–2. [Google Scholar] [CrossRef]
- Walker, B.J.; Abeel, T.; Shea, T.; Priest, M.; Abouelliel, A.; Sakthikumar, S.; Cuomo, C.A.; Zeng, Q.; Wortman, J.; Young, S.K.; et al. Pilon: An Integrated Tool for Comprehensive Microbial Variant Detection and Genome Assembly Improvement. PLoS ONE 2014, 9, e112963. [Google Scholar] [CrossRef] [PubMed]
- Li, H. Aligning sequence reads, clone sequences and assembly contigs with BWA-MEM. arXiv 2013, arXiv:1303.3997. [Google Scholar]
- Langmead, B.; Salzberg, S.L. Fast gapped-read alignment with Bowtie 2. Nat. Methods 2012, 9, 357–359. [Google Scholar] [CrossRef] [PubMed]
- Ter-Hovhannisyan, V.; Lomsadze, A.; Chernoff, Y.O.; Borodovsky, M. Gene prediction in novel fungal genomes using an ab initio algorithm with unsupervised training. Genome Res. 2008, 18, 1979–1990. [Google Scholar] [CrossRef]
- Meyer, F. GenDB--an open source genome annotation system for prokaryote genomes. Nucleic Acids Res. 2003, 31, 2187–2195. [Google Scholar] [CrossRef]
- Takahara, T.; Minamoto, T.; Yamanaka, H.; Doi, H.; Kawabata, Z. Estimation of Fish Biomass Using Environmental DNA. PLoS ONE 2012, 7, e35868. [Google Scholar] [CrossRef] [PubMed]
- Tatusov, R.L.; Fedorova, N.D.; Jackson, J.D.; Jacobs, A.R.; Kiryutin, B.; Koonin, E.V.; Krylov, D.M.; Mazumder, R.; Mekhedov, S.L.; Nikolskaya, A.N.; et al. The COG database: An updated version includes eukaryotes. BMC Bioinform. 2003, 4, 41. [Google Scholar] [CrossRef] [PubMed]
- Kanehisa, M.; Goto, S.; Kawashima, S.; Okuno, Y.; Hattori, M. The KEGG resource for deciphering the genome. Nucleic Acids Res. 2004, 32, D277–D280. [Google Scholar] [CrossRef] [PubMed]
- Boeckmann, B. The SWISS-PROT protein knowledgebase and its supplement TrEMBL in 2003. Nucleic Acids Res. 2003, 31, 365–370. [Google Scholar] [CrossRef] [PubMed]
- Lowe, T.M.; Eddy, S.R. tRNAscan-SE: A Program for Improved Detection of Transfer RNA Genes in Genomic Sequence. Nucleic Acids Res. 1997, 25, 955–964. [Google Scholar] [CrossRef] [PubMed]
- Simão, F.A.; Waterhouse, R.M.; Ioannidis, P.; Kriventseva, E.V.; Zdobnov, E.M. BUSCO: Assessing genome assembly and annotation completeness with single-copy orthologs. Bioinformatics 2015, 31, 3210–3212. [Google Scholar] [CrossRef]
- Blom, J.; Albaum, S.P.; Doppmeier, D.; Pühler, A.; Vorhölter, F.-J.; Zakrzewski, M.; Goesmann, A. EDGAR: A software framework for the comparative analysis of prokaryotic genomes. BMC Bioinform. 2009, 10, 154. [Google Scholar] [CrossRef] [PubMed]
- Blom, J.; Kreis, J.; Spänig, S.; Juhre, T.; Bertelli, C.; Ernst, C.; Goesmann, A. EDGAR 2.0: An enhanced software platform for comparative gene content analyses. Nucleic Acids Res. 2016, 44, W22–W28. [Google Scholar] [CrossRef] [PubMed]
- Wibberg, D.; Rupp, O.; Blom, J.; Jelonek, L.; Kröber, M.; Verwaaijen, B.; Goesmann, A.; Albaum, S.; Grosch, R.; Pühler, A.; et al. Development of a Rhizoctonia solani AG1-IB Specific Gene Model Enables Comparative Genome Analyses between Phytopathogenic R. solani AG1-IA, AG1-IB, AG3 and AG8 Isolates. PLoS ONE 2015, 10, e0144769. [Google Scholar] [CrossRef] [PubMed]
- Edgar, R.C. MUSCLE: Multiple sequence alignment with high accuracy and high throughput. Nucleic Acids Res. 2004, 32, 1792–1797. [Google Scholar] [CrossRef] [PubMed]
- Qin, Q.-L.; Xie, B.-B.; Zhang, X.-Y.; Chen, X.-L.; Zhou, B.-C.; Zhou, J.; Oren, A.; Zhang, Y.-Z. A Proposed Genus Boundary for the Prokaryotes Based on Genomic Insights. J. Bacteriol. 2014, 196, 2210–2215. [Google Scholar] [CrossRef]
- Adamek, M.; Alanjary, M.; Sales-Ortells, H.; Goodfellow, M.; Bull, A.T.; Winkler, A.; Wibberg, D.; Kalinowski, J.; Ziemert, N. Comparative genomics reveals phylogenetic distribution patterns of secondary metabolites in Amycolatopsis species. BMC Genom. 2018, 19, 426. [Google Scholar] [CrossRef] [PubMed]
- Margos, G.; Gofton, A.; Wibberg, D.; Dangel, A.; Marosevic, D.; Loh, S.-M.; Oskam, C.; Fingerle, V. The genus Borrelia reloaded. PLoS ONE 2018, 13, e0208432. [Google Scholar] [CrossRef]
- Camacho, C.; Coulouris, G.; Avagyan, V.; Ma, N.; Papadopoulos, J.; Bealer, K.; Madden, T.L. BLAST+: Architecture and applications. BMC Bioinform. 2009, 10, 421. [Google Scholar] [CrossRef] [PubMed]
- Goris, J.; Konstantinidis, K.T.; Klappenbach, J.A.; Coenye, T.; Vandamme, P.; Tiedje, J.M. DNA–DNA hybridization values and their relationship to whole-genome sequence similarities. Int. J. Syst. Evol. Microbiol. 2007, 57, 81–91. [Google Scholar] [CrossRef]
- Wibberg, D.; Rupp, O.; Jelonek, L.; Kröber, M.; Verwaaijen, B.; Blom, J.; Winkler, A.; Goesmann, A.; Grosch, R.; Pühler, A.; et al. Improved genome sequence of the phytopathogenic fungus Rhizoctonia solani AG1-IB 7/3/14 as established by deep mate-pair sequencing on the MiSeq (Illumina) system. J. Biotechnol. 2015, 203, 19–21. [Google Scholar] [CrossRef] [PubMed]
- Hosseinpour Tehrani, H.; Tharmasothirajan, A.; Track, E.; Blank, L.M.; Wierckx, N. Engineering the morphology and metabolism of pH tolerant Ustilago cynodontis for efficient itaconic acid production. Metab. Eng. 2019, 54, 293–300. [Google Scholar] [CrossRef]
- Zambanini, T.; Sarikaya, E.; Kleineberg, W.; Buescher, J.M.; Meurer, G.; Wierckx, N.; Blank, L.M. Efficient malic acid production from glycerol with Ustilago trichophora TZ1. Biotechnol. Biofuels 2016, 9, 67. [Google Scholar] [CrossRef] [PubMed]
- Becker, J.; Tehrani, H.H.; Ernst, P.; Blank, L.M.; Wierckx, N. An Optimized Ustilago maydis for Itaconic Acid Production at Maximal Theoretical Yield. J. Fungi 2020, 7, 20. [Google Scholar] [CrossRef] [PubMed]
- McTaggart, A.; Shivas, R.G.; Geering, A.D.W.; Callaghan, B.; Vanky, K.; Scharaschkin, T. Soral synapomorphies are significant for the systematics of the Ustilago-Sporisorium-Macalpinomyces complex (Ustilaginaceae). Persoonia—Mol. Phylogeny Evol. Fungi 2012, 29, 63–77. [Google Scholar] [CrossRef] [PubMed]
- Levinson, W.E.; Kurtzman, C.P.; Kuo, T.M. Production of itaconic acid by Pseudozyma antarctica NRRL Y-7808 under nitrogen-limited growth conditions. Enzym. Microb. Technol. 2006, 39, 824–827. [Google Scholar] [CrossRef]
- Hudec, C.; Biessy, A.; Novinscak, A.; St-Onge, R.; Lamarre, S.; Blom, J.; Filion, M. Comparative Genomics of Potato Common Scab-Causing Streptomyces spp. Displaying Varying Virulence. Front. Microbiol. 2021, 12, 716522. [Google Scholar] [CrossRef] [PubMed]
- Piepenbring, M.; Stoll, M.; Oberwinkler, F. The generic position of Ustilago maydis, Ustilago scitaminea, and Ustilago esculenta (Ustilaginales). Mycol. Prog. 2002, 1, 71–80. [Google Scholar] [CrossRef]
- Stoll, M.; Begerow, D.; Oberwinkler, F. Molecular phylogeny of Ustilago, Sporisorium, and related taxa based on combined analyses of rDNA sequences. Mycol. Res. 2005, 109, 342–356. [Google Scholar] [CrossRef]
- Jain, C.; Rodriguez-R, L.M.; Phillippy, A.M.; Konstantinidis, K.T.; Aluru, S. High throughput ANI analysis of 90K prokaryotic genomes reveals clear species boundaries. Nat. Commun. 2018, 9, 5114. [Google Scholar] [CrossRef] [PubMed]
- Wibberg, D.; Genzel, F.; Verwaaijen, B.; Blom, J.; Rupp, O.; Goesmann, A.; Zrenner, R.; Grosch, R.; Pühler, A.; Schlüter, A. Genome Analyses of the Less Aggressive Rhizoctonia solani. J. Fungi 2021, 7, 832. [Google Scholar] [CrossRef] [PubMed]
- Alshahni, M.M.; Yamada, T.; Yo, A.; Murayama, S.Y.; Kuroda, M.; Hoshino, Y.; Ishikawa, J.; Watanabe, S.; Makimura, K. Insight into the draft whole-genome sequence of the dermatophyte Arthroderma vanbreuseghemii. Sci. Rep. 2018, 8, 15127. [Google Scholar] [CrossRef] [PubMed]
- Castro-Chavez, F. The rules of variation: Amino acid exchange according to the rotating circular genetic code. J. Theor. Biol. 2010, 264, 711–721. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Bölker, M.; Basse, C.W.; Schirawski, J. Ustilago maydis secondary metabolism—From genomics to biochemistry. Fungal Genet. Biol. 2008, 45, S88–S93. [Google Scholar] [CrossRef] [PubMed]
- Reyes-Fernández, E.Z.; Shi, Y.-M.; Grün, P.; Bode, H.B.; Bölker, M. An Unconventional Melanin Biosynthesis Pathway in Ustilago maydis. Appl. Environ. Microbiol. 2021, 87, e01510-20. [Google Scholar] [CrossRef]
- Blin, K.; Shaw, S.; Kloosterman, A.M.; Charlop-Powers, Z.; van Wezel, G.P.; Medema, M.H.; Weber, T. antiSMASH 6.0: Improving cluster detection and comparison capabilities. Nucleic Acids Res. 2021, 49, W29–W35. [Google Scholar] [CrossRef]
- Bruce, S.A.; Saville, B.J.; Emery, R.J.N. Ustilago maydis Produces Cytokinins and Abscisic Acid for Potential Regulation of Tumor Formation in Maize. J. Plant Growth Regul. 2011, 30, 51–63. [Google Scholar] [CrossRef]
- Winterberg, B.; Uhlmann, S.; Linne, U.; Lessing, F.; Marahiel, M.A.; Eichhorn, H.; Kahmann, R.; Schirawski, J. Elucidation of the complete ferrichrome A biosynthetic pathway in Ustilago maydis. Mol. Microbiol. 2010, 75, 1260–1271. [Google Scholar] [CrossRef] [PubMed]
- Geiser, E. Itaconic Acid Production by Ustilago Maydis. Ph.D Dissertation, RWTH Aachen University, Aachen, Germany, 2015. [Google Scholar]
- Wierckx, N.; Agrimi, G.; Lübeck, P.S.; Steiger, M.; Mira, N.; Punt, P.J. Metabolic specialization in itaconic acid production: A tale of two fungi. Curr. Opin. Biotechnol. 2020, 62, 153–159. [Google Scholar] [CrossRef] [PubMed]
- Zhu, X.; Guo, Y.; Liu, Z.; Yang, J.; Tang, H.; Wang, Y. Itaconic acid exerts anti-inflammatory and antibacterial effects via promoting pentose phosphate pathway to produce ROS. Sci. Rep. 2021, 11, 18173. [Google Scholar] [CrossRef] [PubMed]
- Van Nguyen, T.; Alfaro, A.C.; Young, T.; Green, S.; Zarate, E.; Merien, F. Itaconic acid inhibits growth of a pathogenic marine Vibrio strain: A metabolomics approach. Sci. Rep. 2019, 9, 5937. [Google Scholar] [CrossRef] [PubMed]
- Loevenich, J. Optimization of Itaconic Acid Production by U. maydis through Metabolic Engineering & Adaptive Laboratory Evolution. Ph.D Dissertation, RWTH Aachen University, Aachen, Germany, 2019. [Google Scholar]
- Saika, A.; Fukuoka, T.; Koike, H.; Yamamoto, S.; Sugahara, T.; Sogabe, A.; Kitamoto, D.; Morita, T. A putative transporter gene PtMMF1-deleted strain produces mono-acylated mannosylerythritol lipids in Pseudozyma tsukubaensis. Appl. Microbiol. Biotechnol. 2020, 104, 10105–10117. [Google Scholar] [CrossRef] [PubMed]
- Hewald, S.; Linne, U.; Scherer, M.; Marahiel, M.A.; Kamper, J.; Bolker, M. Identification of a Gene Cluster for Biosynthesis of Mannosylerythritol Lipids in the Basidiomycetous Fungus Ustilago maydis. Appl. Environ. Microbiol. 2006, 72, 5469–5477. [Google Scholar] [CrossRef]
- Hewald, S.; Josephs, K.; Bolker, M. Genetic Analysis of Biosurfactant Production in Ustilago maydis. Appl. Environ. Microbiol. 2005, 71, 3033–3040. [Google Scholar] [CrossRef]
- Morita, T.; Konishi, M.; Fukuoka, T.; Imura, T.; Kitamoto, D. Identification of Ustilago cynodontis as a new producer of glycolipid biosurfactants, mannosylerythritol lipids, based on ribosomal DNA sequences. J. Oleo Sci. 2008, 57, 549–556. [Google Scholar] [CrossRef]
- Ron, E.Z.; Rosenberg, E. Natural roles of biosurfactants. Minireview. Environ. Microbiol. 2001, 3, 229–236. [Google Scholar] [CrossRef]
- Haskins, R.H.; Thorn, J.A. Biochemistry of the ustilaginales: Vii. antibiotic activity of ustilagic acid. Can. J. Bot. 1951, 29, 585–592. [Google Scholar] [CrossRef]



| Strain No. | Assembly Name | Assembly ACC | Study ID | Sample ID | Contig ACC |
|---|---|---|---|---|---|
| #2169 | Umay_482_v1 | GCA_928722285 | PRJEB50355 | ERS10392836 | CAKMXG010000001-CAKMXG010000046 |
| #2169 | Umay_482_v2 | GCA_928722265 | PRJEB50355 | ERS10392837 | CAKMXF010000001-CAKMXF010000068 |
| #2172 | Umay_485 | GCA_928722245 | PRJEB50356 | ERS10381369 | CAKMXD010000001-CAKMXD010000043 |
| #2136 | Umay_198 | GCA_928743665 | PRJEB50359 | ERS10419843 | CAKMXO010000001-CAKMXO010000107 |
| #2816 | BRIP_26904_a | GCA_928724745 | PRJEB50498 | ERS10395116 | CAKMXJ010000001-CAKMXJ010000042 |
| #2701 | NBRC_100157 | GCA_928724775 | PRJEB50365 | ERS10395119 | CAKMXL010000001-CAKMXL010000048 |
| #2212 | RK_033 | GCA_928724755 | PRJEB50360 | ERS10395351 | CAKMXK010000001-CAKMXK010000041 |
| #2215 | UMa706_1 | GCA_928724875 | PRJEB50363 | ERS10395376 | CAKMXN010000001-CAKMXN010000086 |
| #2215 | UMa706_2 | GCA_928724795 | PRJEB50363 | ERS10395377 | CAKMXM010000001-CAKMXM010000036 |
| #2814 | BRIP_52549_a | GCA_928852645 | PRJEB50497 | ERS10422304 | CAKMXS010000001-CAKMXS010000079 |
| #2821 | BRIP_26929_a | GCA_928722295 | PRJEB50499 | ERS10393563 | CAKMXH010000001-CAKMXH010000039 |
| #2214 | RK031 | GCA_928858565 | PRJEB50362 | ERS10422305 | CAKMXV010000001-CAKMXV010000022 |
| #1946 | NRRL_Y-7808 | GCA_928872045 | PRJEB50496 | ERS10422313 | CAKMYB010000001-CAKMYB010000027 |
| #2706 | NBRC_9727 | GCA_928865425 | PRJEB50366 | ERS10422385 | CAKMXY010000001-CAKMXY010000048 |
| #2826 | BRIP_46795_a | GCA_928869825 | PRJEB50500 | ERS10422474 | CAKMYA010000001-CAKMYA010000069 |
| #2836 | BRIP_60876_a | GCA_928856705 | RJEB50501 | ERS10422499 | CAKMXT010000001-CAKMXT010000026 |
| #2836 * | - | ERZ4998446 | PRJEB50495 | ERS10422500 | ERZ4998446.1-ERZ4998446.260 |
| #2213 | UMa698 | GCA_928991175 | PRJEB50361 | ERS10422257 | CAKMYD010000001-CAKMYD010000051 |
| #2220 | RK_075 | GCA_928722275 | PRJEB50364 | ERS10392838 | CAKMXE010000001-CAKMXE010000037 |
| No. | Organism | Strain | Genome Size (bp) | Contigs | Largest Contig | N50 (bp) | GC (%) | Annotated Genes a |
|---|---|---|---|---|---|---|---|---|
| #2814 | Macalpinomyces mackinlayi | BRIP 52549a | 20,011,713 | 79 | 2,517,462 | 778,176 | 55.2 | 6780 |
| #2816 | Macalpinomyces ordensis | BRIP 26904a | 21,488,978 | 42 | 1,934,029 | 921,621 | 54.4 | 7166 |
| #1946 | Pseudozyma antarctica | NRRLY 7808 | 18,256,718 | 27 | 2,397,276 | 72,913 | 60.8 | 6532 |
| #2212 | Sporisorium exsertum | RK 033 | 19,675,720 | 41 | 1,762,232 | 1,163,924 | 56.7 | 6772 |
| #2213 | Sporisorium scitamineum | UMa698, | 20,280,126 | 51 | 1,999,406 | 881,538 | 54.9 | 6757 |
| #2214 | Sporisorium walkeri | RK 031 | 18,415,360 | 22 | 2,584,725 | 1,158,577 | 53.9 | 6584 |
| #2215 | Ustanciosporium gigantosporum * | UMa706 | 26,778,318 | 86 | 3,628,339 | 1,232,426 | 55.3 | 8712 |
| #2215 | Ustanciosporium gigantosporum | UMa706 | 20,177,783 | 36 | 3,087,333 | 1,238,668 | 52.0 | 6690 |
| #2821 | Ustilago curta | BRIP 26929a | 18,446,555 | 39 | 1,762,630 | 682,062 | 55.2 | 6420 |
| #2706 | Ustilago cynodontis | NBRC 9727 | 23,130,474 | 48 | 2,786,107 | 1,026,819 | 52.1 | 7322 |
| #2826 | Ustilago lituana | BRIP 46795a | 25,770,532 | 69 | 1,993,860 | 819,494 | 54.3 | 7741 |
| #2136 | Ustilago maydis | No. 198 | 21,122,121 | 107 | 2,522,778 | 660,941 | 53.8 | 6807 |
| #2169 | Ustilago maydis | No. 482 | 20,585,367 | 46 | 3,568,257 | 87,601 | 54.0 | 6827 |
| #2169 | Ustilago maydis | No. 482 | 20,591,394 | 68 | 2,499,251 | 674,905 | 53.9 | 6833 |
| #2172 | Ustilago maydis | No. 485 | 20,582,581 | 43 | 2,514,809 | 743,668 | 53.9 | 6777 |
| #2701 | Ustilago trichophora | NBRC 100157 | 20,713,809 | 48 | 1,974,389 | 794,480 | 53.8 | 6585 |
| #2220 | Ustilago vetiveriae | RK 075 | 18,373,116 | 37 | 2,393,662 | 780,848 | 54.8 | 6347 |
| #2836 | Ustilago xerochloae * | BRIP 60876a | 36,081,614 | 315 | 4,586,684 | 648,688 | 54.8 | 11,418 |
| Organism | Strain | NRPS | Terpene | T1PKS | RIPP | Total |
|---|---|---|---|---|---|---|
| Ustilago maydis | No. 485 | 11 | 2 | 1 | 0 | 14 |
| Ustilago maydis | No. 482 1 | 11 | 2 | 1 | 0 | 14 |
| Ustilago maydis | No. 482 2 | 13 | 2 | 1 | 0 | 15 |
| Ustilago vetiveriae | RK 075 | 7 | 2 | 1 | 0 | 10 |
| Ustilago curta | BRIP 26929a | 7 | 3 | 1 | 0 | 11 |
| Macalpinomyces ordensis | BRIP 26904a | 6 | 3 | 1 | 0 | 10 |
| Ustilago trichophora | NBRC 100157 | 7 | 2 | 1 | 0 | 10 |
| Sporisorium exsertum | RK 033 | 8 | 3 | 1 | 0 | 12 |
| Ustanciosporium gigantosporum | UMa706 1 | 7 | 4 | 1 | 1 | 13 |
| Sporisorium scitamineum | UMa698 | 9 | 3 | 1 | 0 | 13 |
| Ustilago xerochloae | BRIP 60876a * | 8 | 2 | 1 | 0 | 11 |
| Ustilago xerochloae | BRIP 60876a * | 5 | 2 | 0 | 0 | 7 |
| Macalpinomyces mackinlayi | BRIP 52549a | 7 | 3 | 2 | 0 | 12 |
| Ustanciosporium gigantosporum | UMa706 2 | 7 | 2 | 1 | 0 | 10 |
| Sporisorium walkeri | RK 031 | 6 | 3 | 1 | 0 | 10 |
| Ustilago maydis | No. 198 | 11 | 2 | 1 | 0 | 14 |
| Ustilago cynodontis | NBRC 9727 | 9 | 2 | 1 | 0 | 12 |
| Ustilago lituana | BRIP 46795a | 6 | 3 | 1 | 0 | 10 |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Ullmann, L.; Wibberg, D.; Busche, T.; Rückert, C.; Müsgens, A.; Kalinowski, J.; Blank, L.M. Seventeen Ustilaginaceae High-Quality Genome Sequences Allow Phylogenomic Analysis and Provide Insights into Secondary Metabolite Synthesis. J. Fungi 2022, 8, 269. https://doi.org/10.3390/jof8030269
Ullmann L, Wibberg D, Busche T, Rückert C, Müsgens A, Kalinowski J, Blank LM. Seventeen Ustilaginaceae High-Quality Genome Sequences Allow Phylogenomic Analysis and Provide Insights into Secondary Metabolite Synthesis. Journal of Fungi. 2022; 8(3):269. https://doi.org/10.3390/jof8030269
Chicago/Turabian StyleUllmann, Lena, Daniel Wibberg, Tobias Busche, Christian Rückert, Andreas Müsgens, Jörn Kalinowski, and Lars M. Blank. 2022. "Seventeen Ustilaginaceae High-Quality Genome Sequences Allow Phylogenomic Analysis and Provide Insights into Secondary Metabolite Synthesis" Journal of Fungi 8, no. 3: 269. https://doi.org/10.3390/jof8030269
APA StyleUllmann, L., Wibberg, D., Busche, T., Rückert, C., Müsgens, A., Kalinowski, J., & Blank, L. M. (2022). Seventeen Ustilaginaceae High-Quality Genome Sequences Allow Phylogenomic Analysis and Provide Insights into Secondary Metabolite Synthesis. Journal of Fungi, 8(3), 269. https://doi.org/10.3390/jof8030269







