Analysis of Stored Wheat Grain-Associated Microbiota Reveals Biocontrol Activity among Microorganisms against Mycotoxigenic Fungi
Abstract
:1. Introduction
2. Materials and Methods
2.1. Wheat Grain Sources
2.2. DNA Extraction, Amplification, and Sequencing
2.3. Bioinformatics
2.4. Isolation and Identification of Microorganisms
2.5. In Vitro Antagonism against Mycotoxigenic Fungi
2.6. Biocontrol Activity of Selected Bacterial Strains and Yeast against Mycotoxigenic Fungi on Wheat Grains
2.7. Aflatoxin-Degradation Assays
2.8. HPLC Analysis
2.9. Statistical Analysis
3. Results and Discussion
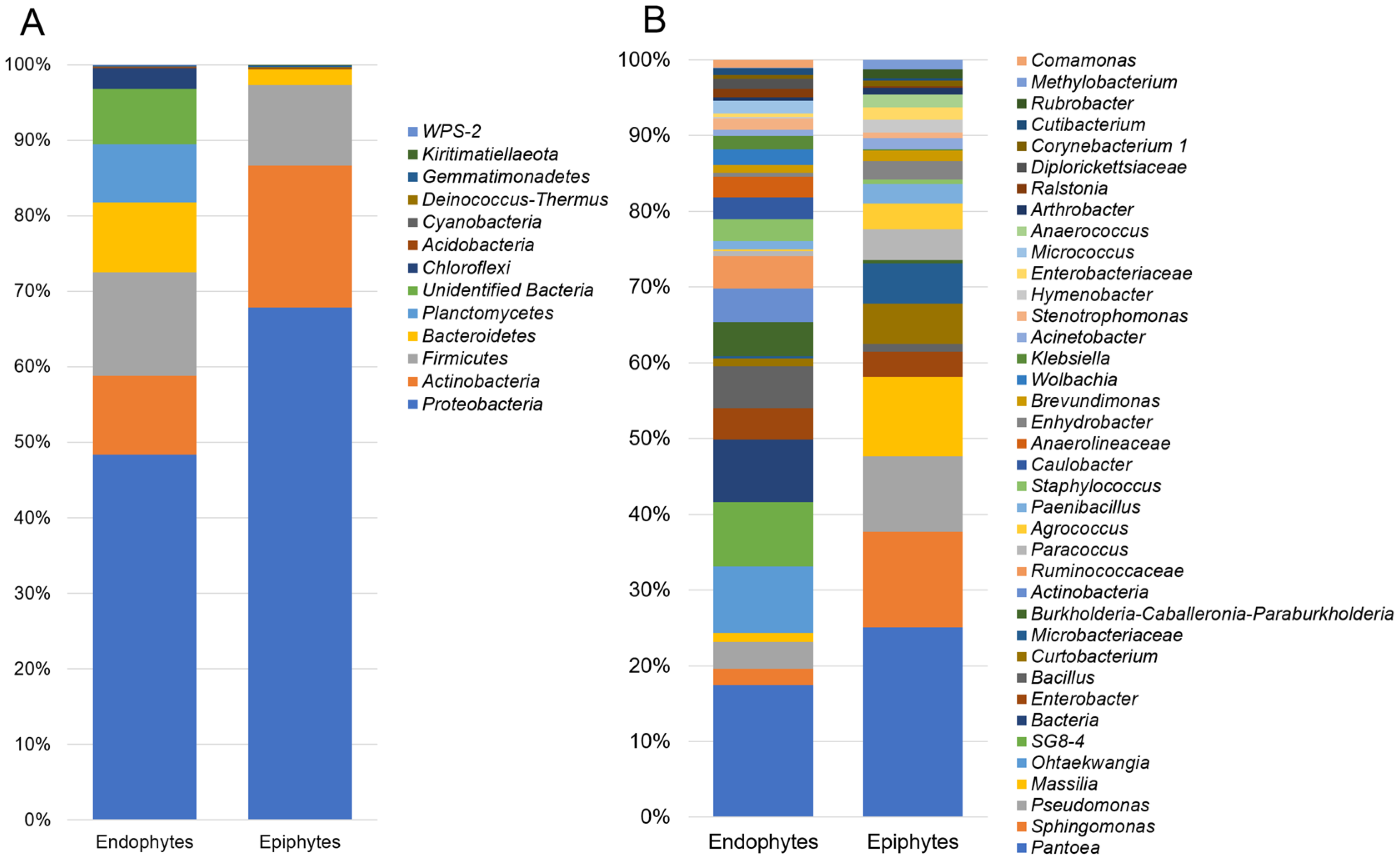
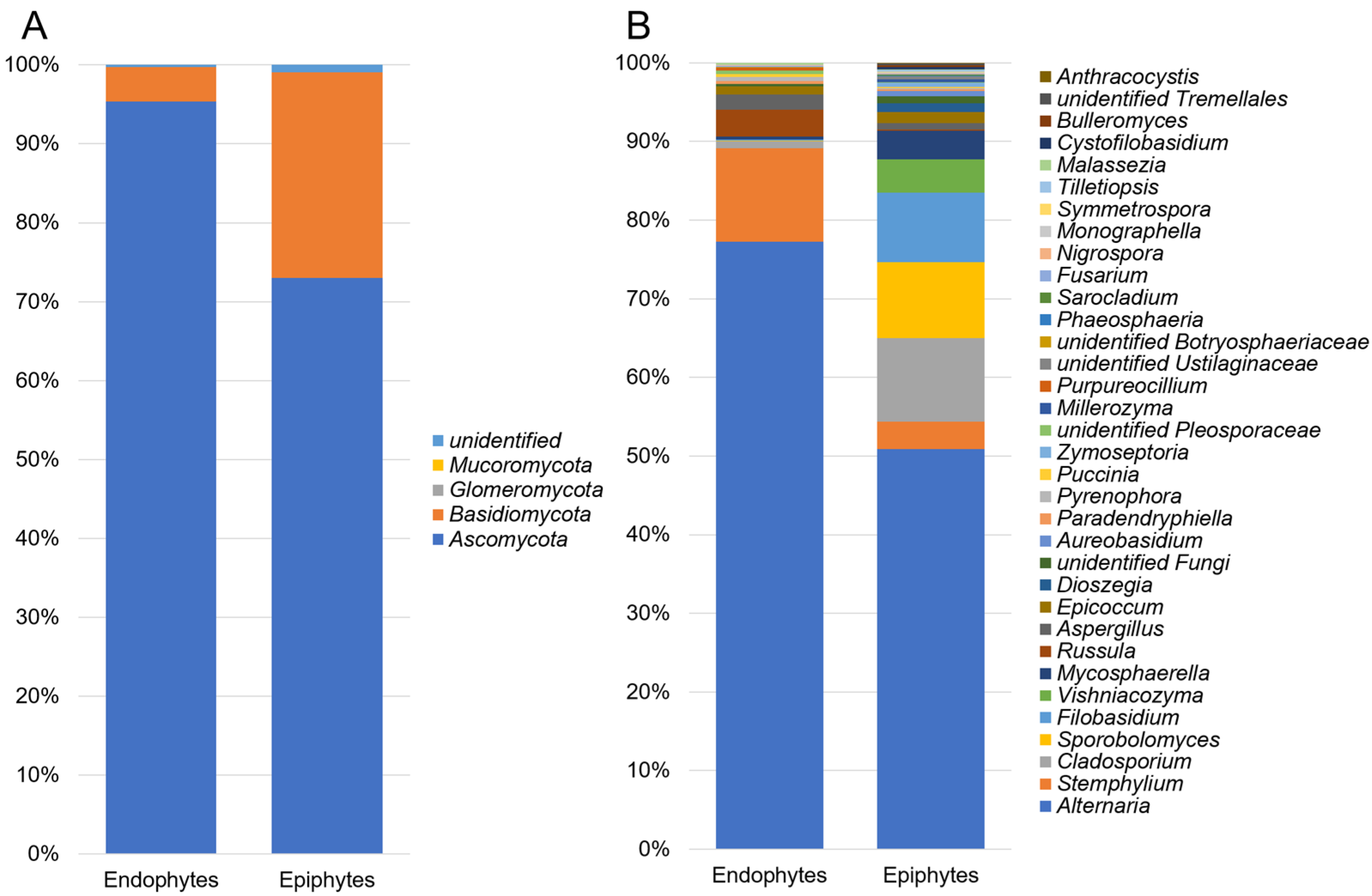
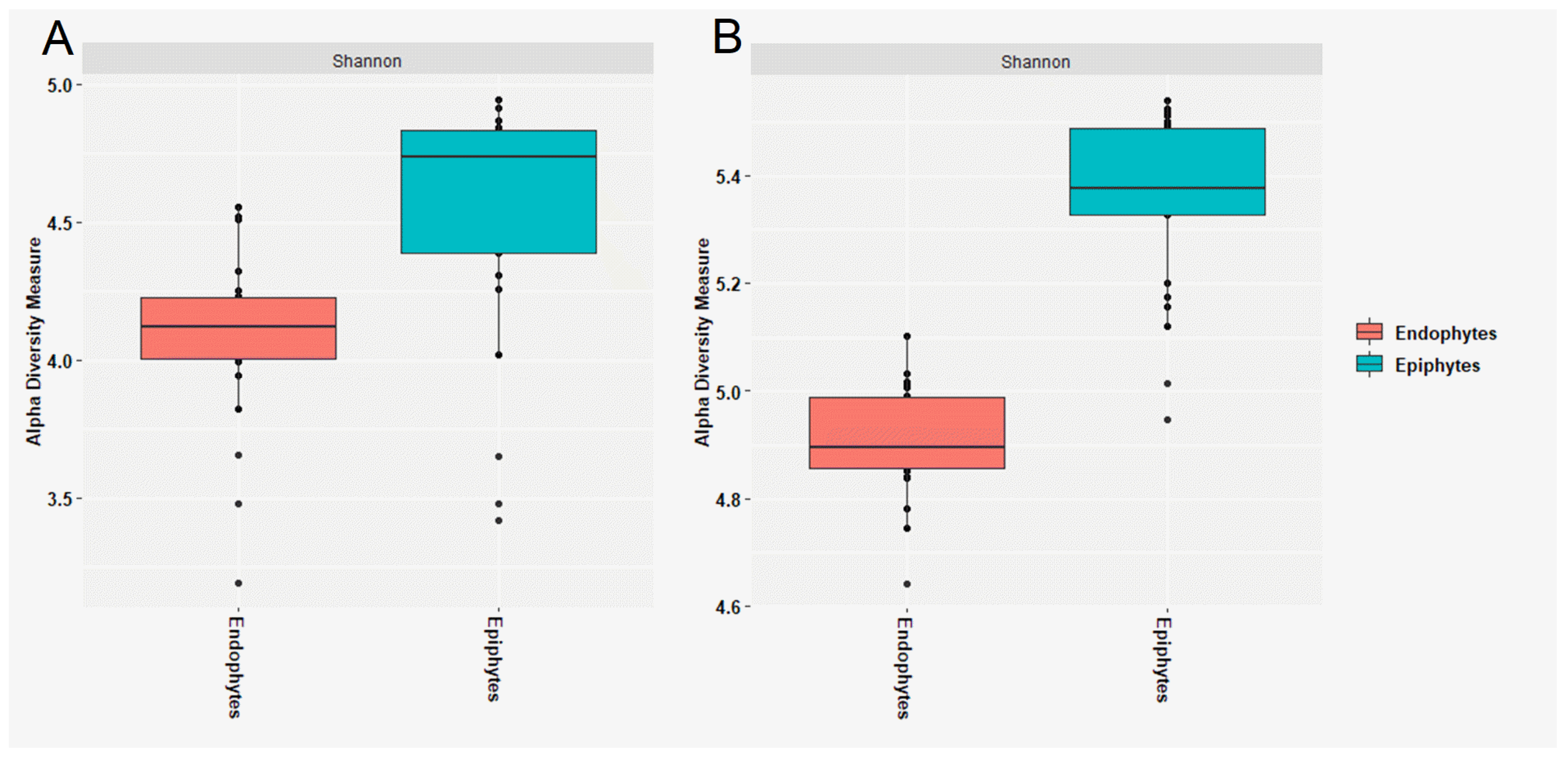
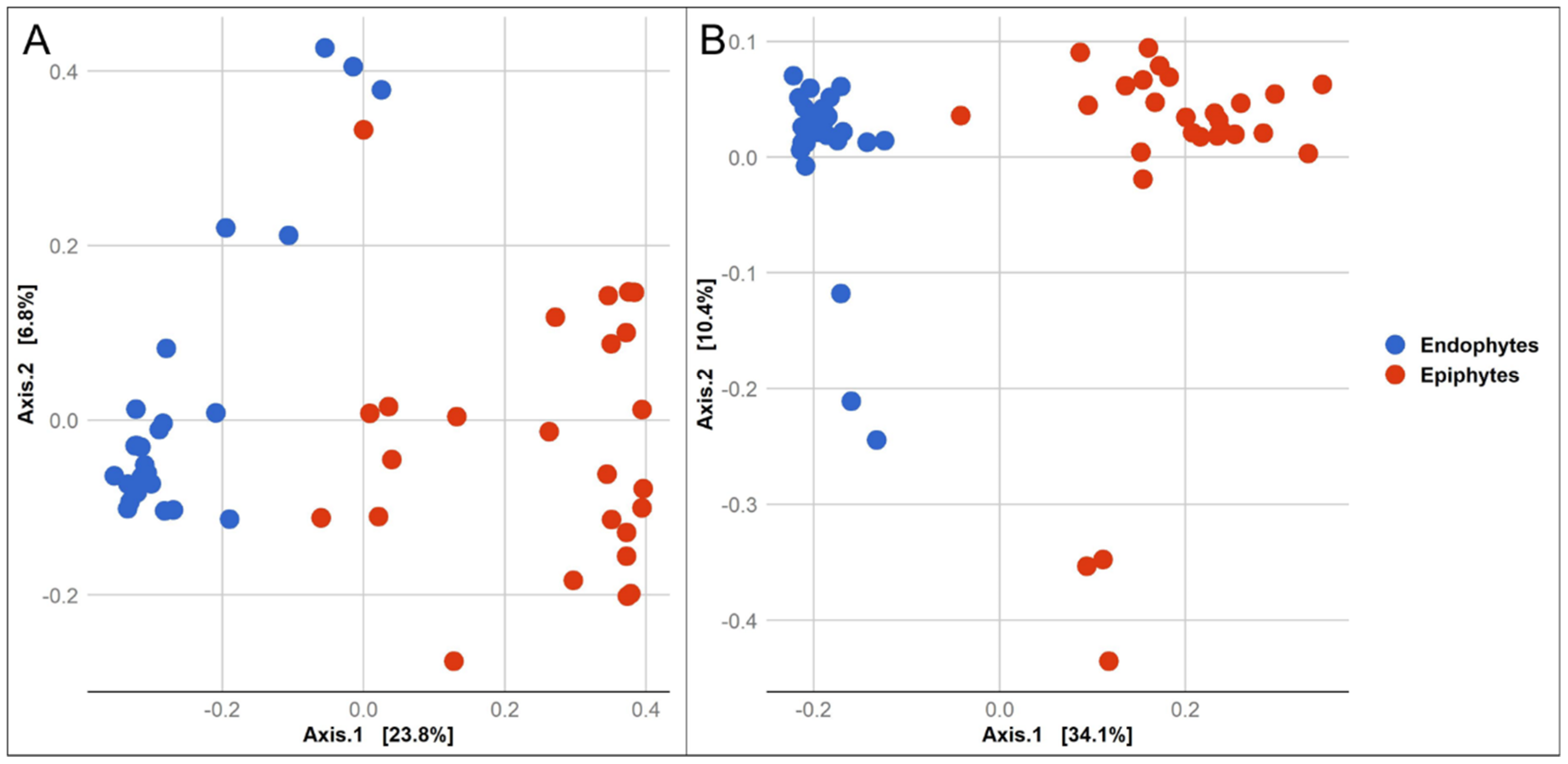
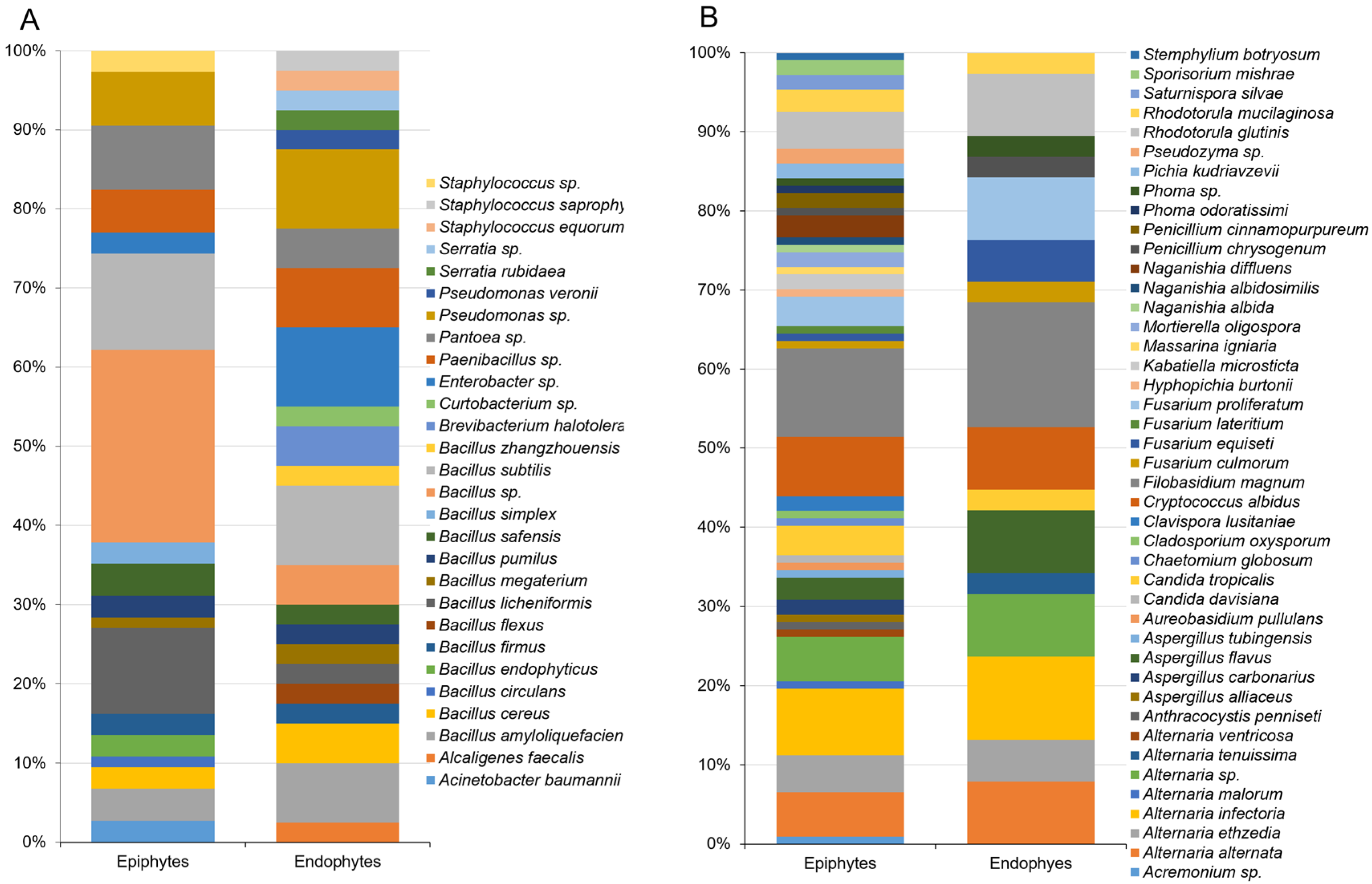
3.1. Composition of Epiphytic and Endophytic Microbiota Associated with Stored Wheat Grain
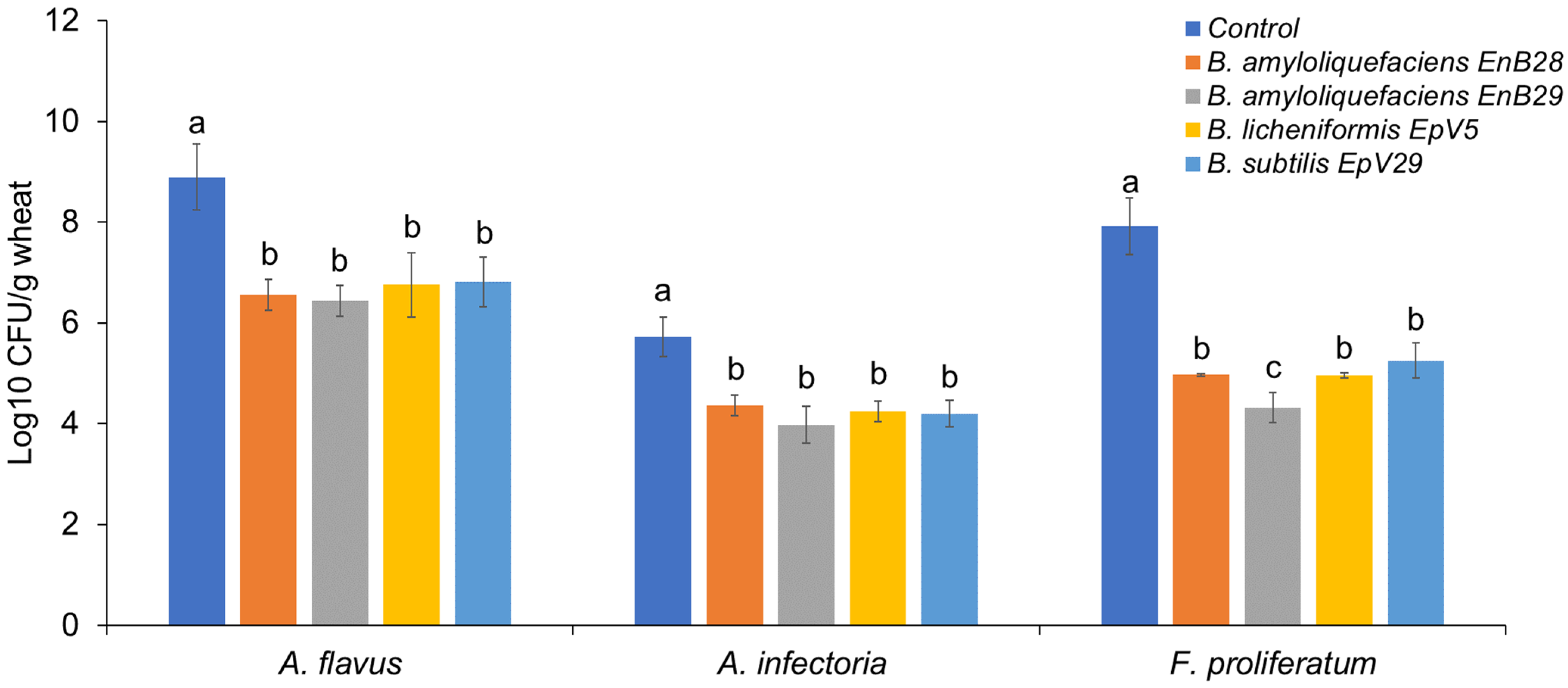
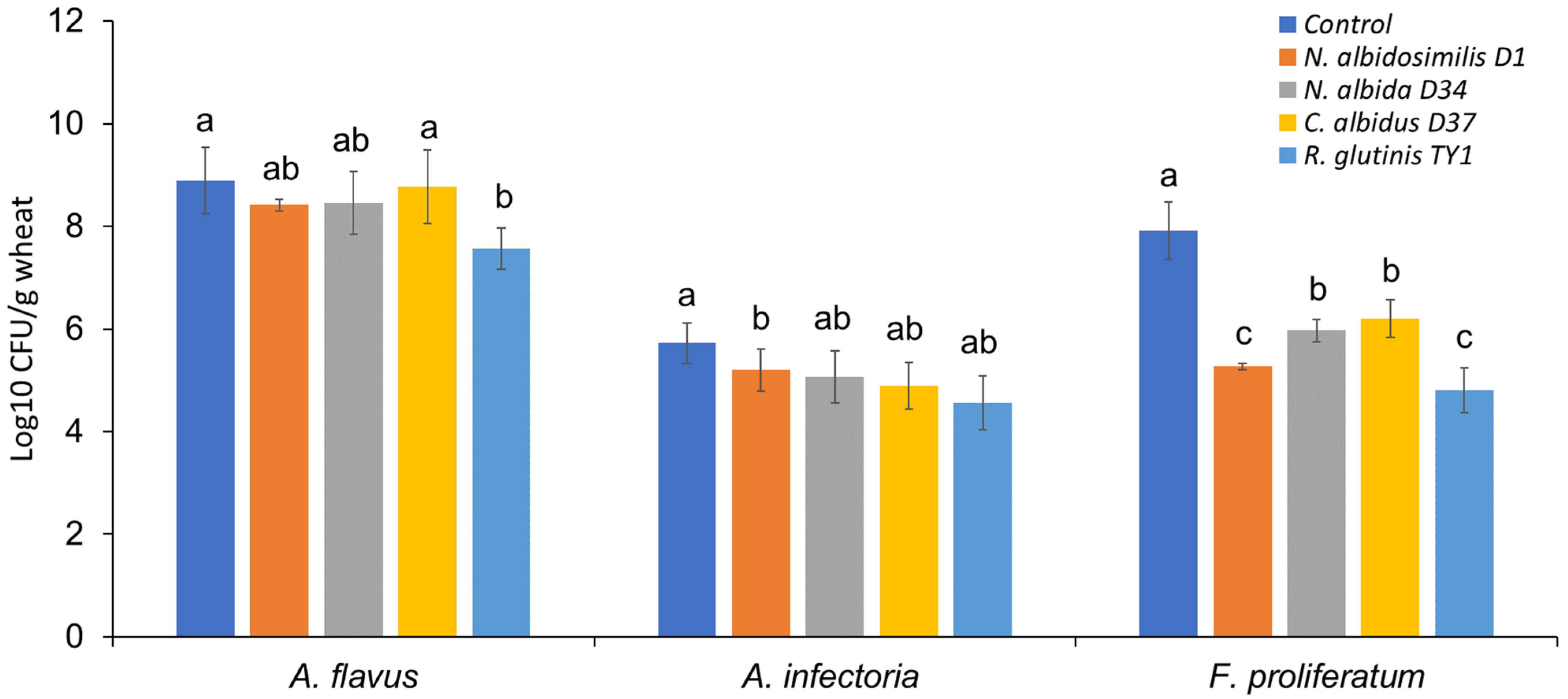
3.2. Microbe Isolation from Stored Grains and the Antagonistic Activity of Selected Isolates against Mycotoxigenic Fungal Pathogens
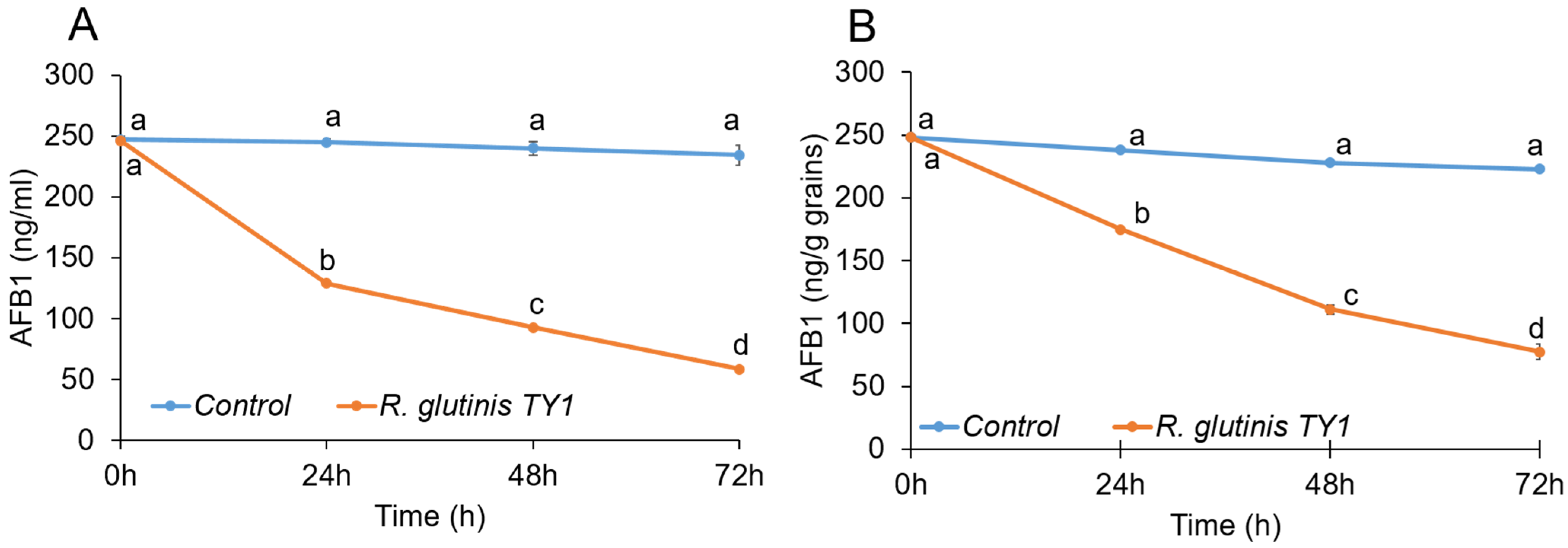
3.3. Mycotoxin-Degrading Activity of the Yeast Rhodotorula glutinis Strain TY1
4. Conclusions
Supplementary Materials
Author Contributions
Funding
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Links, M.G.; Demeke, T.; Gräfenhan, T.; Hill, J.E.; Hemmingsen, S.M.; Dumonceaux, T.J. Simultaneous profiling of seed-associated bacteria and fungi reveals antagonistic interactions between microorganisms within a shared epiphytic microbiome on Triticum and Brassica seeds. New Phytol. 2014, 202, 542–553. [Google Scholar] [CrossRef] [Green Version]
- Placinta, C.M.; D’Mello, C.P.F.; MacDonald, A.M.C. A review of worldwide contamination of cereal grains and animal feed with Fusarium mycotoxins. Anim. Feed Sci. Technol. 1999, 78, 21–37. [Google Scholar] [CrossRef]
- Magan, N.; Aldred, D.; Mylona, K.; Lambert, R.J.W. Limiting mycotoxins in stored wheat. Food Addit. Contam. Part. A 2010, 27, 644–650. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Magan, N.; Hope, R.; Cairns, V.; Aldred, D. Post-harvest fungal ecology: Impact of fungal growth and mycotoxin accumulation in stored grain. Eur. J. Plant. Pathol. 2003, 109, 723–730. [Google Scholar] [CrossRef]
- Ferrigo, D.; Raiola, A.; Causin, R. Fusarium toxins in cereals: Occurrence, legislation, factors promoting the appearance and their management. Molecules 2016, 21, 627. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- McCormick, S.P.; Stanley, A.M.; Stover, N.A.; Alexander, N.J. Trichothecenes: From simple to complex mycotoxins. Toxins 2011, 3, 802–814. [Google Scholar] [CrossRef] [PubMed]
- Lee, H.J.; Ryu, D. Worldwide occurrence of mycotoxins in cereals and cereal-derived food products: Public health perspectives of their co-occurrence. J. Agric. Food Chem. 2017, 65, 7034–7051. [Google Scholar] [CrossRef] [PubMed]
- Streit, E.; Naehrer, K.; Rodrigues, I.; Schatzmayr, G. Mycotoxin occurrence in feed and feed raw materials worldwide: Long-term analysis with special focus on Europe and Asia. J. Sci. Food Agric. 2013, 93, 2892–2899. [Google Scholar] [CrossRef]
- Chen, Y.; Kistler, H.C.; Ma, Z. Fusarium graminearum trichothecene mycotoxins: Biosynthesis, regulation, and management. Annu. Rev. Phytopathol. 2019, 57, 15–39. [Google Scholar] [CrossRef] [Green Version]
- Gdanetz, K.; Trail, F. The wheat microbiome under four management strategies, and potential for endophytes in disease protection. Phytobiomes 2017, 1, 158–168. [Google Scholar] [CrossRef] [Green Version]
- Mousa, W.K.; Shearer, C.; Limay-Rios, V.; Ettinger, C.L.; Eisen, J.A.; Raizada, M.N. Root-hair endophyte stacking in finger millet creates a physicochemical barrier to trap the fungal pathogen Fusarium graminearum. Nat. Microbiol. 2016, 1, 16167. [Google Scholar] [CrossRef] [Green Version]
- Comby, M.; Gacoin, M.; Robineau, M.; Rabenoelina, F.; Ptas, S.; Dupont, J.; Profizi, C.; Baillieul, F. Screening of wheat endophytes as biological control agents against Fusarium head blight using two different in vitro tests. Microbiol. Res. 2017, 202, 11–20. [Google Scholar] [CrossRef]
- Díaz Herrera, S.; Grossi, C.; Zawoznik, M.; Groppa, M.D. Wheat seeds harbour bacterial endophytes with potential as plant growth promoters and biocontrol agents of Fusarium graminearum. Microbiol. Res. 2016, 186, 37–43. [Google Scholar] [CrossRef]
- Baffoni, L.; Gaggia, F.; Dalanaj, N.; Prodi, A.; Nipoti, P.; Pisi, A.; Biavati, B.; Di Gioia, D. Microbial inoculants for the biocontrol of Fusarium spp. in durum wheat. Bmc Microbiol. 2015, 15, 1–10. [Google Scholar] [CrossRef]
- Leah Musyimi, S.; Wanjohi Muthomi, J.; Devi Narla, R.; Maina Wagacha, J. Efficacy of Biological Control and Cultivar Resistance on Fusarium Head Blight and T-2 Toxin Contamination in Wheat. Am. J. Plant Sci. 2012, 3, 599–607. [Google Scholar] [CrossRef] [Green Version]
- Sadhasivam, S.; Britzi, M.; Zakin, V.; Kostyukovsky, M.; Trostanetsky, A.; Quinn, E.; Sionov, E. Rapid detection and identification of mycotoxigenic fungi and mycotoxins in stored wheat grain. Toxins 2017, 9, 302. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Hanshew, A.S.; Mason, C.J.; Raffa, K.F.; Currie, C.R. Minimization of chloroplast contamination in 16S rRNA gene pyrosequencing of insect herbivore bacterial communities. J. Microbiol. Methods 2013, 95, 149–155. [Google Scholar] [CrossRef] [Green Version]
- Scibetta, S.; Schena, L.; Abdelfattah, A.; Pangallo, S.; Cacciola, S.O. Selection and experimental evaluation of universal primers to study the fungal microbiome of higher plants. Phytobiomes J. 2018, 2, 225–236. [Google Scholar] [CrossRef] [Green Version]
- Bolger, A.M.; Lohse, M.; Usadel, B. Trimmomatic: A flexible trimmer for Illumina sequence data. Bioinformatics 2014, 30, 2114–2120. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Zhang, J.; Kobert, K.; Flouri, T.; Stamatakis, A. PEAR: A fast and accurate Illumina Paired-End reAd mergeR. Bioinformatics 2014, 30, 614–620. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Masella, A.P.; Bartram, A.K.; Truszkowski, J.M.; Brown, D.G.; Neufeld, J.D. PANDAseq: Paired-end assembler for Illumina sequences. Bmc Bioinform. 2012, 13, 31. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Edgar, R.C. Search and clustering orders of magnitude faster than BLAST. Bioinformatics 2010, 26, 2460–2461. [Google Scholar] [CrossRef] [Green Version]
- McDonald, D.; Price, M.N.; Goodrich, J.; Nawrocki, E.P.; DeSantis, T.Z.; Probst, A.; Andersen, G.L.; Knight, R.; Hugenholtz, P. An improved Greengenes taxonomy with explicit ranks for ecological and evolutionary analyses of bacteria and archaea. ISME J. 2012, 6, 610–618. [Google Scholar] [CrossRef] [PubMed]
- Rognes, T.; Flouri, T.; Nichols, B.; Quince, C.; Mahé, F. VSEARCH: A versatile open source tool for metagenomics. PeerJ 2016, 4, e2584. [Google Scholar] [CrossRef]
- Caporaso, J.G.; Kuczynski, J.; Stombaugh, J.; Bittinger, K.; Bushman, F.D.; Costello, E.K.; Fierer, N.; Peña, A.G.; Goodrich, J.K.; Gordon, J.I.; et al. QIIME allows analysis of high-throughput community sequencing data. Nat. Methods 2010, 7, 335–336. [Google Scholar] [CrossRef] [Green Version]
- DeSantis, T.Z.; Hugenholtz, P.; Larsen, N.; Rojas, M.; Brodie, E.L.; Keller, K.; Huber, T.; Dalevi, D.; Hu, P.; Andersen, G.L. Greengenes, a chimera-checked 16S rRNA gene database and workbench compatible with ARB. Appl. Env Microbiol. 2006, 72, 5069–5072. [Google Scholar] [CrossRef] [Green Version]
- Abarenkov, K.; Henrik Nilsson, R.; Larsson, K.-H.; Alexander, I.J.; Eberhardt, U.; Erland, S.; Høiland, K.; Kjøller, R.; Larsson, E.; Pennanen, T.; et al. The UNITE database for molecular identification of fungi-recent updates and future perspectives. New Phytol. 2010, 186, 281–285. [Google Scholar] [CrossRef]
- Altschul, S.F.; Gish, W.; Miller, W.; Myers, E.W.; Lipman, D.J. Basic local alignment search tool. J. Mol. Biol. 1990, 215, 403–410. [Google Scholar] [CrossRef]
- Paulson, J.N.; Stine, O.C.; Bravo, H.C.; Pop, M. Differential abundance analysis for microbial marker-gene surveys. Nat. Methods 2013, 10, 1200–1202. [Google Scholar] [CrossRef] [Green Version]
- Bray, J.R.; Curtis, J.T. An ordination of the upland forest communities of southern Wisconsin. Ecol. Monogr. 1957, 27, 325–349. [Google Scholar] [CrossRef]
- Lozupone, C.; Knight, R. UniFrac: A new phylogenetic method for comparing microbial communities. Appl. Env Microbiol. 2005, 71, 8228–8235. [Google Scholar] [CrossRef] [Green Version]
- Kruskal, W.H.; Wallis, W.A. Use of ranks in one-criterion variance analysis. J. Am. Stat. Assoc. 1952, 47, 583–621. [Google Scholar] [CrossRef]
- De, S.; Kaur, G.; Roy, A.; Dogra, G.; Kaushik, R.; Yadav, P.; Singh, R.; Datta, T.K.; Goswami, S.L. A simple method for the efficient isolation of genomic DNA from Lactobacilli isolated from traditional indian fermented milk (dahi). Indian J. Microbiol. 2010, 50, 412–418. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Castoria, R.; Morena, V.; Caputo, L.; Panfili, G.; De Curtis, F.; De Cicco, V. Effect of the biocontrol yeast Rhodotorula glutinis strain LS11 on patulin accumulation in stored apples. Phytopathology 2005, 95, 1271–1278. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Johnston-Monje, D.; Raizada, M.N. Conservation and diversity of seed associated endophytes in zea across boundaries of evolution, ethnography and ecology. PLoS ONE 2011, 6, e20396. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Solanki, M.K.; Abdelfattah, A.; Britzi, M.; Zakin, V.; Wisniewski, M.; Droby, S.; Sionov, E. Shifts in the composition of the microbiota of stored wheat grains in response to fumigation. Front. Microbiol. 2019, 10, 1098. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Truyens, S.; Weyens, N.; Cuypers, A.; Vangronsveld, J. Bacterial seed endophytes: Genera, vertical transmission and interaction with plants. Env Microbiol. Rep. 2015, 7, 40–50. [Google Scholar] [CrossRef]
- Zawoznik, M.S.; Vázquez, S.C.; Herrera, S.M.D.; Groppa, M.D. Search for endophytic diazotrophs in barley seeds. Braz. J. Microbiol. 2014, 45, 621–625. [Google Scholar] [CrossRef]
- Karlsson, I.; Friberg, H.; Kolseth, A.-K.; Steinberg, C.; Persson, P. Organic farming increases richness of fungal taxa in the wheat phyllosphere. Mol. Ecol. 2017, 26, 3424–3436. [Google Scholar] [CrossRef]
- Khan, N.; Martínez-Hidalgo, P.; Ice, T.A.; Maymon, M.; Humm, E.A.; Nejat, N.; Sanders, E.R.; Kaplan, D.; Hirsch, A.M. Antifungal activity of Bacillus species against Fusarium and analysis of the potential mechanisms used in biocontrol. Front. Microbiol. 2018, 9, 2363. [Google Scholar] [CrossRef] [Green Version]
- Zhang, D.; Yu, S.; Yang, Y.; Zhang, J.; Zhao, D.; Pan, Y.; Fan, S.; Yang, Z.; Zhu, J. Antifungal effects of volatiles produced by Bacillus subtilis against Alternaria solani in Potato. Front. Microbiol. 2020, 11, 1196. [Google Scholar] [CrossRef]
- Mannaa, M.; Kim, K.D. Biocontrol Activity of volatile-producing Bacillus megaterium and Pseudomonas protegens against Aspergillus and Penicillium spp. predominant in stored rice grains: Study II. Mycobiology 2018, 46, 52–63. [Google Scholar] [CrossRef] [Green Version]
- Palazzini, J.M.; Alberione, E.; Torres, A.; Donat, C.; Köhl, J.; Chulze, S. Biological control of Fusarium graminearum sensu stricto, causal agent of Fusarium head blight of wheat, using formulated antagonists under field conditions in Argentina. Biol. Control 2016, 94, 56–61. [Google Scholar] [CrossRef]
- Palazzini, J.M.; Ramirez, M.L.; Alberione, E.J.; Torres, A.M.; Chulze, S.N. Osmotic stress adaptation, compatible solutes accumulation and biocontrol efficacy of two potential biocontrol agents on Fusarium head blight in wheat. Biol. Control 2009, 51, 370–376. [Google Scholar] [CrossRef]
- Zalila-Kolsi, I.; Ben Mahmoud, A.; Ali, H.; Sellami, S.; Nasfi, Z.; Tounsi, S.; Jamoussi, K. Antagonist effects of Bacillus spp. strains against Fusarium graminearum for protection of durum wheat (Triticum turgidum L. subsp. durum). Microbiol. Res. 2016, 192, 148–158. [Google Scholar] [CrossRef] [PubMed]
- Crane, J.M.; Bergstrom, G.C. Spatial distribution and antifungal interactions of a Bacillus biological control agent on wheat surfaces. Biol. Control 2014, 78, 23–32. [Google Scholar] [CrossRef]
- Shi, C.; Yan, P.; Li, J.; Wu, H.; Li, Q.; Guan, S. Biocontrol of Fusarium graminearum growth and deoxynivalenol production in wheat kernels with bacterial antagonists. Int. J. Env Res. Public Health 2014, 11, 1094–1105. [Google Scholar] [CrossRef] [Green Version]
- Khan, N.I.; Schisler, D.A.; Boehm, M.J.; Slininger, P.J.; Bothast, R.J. Selection and evaluation of microorganisms for biocontrol of Fusarium head blight of wheat incited by Gibberella zeae. Plant Dis. 2001, 85, 1253–1258. [Google Scholar] [CrossRef] [Green Version]
- Zhao, Y.; Selvaraj, J.N.; Xing, F.; Zhou, L.; Wang, Y.; Song, H.; Tan, X.; Sun, L.; Sangare, L.; Folly, Y.M.E.; et al. Antagonistic action of Bacillus subtilis strain SG6 on Fusarium graminearum. PLoS ONE 2014, 9, 92486. [Google Scholar] [CrossRef]
- Siddiqui, Z.A. Biocontrol of Alternaria triticina by plant growth promoting rhizobacteria on wheat. Arch. Phytopathol. Plant Prot. 2007, 40, 301–308. [Google Scholar] [CrossRef]
- Liu, Q.; Li, W.; Feng, Y.; Tao, C. Efficacy and safety of polymyxins for the treatment of Acinectobacter baumannii infection: A systematic review and meta-analysis. PLoS ONE 2014, 9, e98091. [Google Scholar] [CrossRef] [PubMed]
- Ongena, M.; Jacques, P. Bacillus lipopeptides: Versatile weapons for plant disease biocontrol. Trends Microbiol. 2008, 16, 115–125. [Google Scholar] [CrossRef] [PubMed]
- Cochrane, S.A.; Vederas, J.C. Lipopeptides from Bacillus and Paenibacillus spp.: A Gold Mine of Antibiotic Candidates. Med. Res. Rev. 2016, 36, 4–31. [Google Scholar] [CrossRef] [PubMed]
- Caulier, S.; Nannan, C.; Gillis, A.; Licciardi, F.; Bragard, C.; Mahillon, J. Overview of the antimicrobial compounds produced by members of the Bacillus subtilis group. Front. Microbiol. 2019, 10, 302. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Kusari, S.; Hertweck, C.; Spiteller, M. Chemical ecology of endophytic fungi: Origins of secondary metabolites. Chem. Biol. 2012, 19, 792–798. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Peles, F.; Sipos, P.; Kovács, S.; Gyori, Z.; Pócsi, I.; Pusztahelyi, T. Biological control and mitigation of aflatoxin contamination in commodities. Toxins 2021, 13, 104. [Google Scholar] [CrossRef] [PubMed]
- Ianiri, G.; Idnurm, A.; Wright, S.A.I.; Durán-Patrón, R.; Mannina, L.; Ferracane, R.; Ritieni, A.; Castoria, R. Searching for genes responsible for patulin degradation in a biocontrol yeast provides insight into the basis for resistance to this mycotoxin. Appl. Env Microbiol. 2013, 79, 3101–3115. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Ianiri, G.; Pinedo, C.; Fratianni, A.; Panfili, G.; Castoria, R. Patulin degradation by the biocontrol yeast Sporobolomyces sp. Is an inducible process. Toxins 2017, 9, 61. [Google Scholar] [CrossRef]
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Solanki, M.K.; Abdelfattah, A.; Sadhasivam, S.; Zakin, V.; Wisniewski, M.; Droby, S.; Sionov, E. Analysis of Stored Wheat Grain-Associated Microbiota Reveals Biocontrol Activity among Microorganisms against Mycotoxigenic Fungi. J. Fungi 2021, 7, 781. https://doi.org/10.3390/jof7090781
Solanki MK, Abdelfattah A, Sadhasivam S, Zakin V, Wisniewski M, Droby S, Sionov E. Analysis of Stored Wheat Grain-Associated Microbiota Reveals Biocontrol Activity among Microorganisms against Mycotoxigenic Fungi. Journal of Fungi. 2021; 7(9):781. https://doi.org/10.3390/jof7090781
Chicago/Turabian StyleSolanki, Manoj Kumar, Ahmed Abdelfattah, Sudharsan Sadhasivam, Varda Zakin, Michael Wisniewski, Samir Droby, and Edward Sionov. 2021. "Analysis of Stored Wheat Grain-Associated Microbiota Reveals Biocontrol Activity among Microorganisms against Mycotoxigenic Fungi" Journal of Fungi 7, no. 9: 781. https://doi.org/10.3390/jof7090781
APA StyleSolanki, M. K., Abdelfattah, A., Sadhasivam, S., Zakin, V., Wisniewski, M., Droby, S., & Sionov, E. (2021). Analysis of Stored Wheat Grain-Associated Microbiota Reveals Biocontrol Activity among Microorganisms against Mycotoxigenic Fungi. Journal of Fungi, 7(9), 781. https://doi.org/10.3390/jof7090781







