Nosema Disease of European Honey Bees
Abstract
1. Introduction
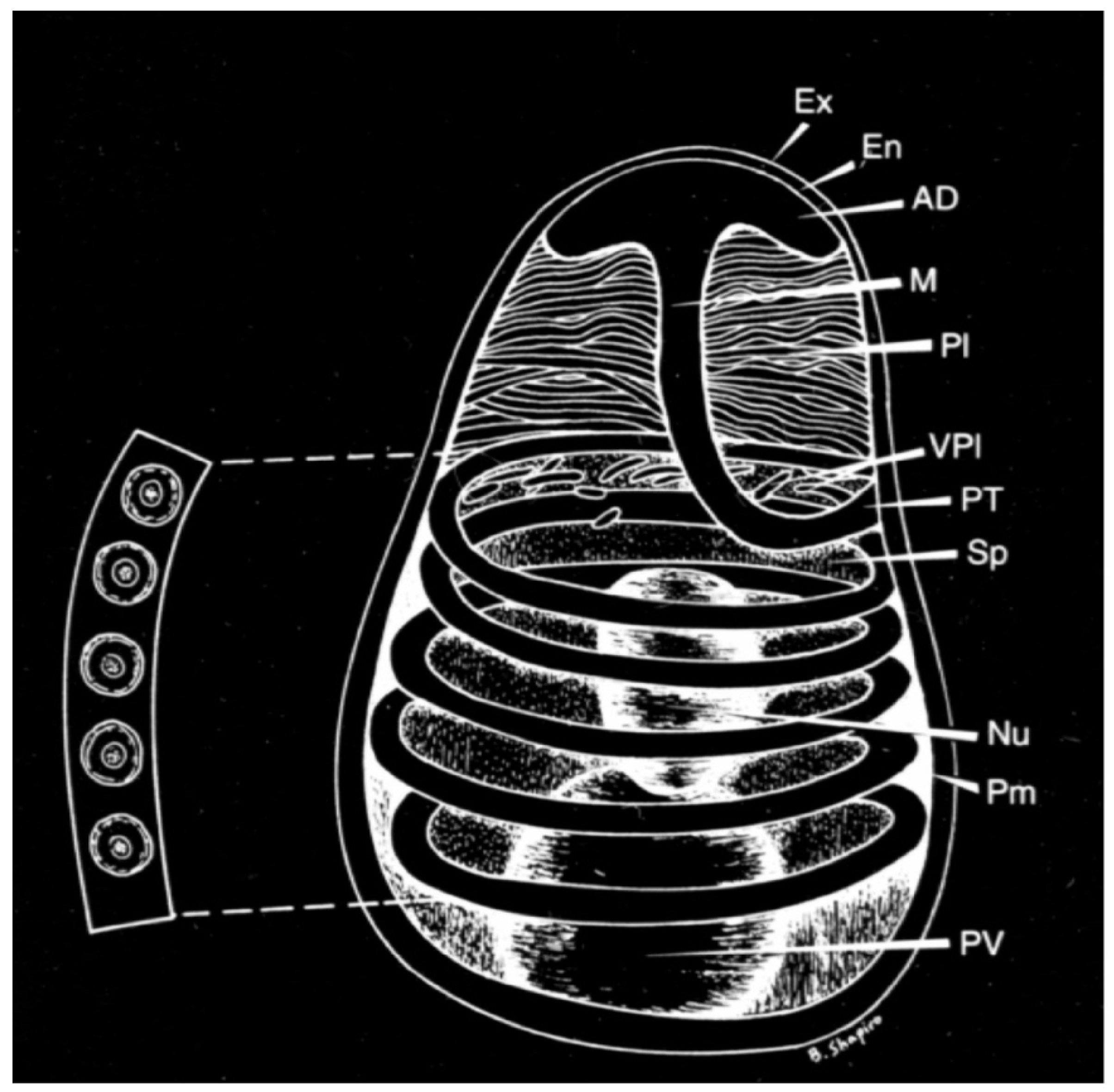
2. Biology and Pathogenesis of Nosema spp.
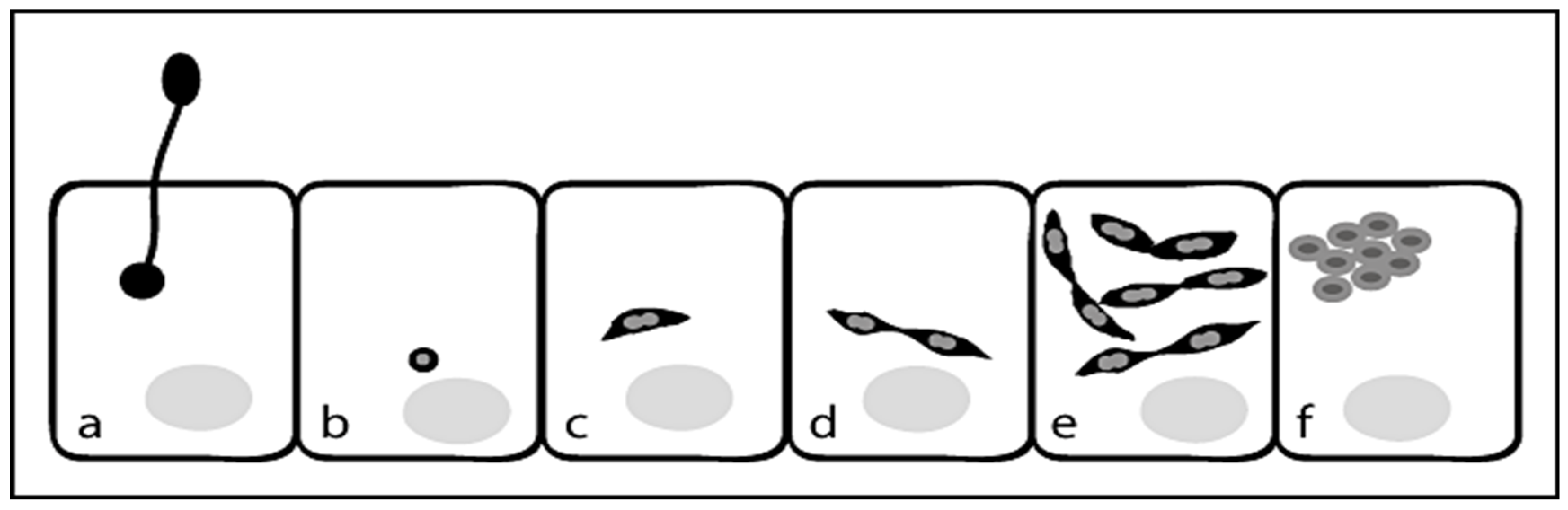
Life Cycle for N. ceranae
3. Transmission
4. Pathogeny in N. apis and N. ceranae
5. Prevalence of N. apis and N. ceranae
6. Laboratory Diagnosis of Nosematosis
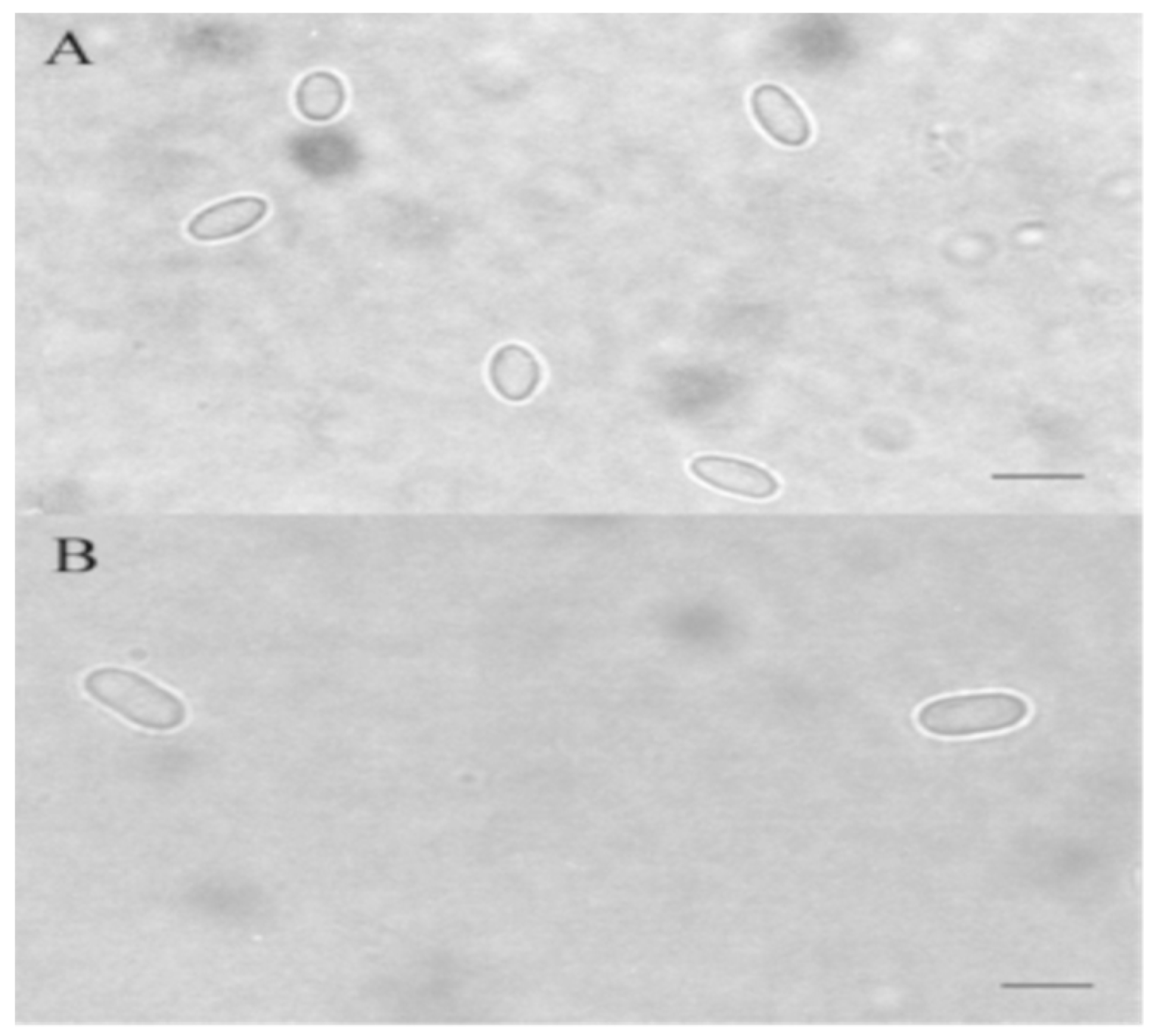
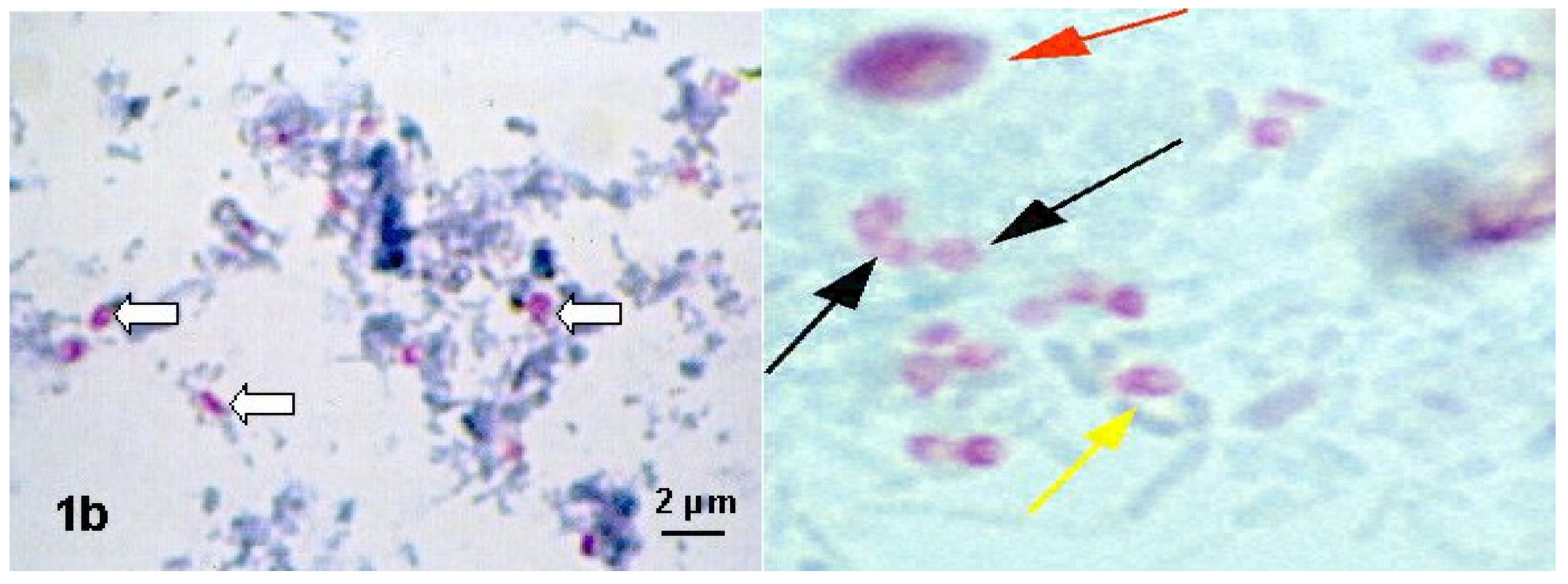
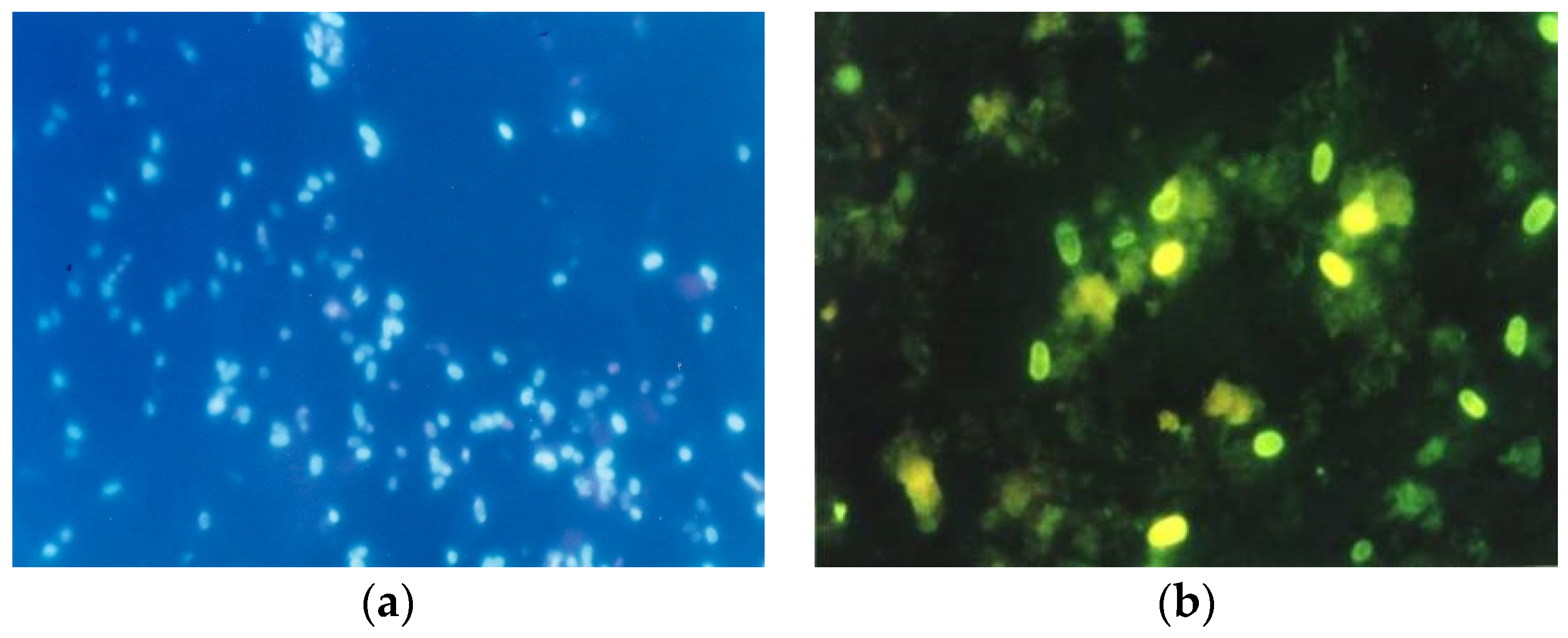
6.1. Microscopic Methods in the Diagnosis of Nosema spp.
6.2. Molecular Detection of Nosema spp. (N. apis, N. ceranae and N. bombi)
7. Diagnostic Procedures Used at Our Workplace
8. Cultivation on Cell Structures
9. Prevention and Treatment of Nosematosis
10. Conclusions
Funding
Conflicts of Interest
References
- Brittain, C.; Williams, N.; Kremen, C.; Klein, A.M. Synergistic effects of non-Apis bees and honey bees for pollination services. Proc. R. Soc. B Biol. Sci. 2013, 280, 1754. [Google Scholar] [CrossRef] [PubMed]
- Goulson, D.; Nicholls, E.; Botías, C.; Rotheray, E.L. Bee declines driven by combined stress from parasites, pesticides, and lack of flowers. Science 2015, 347, 1255957. [Google Scholar] [CrossRef]
- Tesovnik, T.; Zorc, M.; Ristanić, M.; Glavinić, U.; Stevanović, J.; Narat, M.; Stanimirović, Z. Exposure of honey bee larvae to thiamethoxam and its interaction with Nosema ceranae infection in adult honey bees. Environ. Pollut. 2020, 256, 113443. [Google Scholar] [CrossRef] [PubMed]
- Broadrup, R.L.; Mayack, C.H.; Schick, S.J.; Eppley, E.J.; White, H.K.; Macherone, A. Honey bee (Apis mellifera) exposomes and dysregulated metabolic pathways associated with Nosema ceranae infection. PLoS ONE 2019, 14, e0213249. [Google Scholar] [CrossRef]
- Wittner, M.; Weiss, L.M. The Microsporidia and Microsporidiosis; ASM Press: Washington, DC, USA, 1999; ISBN 1-55581-147-7. [Google Scholar]
- Fries, I. Nosema apis—A parasite in the honey bee colony. Bee World 1993, 74, 5–19. [Google Scholar] [CrossRef]
- Fries, I.; Feng, F.; da Silva, A.; Slemenda, S.B.; Pieniazek, N.J. Nosema ceranae n. sp. (Microspora, Nosematidae), morphological and molecular characterization of a microsporidian parasite of the Asian honey bee Apis cerana (Hymenoptera, Apidae). Eur. J. Protistol. 1996, 32, 356–365. [Google Scholar] [CrossRef]
- Chemurot, M.; De Smet, L.; Brunain, M.; De Rycke, R.; de Graaf, D.C. Nosema neumanni n. sp. (Microsporidia, Nosematidae), a new microsporidian parasite of honeybees, Apis mellifera in Uganda. Eur. J. Protistol. 2017, 61, 13–19. [Google Scholar] [CrossRef]
- Fries, I.; Martín-Hernández, R.; Meana, A.; GarcíaPalencia, P.; Higes, M. Natural infections of Nosema ceranae in European honey bees. J. Apicultur. Res. 2006, 45, 230–233. [Google Scholar] [CrossRef]
- Higes, M.; Martín, R.; Meana, A. Nosema ceranae a New Microsporidian Parasite in Honeybees in Europe. J. Invertebr. Pathol. 2006, 92, 93–95. [Google Scholar] [CrossRef]
- Klee, J.; Besana, A.M.; Genersch, E.; Gisder, S.; Nanetti, A.; Tam, D.Q.; Chinh, T.X.; Puerta, F.; Ruz, J.; Kryger, P.; et al. Widespread dispersal of the microsporidian Nosema ceranae, an emergent pathogen of the western honey bee, Apis mellifera. J. Invertebr. Pathol. 2007, 96, 1–10. [Google Scholar] [CrossRef]
- Martin-Hernandez, R.; Meana, A.; Prieto, L.; Salvador, A.M.; Garrido-Bailon, E.; Higes, M. Outcome of colonization of apis mellifera by Nosema ceranae. Appl. Environ. Microbiol. 2007, 73, 6331–6338. [Google Scholar] [CrossRef] [PubMed]
- Botías, C.; Anderson, D.L.; Meana, A.; Garrido-Bailón, E.; Martín-Hernández, R.; Higes, M. Further evidence of an oriental origin for Nosema ceranae (Microsporidia: Nosematidae). J. Invertebr. Pathol. 2012, 110, 108–113. [Google Scholar] [CrossRef]
- Read, A.F.; Taylor, L.H. The ecology of genetically diverse infections. Science 2001, 292, 1099–1102. [Google Scholar] [CrossRef]
- Paxton, R.J.; Klee, J.; Korpela, S.; Fries, I. Nosema ceranae has infected apis mellifera in europe since at least 1998 and may be more virulent than Nosema apis. Apidologie 2007, 38, 558–565. [Google Scholar] [CrossRef]
- Tapaszti, Z.; Forgách, P.; Kövágó, C.; Békési, L.; Bakonyi, T.; Rusvai, M. First detection and dominance of Nosema ceranae in Hungarian honeybee colonies. Acta Vet. Hung. 2009, 57, 383–388. [Google Scholar] [CrossRef]
- Chen, Y.P.; Evans, J.D.; Smith, J.B.; Pettis, J.S. Nosema ceranae is a long-present and wide-spread microsporidian infection of the european honey bee (Apis mellifera) in the united states. J. Invertebr. Pathol. 2008, 97, 186–188. [Google Scholar] [CrossRef] [PubMed]
- Millbrath, M.O.; van Tran, T.; Huang, W.F.; Solter, L.F.; Tarpy, D.R.; Lawrence, F.; Huang, Z.Y. Comparative virulence and competition between Nosema apis and Nosema ceranae in honey bees (Apis mellifera). J. Invertebr. Pathol. 2015, 125, 9–15. [Google Scholar] [CrossRef] [PubMed]
- Cox-Foster, D.L.; Conlan, S.; Holmes, E.; Palacios, G.; Evans, J.D.; Moran, N.A.; Quan, P.L.; Briese, T.; Hornig, M.; Geiser, D.M.; et al. A metagenomic survey of microbes in honey bee colony collapse disorder. Science 2007, 318, 283–287. [Google Scholar] [CrossRef]
- Zander, E.; Bottcher, F.K. Krankheiten der Biene, 7th ed.; Eugen Ulmer: Stuttgart, Germany, 1984. [Google Scholar]
- Bigliardi, E.; Selmi, M.G.; Lupetti, P.; Corona, S.; Gatti, S.; Scaglia, M.; Sacchi, I. Microsporidian spore wall: Ultrastructural findings on Encephalitozoon hellem exospore. J. Eukaryot. Microbiol. 1996, 43, 181–186. [Google Scholar] [CrossRef] [PubMed]
- Keohane, E.M.; Weiss, L.M. The structure, function, and composition of the microsporidian polar tube. In The Microsporidia and Microsporidiosis; Wittner, M., Weiss, L.M., Eds.; ASM Press: Washington, DC, USA, 1999; pp. 196–224. [Google Scholar] [CrossRef]
- Keeling, P.J.; Fast, N. Microsporidia: Biology and evolution of highly reduced intracellular parasites. Annu. Rev. Microbiol. 2002, 56, 93–116. [Google Scholar] [CrossRef]
- Frixione, E.; Ruiz, L.; Santillan, M.; de Vargas, L.V.; Tejero, J.M.; Undeen, A.H. Dynamics of polar filament discharge and sporoplasm expulsion by microsporidian spores. Cell Motil. Cytoskelet. 1992, 22, 38–50. [Google Scholar] [CrossRef]
- Gisder, S.; Möckel, N.; Linde, A.; Genersch, E. A cell culture model for Nosema ceranae and Nosema apis allows new insights into the life cycle of theseimportant honey bee-pathogenic microsporidia. Environ. Microbiol. 2011, 13, 404–413. [Google Scholar] [CrossRef]
- Webster, T.C. Nosema apis spore transmission among honey bees. Am. Bee J. 1993, 133, 869–870. [Google Scholar]
- Vidau, C.; Panek, J.; Texier, C.; Biron, D.G.; Belzunces, L.P.; Le Gall, M.; Broussard, C.; Delbac, F.; El Alaoui, H. Differential proteomic analysis of midguts from Nosema ceranae-infected honeybees reveals manipulation of key host functions. J. Invertebr. Pathol. 2014, 121, 89–96. [Google Scholar] [CrossRef]
- Fernandez, N.; Coineau, Y. Varroa the Serial Bee Killer Mite; Atlantica: Biarritz, France, 2007. [Google Scholar]
- Schmid-Hempel, P. Parasites in Social Insects; Princeton University Press: Princeton, NJ, USA, 1998; pp. 119–123. ISBN 9780691059242. [Google Scholar]
- Veselý, V.; Kubišová, S.; Haragsim, O.; Kamler, K.; Krieg, P.; Škrobal, D.; Ptáček, V.; Titěra, D.; Peroutková, M.; Drobníková, V.; et al. Beekeeping; Brázda: Praha, Czech Republic, 2003; p. 272. (In Czech) [Google Scholar]
- Fries, I.; Chauzat, M.P.; Chen, Y.P.; Doublet, V.; Genersch, E.; Gisder, S.; Higes, M.; McMahon, D.P.; Martín-Hernández, R.; Natsopoulou, M.; et al. Standard methods for Nosema research. J. Apic. Res. 2013, 52, 1. [Google Scholar] [CrossRef]
- Fries, I.; Ekbohm, G.; Villumstad, E. Nosema apis, sampling techniques andhoney yield. J. Apicult. Res. 1984, 23, 102–105. [Google Scholar] [CrossRef]
- Nosemosis of Honey Bees. In OIE Terrestrial Manual; OIE: Paris, France, 2013; Chapter 2.2.4.
- Grozinger, C.; Robinson, G.E. The power and promise of applying genomics tohoney bee health. Curr. Opin. Insect Sci. 2015, 10, 124–132. [Google Scholar] [CrossRef]
- Botías, C.; Martín-Hernández, R.; Barrios, L.; Meana, A.; Higes, M. Nosema spp. Infection and its negative effects on honey bees (Apis mellifera iberiensis) at the colonylevel. Vet. Res. 2013, 10, 25. Available online: http://www.veterinaryresearch.org/content/44/1/25 (accessed on 10 April 2013). [CrossRef] [PubMed]
- Varis, A.L.; Ball, B.V.; Allen, M. The incidence of pathogens in honey bee (Apismellifera L.) colonies in Finland and Great Britain. Apidologie 1992, 23, 133–137. [Google Scholar] [CrossRef]
- Gisder, S.; Schüler, V.; Horchler, L.L.; Groth, D.; Genersch, E. Long-Term Temporal Trends of Nosema spp. Infection Prevalence in Northeast Germany: Continuous Spread of Nosema ceranae, an Emerging Pathogen of Honey Bees (Apis mellifera), but No General Replacement of Nosema Apis. Front. Cell. Infect. Microbiol. 2017, 7, 301. [Google Scholar] [CrossRef]
- Kamler, F.; Titěra, D.; Kamler, M. Rozšíření, Patogeneze a Návrh Opatření Vchovech Včel Ohrožených Mikrosporidií Nosema Ceranae; Závěrečná Zpráva za Rok; APIC: Arlington, VA, USA, 2011; p. 20. (In Slovak) [Google Scholar]
- Staroň, M.; Jurovčíková, J.; Čermáková, T.; Staroňová, D. Scientific note on incidence of Nosema apis and Nosema ceranae in Slovakia during the years 2009 and 2010. Slovak J. Anim. Sci. 2012, 45, 36–38. [Google Scholar]
- Csáki, T.; Heltai, M.; Markolt, F.; Kovács, B.; Békési, L.; Ladányi, M.; Péntek-Zakar, E.; Meana, A.; Botías, C.; Martín-Henández, R.; et al. Permanent prevalence of Nosema ceranae in honey bees (Apis mellifera) in Hungary. Acta Vet. Hung. 2015, 63, 358–369. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Papini, R.; Mancianti, F.; Canovai, R.; Cosci, F.; Rocchigiani, G.; Benelli, G.; Canale, A. Prevalence of the microsporidian Nosema ceranae in honeybee (Apis mellifera) apiaries in Central Italy Saudi. J. Biol. Sci. 2017, 24, 979–982. [Google Scholar] [CrossRef]
- Shumkova, R.; Georgieva, A.; Radoslavov, G.; Sirakova, D.; Dzhebir, G.; Neov, B.; Bouga, M.; Hristov, P. The first report of the prevalence of Nosema ceranae in Bulgaria. PeerJ 2018, 6, e4252. [Google Scholar] [CrossRef]
- Matthijs, S.; De Waele, V.; Vandenberge, V.; Verhoeven, B.; Evers, J.; Brunain, M.; Saegerman, C.; De Winter, P.J.J.; Roels, S.; de Graaf, D.C.; et al. Nationwide Screening for Bee Viruses and Parasites in Belgian Honey Bees. Viruses 2020, 12, 890. [Google Scholar] [CrossRef]
- Porrini, C.; Mutinelli, F.; Bortolotti, L.; Granato, A. The Status of Honey Bee Health in Italy: Results from the Nationwide Bee Monitoring Network. PLoS ONE 2016, 11, e0155411. [Google Scholar] [CrossRef]
- van Gool, T.; Snijders, F.; Reiss, P.; Eeftinck Schattenkerk, J.K.; van den Bergh Weerman, M.A.; Bartelsman, J.F.; Bruins, J.J.; Canning, E.U.; Dankert, J. Diagnosis of intestinal and disseminated microsporidial infections in patients with HIV by a new rapid fluorescence technique. J. Clin. Pathol. 1993, 46, 694–699. [Google Scholar] [CrossRef]
- Chapter 3.02.04 Nosemosis 2018. In Terrestrial Manual of the OIE; Terrestrial Animal Health Code—Contents (oie.int); OIE: Paris, France, 2018.
- Weber, R.; Bryan, R.T.; Owen, R.L.; Wilcox, C.M.; Gorelkin, L.; Visvesvara, G.S. Improved light-microscopical detection of microsporidia spores in stool and duodenal aspirates. The Enteric Opportunistic Infections Working Group. N. Engl. J. Med. 1992, 326, 161–166. [Google Scholar] [CrossRef]
- Vávra, J.; Dahbiová, R.; Hollister, W.S.; Canning, E.U. Staining of microsporidian spores by optical brighteners with remarks on the use of brighteners for the diagnosis of AIDS associated human microsporidioses. Folia Parasitol. 1993, 40, 267–272. [Google Scholar] [PubMed]
- Chen, Y.P.; Huang, Z.Y. Nosema ceranae, a newly identified pathogen of Apis mellifera in the USA and Asia. Apidologie 2010, 41, 364–374. [Google Scholar] [CrossRef]
- Bokaie, S.; Sharifi, L.; Mehrabadi, M. Prevalence and epizootical aspects of varroasis in Golestan province, northern Iran. J. Arthropod-Borne Dis. 2014, 8, 102–107. [Google Scholar] [PubMed]
- Ryan, N.J.; Sutherland, G.; Coughlan, K.; Globan, M.; Doultree, J.; Marshall, J.; Baird, R.W.; Pedersen, J.; Dwyer, B. A new trichrome-blue stain for detection of microsporidial species in urine, stool, and nasopharyngeal specimens. J. Clin. Microbiol. 1993, 31, 3264–3269. [Google Scholar] [CrossRef]
- Fedorko, D.P.; Hijazi, Y.M. Application of molecular techniques to the diagnosis of microsporidial infection. Emerg. Infect. Dis. 1996, 2, 183–191. [Google Scholar] [CrossRef]
- Burges, H.D.; Canning, E.U.; Hulls, I.K. Ultrastructure of Nosema oryzaephili and the taxonomic value of the polar filament. J. Invertebr. Pathol. 1974, 23, 135–139. [Google Scholar] [CrossRef]
- Evans, J.D.; Schwarz, R.S.; Chen, Y.P.; Budge, G.; Cornman, R.S.; Rua, P.D.; la Miranda, J.R.; de Foret, S.; Foster, L.; Gauthier, L.; et al. Standard methods for molecular research in Apis mellifera. J. Agricult. Res. 2013, 52, 1–54. [Google Scholar] [CrossRef]
- Plischuk, S.; Martín-Hernández, R.; Prieto, L.; Lucía, M.; Botías, C.; Meana, A.; Abrahamovich, A.H.; Lange, C.; Higes, M. South American native bumblebees (Hymenoptera: Apidae) infected by Nosema ceranae (Microsporidia), an emerging pathogen of honeybees (Apis mellifera). Environ. Microbiol. Rep. 2009, 1, 131–135. [Google Scholar] [CrossRef] [PubMed]
- Klee, J.; Tay, W.T.; Paxton, R.J. Specific and sensitive detection of Nosema bombi (Microsporidia: Nosematidae) in bumble bees (Bombus spp.; Hymenoptera: Apidae) by PCR of partial rRNA gene sequences. J. Invertebr. Pathol. 2006, 91, 98–104. [Google Scholar] [CrossRef]
- Erler, S.; Lommatzsch, S.; Lattorff, H.M.G. Comparative analysis of detection limits and specificity of molecular diagnostic markers for three pathogens (Microsporidia, Nosema spp.) in the key pollinators Apis mellifera and Bombus Terrestris. Parasitol. Res. 2012, 110, 1403–1410. [Google Scholar] [CrossRef]
- Webster, T.C.; Pomper, K.W.; Hunt, G.; Thacker, E.M.; Jones, S.C. Nosema apis infection in worker and queen Apis mellifera. Apidologie 2004, 35, 49–54. [Google Scholar] [CrossRef]
- Forsgren, E.; Fries, I. Comparative Virulence of Nosema Ceranae and Nosema Apis in Individual European Honey Bees. Vet. Parasitol. 2010, 170, 212–217. [Google Scholar] [CrossRef] [PubMed]
- Stevanovic, J.; Stanimirovic, Z.; Genersch, E.; Kovacevic, S.R.; Ljubenkovic, J.; Radavic, M.; Aleksic, N. Dominance of Nosema ceranae in honey bees in the Balkan countries in the absence of symptoms of colony collapse disorder. Apidologie 2011, 42, 49. [Google Scholar] [CrossRef]
- Tay, W.T.; O’Mahoney, E.M.; Paxton, R.J. Complete rRNA gene sequences reveal that the microsporidium Nosema bombi infects diverse bumble bee (Bombus spp.) hosts and contains multiple polymorphic sites. J. Eukaryot. Microbiol. 2005, 52, 505–513. [Google Scholar] [CrossRef]
- Chen, Y.; Evans, J.D.; Zhou, L.; Boncristiani, H.; Kimura, K.; Xiao, T.; Litkowski, A.M.; Pettis, J.S. Asymmetrical coexistence of Nosema ceranae and Nosema apis in honey bees. J. Invertebr. Pathol. 2009, 101, 204–209. [Google Scholar] [CrossRef]
- Carreck, N.L.; Ratnieks, F.L.W. Will neonicotinoid moratorium save the bees? Res. Fortn. 2013, 415, 20–22. [Google Scholar] [CrossRef]
- Goodwin, R.H.; Tompkins, G.J.; McCawley, P. Gypsy moth cell lines divergent in viral susceptibility. I. Culture and identification. Vitro 1978, 14, 485–494. [Google Scholar] [CrossRef]
- Gisder, S.; Hedtke, K.; Möckel, N.; Frielitz, M.C.; Linde, A.; Genersch, E. Five-year cohort study of Nosema spp. in Germany: Does climate shape virulence and assertiveness of Nosema ceranae? Appl. Enviro. Microbiol. 2010, 76, 3032–3038. [Google Scholar] [CrossRef] [PubMed]
- Holt, H.L.; Aronstein, K.A.; Grozinger, C.M. Chronic parasitization by Nosema microsporidia causes global expression changes in core nutritional, metabolic andbehavioral pathways in honey bee workers (Apis mellifera). BMC Genom. 2013, 14, 799. [Google Scholar] [CrossRef]
- Schmid-Hempel, R.; Tognazzo, M. Molecular divergence defines two distinctlineages of Crithidia bombi (Trypanosomatidae), parasites of bumblebees. J. Eukaryot. Microbiol. 2010, 57, 337–345. [Google Scholar] [CrossRef] [PubMed]
- Bradbear, N. Bees and Their Role in Forest Livelihoods; Food and Agriculture Organization of the United Nations: Rome, Italy, 2009; p. 204. [Google Scholar]
- Muñoz, I.; Cepero, A.; Pinto, M.A.; Martín-Hernández, R.; Higes, M.; De la Rúa, P. Presence of Nosema ceranae associated with honeybee queen introductions. Infect. Genet. Evol. 2014, 23, 161–168. [Google Scholar] [CrossRef] [PubMed]
- Evans, J.D.; Aronstein, K.; Chen, Y.P.; Hetru, C.; Imler, J.L.; Jiang, H.; Kanost, M.; Thompson, G.J.; Zou, Z.; Hultmark, D. Immune pathways and defence mechanisms in honeybees Apis mellifera. Insect Mol. Biol. 2006, 15, 645–656. [Google Scholar] [CrossRef]
| Nosema spp. | N. apis | N. ceranae | N. apis + N. ceranae (Mixed Infection) | |
|---|---|---|---|---|
| N = 4010 | 2167 | 678 | 1134 | 356 |
| prevalence | 54.0% | 31.3% | 52.3% | 16.4% |
| Fragment | (bp) | |||||||
|---|---|---|---|---|---|---|---|---|
| Title | Source | Sequence | Locus | PCR | N. apis | N. bombi | N. ceranae | |
| 218MITOC | [12] | fwd rev | CGGCGACGATGTGATATGAAAATATTAA CCCGGTCATTCTCAAACAAAAAACCG | SSU rRNA | qPCR | 218–219 | ||
| 321APIS | [12] | fwd rev | GGGGGCATGTCTTTGACGTACTATGTA GGGGGGCGTTTAAAATGTGAAACAACTATG | SSU rRNA | qPCR | 321 | ||
| BOMBICAR | [1] | fwd rev | GGCCCATGCATGTTTTTGAAGATTATTAT CTACACTTTAACGTAGTTATCTGCGG | SSU rRNA | PCR | 101 | ||
| ITS | [55] | fwd rev | GATATAAGTCGTAACATGGTTGCT CATCGTTATGGTATCCTATTGATC | ITS region | PCR | 120 | 120 | 120 |
| N.b.a | [56] | fwd rev | TGCGGCTTAATTTGACTC GGGTAATGACATACAAACAAAC | SSU rRNA/ITS | PCR | 511 | ||
| Nbombi-SSU-J | [57] | fwd rev | CCATGCATGTTTTTGAAGATTATTAT CATATATTTTTAAAATATGAAACAATAA | SSU rRNA | PCR | 323 | ||
| NOS | [10] | fwd rev | TGCCGACGATGTGATATGAG CACAGCATCCATTGAAAACG | SSU rRNA | PCR | 240 | ||
| NosA | [58] | fwd rev | CCGACGATGTGATATGAGATG CACTATTATCATCCTCAGATCATA | SSU rRNA | PCR | 209 | ||
| SSU-res | [11] | fwd rev | GCCTGACGTAGACGCTATTC GTATTACCGCGGCTGCTGG | SSU rRNA | PCR | 402 | 402 | 402 |
| NaFor | [59] | Fwd (a) | CTAGTATATTTGAATATTTGTTTACAATGG b | LSU rRNA | qPCR | 278 | ||
| NcFor | Fwd (c) | TATTGTAGAGAGGTGGGAGATT | 316 | |||||
| UnivRev | Urev | GTCGCTATGATCGCTTGCC | ||||||
| Nosema | [17] | fwd rev | GGCAGTTATGGGAAGTAACA GGTCGTCACATTTCATCTCT | SSU-rRNA | Generic | 208 | 212 | |
| N. ceranae | [17] | fwd rev | CGGATAAAAGAGTCCGTTACC TGAGCAGGGTTCTAGGGAT | SSU-rRNA | PCR | 250 | ||
| N. apis | [17] | fwd rev | CCATTGCCGGATAAGAGAGT CACGCATTGCTGCATCATTGAC | SSU-rRNA | PCR | 401 | ||
| Nos-16S | [60] | fwd rev | CGTAGACGCTATTCCCTAAGATT CTCCCAACTATACAGTACACCTCATA | SSU-rRNA | PCR | 488 | 488 | |
| Mnceranae-F | [31] | fwd | CGTTAAAGTGTAGATAAGATGTT | SSU-rRNA | PCR | |||
| Mnapis-F | fwd | GCATGTCTTTGACGTACTATG | 143 | |||||
| Mnbombi-F | fwd | TTTATTTTATGTRYACMGCAG | 171 | |||||
| Muniv-R | Urev | GACTTAGTAGCCGTCTCTC | 224 | |||||
| SSUrRNA-f1b | [61] | Ufwd | CACCAGGTTGATTCTGCCT | SSU-rRNA | Generic | Ca. | ||
| SSUrRNA-r1b | Urev | TGTTCGTCCAGTCAGGGTCGTCA | ||||||
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Galajda, R.; Valenčáková, A.; Sučik, M.; Kandráčová, P. Nosema Disease of European Honey Bees. J. Fungi 2021, 7, 714. https://doi.org/10.3390/jof7090714
Galajda R, Valenčáková A, Sučik M, Kandráčová P. Nosema Disease of European Honey Bees. Journal of Fungi. 2021; 7(9):714. https://doi.org/10.3390/jof7090714
Chicago/Turabian StyleGalajda, Richard, Alexandra Valenčáková, Monika Sučik, and Petra Kandráčová. 2021. "Nosema Disease of European Honey Bees" Journal of Fungi 7, no. 9: 714. https://doi.org/10.3390/jof7090714
APA StyleGalajda, R., Valenčáková, A., Sučik, M., & Kandráčová, P. (2021). Nosema Disease of European Honey Bees. Journal of Fungi, 7(9), 714. https://doi.org/10.3390/jof7090714






