Mass Spectrometry-Based Proteomic and Immunoproteomic Analyses of the Candida albicans Hyphal Secretome Reveal Diagnostic Biomarker Candidates for Invasive Candidiasis
Abstract
1. Introduction
2. Materials and Methods
2.1. Study Population and Serum Samples
2.2. Isolation of C. albicans Hyphal Secreted Proteins
2.3. Protein Separation by Sodium Dodecyl Sulfate-Polyacrylamide gel Electrophoresis (SDS-PAGE)
2.4. Protein Identification by Liquid Chromatography–Tandem Mass Spectrometry (LC-MS/MS) Analysis
2.5. Indirect Enzyme-Linked Immunosorbent Assay (ELISA)
2.6. Immunoproteomic Analysis or SERPA
2.7. Bioinformatic Analysis
3. Results
3.1. Isolation of the C. albicans Hyphal Secretome
3.2. Gel-Free LC-MS/MS Analysis of the C. albicans Hyphal Secretome
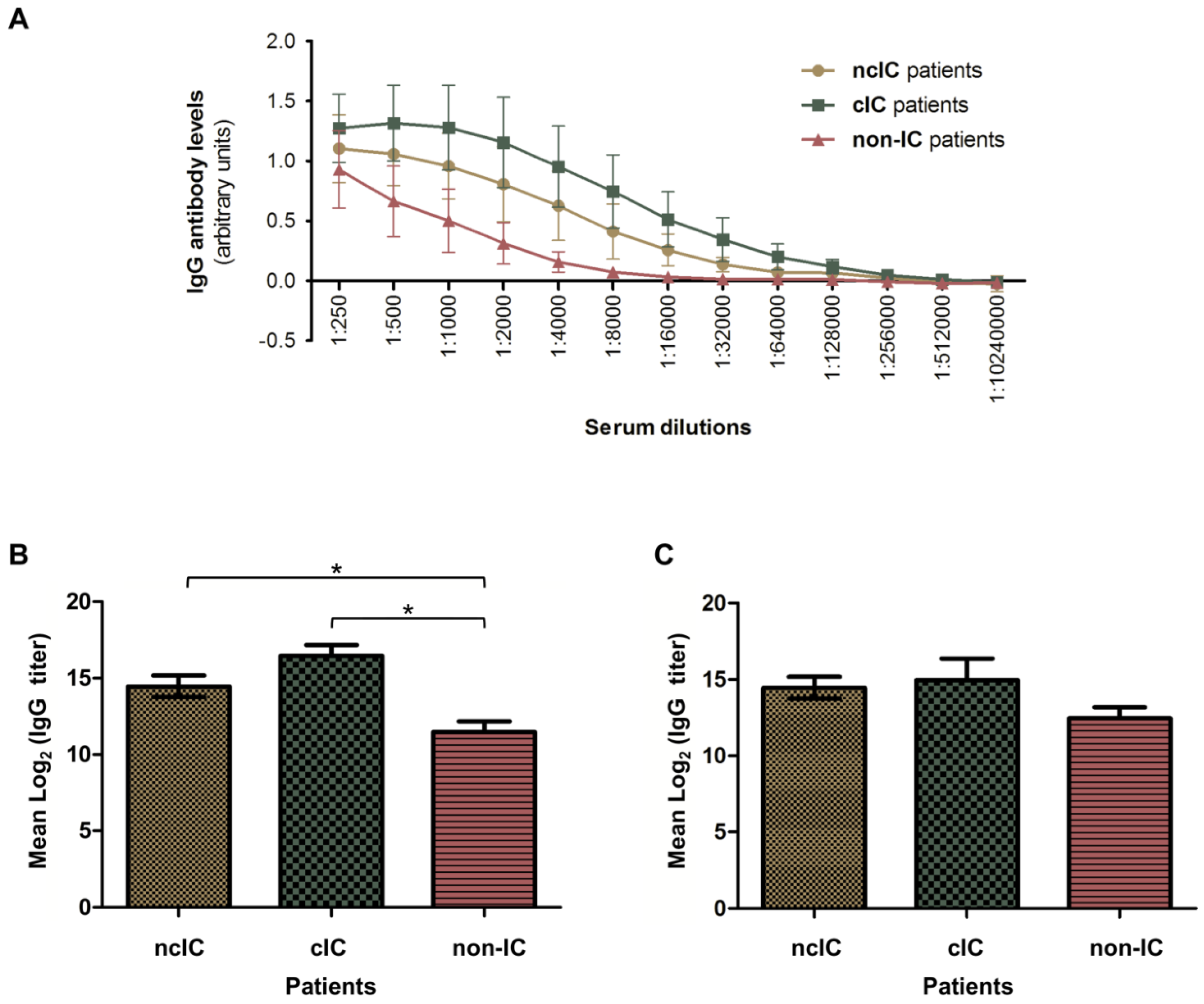
3.3. Serologic Responses to the C. albicans Hyphal Secretome in ncIC, cIC and Non-IC Patients
4. Discussion
4.1. The C. albicans Hyphal Secretome Comprises Proteins Involved in Interaction with Host and Immunogenic Proteins
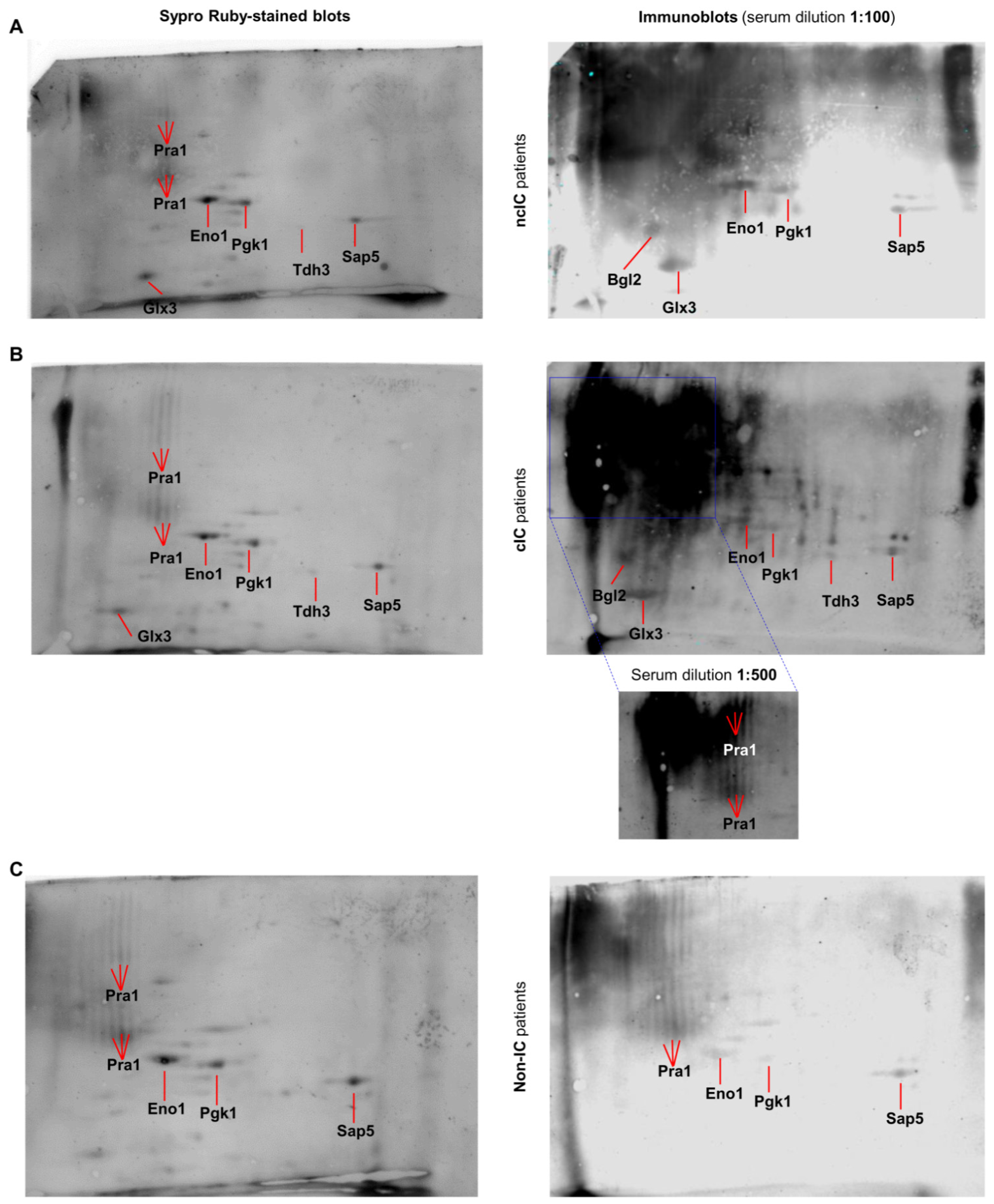
4.2. The C. albicans Hyphal Immunosecretome Recognized by IgG Antibodies Differs between IC and Non-IC Patients
4.3. IgG Antibodies to Bgl2, Eno1, Pgk1, Glx3, Pra1, Sap5 and Tdh3 Are IC Diagnostic Biomarker Candidates
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Sudbery, P.; Gow, N.; Berman, J. The distinct morphogenic states of Candida albicans. Trends Microbiol. 2004, 12, 317–324. [Google Scholar] [CrossRef]
- Höfs, S.; Mogavero, S.; Hube, B. Interaction of Candida albicans with host cells: Virulence factors, host defense, escape strategies, and the microbiota. J. Microbiol. 2016, 54, 149–169. [Google Scholar] [CrossRef] [PubMed]
- Gow, N.A.; Hube, B. Importance of the Candida albicans cell wall during commensalism and infection. Curr. Opin. Microbiol. 2012, 15, 406–412. [Google Scholar] [CrossRef] [PubMed]
- Wilson, D.; Naglik, J.R.; Hube, B. The missing link between Candida albicans hyphal morphogenesis and host cell damage. PLoS Pathog. 2016, 12. [Google Scholar] [CrossRef] [PubMed]
- Pappas, P.G.; Lionakis, M.S.; Arendrup, M.C.; Ostrosky-Zeichner, L.; Kullberg, B.J. Invasive candidiasis. Nat. Rev. Dis. Primers 2018, 4, 18026. [Google Scholar] [CrossRef]
- Kullberg, B.J.; Arendrup, M.C. Invasive candidiasis. N. Engl. J. Med. 2015, 373, 1445–1456. [Google Scholar] [CrossRef]
- Pitarch, A.; Nombela, C.; Gil, C. Diagnosis of invasive candidiasis: From gold standard methods to promising leading-edge technologies. Curr. Top. Med. Chem. 2018, 18, 1375–1392. [Google Scholar] [CrossRef]
- Nucci, M.; Anaissie, E. Revisiting the source of candidemia: Skin or gut? Clin. Infect. Dis. 2001, 33, 1959–1967. [Google Scholar] [CrossRef]
- Phua, A.I.-H.; Hon, K.Y.; Holt, A.; O’Callaghan, M.; Bihari, S. Candida catheter-related bloodstream infection in patients on home parenteral nutrition—Rates, risk factors, outcomes, and management. Clin. Nutr. ESPEN 2019, 31, 1–9. [Google Scholar] [CrossRef]
- Magill, S.S.; Edwards, J.R.; Bamberg, W.; Beldavs, Z.G.; Dumyati, G.; Kainer, M.A.; Lynfield, R.; Maloney, M.; McAllister-Hollod, L.; Nadle, J.; et al. Multistate point-prevalence survey of health care-associated infections. N. Engl. J. Med. 2014, 370, 1198–1208. [Google Scholar] [CrossRef]
- Chaves, F.; Garnacho-Montero, J.; Del Pozo, J.L.; Bouza, E.; Capdevila, J.A.; de Cueto, M.; Dominguez, M.A.; Esteban, J.; Fernandez-Hidalgo, N.; Fernandez Sampedro, M.; et al. Diagnosis and treatment of catheter-related bloodstream infection: Clinical guidelines of the Spanish Society of Infectious Diseases and Clinical Microbiology and (SEIMC) and the Spanish Society of Spanish Society of Intensive and Critical Care Medicine and Coronary Units (SEMICYUC). Med. Intensiva 2018, 42, 5–36. [Google Scholar]
- Clancy, C.J.; Nguyen, M.H. Finding the “missing 50%” of invasive candidiasis: How nonculture diagnostics will improve understanding of disease spectrum and transform patient care. Clin. Infect. Dis. 2013, 56, 1284–1292. [Google Scholar] [CrossRef]
- Posch, W.; Heimdörfer, D.; Wilflingseder, D.; Lass-Flörl, C. Invasive candidiasis: Future directions in non-culture based diagnosis. Expert Rev. Anti Infect. Ther. 2017, 15, 829–838. [Google Scholar] [CrossRef]
- Pfaller, M.A.; Castanheira, M. Nosocomial candidiasis: Antifungal stewardship and the importance of rapid diagnosis. Med. Mycol. 2015, 54, 1–22. [Google Scholar] [CrossRef]
- Clancy, C.J.; Nguyen, M.H. T2 magnetic resonance for the diagnosis of bloodstream infections: Charting a path forward. J. Antimicrob. Chemother. 2018, 73, iv2–iv5. [Google Scholar] [CrossRef] [PubMed]
- Mylonakis, E.; Clancy, C.J.; Ostrosky-Zeichner, L.; Garey, K.; Alangaden, G.J.; Vazquez, J.A.; Groeger, J.S.; Judson, M.A.; Vinagre, Y.-M.; Heard, S.O.; et al. T2 magnetic resonance assay for the rapid diagnosis of candidemia in whole blood: A clinical trial. Clin. Infect. Dis. 2015, 60, 892–899. [Google Scholar] [CrossRef] [PubMed]
- Neely, L.A.; Audeh, M.; Phung, N.A.; Min, M.; Suchocki, A.; Plourde, D.; Blanco, M.; Demas, V.; Skewis, L.R.; Anagnostou, T.; et al. T2 magnetic resonance enables nanoparticle-mediated rapid detection of candidemia in whole blood. Sci. Transl. Med. 2013, 5, 182ra54. [Google Scholar] [CrossRef] [PubMed]
- Krah, A.; Jungblut, P.R. Immunoproteomics. Methods Mol. Med. 2004, 94, 19–32. [Google Scholar] [CrossRef]
- Dea-Ayuela, M.A.; Ubeira, F.M.; Pitarch, A.; Gil, C.; Martinez-Fernandez, A.R.; Bolas, F. A comparison of antigenic peptides in muscle larvae of several Trichinella species by two-dimensional Western-blot analysis with monoclonal antibodies. Parasite 2001, 8, S117–S119. [Google Scholar] [CrossRef]
- Jungblut, P.R.; Bumann, D. Immunoproteome of Helicobacter pylori. Methods Enzymol. 2002, 358, 307–316. [Google Scholar]
- Pedersen, S.K.; Sloane, A.J.; Prasad, S.S.; Sebastian, L.T.; Lindner, R.A.; Hsu, M.; Robinson, M.; Bye, P.T.; Weinberger, R.P.; Harry, J.L. An immunoproteomic approach for identification of clinical biomarkers for monitoring disease: Application to cystic fibrosis. Mol. Cell Proteomics 2005, 4, 1052–1060. [Google Scholar] [CrossRef] [PubMed]
- Thomas, D.P.; Pitarch, A.; Monteoliva, L.; Gil, C.; Lopez-Ribot, J. Proteomics to study Candida albicans biology and pathogenicity. Infect. Disord. Drug Targets 2006, 6, 335–341. [Google Scholar] [CrossRef] [PubMed]
- Pitarch, A.; Sánchez, M.; Nombela, C.; Gil, C. Analysis of the Candida albicans proteome. I. Strategies and applications. J. Chromatogr. B Analyt. Technol. Biomed. Life Sci. 2003, 787, 101–128. [Google Scholar] [CrossRef]
- Pitarch, A.; Jiménez, A.; Nombela, C.; Gil, C. Decoding serological response to Candida cell wall immunome into novel diagnostic, prognostic, and therapeutic candidates for systemic candidiasis by proteomic and bioinformatic analyses. Mol. Cell. Proteomics 2006, 5, 79–96. [Google Scholar] [CrossRef]
- Pitarch, A.; Abian, J.; Carrascal, M.; Sánchez, M.; Nombela, C.; Gil, C. Proteomics—based identification of novel Candida albicans antigens for diagnosis of systemic candidiasis in patients with underlying hematological malignancies. Proteomics 2004, 4, 3084–3106. [Google Scholar] [CrossRef] [PubMed]
- Pitarch, A.; Nombela, C.; Gil, C. Contributions of proteomics to diagnosis, treatment, and prevention of candidiasis. Methods Biochem. Anal. 2006, 49, 331–361. [Google Scholar] [CrossRef] [PubMed]
- Pitarch, A.; Nombela, C.; Gil, C. Prediction of the clinical outcome in invasive candidiasis patients based on molecular fingerprints of five anti-Candida antibodies in serum. Mol. Cell. Proteomics 2011, 10, M110:004010. [Google Scholar] [CrossRef] [PubMed]
- Pitarch, A.; Nombela, C.; Gil, C. Serum antibody signature directed against Candida albicans Hsp90 and enolase detects invasive candidiasis in non-neutropenic patients. J. Proteome Res. 2014, 13, 5165–5184. [Google Scholar] [CrossRef] [PubMed]
- Pitarch, A.; Nombela, C.; Gil, C. Seroprofiling at the Candida albicans protein species level unveils an accurate molecular discriminator for candidemia. J. Proteomics 2016, 134, 144–162. [Google Scholar] [CrossRef]
- Huertas, B.; Prieto, D.; Pitarch, A.; Gil, C.; Pla, J.; Díez-Orejas, R. Serum antibody profile during colonization of the mouse gut by Candida albicans: Relevance for protection during systemic infection. J. Proteome Res. 2017, 16, 335–345. [Google Scholar] [CrossRef]
- Pitarch, A.; Nombela, C.; Gil, C. The Candida immunome as a mine for clinical biomarker development for invasive candidiasis: From biomarker discovery to assay validation. In Pathogenic Fungi: Insights in Molecular Biology Wymondham; San-Blas, G., Calderone, R., Eds.; Caister Academic Press: Norfolk, UK, 2008; pp. 103–142. [Google Scholar]
- Pitarch, A.; Diez-Orejas, R.; Molero, G.; Pardo, M.; Sanchez, M.; Gil, C.; Nombela, C. Analysis of the serologic response to systemic Candida albicans infection in a murine model. Proteomics 2001, 1, 550–559. [Google Scholar] [CrossRef]
- Luo, T.; Krüger, T.; Knüpfer, U.; Kasper, L.; Wielsch, N.; Hube, B.; Kortgen, A.; Bauer, M.; Giamarellos-Bourboulis, E.J.; Dimopoulos, G.; et al. Immunoproteomic analysis of antibody responses to extracellular proteins of Candida albicans revealing the importance of glycosylation for antigen recognition. J. Proteome Res. 2016, 15, 2394–2406. [Google Scholar] [CrossRef]
- Klis, F.M.; Brul, S. Adaptations of the secretome of Candida albicans in response to host-related environmental conditions. Eukaryot. Cell 2015, 14, 1165–1172. [Google Scholar] [CrossRef] [PubMed]
- Gil-Bona, A.; Amador-García, A.; Gil, C.; Monteoliva, L. The external face of Candida albicans: A proteomic view of the cell surface and the extracellular environment. J. Proteomics 2018, 180, 70–79. [Google Scholar] [CrossRef] [PubMed]
- Citiulo, F.; Jacobsen, I.D.; Miramón, P.; Schild, L.; Brunke, S.; Zipfel, P.; Brock, M.; Hube, B.; Wilson, D. Candida albicans scavenges host zinc via Pra1 during endothelial invasion. PLoS Pathog. 2012, 8, e1002777. [Google Scholar] [CrossRef]
- Wu, H.; Downs, D.; Ghosh, K.; Ghosh, A.K.; Staib, P.; Monod, M.; Tang, J. Candida albicans secreted aspartic proteases 4–6 induce apoptosis of epithelial cells by a novel Trojan horse mechanism. FASEB J. 2013, 27, 2132–2144. [Google Scholar] [CrossRef]
- Nombela, C.; Gil, C.; Chaffin, W.L. Non-conventional protein secretion in yeast. Trends Microbiol. 2006, 14, 15–21. [Google Scholar] [CrossRef]
- Gil-Bona, A.; Llama-Palacios, A.; Parra, C.M.; Vivanco, F.; Nombela, C.; Monteoliva, L.; Gil, C. Proteomics unravels extracellular vesicles as carriers of classical cytoplasmic proteins in Candida albicans. J. Proteome Res. 2015, 14, 142–153. [Google Scholar] [CrossRef]
- Satala, D.; Satala, G.; Karkowska-Kuleta, J.; Bukowski, M.; Kluza, A.; Rapala-Kozik, M.; Kozik, A. Structural insights into the interactions of candidal enolase with human vitronectin, fibronectin and plasminogen. Int. J. Mol. Sci. 2020, 21, 7843. [Google Scholar] [CrossRef] [PubMed]
- Pärnänen, P.; Sorsa, T.; Tervahartiala, T.; Nikula-Ijäs, P. Isolation, characterization and regulation of moonlighting proteases from Candida glabrata cell wall. Microb. Pathog. 2020, 149, 104547. [Google Scholar] [CrossRef]
- Nimrichter, L.; De Souza, M.M.; Del Poeta, M.; Nosanchuk, J.D.; Joffe, L.; Tavares, P.D.M.; Rodrigues, M. Extracellular vesicle-associated transitory cell wall components and their impact on the interaction of fungi with host cells. Front. Microbiol. 2016, 7, 1034. [Google Scholar] [CrossRef]
- Zarnowski, R.; Sanchez, H.; Covelli, A.S.; Dominguez, E.; Jaromin, A.; Bernhardt, J.; Mitchell, K.F.; Heiss, C.; Azadi, P.; Mitchell, A.; et al. Candida albicans biofilm–induced vesicles confer drug resistance through matrix biogenesis. PLoS Biol. 2018, 16, e2006872. [Google Scholar] [CrossRef]
- Yin, Q.Y. Exploring the Fungal Wall Proteome by Mass Spectrometry; University of Amsterdam: Amsterdam, The Netherlands, 2008. [Google Scholar]
- Lee, K.L.; Buckley, H.R.; Campbell, C.C. An amino acid liquid synthetic medium for the development of mycelial and yeast forms of Candida albicans. Sabouraudia 1975, 13, 148–153. [Google Scholar] [CrossRef]
- Pitarch, A.; Nombela, C.; Gil, C. Top-down characterization data on the speciation of the Candida albicans immunome in candidemia. Data Brief. 2016, 6, 257–261. [Google Scholar] [CrossRef]
- Pitarch, A.; Nombela, C.; Gil, C. Identification of the Candida albicans immunome during systemic infection by mass spectrometry. Methods Mol. Biol. 2009, 470, 187–235. [Google Scholar] [CrossRef]
- Valdés, I.; Pitarch, A.; Gil, C.; Bermúdez, A.; Llorente, M.; Nombela, C.; Méndez, E. Novel procedure for the identification of proteins by mass fingerprinting combining two-dimensional electrophoresis with fluorescent SYPRO Red staining. J. Mass Spectrom. 2000, 35, 672–682. [Google Scholar] [CrossRef]
- Pitarch, A.; Nombela, C.; Gil, C. Reliability of antibodies to Candida methionine synthase for diagnosis, prognosis and risk stratification in systemic candidiasis: A generic strategy for the prototype development phase of proteomic markers. Proteomics Clin. Appl. 2007, 1, 1221–1242. [Google Scholar] [CrossRef] [PubMed]
- Pitarch, A.; Pardo, M.; Jimenez, A.; Pla, J.; Gil, C.; Sanchez, M.; Nombela, C. Two-dimensional gel electrophoresis as analytical tool for identifying Candida albicans immunogenic proteins. Electrophoresis 1999, 20, 1001–1010. [Google Scholar] [CrossRef]
- Pitarch, A.; Nombela, C.; Gil, C. Proteomic profiling of serologic response to Candida albicans during host-commensal and host-pathogen interactions. Methods Mol. Biol. 2009, 470, 369–411. [Google Scholar] [CrossRef] [PubMed]
- Zybailov, B.; Mosley, A.L.; Sardiu, M.E.; Coleman, M.K.; Florens, L.; Washburn, M.P. Statistical analysis of membrane proteome expression changes in Saccharomyces cerevisiae. J. Proteome Res. 2006, 5, 2339–2347. [Google Scholar] [CrossRef] [PubMed]
- Martínez, J.P.; Gil, M.L.; López-Ribot, J.L.; Chaffin, W.L. Serologic response to cell wall mannoproteins and proteins of Candida albicans. Clin. Microbiol. Rev. 1998, 11, 121–141. [Google Scholar] [CrossRef] [PubMed]
- Mochon, A.B.; Jin, Y.; Kayala, M.A.; Wingard, J.R.; Clancy, C.J.; Nguyen, M.H.; Felgner, P.; Baldi, P.; Liu, H. Serological profiling of a Candida albicans protein microarray reveals permanent host-pathogen interplay and stage-specific responses during candidemia. PLoS Pathog. 2010, 6, e1000827. [Google Scholar] [CrossRef]
- Nantel, A.; Dignard, D.; Bachewich, C.; Harcus, D.; Marcil, A.; Bouin, A.-P.; Sensen, C.W.; Hogues, H.; Hoog, M.V.H.; Gordon, P.; et al. Transcription profiling of Candida albicans cells undergoing the yeast-to-hyphal transition. Mol. Biol. Cell 2002, 13, 3452–3465. [Google Scholar] [CrossRef]
- Pitarch, A.; Sánchez, M.; Nombela, C.; Gil, C. Sequential fractionation and two-dimensional gel analysis unravels the complexity of the dimorphic fungus Candida albicans cell wall proteome. Mol. Cell. Proteomics 2002, 1, 967–982. [Google Scholar] [CrossRef]
- Naglik, J.R.; Challacombe, S.J.; Hube, B. Candida albicans secreted aspartyl proteinases in virulence and pathogenesis. Microbiol. Mol. Biol. Rev. 2003, 67, 400–428. [Google Scholar] [CrossRef]
- Chaffin, W.L. Candida albicans cell wall proteins. Microbiol. Mol. Biol. Rev. 2008, 72, 495–544. [Google Scholar] [CrossRef]
- Martinez, R.; Monteoliva, L.; Diez-Orejas, R.; Nombela, C.; Gil, C. The GPI-anchored protein CaEcm33p is required for cell wall integrity, morphogenesis and virulence in Candida albicans. Microbiology 2004, 150, 3341–3354. [Google Scholar] [CrossRef]
- Gil-Bona, A.; Reales-Calderon, J.A.; Giraldo, C.M.P.; Martinez, R.; Monteoliva, L.; Gil, C. The cell wall protein Ecm33 of Candida albicans is involved in chronological life span, morphogenesis, cell wall regeneration, stress tolerance, and host–cell interaction. Front. Microbiol. 2016, 7, 64. [Google Scholar] [CrossRef]
- Gil-Bona, A.; Monteoliva, L.; Gil García, C. Global proteomic profiling of the secretome of Candida albicans ecm33 cell wall mutant reveals the involvement of Ecm33 in Sap2 secretion. J. Proteome Res. 2015, 14, 4270–4281. [Google Scholar] [CrossRef]
- Martinez-Lopez, R.; Park, H.; Myers, C.L.; Gil, C.; Filler, S.G. Candida albicans Ecm33p is important for normal cell wall architecture and interactions with host cells. Eukaryot. Cell 2006, 5, 140–147. [Google Scholar] [CrossRef]
- Monteoliva, L.; Martinez, R.; Pitarch, A.; Hernaez, M.L.; Serna, A.; Nombela, C.; Albar, J.P.; Gil, C. Quantitative proteome and acidic subproteome profiling of Candida albicans yeast-to-hypha transition. J. Proteome Res. 2011, 10, 502–517. [Google Scholar] [CrossRef]
- Ene, I.V.; Heilmann, C.J.; Sorgo, A.G.; Walker, L.A.; de Koster, C.G.; Munro, C.A.; Klis, F.M.; Brown, A.J. Carbon source-induced reprogramming of the cell wall proteome and secretome modulates the adherence and drug resistance of the fungal pathogen Candida albicans. Proteomics 2012, 12, 3164–3179. [Google Scholar] [CrossRef]
- de Groot, P.W.; de Boer, A.D.; Cunningham, J.; Dekker, H.L.; de Jong, L.; Hellingwerf, K.J.; de Koster, C.; Klis, F.M. Proteomic analysis of Candida albicans cell walls reveals covalently bound carbohydrate-active enzymes and adhesins. Eukaryot. Cell 2004, 3, 955–965. [Google Scholar] [CrossRef]
- Pitarch, A.; Nombela, C.; Gil, C. Cell wall fractionation for yeast and fungal. Methods Mol. Biol. 2008, 425, 217–239. [Google Scholar] [CrossRef] [PubMed]
- Sorgo, A.G.; Heilmann, C.J.; Dekker, H.L.; Brul, S.; De Koster, C.G.; Klis, F.M. Mass spectrometric analysis of the secretome of Candida albicans. Yeast 2010, 27, 661–672. [Google Scholar] [CrossRef]
- Wolf, J.M.; Espadas, J.; Luque-Garcia, J.L.; Reynolds, T.; Casadevall, A. Lipid biosynthetic genes affect Candida albicans extracellular vesicle morphology, cargo, and immunostimulatory properties. Eukaryot. Cell 2015, 14, 745–754. [Google Scholar] [CrossRef] [PubMed]
- Gil-Navarro, I.; Gil, M.L.; Casanova, M.; O’Connor, J.E.; Martínez, J.P.; Gozalbo, D. The glycolytic enzyme glyceraldehyde-3-phosphate dehydrogenase of Candida albicans is a surface antigen. J. Bacteriol. 1997, 179, 4992–4999. [Google Scholar] [CrossRef]
- Karkowska-Kuleta, J.; Kozik, A. Moonlighting proteins as virulence factors of pathogenic fungi, parasitic protozoa and multicellular parasites. Mol. Oral Microbiol. 2014, 29, 270–283. [Google Scholar] [CrossRef]
- Heilmann, C.J.; Sorgo, A.G.; Mohammadi, S.; Sosinska, G.J.; De Koster, C.G.; Brul, S.; De Koning, L.J.; Klis, F.M. Surface stress induces a conserved cell wall stress response in the pathogenic fungus Candida albicans. Eukaryot. Cell 2012, 12, 254–264. [Google Scholar] [CrossRef]
- Gil-Bona, A.; Parra-Giraldo, C.M.; Hernáez, M.L.; Reales-Calderon, J.A.; Solis, N.V.; Filler, S.G.; Monteoliva, L.; Gil, C. Candida albicans cell shaving uncovers new proteins involved in cell wall integrity, yeast to hypha transition, stress response and host–pathogen interaction. J. Proteomics 2015, 127, 340–351. [Google Scholar] [CrossRef]
- Marin, E.; Parragiraldo, C.M.; Hernández-Haro, C.; Hernaez, M.L.; Nombela, C.; Monteoliva, L.; Gil, C. Candida albicans shaving to profile human serum proteins on hyphal surface. Front. Microbiol. 2015, 6, 1343. [Google Scholar] [CrossRef]
- Hernáez, M.L.; Ximénez-Embún, P.; Martínez-Gomariz, M.; Gutiérrez-Blázquez, M.D.; Nombela, C.; Gil, C. Identification of Candida albicans exposed surface proteins in vivo by a rapid proteomic approach. J. Proteomics 2010, 73, 1404–1409. [Google Scholar] [CrossRef] [PubMed]
- Martinez-Gomariz, M.; Perumal, P.; Mekala, S.; Nombela, C.; Chaffin, W.L.; Gil, C. Proteomic analysis of cytoplasmic and surface proteins from yeast cells, hyphae, and biofilms of Candida albicans. Proteomics 2009, 9, 2230–2252. [Google Scholar] [CrossRef]
- Vialás, V.; Perumal, P.; Gutierrez, D.; Ximénez-Embún, P.; Nombela, C.; Gil, C.; Chaffin, W.L. Cell surface shaving of Candida albicans biofilms, hyphae, and yeast form cells. Proteomics 2012, 12, 2331–2339. [Google Scholar] [CrossRef]
- Pardo, M.; Ward, M.; Pitarch, A.; Sanchez, M.; Nombela, C.; Blackstock, W.; Gil, C. Cross-species identification of novel Candida albicans immunogenic proteins by combination of two-dimensional polyacrylamide gel electrophoresis and mass spectrometry. Electrophoresis 2000, 21, 2651–2659. [Google Scholar] [CrossRef]
- Pitarch, A.; Jiménez, A.; Nombela, C.; Gil, C. Serological proteome analysis to identify systemic candidiasis patients in the intensive care unit: Analytical, diagnostic and prognostic validation of anti-Candida enolase antibodies on quantitative clinical platforms. Proteomics Clin. Appl. 2008, 2, 596–618. [Google Scholar] [CrossRef]
- Gomez, M.J.; Torosantucci, A.; Arancia, S.; Maras, B.; Parisi, L.; Cassone, A. Purification and biochemical characterization of a 65-kilodalton mannoprotein (MP65), a main target of anti-Candida cell-mediated immune responses in humans. Infect. Immun. 1996, 64, 2577–2584. [Google Scholar] [CrossRef]
- Torosantucci, A.; Tumbarello, M.; Bromuro, C.; Chiani, P.; Posteraro, B.; Sanguinetti, M.; Cauda, R.; Cassone, A. Antibodies against a beta-glucan-protein complex of Candida albicans and its potential as indicator of protective immunity in candidemic patients. Sci. Rep. 2017, 7, 2722. [Google Scholar] [CrossRef]
- Viudes, A.; Lazzell, A.; Perea, S.; Kirkpatrick, W.R.; Peman, J.; Patterson, T.F.; Martinez, J.P.; Lopez-Ribot, J.L. The C-terminal antibody binding domain of Candida albicans mp58 represents a protective epitope during candidiasis. FEMS Microbiol. Lett. 2004, 232, 133–138. [Google Scholar] [CrossRef]
- Yang, Y.; Thannhauser, T.W.; Li, L.; Zhang, S. Development of an integrated approach for evaluation of 2-D gel image analysis: Impact of multiple proteins in single spots on comparative proteomics in conventional 2-D gel/MALDI workflow. Electrophoresis 2007, 28, 2080–2094. [Google Scholar] [CrossRef]
- Chou, H.; Tam, M.F.; Chang, C.Y.; Lai, H.Y.; Huang, M.H.; Chou, C.T.; Lee, S.S.; Shen, H.D. Characterization of a novel Candida albicans 29 kDa IgE-binding protein—Purification, cDNA isolation and heterologous expression of Cand a 3. Allergy 2003, 58, 1157–1164. [Google Scholar] [CrossRef]
- Ardizzoni, A.; Posteraro, B.; Baschieri, M.C.; Bugli, F.; Sáez-Rosòn, A.; Manca, L.; Cacaci, M.; Sterbini, F.P.; De Waure, C.; Sevilla, M.; et al. An antibody reactivity-based assay for diagnosis of invasive candidiasis using protein array. Int. J. Immunopathol. Pharmacol. 2014, 27, 403–412. [Google Scholar] [CrossRef] [PubMed]
- He, Z.X.; Chen, J.; Li, W.; Cheng, Y.; Zhang, H.P.; Zhang, L.N.; Hou, T.W. Serological response and diagnostic value of recombinant Candida cell wall protein enolase, phosphoglycerate kinase, and beta-glucosidase. Front. Microbiol. 2015, 6, 920. [Google Scholar] [CrossRef] [PubMed]
- Xin, H.; Dziadek, S.; Bundle, D.R.; Cutler, J.E. Synthetic glycopeptide vaccines combining beta-mannan and peptide epitopes induce protection against candidiasis. Proc. Natl. Acad. Sci. USA 2008, 105, 13526–13531. [Google Scholar] [CrossRef]
- Xin, H.; Cutler, J.E. Vaccine and monoclonal antibody that enhance mouse resistance to candidiasis. Clin. Vaccine Immunol. 2011, 18, 1656–1667. [Google Scholar] [CrossRef]
- Li, W.Q.; Hu, X.C.; Zhang, X.; Ge, Y.; Zhao, S.; Hu, Y.; Ashman, R.B. Immunisation with the glycolytic enzyme enolase confers effective protection against Candida albicans infection in mice. Vaccine 2011, 29, 5526–5533. [Google Scholar] [CrossRef] [PubMed]
- Sarthy, A.V.; McGonigal, T.; Coen, M.; Frost, D.J.; Meulbroek, J.A.; Goldman, R.C. Phenotype in Candida albicans of a disruption of the BGL2 gene encoding a 1,3-beta-glucosyltransferase. Microbiology 1997, 143, 367–376. [Google Scholar] [CrossRef]
- Taff, H.T.; Nett, J.E.; Zarnowski, R.; Ross, K.M.; Sanchez, H.; Cain, M.T.; Hamaker, J.; Mitchell, A.P.; Andes, D.R. A Candida biofilm-induced pathway for matrix glucan delivery: Implications for drug resistance. PLoS Pathog. 2012, 8, e1002848. [Google Scholar] [CrossRef]
- Silva, R.; Padovan, A.C.B.; Pimenta, D.C.; Ferreira, R.C.; Da Silva, C.V.; Briones, M.R.S. Extracellular enolase of Candida albicans is involved in colonization of mammalian intestinal epithelium. Front. Cell. Infect. Microbiol. 2014, 4, 66. [Google Scholar] [CrossRef]
- Jong, A.Y.; Chen, S.H.M.; Stins, M.F.; Kim, K.S.; Tuan, T.-L.; Huang, S.-H. Binding of Candida albicans enolase to plasmin(ogen) results in enhanced invasion of human brain microvascular endothelial cells. J. Med. Microbiol. 2003, 52, 615–622. [Google Scholar] [CrossRef]
- Pitarch, A.; Nombela, C.; Gil, C. Candida albicans biology and pathogenicity: Insights from proteomics. Methods Biochem. Anal. 2006, 49, 285–330. [Google Scholar] [CrossRef]
- Crowe, J.D.; Sievwright, I.K.; Auld, G.C.; Moore, N.R.; Gow, N.A.R.; Booth, N.A. Candida albicans binds human plasminogen: Identification of eight plasminogen-binding proteins. Mol. Microbiol. 2003, 47, 1637–1651. [Google Scholar] [CrossRef]
- Alloush, H.M.; Lopez-Ribot, J.; Masten, B.J.; Chaffin, W.L. 3-Phosphoglycerate kinase: A glycolytic enzyme protein present in the cell wall of Candida albicans. Microbiology 1997, 143, 321–330. [Google Scholar] [CrossRef]
- Hasim, S.; Hussin, N.A.; Alomar, F.; Bidasee, K.R.; Nickerson, K.W.; Wilson, M.A. A Glutathione-independent glyoxalase of the DJ-1 superfamily plays an important role in managing metabolically generated methylglyoxal in Candida albicans. J. Biol. Chem. 2014, 289, 1662–1674. [Google Scholar] [CrossRef] [PubMed]
- Cabello, L.; Gómez-Herreros, E.; Fernández-Pereira, J.; Maicas, S.; Martínez-Esparza, M.C.; De Groot, P.W.J.; Valentín, E. Deletion of GLX3 in Candida albicans affects temperature tolerance, biofilm formation and virulence. FEMS Yeast Res. 2018, 19. [Google Scholar] [CrossRef]
- Joo, M.Y.; Song, E.S.; Kee, S.J.; Shin, J.H.; Jang, H.-C.; Suh, S.P.; Ryang, D.W. Expression of SAP5 and SAP9 in Candida albicans biofilms: Comparison of bloodstream isolates with isolates from other sources. Med. Mycol. 2013, 51, 892–896. [Google Scholar] [CrossRef]
- Luo, S.; Dasari, P.; Reiher, N.; Hartmann, A.; Jacksch, S.; Wende, E.; Barz, D.; Niemiec, M.J.; Jacobsen, I.; Beyersdorf, N.; et al. The secreted Candida albicans protein Pra1 disrupts host defense by broadly targeting and blocking complement C3 and C3 activation fragments. Mol. Immunol. 2018, 93, 266–277. [Google Scholar] [CrossRef]
- Hiller, E.; Heine, S.; Brunner, H.; Rupp, S. Candida albicans Sun41p, a putative glycosidase, is involved in morphogenesis, cell wall biogenesis, and biofilm formation. Eukaryot. Cell 2007, 6, 2056–2065. [Google Scholar] [CrossRef] [PubMed][Green Version]
- McCreath, K.J.; Specht, C.A.; Robbins, P.W. Molecular cloning and characterization of chitinase genes from Candida albicans. Proc. Natl. Acad. Sci. USA 1995, 92, 2544–2548. [Google Scholar] [CrossRef]
- Bromuro, C.; Torosantucci, A.; Gomez, M.; Urbani, F.; Cassone, A. Differential release of an immunodominant 65 kDa mannoprotein antigen from yeast and mycelial forms of Candida albicans. Med. Mycol. 1994, 32, 447–459. [Google Scholar] [CrossRef] [PubMed]
- Lisowska, E. The role of glycosylation in protein antigenic properties. Cell. Mol. Life Sci. 2002, 59, 445–455. [Google Scholar] [CrossRef] [PubMed]

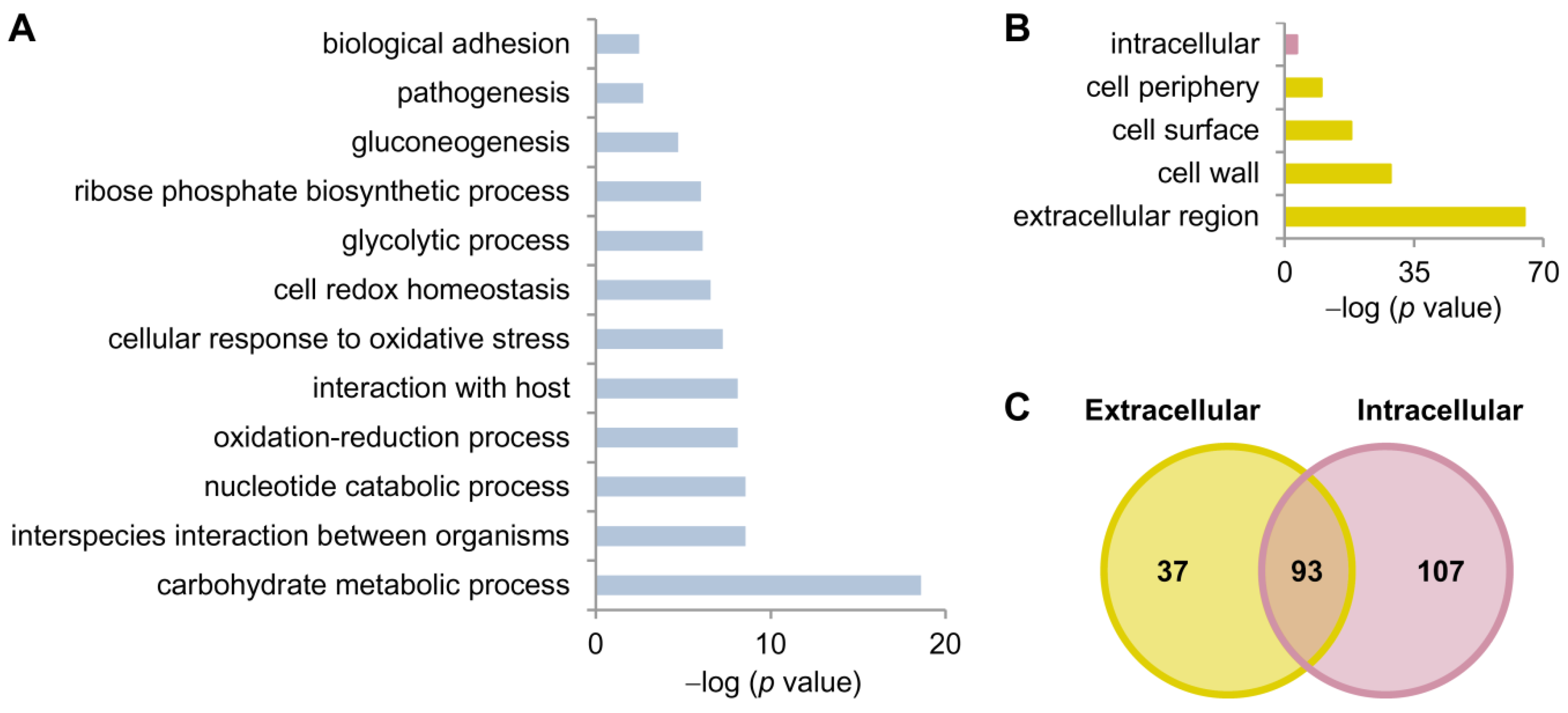
| Cellular Location | No. of Identified Proteins | Protein Names |
|---|---|---|
| Extracellular region, cell surface, cell wall, and cell periphery | 37 | Ade8, Als1, Bgl2, Cht1, Cht3, Cip1, Coi1, Ece1, Ecm33, Gcy1, Grp2, Hex1, Kre9, Mal2, Mdg1, Met15, Mp65, Nit3, Ofr1, Orf19.1394, Orf19.3053, Orf19.6809, Orf19.6867, Orf19.7322, Pga12, Pga4, Pra1, Pst2, Rbe1, Rbp1, Rhd3, Sah1, Sap10, Sap8, Slk19, Sol3, Xyl2 |
| Intracellular region | 107 | Ade17, Aha1, Ams1, Anb1, Arf2, Arg1, Arg3, Arg4, Aro2, Asc1, Bfr1, Bud7, Cit1, Cmd1, Cpr6, Crm1, Cyc1, Dtd2, Ecm4, Egd1, Eif4e, Erg10, Erg20, Etr1, Fbp1, Fum11, Gcv3, Grs1, Grx3, Het1, His1, Hom2, Hta3, Hxk2, Idh1, Kex1, Krs1, Lat1, Lsc1, Lys21, Mca1, Mcr1, Mdh1, Mmd1, Mrf1, Mxr1, Npt1, Ntf2, Orf19.1355, Orf19.1448.1, Orf19.1738.1, Orf19.1815, Orf19.2930, Orf19.3319, Orf19.3681, Orf19.3932, Orf19.4150, Orf19.4382, Orf19.4898, Orf19.518, Orf19.5322, Orf19.5943.1, Orf19.5961, Orf19.6559, Orf19.6596, Orf19.6701, Orf19.6872, Orf19.7152, Orf19.7196, Orf19.7214, Orf19.7297, Orf19.7330, Orf19.7368, Orf19.7404, Orf19.7531, Orf19.7578, Orf19.86, Orf19.904, Pol30, Rdi1, Rnr21, Rpl10a, Rpl12, Rpl30, Rpp0, Rps12, Rps19a, Rps21b, Rps22a, Sbp1, Sec14, Skp1, Sno1, Snz1, Sod2, Sod3, Sub2, Sui1, Tfs1, Thi4, Tif1, Tub1, Tub2, Tup1, Uba1, Yhb1, Ykt6 |
| Both locations | 93 | Aat21, Abp1, Acb1, Aco1, Ahp1, Ape3, Atp1, Atp2, Bmh1, Cam1, Cat1, Cdc19, Cdc3, Cof1, Cyp1, Cyp5, Eft2, Egd2, Emp24, Eng1, Eno1, Fba1, Fdh1, Gdh3, Gnd1, Gpd2, Gph1, Gpm1, Gsp1, Hem13, Hsp12, Hsp70, Hsp90, Ino1, Ipp1, Kel1, Leu2, Lpd1, Lsp1, Mdh1-1, Met6, Mir1, Mlc1, Mnt1, Orf19.1085, Orf19.1862, Orf19.1946, Orf19.3915, Orf19.4395, Orf19.4597, Orf19.5342, Pdc11, Pet9, Pfy1, Pgi1, Pgk1, Phr2, Pin3, Pmm1, Por1, Prx1, Pst3, Rho1, Rib3, Rpl14, Rpl6, Rps20, Sam2, Sap4, Sap5, Sec4, Sim1, Smt3, Sod1, Spe3, Ssa2, Ssb1, Tal1, Tdh3, Tma19, Tpi1, Tpm2, Trr1, Trx1, Tsa1, Ttr1, Ubi3, Ugp1, Utr2, Vps21, Xog1, Ynk1, Ypt1 |
| Immunoreactive C. albicans Hyphal Secreted Proteins | IgG Antibody- Reactivity Levels a | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Protein Name | Protein Description | LC-MS/MS | MALDI-TOF-MS | ncIC | cIC | Non- IC | ||||
| No. of Peptides | Ranking According to NSAF | NSAF | No. of Matched/ Unmatched Peptides | Mascot Score | % of Sequence Coverage | |||||
| Bgl2 | Cell wall 1,3-beta- glucosyltransferase | 8 | 55 | 0.004 | 10/58 | 213 | 20 | +++ | +++ | − |
| Eno1 | Enolase | 30 | 1 | 0.08 | 9/61 | 157 | 32 | ++++ | +++ | + |
| Glx3 | Glutathione- independent glyoxalase | NI b | NI b | NI b | 12/56 | 276 | 61 | +++ | +++ | − |
| Pgk1 | Phosphoglycerate kinase | 38 | 2 | 0.04 | 21/47 | 460 | 48 | +++ | ++ | + |
| Pra1 | pH regulated antigen | 10 | 12 | 0.01 | 11/57 | 240 | 29 | ND | +++++ | +++ |
| 9/59 | 218 | 16 | ||||||||
| 10/58 | 229 | 20 | ||||||||
| 7/61 | 75 | 8 | ||||||||
| 7/60 | 115 | 11 | ||||||||
| 5/63 | 96 | 7 | ||||||||
| Sap5 | Secreted aspartyl proteinase | 26 | 3 | 0.03 | 14/54 | 272 | 39 | ++ | +++ | ++ |
| 28/43 | 815 | 60 | ||||||||
| 39/38 | 1220 | 67 | ||||||||
| 6/62 | 60 | 18 | ||||||||
| Tdh3 | Glyceraldehyde-3- phosphate | 27 | 9 | 0.02 | 16/53 | 140 | 51 | − | + | − |
| Standard Name | Description | NetNGlyc 1.0 Server Prediction | NetNGlyc 4.0 Server Prediction | NSAF | No. Peptides | Mr |
|---|---|---|---|---|---|---|
| Rbt4 | Pry family protein | no | yes | 0.19 | 11 | 37.4 |
| Mp65 | Cell surface mannoprotein | no | yes | 0.15 | 13 | 39.3 |
| Sun41 | Cell wall glycosidase | yes | yes | 0.12 | 11 | 43.7 |
| Tos1 | Protein similar to alpha agglutinin anchor subunit | no | yes | 0.10 | 15 | 49.4 |
| Ecm33 | GPI-anchored cell wall protein | yes | yes | 0.03 | 6 | 43.5 |
| Cht3 | Major chitinase | no | yes | 0.03 | 7 | 60.0 |
| Pra1 | Cell surface protein that sequesters zinc from host tissue | yes | yes | 0.02 | 7 | 33.1 |
| Glx3 | Glutathione-independent glyoxalase | yes | no | 0.02 | 12 | 25.8 |
| Sim1 | Adhesin-like protein | yes | yes | 0.01 | 9 | 39.4 |
| Scw11 | Cell wall protein | yes | yes | 0.01 | 10 | 54.4 |
| Standard Namea | Description a | Cell Localizationb | Signal Peptide a | References c |
|---|---|---|---|---|
| Aco1 | Aconitase | Shared | No | [25,77] |
| Ade17 | 5-Aminoimidazole-4-carboxamide ribotide transformylase | Intracellular | No | [25] |
| Als1 | Cell-surface adhesin | Extracellular | yes | [54] |
| Asc1 | 40S ribosomal subunit similar to G-beta subunits | Intracellular | No | [25] |
| Atp1 | ATP synthase alpha subunit | Shared | No | [25] |
| Atp2 | F1 beta subunit of F1F0 ATPase complex | Shared | No | [25] |
| Bgl2 | Cell wall 1,3-beta-glucosyltransferase | Extracellular | Yes | [24] |
| Cdc19 | Pyruvate kinase at yeast cell surface | Shared | No | [77] |
| Ece1 | Extent of cell elongation protein | Extracellular | yes | [54] |
| Ecm33 | GPI-anchored cell wall protein | Extracellular | yes | [54] |
| Eft2 | Elongation factor 2 | Shared | No | [77] |
| Eno1 | Enolase | Shared | No | [50,53,78] |
| Fba1 | Fructose-bisphosphate aldolase | Shared | No | [25] |
| Gnd1 | 6-phosphogluconate dehydrogenase | Shared | No | [29] |
| Gpm1 | Phosphoglycerate mutase | Shared | No | [25,77] |
| Grp2 | NAD(H)-linked methylglyoxal oxidoreductase involved in regulation of methylglyoxal and pyruvate levels | Extracellular | No | [25] |
| Hem13 | Coproporphyrinogen III oxidase | Shared | No | [25] |
| Hsp70 | Putative Hsp70 chaperone | Shared | No | [25,53] |
| Hsp90 | Essential chaperone | Shared | No | [25,53] |
| Hxk2 | Hexokinase II | Intracellular | No | [25] |
| Ino1 | Inositol-1-phosphate synthase | Shared | No | [25] |
| Ipp1 | Putative inorganic pyrophosphatase | Shared | No | [25] |
| Mdh1 | Mitochondrial malate dehydrogenase | Intracellular | Yes | [25] |
| Mdh1-1 | Predicted malate dehydrogenase precursor | Shared | No | |
| Met6 | 5-methyltetrahydropteroyltriglutamate-homocysteine methyltransferase (methionine synthase) | Shared | No | [49,77] |
| Mp65 | Cell surface mannoprotein | Extracellular | Yes | [79,80] |
| Msi3 | Essential HSP70 family protein | Intracellular | No | |
| orf19.7196 | Putative vacuolar protease | Intracellular | Yes | [33] |
| orf19.7214 | Glucan 1,3-beta-glucosidase | Intracellular | No | [54] |
| Pdc11 | Pyruvate decarboxylase | Shared | No | [25] |
| Pga4 | GPI-anchored cell surface protein | Extracellular | Yes | [54] |
| Pgi1 | Glucose-6-phosphate isomerase | Shared | No | [25] |
| Pgk1 | Phosphoglycerate kinase | Shared | No | [50,53] |
| Por1 | Mitochondrial outer membrane porin | Shared | No | [25] |
| Pra1 | pH regulated antigen | Extracellular | Yes | [54,81] |
| Prx1 | Thioredoxin peroxidase | Shared | No | [33] |
| Sah1 | S-adenosyl-L-homocysteine hydrolase | Extracellular | No | [25] |
| Slk19 | Alkaline-induced protein of plasma membrane | Extracellular | no | [54] |
| Ssa2 | HSP70 family chaperone | Shared | No | [25,54] |
| Ssb1 | HSP70 family heat shock protein | Shared | No | [25,32] |
| Tal1 | Transaldolase | Shared | No | [33] |
| Tdh3 | NAD-linked glyceraldehyde-3-phosphate dehydrogenase | Shared | No | [25,50,69] |
| Tif | Translation initiation factor | Intracellular | No | [25] |
| Tpi1 | Triose-phosphate isomerase | Shared | No | [25,32] |
| Tsa1 | TSA/alkyl hydroperoxide peroxidase C (AhPC) family protein | Shared | No | [27] |
| Utr2 | Putative GPI anchored cell wall glycosidase | Extracellular | yes | [54] |
| Xog1 | Exo-1,3-beta-glucanase | Shared | Yes | [54] |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Vaz, C.; Pitarch, A.; Gómez-Molero, E.; Amador-García, A.; Weig, M.; Bader, O.; Monteoliva, L.; Gil, C. Mass Spectrometry-Based Proteomic and Immunoproteomic Analyses of the Candida albicans Hyphal Secretome Reveal Diagnostic Biomarker Candidates for Invasive Candidiasis. J. Fungi 2021, 7, 501. https://doi.org/10.3390/jof7070501
Vaz C, Pitarch A, Gómez-Molero E, Amador-García A, Weig M, Bader O, Monteoliva L, Gil C. Mass Spectrometry-Based Proteomic and Immunoproteomic Analyses of the Candida albicans Hyphal Secretome Reveal Diagnostic Biomarker Candidates for Invasive Candidiasis. Journal of Fungi. 2021; 7(7):501. https://doi.org/10.3390/jof7070501
Chicago/Turabian StyleVaz, Catarina, Aida Pitarch, Emilia Gómez-Molero, Ahinara Amador-García, Michael Weig, Oliver Bader, Lucía Monteoliva, and Concha Gil. 2021. "Mass Spectrometry-Based Proteomic and Immunoproteomic Analyses of the Candida albicans Hyphal Secretome Reveal Diagnostic Biomarker Candidates for Invasive Candidiasis" Journal of Fungi 7, no. 7: 501. https://doi.org/10.3390/jof7070501
APA StyleVaz, C., Pitarch, A., Gómez-Molero, E., Amador-García, A., Weig, M., Bader, O., Monteoliva, L., & Gil, C. (2021). Mass Spectrometry-Based Proteomic and Immunoproteomic Analyses of the Candida albicans Hyphal Secretome Reveal Diagnostic Biomarker Candidates for Invasive Candidiasis. Journal of Fungi, 7(7), 501. https://doi.org/10.3390/jof7070501






