Structural, Evolutionary, and Functional Analysis of the Protein O-Mannosyltransferase Family in Pathogenic Fungi
Abstract
1. Introduction
2. Protein Glycosylation in Fungi
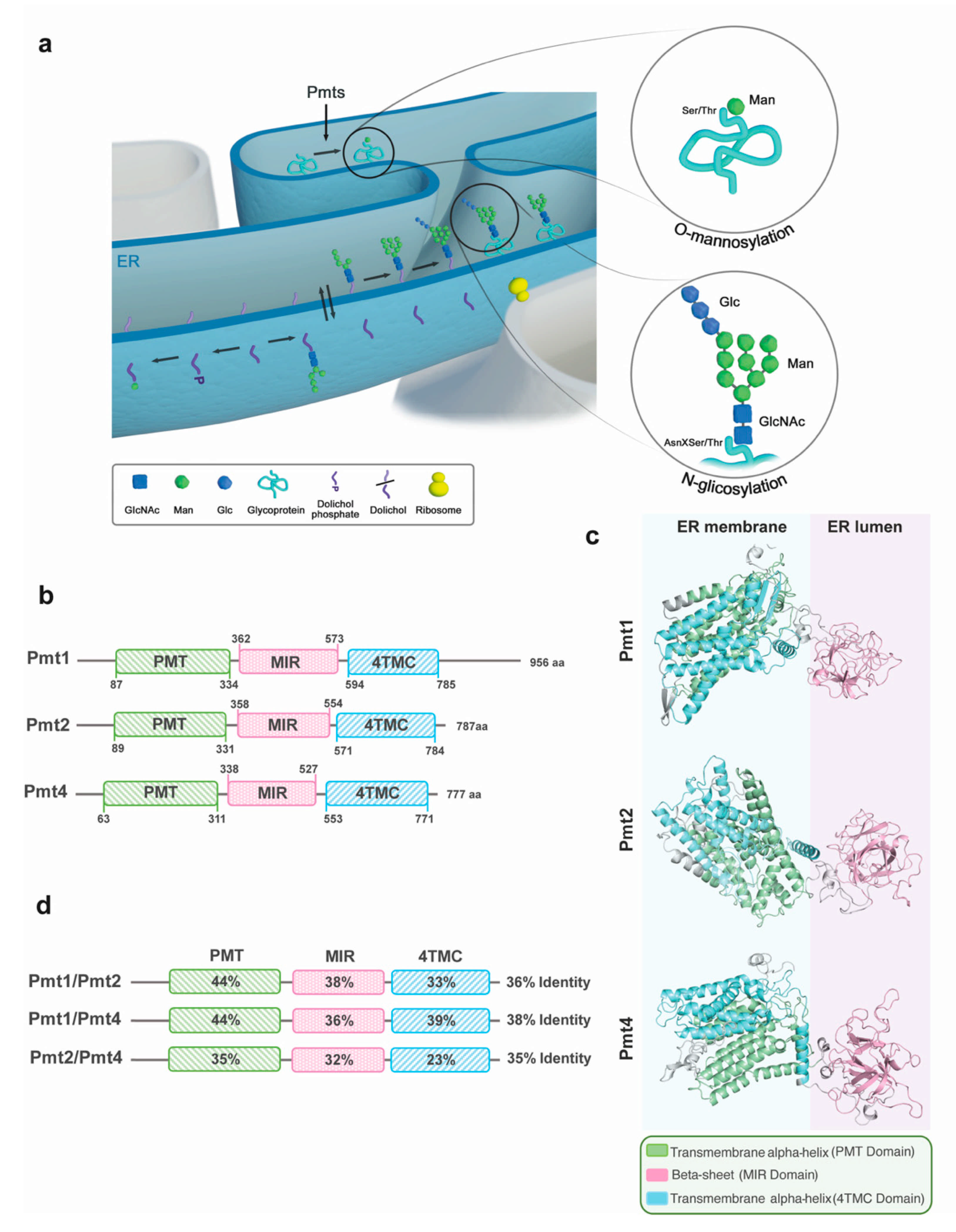
2.1. N-Glycosylation
2.2. O-Mannosylation and the Pmt Family
3. The Structure of Pmts
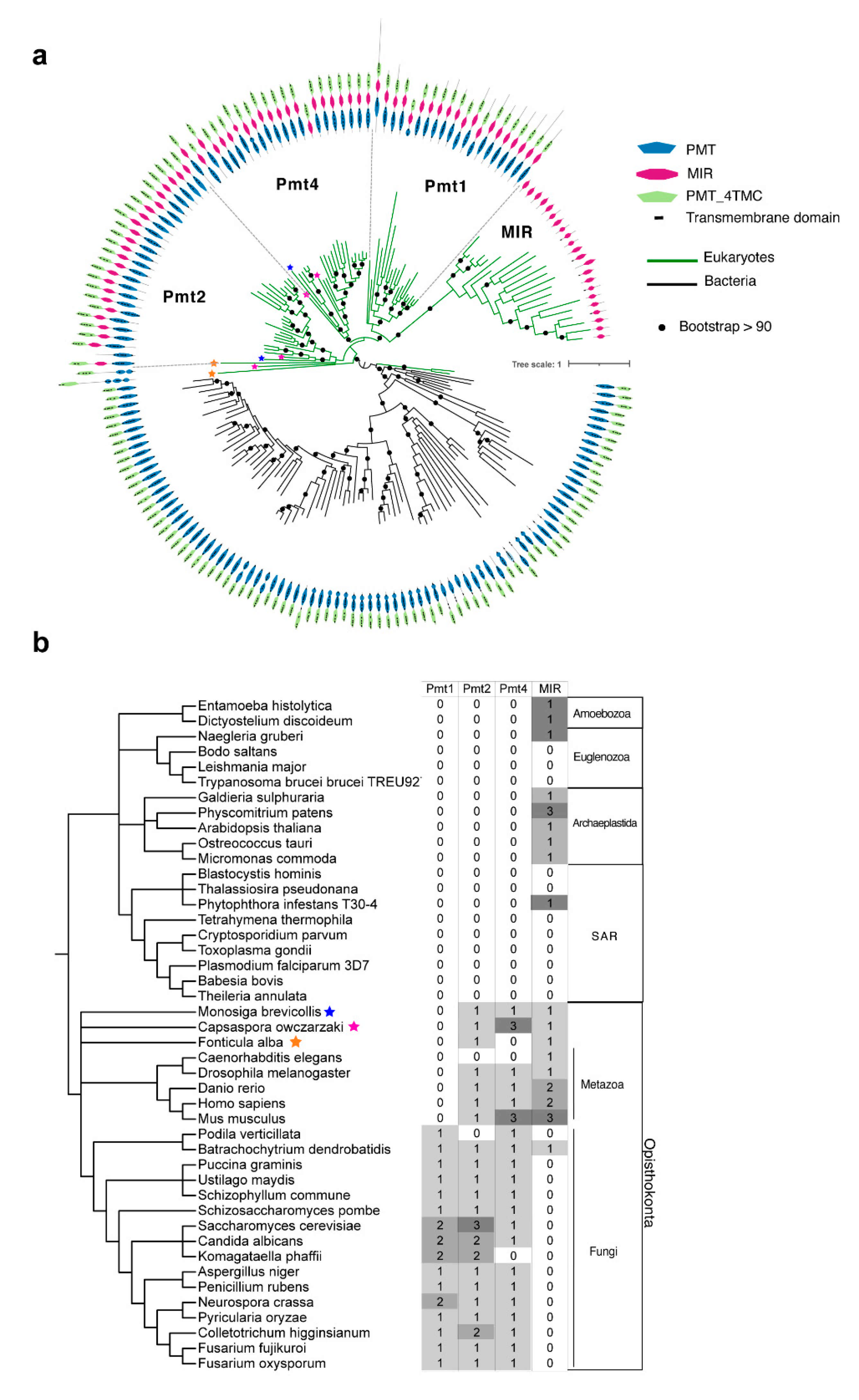
4. Evolution of the Pmt Protein Family
4.1. Bacterial and Eukaryotic Contributions to the Origin and Diversification of Pmts in Opisthokonts
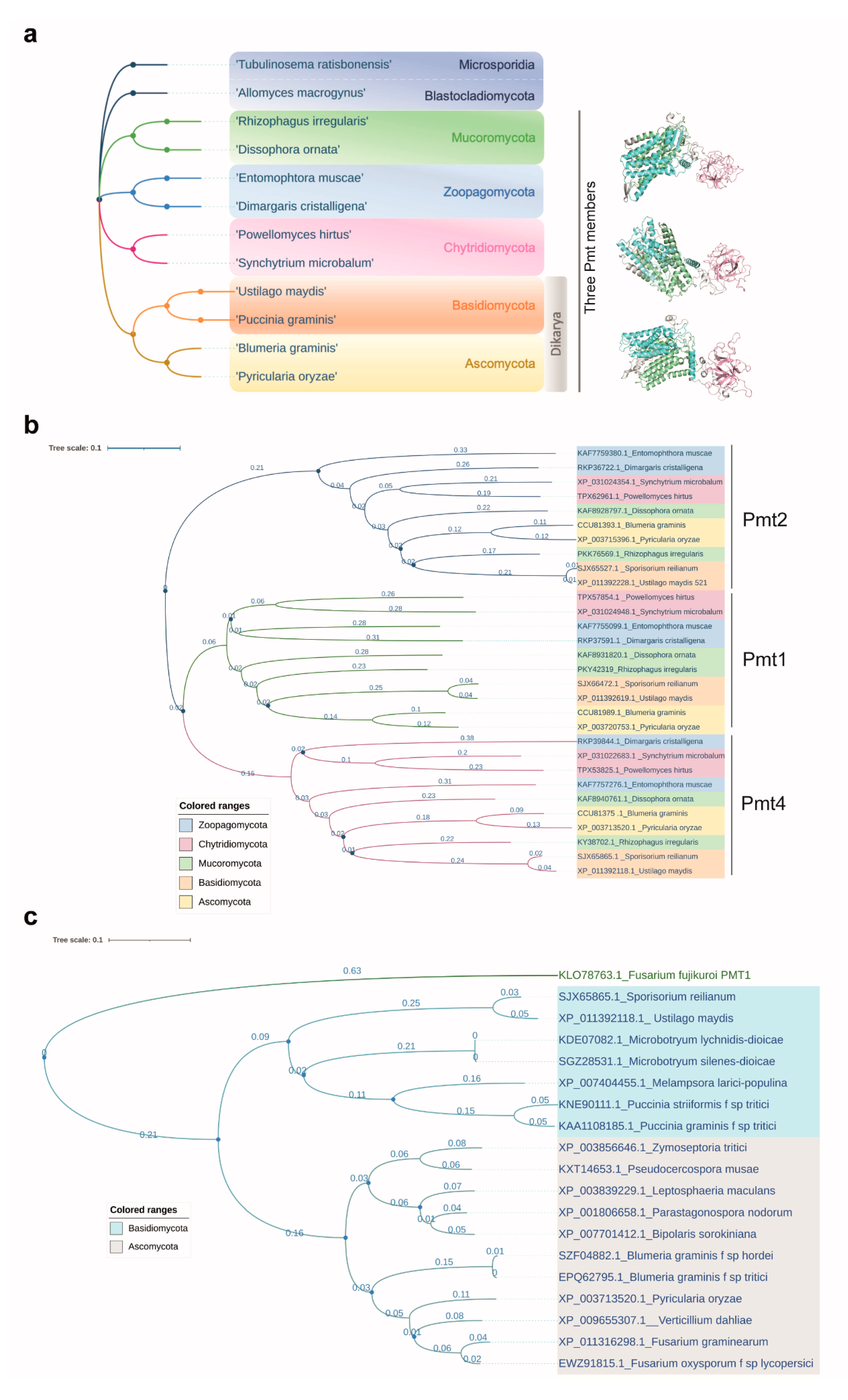
4.2. The Origin of Fungi-Specific Pmt1 Might Be Pmt4
5. Variability in the Number of Pmt Family Members
5.1. The Pmt Family in Animals Consists of Two Members
5.2. The Addition of a Third Pmt in Fungi
6. The Role of Pmts in Fungal Pathogenesis: Pmt4 as the Key Protein Factor in Virulence
6.1. Pmt1, the Most Dispensable of the Pmts
6.2. Pmt2 and Its Essential Role for Cell Viability
6.3. Pmt4, the Key Pmt Protein in Virulence
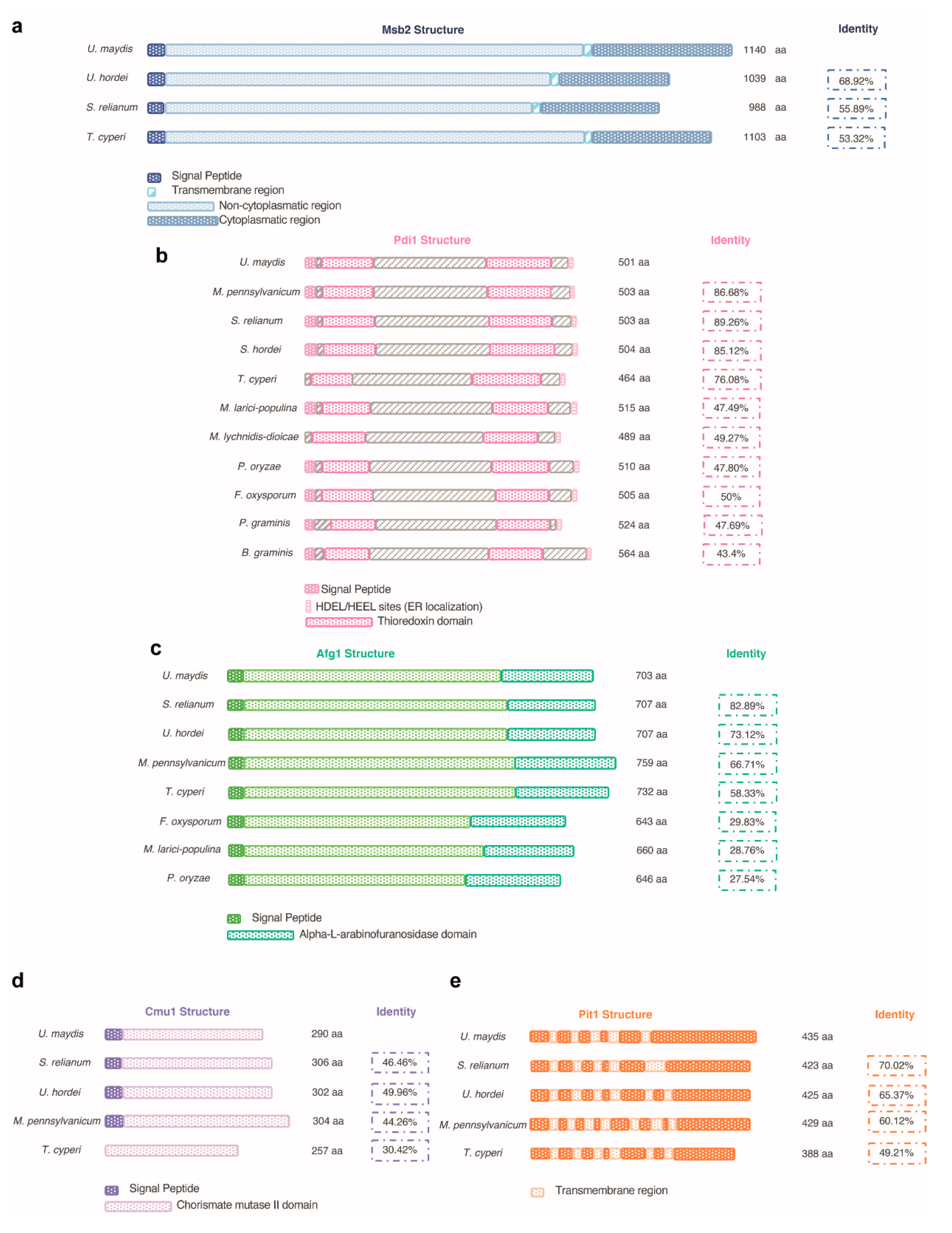
7. What Pmt4 Substrates Might Justify the Virulence Defects Observed in pmt4 Mutants in U. maydis?
8. Conclusions
9. Methods
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Varki, A.; Kornfeld, S. Historical background and overview. In Essentials of Glycobiology; Varki, A., Cummings, R.D., Esko, J.D., Stanley, P., Hart, G.W., Aebi, M., Darvill, A.G., Kinoshita, T., Packer, N.H., Eds.; Cold Spring Harbor: New York, NY, USA, 2015; pp. 1–18. [Google Scholar] [CrossRef]
- Xu, C.; Ng, D.T. Glycosylation-directed quality control of protein folding. Nat. Rev. Mol. Cell Biol. 2015, 16, 742–752. [Google Scholar] [CrossRef] [PubMed]
- Varki, A. Biological roles of glycans. Glycobiology 2017, 27, 3–49. [Google Scholar] [CrossRef] [PubMed]
- Reily, C.; Stewart, T.J.; Renfrow, M.B.; Novak, J. Glycosylation in health and disease. Nat. Rev. Nephrol. 2019, 15, 346–366. [Google Scholar] [CrossRef] [PubMed]
- Schjoldager, K.T.; Narimatsu, Y.; Joshi, H.J.; Clausen, H. Global view of human protein glycosylation pathways and functions. Nat. Rev. Mol. Cell Biol. 2020, 21, 729–749. [Google Scholar] [CrossRef]
- Hansen, L.; Lind-Thomsen, A.; Joshi, H.J.; Pedersen, N.B.; Have, C.T.; Kong, Y.; Wang, S.; Sparso, T.; Grarup, N.; Vester-Christensen, M.B.; et al. A glycogene mutation map for discovery of diseases of glycosylation. Glycobiology 2015, 25, 211–224. [Google Scholar] [CrossRef] [PubMed]
- Freeze, H.H.; Eklund, E.A.; Ng, B.G.; Patterson, M.C. Neurological aspects of human glycosylation disorders. Annu. Rev. Neurosci. 2015, 38, 105–125. [Google Scholar] [CrossRef]
- Ng, B.G.; Freeze, H.H. Perspectives on Glycosylation and Its Congenital Disorders. Trends Genet. 2018, 34, 466–476. [Google Scholar] [CrossRef]
- Monticelli, M.; Ferro, T.; Jaeken, J.; Dos Reis Ferreira, V.; Videira, P.A. Immunological aspects of congenital disorders of glycosylation (CDG): A review. J. Inherit. Metab. Dis. 2016, 39, 765–780. [Google Scholar] [CrossRef]
- Pinho, S.S.; Reis, C.A. Glycosylation in cancer: Mechanisms and clinical implications. Nat. Rev. Cancer 2015, 15, 540–555. [Google Scholar] [CrossRef]
- Corfield, A.P.; Berry, M. Glycan variation and evolution in the eukaryotes. Trends Biochem. Sci. 2015, 40, 351–359. [Google Scholar] [CrossRef]
- Calo, D.; Kaminski, L.; Eichler, J. Protein glycosylation in Archaea: Sweet and extreme. Glycobiology 2010, 20, 1065–1076. [Google Scholar] [CrossRef] [PubMed]
- Nothaft, H.; Szymanski, C.M. Protein glycosylation in bacteria: Sweeter than ever. Nat. Rev. Microbiol. 2010, 8, 765–778. [Google Scholar] [CrossRef] [PubMed]
- Fernandez-Alvarez, A.; Marin-Menguiano, M.; Lanver, D.; Jimenez-Martin, A.; Elias-Villalobos, A.; Perez-Pulido, A.J.; Kahmann, R.; Ibeas, J.I. Identification of O-mannosylated virulence factors in Ustilago maydis. PLoS Pathog. 2012, 8, e1002563. [Google Scholar] [CrossRef] [PubMed]
- Schirawski, J.; Bohnert, H.U.; Steinberg, G.; Snetselaar, K.; Adamikowa, L.; Kahmann, R. Endoplasmic reticulum glucosidase II is required for pathogenicity of Ustilago maydis. Plant Cell 2005, 17, 3532–3543. [Google Scholar] [CrossRef]
- Fernandez-Alvarez, A.; Elias-Villalobos, A.; Ibeas, J.I. The O-mannosyltransferase PMT4 is essential for normal appressorium formation and penetration in Ustilago maydis. Plant Cell 2009, 21, 3397–3412. [Google Scholar] [CrossRef]
- Fernandez-Alvarez, A.; Elias-Villalobos, A.; Jimenez-Martin, A.; Marin-Menguiano, M.; Ibeas, J.I. Endoplasmic reticulum glucosidases and protein quality control factors cooperate to establish biotrophy in Ustilago maydis. Plant Cell 2013, 25, 4676–4690. [Google Scholar] [CrossRef]
- Loibl, M.; Strahl, S. Protein O-mannosylation: What we have learned from baker’s yeast. Biochim. Biophys. Acta 2013, 1833, 2438–2446. [Google Scholar] [CrossRef]
- Aebi, M. N-linked protein glycosylation in the ER. Biochim. Biophys. Acta 2013, 1833, 2430–2437. [Google Scholar] [CrossRef]
- Lehle, L.; Strahl, S.; Tanner, W. Protein glycosylation, conserved from yeast to man: A model organism helps elucidate congenital human diseases. Angew. Chem. Int. Ed. Engl. 2006, 45, 6802–6818. [Google Scholar] [CrossRef]
- Spiro, R.G. Protein glycosylation: Nature, distribution, enzymatic formation, and disease implications of glycopeptide bonds. Glycobiology 2002, 12, 43R–56R. [Google Scholar] [CrossRef]
- Schwarz, F.; Aebi, M. Mechanisms and principles of N-linked protein glycosylation. Curr. Opin. Struct. Biol. 2011, 21, 576–582. [Google Scholar] [CrossRef]
- Breitling, J.; Aebi, M. N-linked protein glycosylation in the endoplasmic reticulum. Cold Spring Harb Perspect. Biol. 2013, 5, a013359. [Google Scholar] [CrossRef]
- Kaplan, H.A.; Welply, J.K.; Lennarz, W.J. Oligosaccharyl transferase: The central enzyme in the pathway of glycoprotein assembly. Biochim. Biophys. Acta 1987, 906, 161–173. [Google Scholar] [CrossRef]
- Burda, P.; Aebi, M. The ALG10 locus of Saccharomyces cerevisiae encodes the alpha-1,2 glucosyltransferase of the endoplasmic reticulum: The terminal glucose of the lipid-linked oligosaccharide is required for efficient N-linked glycosylation. Glycobiology 1998, 8, 455–462. [Google Scholar] [CrossRef]
- Burda, P.; Aebi, M. The dolichol pathway of N-linked glycosylation. Biochim. Biophys. Acta 1999, 1426, 239–257. [Google Scholar] [CrossRef]
- Burda, P.; te Heesen, S.; Brachat, A.; Wach, A.; Dusterhoft, A.; Aebi, M. Stepwise assembly of the lipid-linked oligosaccharide in the endoplasmic reticulum of Saccharomyces cerevisiae: Identification of the ALG9 gene encoding a putative mannosyl transferase. Proc. Natl. Acad. Sci. USA 1996, 93, 7160–7165. [Google Scholar] [CrossRef]
- Knauer, R.; Lehle, L. The oligosaccharyltransferase complex from yeast. Biochim. Biophys. Acta 1999, 1426, 259–273. [Google Scholar] [CrossRef]
- Jones, J.; Krag, S.S.; Betenbaugh, M.J. Controlling N-linked glycan site occupancy. Biochim. Biophys. Acta 2005, 1726, 121–137. [Google Scholar] [CrossRef]
- Helenius, J.; Aebi, M. Transmembrane movement of dolichol linked carbohydrates during N-glycoprotein biosynthesis in the endoplasmic reticulum. Semin. Cell Dev. Biol. 2002, 13, 171–178. [Google Scholar] [CrossRef]
- Miwa, H.E.; Gerken, T.A.; Jamison, O.; Tabak, L.A. Isoform-specific O-glycosylation of osteopontin and bone sialoprotein by polypeptide N-acetylgalactosaminyltransferase-1. J. Biol. Chem. 2010, 285, 1208–1219. [Google Scholar] [CrossRef]
- Koomey, M. O-linked protein glycosylation in bacteria: Snapshots and current perspectives. Curr. Opin. Struct. Biol. 2019, 56, 198–203. [Google Scholar] [CrossRef]
- West, C.M.; Kim, H.W. Nucleocytoplasmic O-glycosylation in protists. Curr. Opin. Struct. Biol. 2019, 56, 204–212. [Google Scholar] [CrossRef]
- Lommel, M.; Strahl, S. Protein O-mannosylation: Conserved from bacteria to humans. Glycobiology 2009, 19, 816–828. [Google Scholar] [CrossRef]
- Goto, M. Protein O-glycosylation in fungi: Diverse structures and multiple functions. Biosci. Biotechnol. Biochem. 2007, 71, 1415–1427. [Google Scholar] [CrossRef]
- Neubert, P.; Strahl, S. Protein O-mannosylation in the early secretory pathway. Curr. Opin. Cell Biol. 2016, 41, 100–108. [Google Scholar] [CrossRef]
- Marin-Menguiano, M.; Moreno-Sanchez, I.; Barrales, R.R.; Fernandez-Alvarez, A.; Ibeas, J.I. N-glycosylation of the protein disulfide isomerase Pdi1 ensures full Ustilago maydis virulence. PLoS Pathog. 2019, 15, e1007687. [Google Scholar] [CrossRef]
- Pan, Y.; Pan, R.; Tan, L.; Zhang, Z.; Guo, M. Pleiotropic roles of O-mannosyltransferase MoPmt4 in development and pathogenicity of Magnaporthe oryzae. Curr. Genet. 2019, 65, 223–239. [Google Scholar] [CrossRef]
- Zhao, G.; Xu, Y.; Ouyang, H.; Luo, Y.; Sun, S.; Wang, Z.; Yang, J.; Jin, C. Protein O-mannosylation affects protein secretion, cell wall integrity and morphogenesis in Trichoderma reesei. Fungal Genet. Biol. 2020, 144, 103440. [Google Scholar] [CrossRef]
- Zhao, T.; Tian, H.; Xia, Y.; Jin, K. MaPmt4, a protein O-mannosyltransferase, contributes to cell wall integrity, stress tolerance and virulence in Metarhizium acridum. Curr. Genet. 2019, 65, 1025–1040. [Google Scholar] [CrossRef]
- Xu, Y.; Zhou, H.; Zhao, G.; Yang, J.; Luo, Y.; Sun, S.; Wang, Z.; Li, S.; Jin, C. Genetical and O-glycoproteomic analyses reveal the roles of three protein O-mannosyltransferases in phytopathogen Fusarium oxysporum f.sp. cucumerinum. Fungal Genet. Biol. 2020, 134, 103285. [Google Scholar] [CrossRef]
- VanderVen, B.C.; Harder, J.D.; Crick, D.C.; Belisle, J.T. Export-mediated assembly of mycobacterial glycoproteins parallels eukaryotic pathways. Science 2005, 309, 941–943. [Google Scholar] [CrossRef] [PubMed]
- Matei, A.; Doehlemann, G. Cell biology of corn smut disease-Ustilago maydis as a model for biotrophic interactions. Curr. Opin. Microbiol. 2016, 34, 60–66. [Google Scholar] [CrossRef] [PubMed]
- Mendoza-Mendoza, A.; Berndt, P.; Djamei, A.; Weise, C.; Linne, U.; Marahiel, M.; Vranes, M.; Kamper, J.; Kahmann, R. Physical-chemical plant-derived signals induce differentiation in Ustilago maydis. Mol. Microbiol. 2009, 71, 895–911. [Google Scholar] [CrossRef] [PubMed]
- Lanver, D.; Tollot, M.; Schweizer, G.; Lo Presti, L.; Reissmann, S.; Ma, L.S.; Schuster, M.; Tanaka, S.; Liang, L.; Ludwig, N.; et al. Ustilago maydis effectors and their impact on virulence. Nat. Rev. Microbiol. 2017, 15, 409–421. [Google Scholar] [CrossRef]
- Ponting, C.P. Novel repeats in ryanodine and IP3 receptors and protein O-mannosyltransferases. Trends Biochem. Sci. 2000, 25, 48–50. [Google Scholar] [CrossRef]
- Albuquerque-Wendt, A.; Hutte, H.J.; Buettner, F.F.R.; Routier, F.H.; Bakker, H. Membrane Topological Model of Glycosyltransferases of the GT-C Superfamily. Int. J. Mol. Sci. 2019, 20, 4842. [Google Scholar] [CrossRef]
- Bai, L.; Kovach, A.; You, Q.; Kenny, A.; Li, H. Structure of the eukaryotic protein O-mannosyltransferase Pmt1-Pmt2 complex. Nat. Struct. Mol. Biol. 2019, 26, 704–711. [Google Scholar] [CrossRef]
- Chiapparino, A.; Grbavac, A.; Jonker, H.R.; Hackmann, Y.; Mortensen, S.; Zatorska, E.; Schott, A.; Stier, G.; Saxena, K.; Wild, K.; et al. Functional implications of MIR domains in protein O-mannosylation. Elife 2020, 9. [Google Scholar] [CrossRef]
- Girrbach, V.; Strahl, S. Members of the evolutionarily conserved PMT family of protein O-mannosyltransferases form distinct protein complexes among themselves. J. Biol. Chem. 2003, 278, 12554–12562. [Google Scholar] [CrossRef]
- Akasaka-Manya, K.; Manya, H.; Nakajima, A.; Kawakita, M.; Endo, T. Physical and functional association of human protein O-mannosyltransferases 1 and 2. J. Biol. Chem. 2006, 281, 19339–19345. [Google Scholar] [CrossRef]
- Kriangkripipat, T.; Momany, M. Aspergillus nidulans Pmts form heterodimers in all pairwise combinations. FEBS Open Bio 2014, 4, 335–341. [Google Scholar] [CrossRef]
- Mouyna, I.; Kniemeyer, O.; Jank, T.; Loussert, C.; Mellado, E.; Aimanianda, V.; Beauvais, A.; Wartenberg, D.; Sarfati, J.; Bayry, J.; et al. Members of protein O-mannosyltransferase family in Aspergillus fumigatus differentially affect growth, morphogenesis and viability. Mol. Microbiol. 2010, 76, 1205–1221. [Google Scholar] [CrossRef]
- Kriangkripipat, T.; Momany, M. Aspergillus nidulans protein O-mannosyltransferases play roles in cell wall integrity and developmental patterning. Eukaryot. Cell 2009, 8, 1475–1485. [Google Scholar] [CrossRef]
- Olson, G.M.; Fox, D.S.; Wang, P.; Alspaugh, J.A.; Buchanan, K.L. Role of protein O-mannosyltransferase Pmt4 in the morphogenesis and virulence of Cryptococcus neoformans. Eukaryot. Cell 2007, 6, 222–234. [Google Scholar] [CrossRef]
- Guo, M.; Tan, L.; Nie, X.; Zhu, X.; Pan, Y.; Gao, Z. The Pmt2p-Mediated Protein O-Mannosylation Is Required for Morphogenesis, Adhesive Properties, Cell Wall Integrity and Full Virulence of Magnaporthe oryzae. Front. Microbiol. 2016, 7, 630. [Google Scholar] [CrossRef]
- Wang, J.J.; Qiu, L.; Chu, Z.J.; Ying, S.H.; Feng, M.G. The connection of protein O-mannosyltransferase family to the biocontrol potential of Beauveria bassiana, a fungal entomopathogen. Glycobiology 2014, 24, 638–648. [Google Scholar] [CrossRef][Green Version]
- He, Z.; Luo, L.; Keyhani, N.O.; Yu, X.; Ying, S.; Zhang, Y. The C-terminal MIR-containing region in the Pmt1 O-mannosyltransferase restrains sporulation and is dispensable for virulence in Beauveria bassiana. Appl. Microbiol. Biotechnol. 2017, 101, 1143–1161. [Google Scholar] [CrossRef]
- Gonzalez, M.; Brito, N.; Frias, M.; Gonzalez, C. Botrytis cinerea protein O-mannosyltransferases play critical roles in morphogenesis, growth, and virulence. PLoS ONE 2013, 8, e65924. [Google Scholar] [CrossRef]
- Prill, S.K.; Klinkert, B.; Timpel, C.; Gale, C.A.; Schroppel, K.; Ernst, J.F. PMT family of Candida albicans: Five protein mannosyltransferase isoforms affect growth, morphogenesis and antifungal resistance. Mol. Microbiol. 2005, 55, 546–560. [Google Scholar] [CrossRef] [PubMed]
- Harries, E.; Gandia, M.; Carmona, L.; Marcos, J.F. The Penicillium digitatum protein O-mannosyltransferase Pmt2 is required for cell wall integrity, conidiogenesis, virulence and sensitivity to the antifungal peptide PAF26. Mol. Plant. Pathol. 2015, 16, 748–761. [Google Scholar] [CrossRef]
- Mahne, M.; Tauch, A.; Puhler, A.; Kalinowski, J. The Corynebacterium glutamicum gene pmt encoding a glycosyltransferase related to eukaryotic protein-O-mannosyltransferases is essential for glycosylation of the resuscitation promoting factor (Rpf2) and other secreted proteins. FEMS Microbiol. Lett. 2006, 259, 226–233. [Google Scholar] [CrossRef] [PubMed]
- Liu, C.F.; Tonini, L.; Malaga, W.; Beau, M.; Stella, A.; Bouyssie, D.; Jackson, M.C.; Nigou, J.; Puzo, G.; Guilhot, C.; et al. Bacterial protein-O-mannosylating enzyme is crucial for virulence of Mycobacterium tuberculosis. Proc. Natl. Acad. Sci. USA 2013, 110, 6560–6565. [Google Scholar] [CrossRef] [PubMed]
- Wehmeier, S.; Varghese, A.S.; Gurcha, S.S.; Tissot, B.; Panico, M.; Hitchen, P.; Morris, H.R.; Besra, G.S.; Dell, A.; Smith, M.C. Glycosylation of the phosphate binding protein, PstS, in Streptomyces coelicolor by a pathway that resembles protein O-mannosylation in eukaryotes. Mol. Microbiol. 2009, 71, 421–433. [Google Scholar] [CrossRef] [PubMed]
- Haines, N.; Seabrooke, S.; Stewart, B.A. Dystroglycan and protein O-mannosyltransferases 1 and 2 are required to maintain integrity of Drosophila larval muscles. Mol. Biol. Cell 2007, 18, 4721–4730. [Google Scholar] [CrossRef] [PubMed]
- Martin-Blanco, E.; Garcia-Bellido, A. Mutations in the rotated abdomen locus affect muscle development and reveal an intrinsic asymmetry in Drosophila. Proc. Natl. Acad. Sci. USA 1996, 93, 6048–6052. [Google Scholar] [CrossRef]
- Baker, R.; Nakamura, N.; Chandel, I.; Howell, B.; Lyalin, D.; Panin, V.M. Protein O-Mannosyltransferases Affect Sensory Axon Wiring and Dynamic Chirality of Body Posture in the Drosophila Embryo. J. Neurosci. 2018, 38, 1850–1865. [Google Scholar] [CrossRef]
- Willer, T.; Prados, B.; Falcon-Perez, J.M.; Renner-Muller, I.; Przemeck, G.K.; Lommel, M.; Coloma, A.; Valero, M.C.; de Angelis, M.H.; Tanner, W.; et al. Targeted disruption of the Walker-Warburg syndrome gene Pomt1 in mouse results in embryonic lethality. Proc. Natl. Acad. Sci. USA 2004, 101, 14126–14131. [Google Scholar] [CrossRef]
- Beltran-Valero de Bernabe, D.; Currier, S.; Steinbrecher, A.; Celli, J.; van Beusekom, E.; van der Zwaag, B.; Kayserili, H.; Merlini, L.; Chitayat, D.; Dobyns, W.B.; et al. Mutations in the O-mannosyltransferase gene POMT1 give rise to the severe neuronal migration disorder Walker-Warburg syndrome. Am. J. Hum. Genet. 2002, 71, 1033–1043. [Google Scholar] [CrossRef]
- Akasaka-Manya, K.; Manya, H.; Endo, T. Mutations of the POMT1 gene found in patients with Walker-Warburg syndrome lead to a defect of protein O-mannosylation. Biochem. Biophys. Res. Commun. 2004, 325, 75–79. [Google Scholar] [CrossRef]
- Gentzsch, M.; Tanner, W. The PMT gene family: Protein O-glycosylation in Saccharomyces cerevisiae is vital. EMBO J. 1996, 15, 5752–5759. [Google Scholar] [CrossRef]
- Strahl-Bolsinger, S.; Scheinost, A. Transmembrane topology of pmt1p, a member of an evolutionarily conserved family of protein O-mannosyltransferases. J. Biol. Chem. 1999, 274, 9068–9075. [Google Scholar] [CrossRef]
- Nett, J.H.; Cook, W.J.; Chen, M.T.; Davidson, R.C.; Bobrowicz, P.; Kett, W.; Brevnova, E.; Potgieter, T.I.; Mellon, M.T.; Prinz, B.; et al. Characterization of the Pichia pastoris protein-O-mannosyltransferase gene family. PLoS ONE 2013, 8, e68325. [Google Scholar] [CrossRef]
- Willer, T.; Brandl, M.; Sipiczki, M.; Strahl, S. Protein O-mannosylation is crucial for cell wall integrity, septation and viability in fission yeast. Mol. Microbiol. 2005, 57, 156–170. [Google Scholar] [CrossRef]
- Willger, S.D.; Ernst, J.F.; Alspaugh, J.A.; Lengeler, K.B. Characterization of the PMT gene family in Cryptococcus neoformans. PLoS ONE 2009, 4, e6321. [Google Scholar] [CrossRef]
- Liu, C.; Talbot, N.J.; Chen, X.L. Protein glycosylation during infection by plant pathogenic fungi. New Phytol. 2021. [Google Scholar] [CrossRef]
- Fernandez-Alvarez, A.; Elias-Villalobos, A.; Ibeas, J.I. The requirement for protein O-mannosylation for Ustilago maydis virulence seems to be linked to intrinsic aspects of the infection process rather than an altered plant response. Plant. Signal. Behav. 2010, 5, 412–414. [Google Scholar] [CrossRef]
- Hutzler, J.; Schmid, M.; Bernard, T.; Henrissat, B.; Strahl, S. Membrane association is a determinant for substrate recognition by PMT4 protein O-mannosyltransferases. Proc. Natl. Acad. Sci. USA 2007, 104, 7827–7832. [Google Scholar] [CrossRef]
- Lanver, D.; Mendoza-Mendoza, A.; Brachmann, A.; Kahmann, R. Sho1 and Msb2-related proteins regulate appressorium development in the smut fungus Ustilago maydis. Plant. Cell 2010, 22, 2085–2101. [Google Scholar] [CrossRef]
- Liu, W.; Zhou, X.; Li, G.; Li, L.; Kong, L.; Wang, C.; Zhang, H.; Xu, J.R. Multiple plant surface signals are sensed by different mechanisms in the rice blast fungus for appressorium formation. PLoS Pathog. 2011, 7, e1001261. [Google Scholar] [CrossRef]
- Lanver, D.; Berndt, P.; Tollot, M.; Naik, V.; Vranes, M.; Warmann, T.; Munch, K.; Rossel, N.; Kahmann, R. Plant surface cues prime Ustilago maydis for biotrophic development. PLoS Pathog. 2014, 10, e1004272. [Google Scholar] [CrossRef]
- Doehlemann, G.; Reissmann, S.; Assmann, D.; Fleckenstein, M.; Kahmann, R. Two linked genes encoding a secreted effector and a membrane protein are essential for Ustilago maydis-induced tumour formation. Mol. Microbiol. 2011, 81, 751–766. [Google Scholar] [CrossRef]
- Djamei, A.; Schipper, K.; Rabe, F.; Ghosh, A.; Vincon, V.; Kahnt, J.; Osorio, S.; Tohge, T.; Fernie, A.R.; Feussner, I.; et al. Metabolic priming by a secreted fungal effector. Nature 2011, 478, 395–398. [Google Scholar] [CrossRef] [PubMed]
- Potter, S.C.; Luciani, A.; Eddy, S.R.; Park, Y.; Lopez, R.; Finn, R.D. HMMER web server: 2018 update. Nucleic Acids Res. 2018, 46, W200–W204. [Google Scholar] [CrossRef] [PubMed]
- Fu, L.; Niu, B.; Zhu, Z.; Wu, S.; Li, W. CD-HIT: Accelerated for clustering the next-generation sequencing data. Bioinformatics 2012, 28, 3150–3152. [Google Scholar] [CrossRef] [PubMed]
- Katoh, K.; Standley, D.M. MAFFT multiple sequence alignment software version 7: Improvements in performance and usability. Mol. Biol. Evol. 2013, 30, 772–780. [Google Scholar] [CrossRef] [PubMed]
- Capella-Gutierrez, S.; Silla-Martinez, J.M.; Gabaldon, T. trimAl: A tool for automated alignment trimming in large-scale phylogenetic analyses. Bioinformatics 2009, 25, 1972–1973. [Google Scholar] [CrossRef]
- Nguyen, L.T.; Schmidt, H.A.; von Haeseler, A.; Minh, B.Q. IQ-TREE: A fast and effective stochastic algorithm for estimating maximum-likelihood phylogenies. Mol. Biol. Evol. 2015, 32, 268–274. [Google Scholar] [CrossRef]
- Hoang, D.T.; Chernomor, O.; von Haeseler, A.; Minh, B.Q.; Vinh, L.S. UFBoot2: Improving the Ultrafast Bootstrap Approximation. Mol. Biol. Evol. 2018, 35, 518–522. [Google Scholar] [CrossRef]
- Kalyaanamoorthy, S.; Minh, B.Q.; Wong, T.K.F.; von Haeseler, A.; Jermiin, L.S. ModelFinder: Fast model selection for accurate phylogenetic estimates. Nat. Methods 2017, 14, 587–589. [Google Scholar] [CrossRef]
- Letunic, I.; Bork, P. Interactive Tree Of Life (iTOL) v4: Recent updates and new developments. Nucleic Acids Res. 2019, 47, W256–W259. [Google Scholar] [CrossRef]
- Finn, R.D.; Bateman, A.; Clements, J.; Coggill, P.; Eberhardt, R.Y.; Eddy, S.R.; Heger, A.; Hetherington, K.; Holm, L.; Mistry, J.; et al. Pfam: The protein families database. Nucleic Acids Res. 2014, 42, D222–D230. [Google Scholar] [CrossRef] [PubMed]

| Fungus | Viability | Cellular and Hyphal Growth | Virulence | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| ∆pmt1 | ∆pmt2 | ∆pmt4 | ∆pmt1/4 | ∆pmt1 | ∆pmt2 | ∆pmt4 | ∆pmt1/4 | ∆pmt1 | ∆pmt2 | ∆pmt4 | ∆pmt1/4 | |
| Ustilago maydis [16] | V | L | V | V | NA | L | A | A | NA | L | A | A |
| Aspergillus fumigatus [53] | V | L | V | L | NA | L | A | L | NA | L | A | L |
| Aspergillus nidulans [54] | V | V | V | NT | NA | NA | A | NT | NA | NA | NA | NT |
| Cryptococcus neoformans [55] | V | L | V | L | NA | L | A | L | A | L | A | L |
| Magnaporthe oryzae [38,56] | V | V | V | NT | NA | NA | A | NT | NA | A | A | NT |
| Beauveria bassiana [57,58] | V | L | V | NT | NA | L | NA | NT | A | L | A | NT |
| Botrytis cinerea [59] | V | V | V | NT | NA | A | A | NT | NA | A | A | NT |
| Candida albicans [60] | V | L | V | L | A | L | A | NA | NA | L | A | NA |
| Fusarium oxysporum [41] | V | L | V | V | NA | L | A | NA | NA | L | A | NA |
| Penicillium digitatum [61] | NT | V | NT | NT | NT | A | NT | NT | NT | A | NT | NT |
| Metarhizium acridum [40] | NT | NT | V | NT | NT | NT | A | NT | NT | NT | A | NT |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Pejenaute-Ochoa, M.D.; Santana-Molina, C.; Devos, D.P.; Ibeas, J.I.; Fernández-Álvarez, A. Structural, Evolutionary, and Functional Analysis of the Protein O-Mannosyltransferase Family in Pathogenic Fungi. J. Fungi 2021, 7, 328. https://doi.org/10.3390/jof7050328
Pejenaute-Ochoa MD, Santana-Molina C, Devos DP, Ibeas JI, Fernández-Álvarez A. Structural, Evolutionary, and Functional Analysis of the Protein O-Mannosyltransferase Family in Pathogenic Fungi. Journal of Fungi. 2021; 7(5):328. https://doi.org/10.3390/jof7050328
Chicago/Turabian StylePejenaute-Ochoa, María Dolores, Carlos Santana-Molina, Damien P. Devos, José Ignacio Ibeas, and Alfonso Fernández-Álvarez. 2021. "Structural, Evolutionary, and Functional Analysis of the Protein O-Mannosyltransferase Family in Pathogenic Fungi" Journal of Fungi 7, no. 5: 328. https://doi.org/10.3390/jof7050328
APA StylePejenaute-Ochoa, M. D., Santana-Molina, C., Devos, D. P., Ibeas, J. I., & Fernández-Álvarez, A. (2021). Structural, Evolutionary, and Functional Analysis of the Protein O-Mannosyltransferase Family in Pathogenic Fungi. Journal of Fungi, 7(5), 328. https://doi.org/10.3390/jof7050328






