Improvements in the Resistance of the Banana Species to Fusarium Wilt: A Systematic Review of Methods and Perspectives
Abstract
1. Introduction
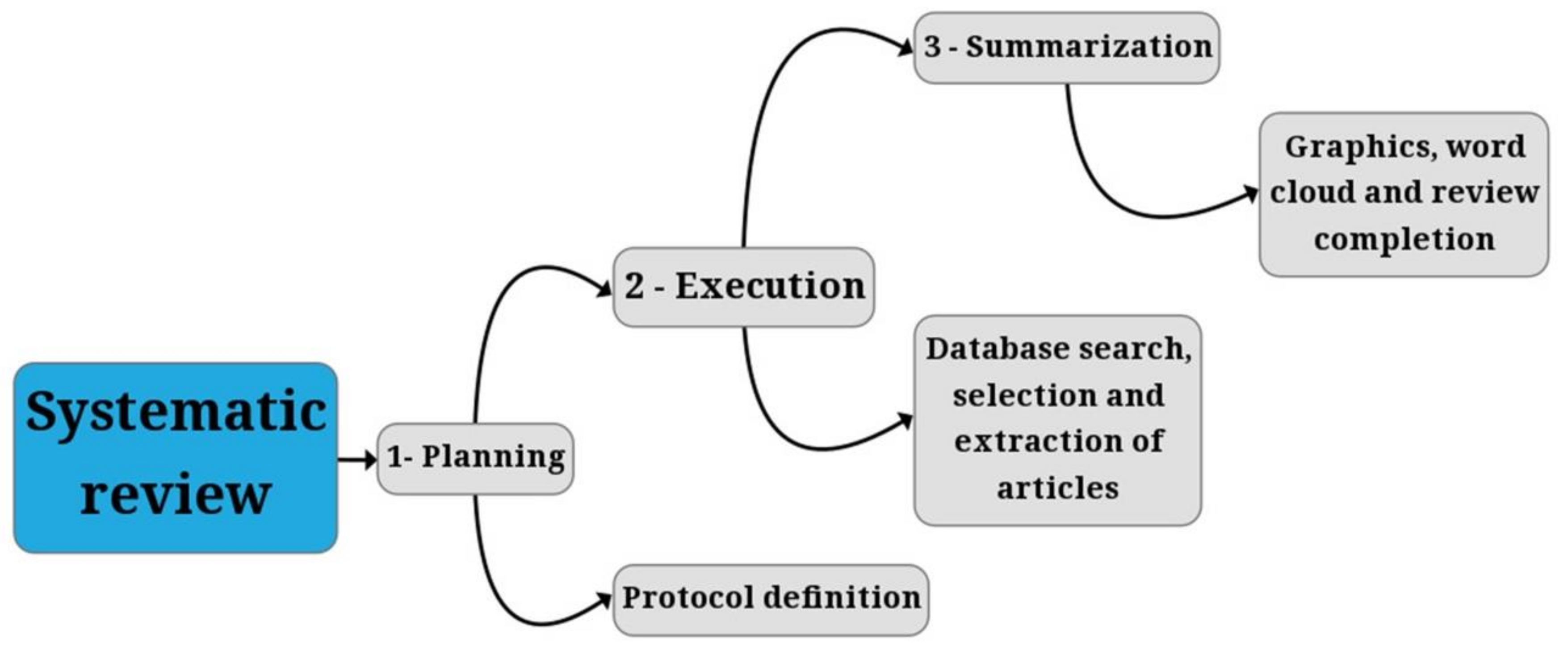
2. Materials and Methods
2.1. Planning
2.2. Execution: Search
2.3. Execution: Selection and Extraction
2.4. Analysis of the Articles
3. Results
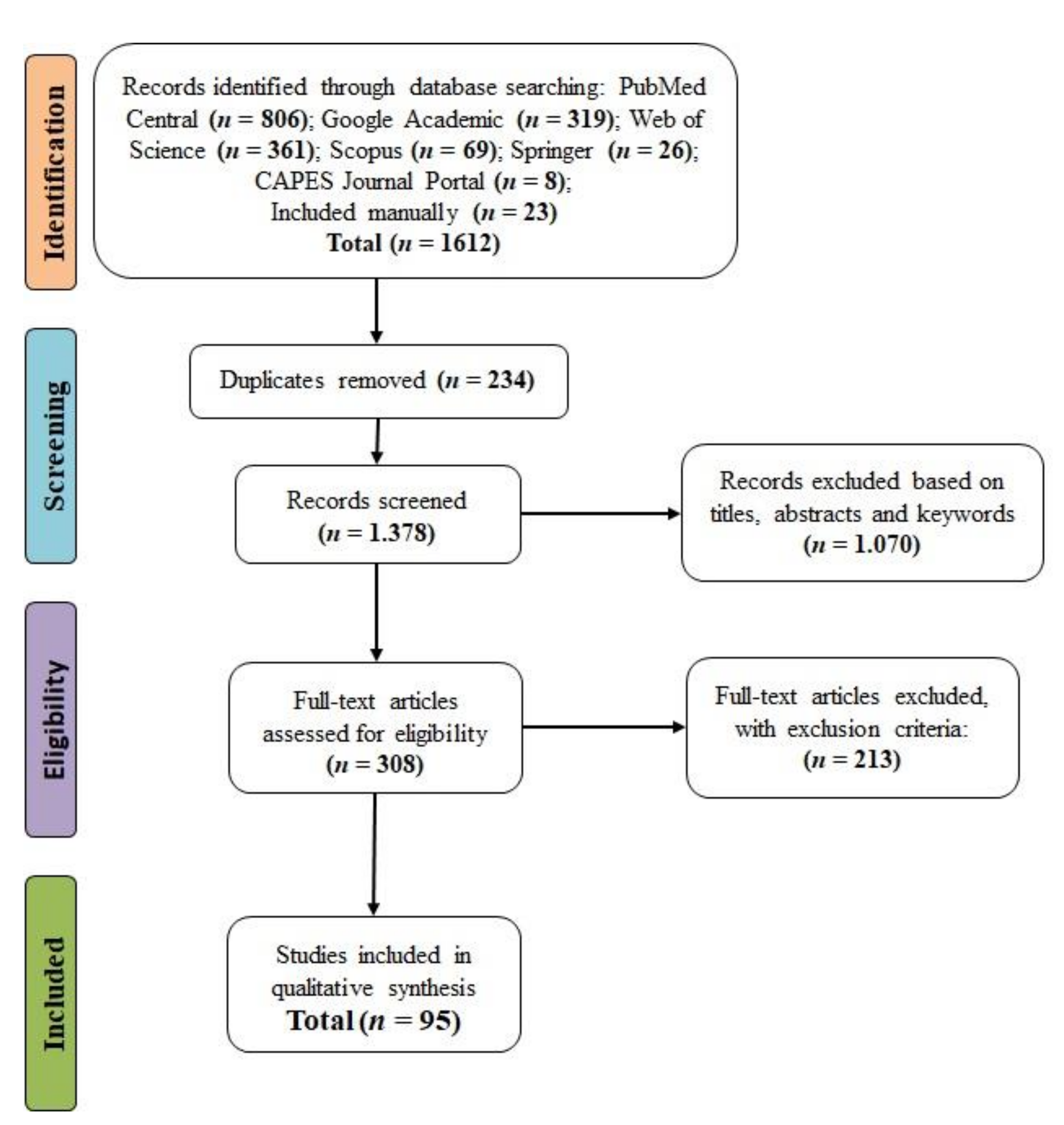
3.1. Screening of Studies
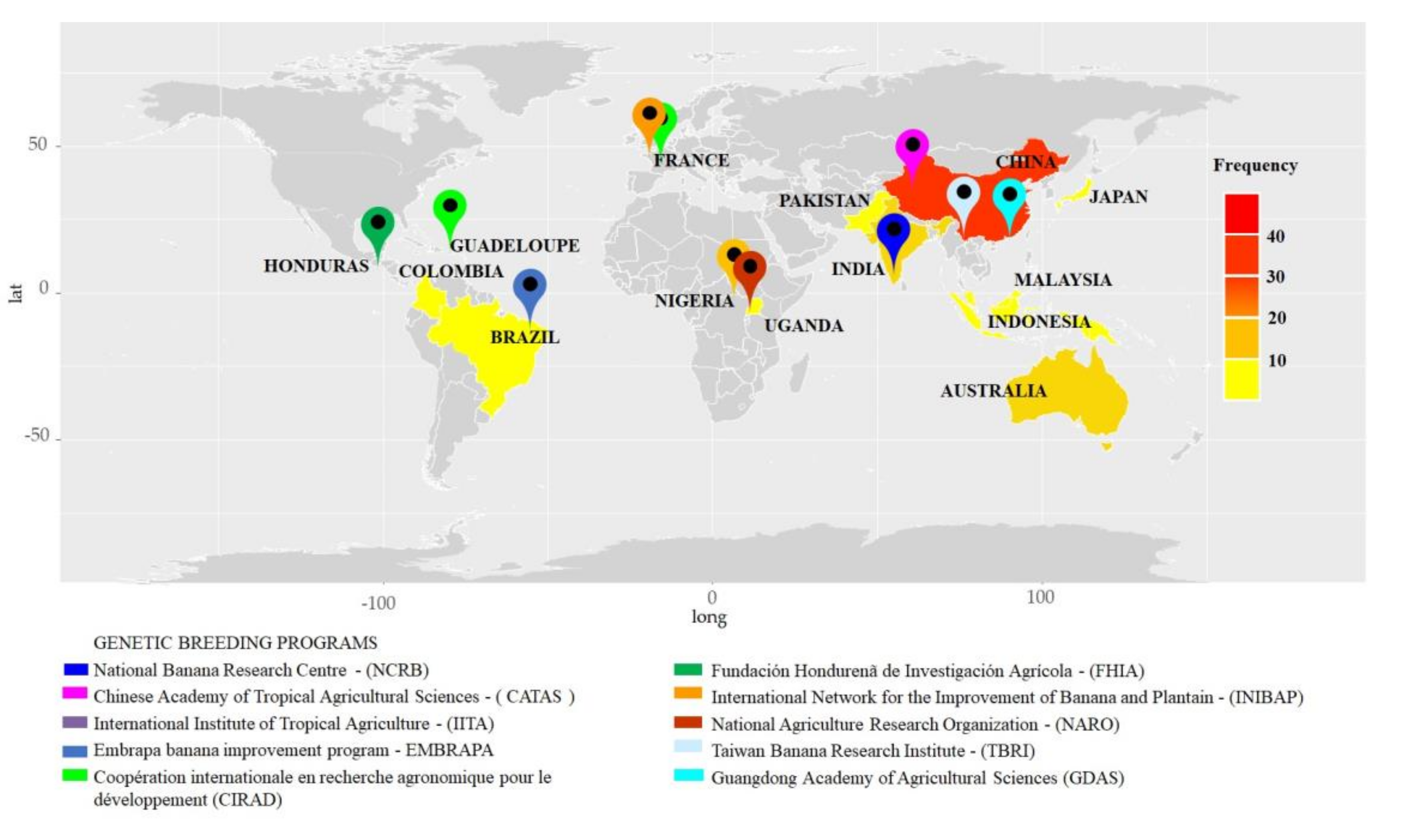
3.2. Known Origin Sites
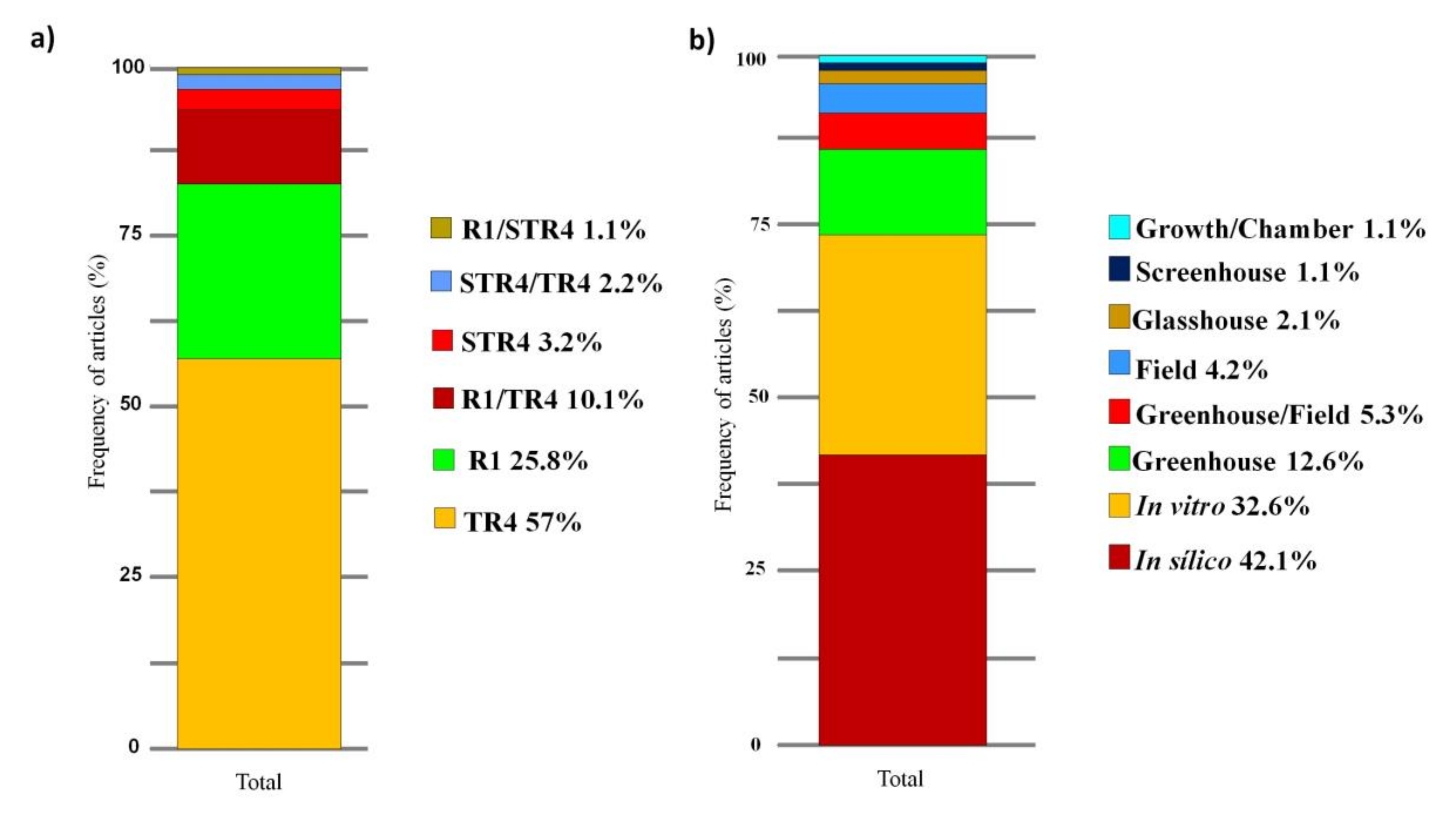
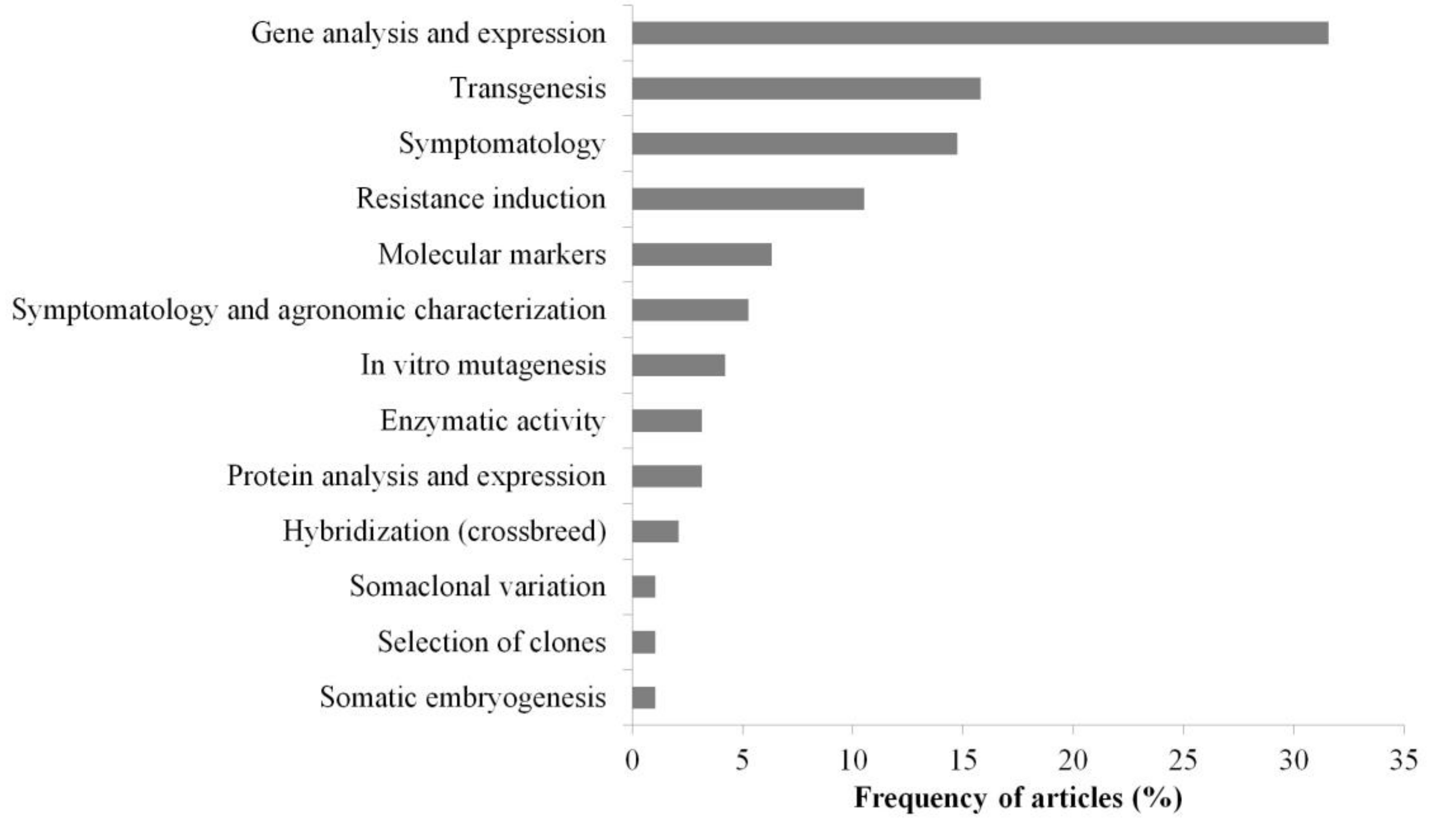
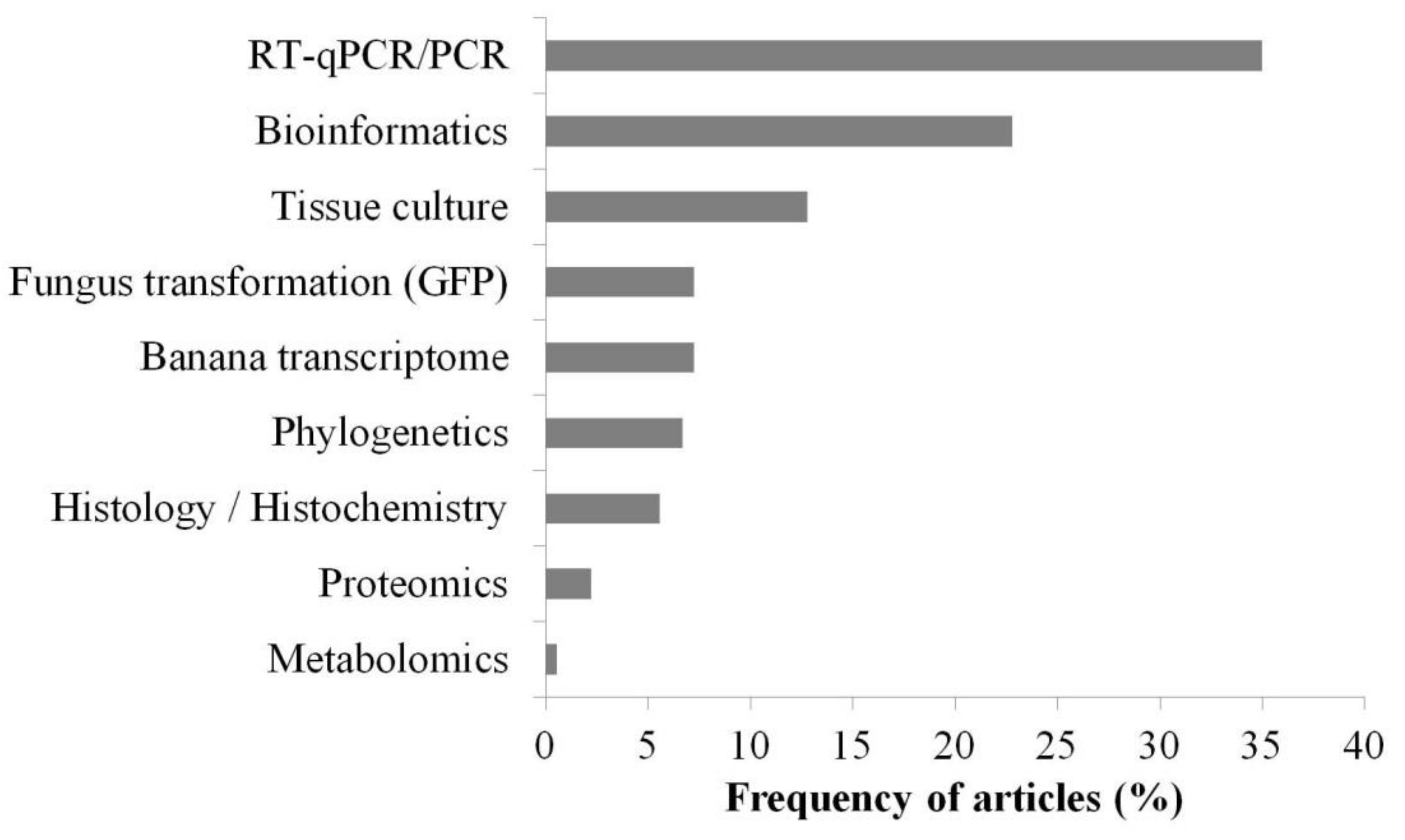
3.3. Main Methods and Tools
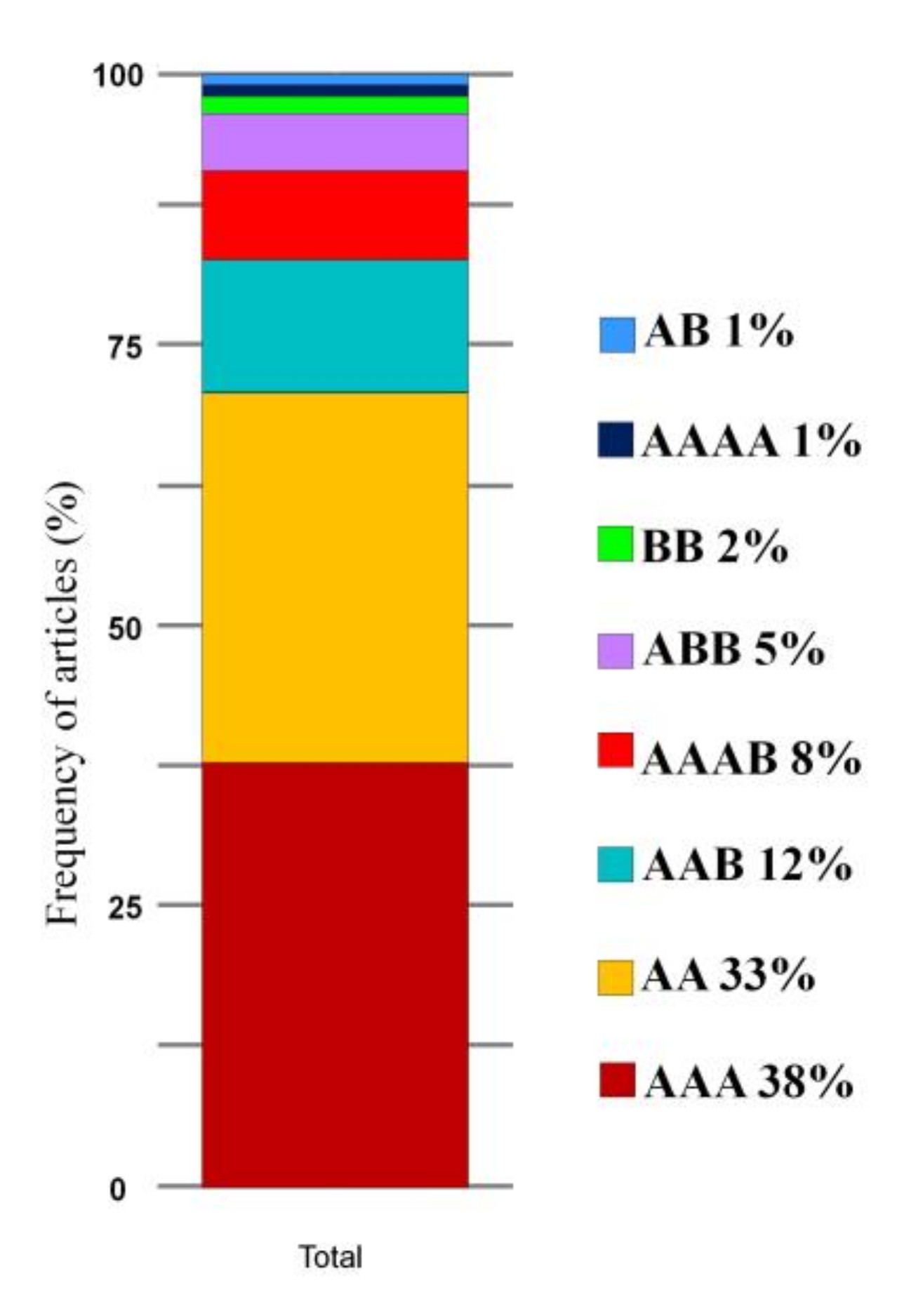
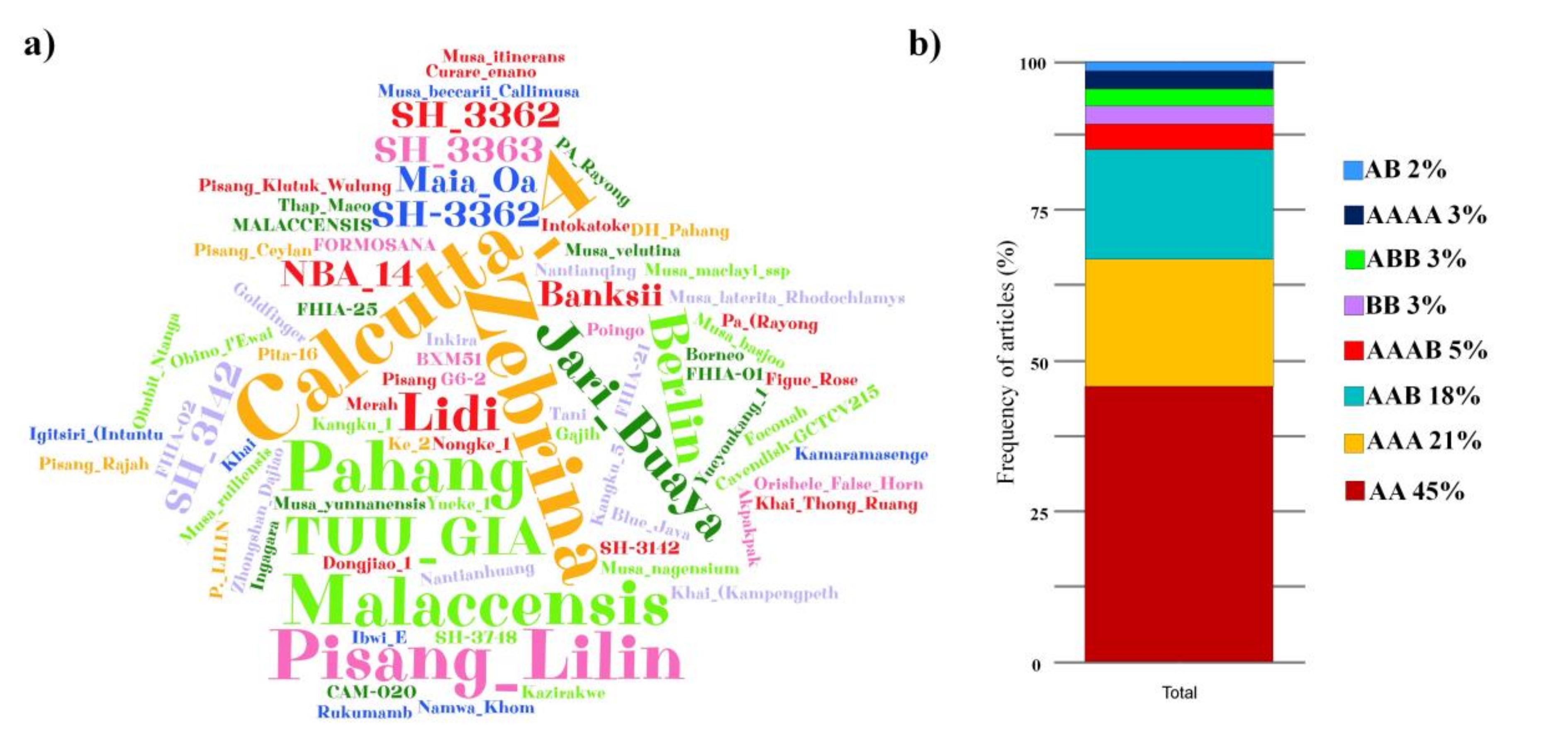
3.4. Resistance Sources
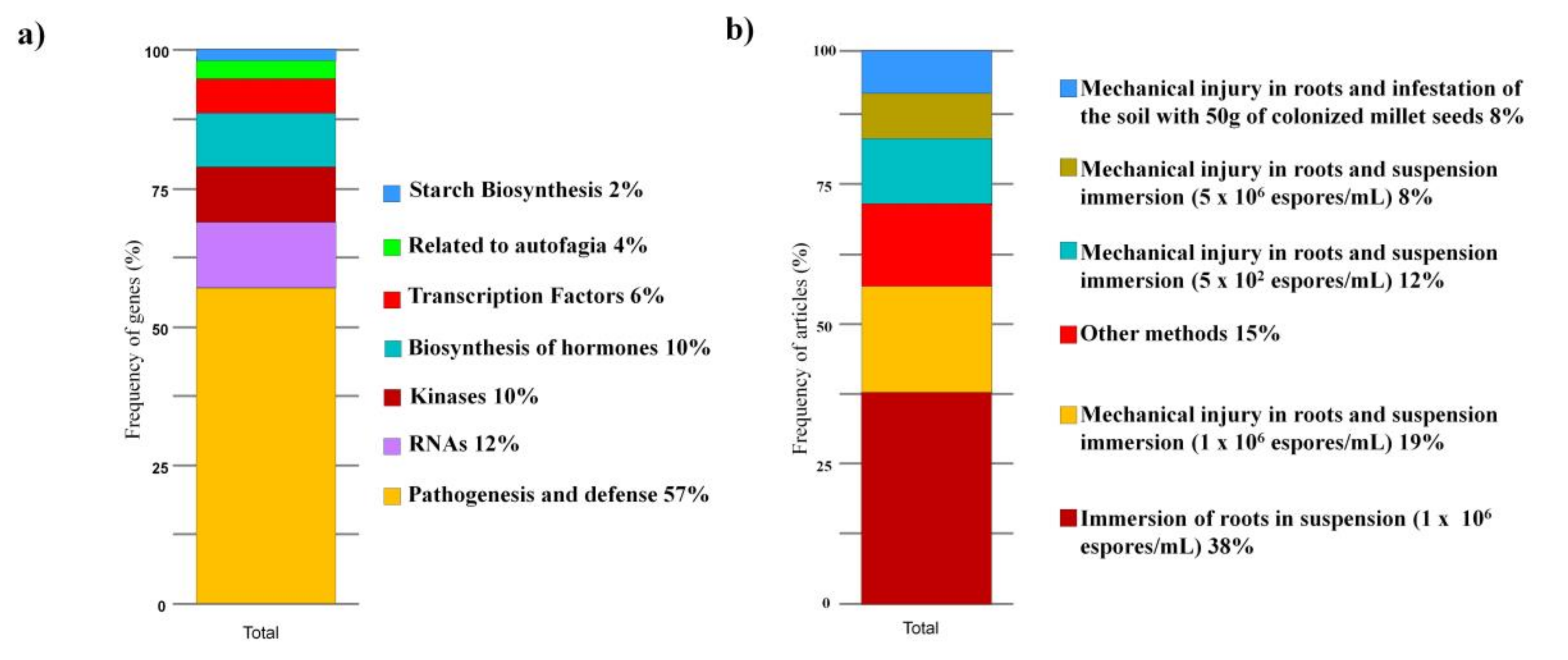
3.5. Gene Expression Analysis
3.6. Studies on the Achievement and Evaluation of Hybrids and on Genetic Inheritance of Musa spp.
3.7. Transgenesis
3.8. Induction of Resistance
4. Discussion
4.1. Screening of the Studies
4.2. Locations of Knowledge Generation
4.3. Gene Expression Analysis
4.4. Studies on Resistance Sources
4.5. Main Methods and Tools Adopted
4.6. Perspectives
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- FAOSTAT. Available online: http://www.fao.org/faostat/en/#data/QC (accessed on 4 June 2020).
- FAO. Available online: http://www.fao.org/fileadmin/templates/est/COMM_MARKETS_MONITORING/Bananas/Documents/Banana_Market_Review_Prelim_Results_2018.pdf. (accessed on 4 June 2020).
- Adeniji, T.; Tenkouano, A.; Ezurike, J.; Ariyo, C.; Vroh-Bi, I. Value-adding post harvest processing of cooking bananas (Musa spp. AAB and ABB genome groups). Afri. J. Biotech. 2010, 9, 9135–9141. [Google Scholar]
- Dotto, J.; Matemu, A.O.; Ndakidemi, P.A. Potential of cooking bananas in addressing food security in East Africa. Int. J. Biol. Sci. 2018, 13, 278–294. [Google Scholar]
- Viljoen, A.; Mostert, D.; Chiconela, T.; Beukes, I.; Fraser, C.; Dwyer, J.; Amugoli, O.M. Occurrence and spread of the banana fungus Fusarium oxysporum f. sp. cubense TR4 in Mozambique. S. Afr. J. Sci. 2020, 116, 11–12. [Google Scholar] [CrossRef]
- Dita, M.A.; Garming, H.; Van den Bergh, I.; Staver, C.; Lescot, T. Banana in Latin America and the Caribbean: Current state, challenges and perspectives. Acta Hortic. 2011, 986, 365–380. [Google Scholar] [CrossRef]
- Lescot, T. Close-up Banana: Statistics. FruiTrop 2011, 189, 59–62. [Google Scholar]
- Gomes, E.W.F.; Willadino, L.; Martins, L.S.S.; Camara, T.R. The effects of salinity on five banana genotypes (Musa spp). In Plant Nutrition; Horst, W.J., Schenk, M.K., Eds.; Springer: Dordrecht, The Netherlands, 2001; Volume 92, pp. 410–411. [Google Scholar] [CrossRef]
- Willadino, L.; Camara, T.R.; Ribeiro, M.B.; Amaral, D.O.J.; Suassuna, F.; Silva, M.V.D. Mechanisms of tolerance to salinity in banana: Physiological, biochemical, and molecular aspects. Rev. Bras. de Frutic. 2017, 39, 1–8. [Google Scholar] [CrossRef]
- Said, E.M.; Mahmoud, R.A.; Al-Akshar, R.; Safwat, G. Drought stress tolerance and enhancement of banana plantlets in vitro. Austin J. Biotechnol. Bioeng. 2015, 2, 1040. [Google Scholar]
- Marssaro, A.L.; Morais-Lino, L.S.; Cruz, J.L.; Ledo, C.A.S.; Santos-Serejo, J.A. Simulation of in vitro water deficit for selecting drought-tolerant banana genotypes. Pesqui. Agropecu. Bras. 2017, 52, 1301–1304. [Google Scholar] [CrossRef][Green Version]
- Nansamba, M.; Sibiya, J.; Tumuhimbise, R.; Karamura, D.; Kubiriba, J.; Karamura, E. Karamura, E. Breeding banana (Musa spp.) for drought tolerance: A review. Plant Breed. 2020, 139, 685–696. [Google Scholar] [CrossRef]
- Vu, T.; Sikora, R.; Hauschild, R. Fusarium oxysporum endophytes induced systemic resistance against Radopholus similis on banana. Nematology 2006, 8, 847–852. [Google Scholar] [CrossRef]
- Gaidashova, S.V.; Uwimpuhwe, B.; Karamura, E.B. Identificatio of banana varieties with resistance to nematodes in Rwanda. Afr. Crop Sci. J. 2008, 16, 27–33. [Google Scholar] [CrossRef]
- Tripathi, L.; Atkinson, H.; Roderick, H.; Kubiriba, J.; Tripathi, J.N. Genetically engineered bananas resistant to Xanthomonas wilt disease and nematodes. Food Energy Secur. 2017, 6, 37–47. [Google Scholar] [CrossRef]
- Njau, N.; Mwangi, M.; Gathu, R.; Mbaka, J.; Muasya, R. Banana weevil (Cosmopolites sordidus) reduces availability of corms for seedling production through macropropagation technology. J. Anim. Plant Sci. 2011, 12, 1537–1542. [Google Scholar]
- Arinaitwe, I.K.; Hilman, E.; Ssali, R.; Barekye, A.; Kubiriba, J.; Kagezi, G.; Talwana, H.; Nankinga, C.; Ragama, P.E.; Tushemereirwe, W.K.; et al. Response of banana hybrids to the banana weevil (Cosmopolites sordidus Germar)(Coleoptera: Curculionidae) in Uganda. Uganda J. Agric. Sci. 2014, 15, 73–85. [Google Scholar]
- Twesigye, C.K.; Ssekatawa, K.; Kiggundu, A.; Tushemereirwe, W.; Matovu, E.; Karamura, E. Corm damage caused by banana weevils Cosmopolites sordidus (Germar) collected from different banana growing regions in Uganda. Agric. Food Secur. 2018, 7, 1–8. [Google Scholar] [CrossRef]
- Monteiro, J.D.; Santos, M.; Santos, J.R.P.; Cares, J.E.; Marchão, R.L.; Amorim, E.P.; Costa, D.D.C. Identification of plant parasitic nematodes in triploid and tetraploid bananas in brazil. Rev. Caatinga 2020, 33, 865–877. [Google Scholar] [CrossRef]
- Galvez, L.C.; Barbosa, C.F.C.; Koh, R.B.L.; Aquino, V.M. Loop-mediated isothermal amplification (LAMP) assays for the detection of abaca bunchy top virus and banana bunchy top virus in abaca. Crop. Prot. 2020, 131, 105101. [Google Scholar] [CrossRef]
- Sairam, S.; Selvarajan, R.; Handanahalli, S.S.; Venkataraman, S.; Sairam, S.; Selvarajan, R.; Handanahalli, S.S.; Venkataraman, S. Towards understanding the structure of the capsid of Banana Bunchy Top Virus. BioRxiv 2020. [Google Scholar] [CrossRef]
- Tripathi, L.; Tripathi, J.N.; Tushemereirwe, W.K. Strategies for resistance to bacterial wilt disease of bananas through genetic engineering. Afr. J. Biotechnol. 2004, 3, 688–692. [Google Scholar]
- Nakato, V.; Mahuku, G.; Coutinho, T. Xanthomonas campestris pv. musacearum: A major constraint to banana, plantain and enset production in central and east Africa over the past decade. Mol. Plant. Pathol. 2018, 19, 525–536. [Google Scholar] [CrossRef]
- Geberewold, A.Z. Review on impact of banana bacterial wilt (Xhantomonas campestris pv. Musacerum) in East and Central Africa. Cogent Food Agric. 2019, 5, 1586075. [Google Scholar] [CrossRef]
- Studholme, D.J.; Wicker, E.; Abrare, S.M.; Aspin, A.; Bogdanove, A.; Broders, K.; Dubrow, Z.; Grant, M.; Jones, J.B.; Karamura, G.; et al. Transfer of Xanthomonas campestris pv. arecae and X. campestris pv. musacearum to X. vasicola (Vauterin) as X. vasicola pv. arecae comb. nov. and X. vasicola pv. musacearum comb. nov. and description of X. vasicola pv. vasculorum pv. nov. Phytopathology 2020, 110, 1153–1160. [Google Scholar] [CrossRef] [PubMed]
- Timm, S.E.; Pardo, H.L.; Coello, P.R.; Navarrete, C.T.; Villegas, N.O.; Ordóñez, S.E. Identification of differentially-expressed genes in response to Mycosphaerella fijiensis in the resistant Musa accession ‘Calcutta-4’using suppression subtractive hybridization. PLoS ONE 2016, 11, 1–17. [Google Scholar] [CrossRef][Green Version]
- Arango Isaza, R.E.; Diaz-Trujillo, C.; Dhillon, B.; Aerts, A.; Carlier, J.; Crane, C.F.; Jong, T.V.; Vries, I.; Dietrich, R.; Farmer, A.D.; et al. Combating a global threat to a clonal crop: Banana black Sigatoka pathogen Pseudocercospora fijiensis (synonym Mycosphaerella fijiensis) genomes reveal clues for disease control. Plos Genet. 2016, 12, e1005876. [Google Scholar] [CrossRef]
- Vázquez-Euán, R.; Chi-Manzanero, B.; Hernández-Velázquez, I.; Tzec-Simá, M.; Islas-Flores, I.; Martínez-Bolaños, L.; Canto-Canché, B. Identification of new hosts of pseudocercospora fijiensis suggests innovative pest management programs for black sigatoka disease in banana plantations. Agronomy 2019, 9, 666. [Google Scholar] [CrossRef]
- Nascimento, F.D.S.; Sousa, Y.M.; Rocha, A.D.J.; Ferreira, C.F.; Haddad, F.; Amorim, E.P. Sources of black Sigatoka resistance in wild banana diploids. Rev. Bras. Frutic 2020, 42. [Google Scholar] [CrossRef]
- Dita, M.; Barquero, M.; Heck, D.; Mizubuti, E.S.; Staver, C.P. Fusarium wilt of banana: Current knowledge on epidemiology and research needs toward sustainable disease management. Front. Plant. Sci. 2018, 9, 1468. [Google Scholar] [CrossRef]
- Arinaitwe, I.K.; Teo, C.H.; Kayat, F.; Tumuhimbise, R.; Uwimana, B.; Kubiriba, J.; Swennen, R.; Harikrishna, J.A.; Othman, R.Y. Evaluation of banana germplasm and genetic analysis of an F1 population for resistance to Fusarium oxysporum f. sp. cubense race 1. Euphytica 2019, 215, 175. [Google Scholar] [CrossRef]
- Sun, J.; Zhang, J.; Fang, H.; Peng, L.; Wei, S.; Li, C.; Lu, J. Comparative transcriptome analysis reveals resistance-related genes and pathways in Musa acuminata banana ‘Guijiao 9’ in response to Fusarium wilt. Plant. Physiol. Biochem. 2019, 141, 83–94. [Google Scholar] [CrossRef]
- Gonçalves, Z.S.; Haddad, F.; Amorim, V.B.O.; Ferreira, C.F.; Oliveira, S.A.S.; Amorim, E.P. Agronomic characterization and identification of banana genotypes resistant to Fusarium wilt race 1. Eur. J. Plant. Pathol. 2019, 155, 1093–1103. [Google Scholar] [CrossRef]
- Ploetz, R.C.; Kema, G.H.; Ma, L.J. Impact of diseases on export and smallholder production of banana. Annu. Rev. Phytopathol. 2015, 53, 269–288. [Google Scholar] [CrossRef]
- Pegg, K.G.; Coates, L.M.; O’Neill, W.T.; Turner, D.W. The epidemiology of Fusarium wilt of banana. Front. Plant. Sci. 2019, 10, 1395. [Google Scholar] [CrossRef]
- Ploetz, R.C. Fusarium wilt. In Handbook of Diseases of Banana, Abacá and Ense; Jones, D., Ed.; CABI Publishing: Wallingford, UK, 2019; pp. 207–228. [Google Scholar]
- Li, C.; Shao, J.; Wang, Y.; Li, W.; Guo, D.; Yan, B.; Xia, Y.; Peng, M. Analysis of banana transcriptome and global gene expression profiles in banana roots in response to infection by race 1 and tropical race 4 of Fusarium oxysporum f. sp. cubense. BMC Genom. 2013, 14, 851. [Google Scholar] [CrossRef]
- Lopes, O.P.; Maia, V.M.; Xavier, A.A.; Costa, M.R.D.; Rodrigues, M.G.V. Diversidade genética, crescimento e produção de genótipos de bananeira ‘Prata-Anã’ em área com mal do panamá. Rev. Bras. Frutic. 2014, 36, 924–939. [Google Scholar] [CrossRef][Green Version]
- Stover, R.H. Fusarial Wilt (Panama Disease) of Bananas and Other Musa Species; Commonwealth Mycological Institute: Kew, UK, 1962; p. 117. [Google Scholar]
- Ploetz, R.C. Fusarium wilt of banana is caused by several pathogens referred to as Fusarium oxysporum f. sp. cubense. Phytopathology 2006, 96, 653–656. [Google Scholar] [CrossRef]
- Ploetz, R.; Freeman, S.; Konkol, J.; Al-Abed, A.; Naser, Z.; Shalan, K.; Israeli, Y. Tropical race 4 of Panama disease in the Middle East. Phytoparasitica 2015, 43, 283–293. [Google Scholar] [CrossRef]
- García-Bastidas, F.A.; Quintero-Vargas, J.C.; Ayala-Vasquez, M.; Schermer, T.; Seidl, M.F.; Santos-Paiva, M.; Noguera, A.M.; Aguilera-Galvez, C.; Wittenberg, A.; Hofstede, R.; et al. First report of Fusarium wilt Tropical Race 4 in Cavendish bananas caused by Fusarium odoratissimum in Colombia. Plant. Dis. 2020, 104, 994. [Google Scholar] [CrossRef]
- FAO. Available online: http://www.fao.org/3/ca6911en/CA6911EN_TR4EN.pdf (accessed on 4 June 2020).
- Thangavelu, R.; Edwin Raj, E.; Loganathan, M.; Pushpakanth, P.; Uma, S. Draft genome of Fusarium oxysporum f. sp. cubense strain Tropical Race-4 infecting Cavendish (AAA) group of banana in India. Plant. Dis. 2020, 105, 481–483. [Google Scholar] [CrossRef]
- Gang, G.; Bizun, W.; Weihong, M.; Xiaofen, L.; Xiaolin, Y.; Chaohua, Z.; Jianhong, M.; Huicai, Z. Biocontrol of Fusarium wilt of banana: Key influence factors and strategies. Afr. J. Microbiol. Res. 2013, 7, 4835–4843. [Google Scholar] [CrossRef]
- Warman, N.M.; Aitken, E.A. The movement of Fusarium oxysporum f. sp. cubense (sub-tropical race 4) in susceptible cultivars of banana. Front. Plant. Sci 2018, 9, 1748. [Google Scholar] [CrossRef]
- Mostert, D.; Molina, A.B.; Daniells, J.; Fourie, G.; Hermanto, C.; Chao, C.P.; Fabregar, E.; Sinohin, V.G.S.; Masdek, N.; Thangavelu, R.A.; et al. The distribution and host range of the banana Fusarium wilt fungus, Fusarium oxysporum f. sp. cubense, in Asia. PLoS ONE 2017, 12, e0181630. [Google Scholar] [CrossRef]
- Costa, S.N.; Bragança, C.A.D.; Ribeiro, L.R.; Amorim, E.P.; Oliveira, S.A.S.; Dita, M.A.; Laranjeira, F.F.; Haddad, F. Genetic structure of Fusarium oxysporum f. sp. cubense in different regions from Brazil. Plant. Pathol. 2015, 64, 137–146. [Google Scholar] [CrossRef]
- Fourie, G.; Steenkamp, E.T.; Ploetz, R.C.; Gordon, T.R.; Viljoen, A. Current status of the taxonomic position of Fusarium oxysporum formae specialis cubense within the Fusarium oxysporum complex. Infect. Genet. Evol. 2011, 11, 533–542. [Google Scholar] [CrossRef] [PubMed]
- Amorim, E.P.; Amorim, V.B.O.; Silva, M.S.; Haddad, F.; Ferreira, C.F.; Santos-Serejo, J.A. Developing hybrid banana varieties with improved properties. In Achieving Sustainable Cultivation of Bananas: Germplasm and Genetic Improvement; Kema, G.H.J., Drenth, A., Eds.; Burleigh Dodds Science Publishing: Cambridge, UK, 2021; Volume 2, pp. 1–17. ISBN 978 1 78676 344 0. [Google Scholar]
- Wang, F.; Xia, L.; Shun, L.V.; Xu, C.; Niu, Y.; Liu, W.; Bei, H.U. Development of a mitochondrial SCAR marker related to susceptibility of banana (Musa AAA Cavendish) to Fusarium oxysporum f. sp. cubense race 4. Not. Bot. Horti. Agrobot. Cluj-Napoca 2018, 46, 509–516. [Google Scholar] [CrossRef]
- Ghag, S.B.; Shekhawat, U.K.; Ganapathi, T.R. Fusarium wilt of banana: Biology, epidemiology and management. Int. J. Pest. Manag. 2015, 61, 250–263. [Google Scholar] [CrossRef]
- Ploetz, R.C. Management of Fusarium wilt of banana: A review with special reference to tropical race 4. J. Crop. Prot 2015, 73, 7–15. [Google Scholar] [CrossRef]
- Thangavelu, R.; Loganathan, M.; Arthee, R.; Prabakaran, M.; Uma, S. Fusarium wilt: A threat to banana cultivation and its management. Cab Rev. 2020, 15, 1–24. [Google Scholar] [CrossRef]
- Bubici, G.; Kaushal, M.; Prigigallo, M.I.; Cabanás, C.G.L.; Mercado-Blanco, J. Biological control agents against Fusarium wilt of banana. Front. Microbiol. 2019, 10, 616. [Google Scholar] [CrossRef]
- Siamak, S.B.; Zheng, S. Banana Fusarium wilt (Fusarium oxysporum f. sp. cubense) control and resistance, in the context of developing wilt-resistant bananas within sustainable production systems. Hortic. Plant J. 2018, 4, 208–218. [Google Scholar] [CrossRef]
- Sipen, P.; Chubo, J.K.; King, P.J.; Huat, O.K.; Davey, M.R. Genetic improvement of banana using conventional and in vitro technologies. J. Crop. Improv. 2011, 25, 697–727. [Google Scholar] [CrossRef]
- Dash, P.K.; Rai, R. Translating the “Banana genome” to delineate stress resistance, dwarfing, parthenocarpy and mechanisms of fruit ripening. Front. Plant. Sci. 2016, 7, 1543. [Google Scholar] [CrossRef]
- Newman, M.; Gough, D. Systematic Reviews in Educational Research: Methodology, Perspectives and Application. In Systematic Reviews in Educational Research; Zawacki-Richter, O., Bedenlier, S., Buntins, K., Kerres, M., Bond, M., Eds.; Springer VS: Wiesbaden, Germany, 2020; pp. 3–22. [Google Scholar]
- Silva, F.A.F.; Brito, B.B.; Santos, M.L.C.; Marques, H.S.; Silva, R.T., Jr.; Carvalho, L.S.D.; Vieira, E.S.; Oliveira, M.V.; Melo, F.F.D. COVID-19 gastrointestinal manifestations: A systematic review. Rev. Soc. Bras. Med. Trop. 2020, 53, e20200714. [Google Scholar] [CrossRef]
- Munn, Z.; Stern, C.; Aromataris, E.; Lockwood, C.; Jordan, Z. What kind of systematic review should I conduct? A proposed typology and guidance for systematic reviewers in the medical and health sciences. BMC Med. Res. Methodol. 2018, 18, 1–9. [Google Scholar] [CrossRef]
- Santos, A.S.; Amorim, E.P.; Ferreira, C.F.; Pirovani, C.P. Water stress in Musa spp.: A systematic review. PLoS ONE 2018, 13, e0208052. [Google Scholar] [CrossRef]
- Falcomer, A.L.; Riquette, R.F.R.; Lima, B.R.; Ginani, V.C.; Zandonadi, R.P. Health benefits of green banana consumption: A systematic review. Nutrients 2019, 11, 1222. [Google Scholar] [CrossRef]
- Maryani, N.; Lombard, L.; Poerba, Y.S.; Subandiyah, S.; Crous, P.W.; Kema, G.H.J. Phylogeny and genetic diversity of the banana Fusarium wilt pathogen Fusarium oxysporum f. sp. cubense in the Indonesian centre of origin. Stud. Myco. 2019, 92, 155–194. [Google Scholar] [CrossRef]
- Orjeda, G. Evaluation of Musa germplasm for resistance to Sigatoka diseases and Fusarium wilt. In INIBAP Technical Guidelines 3; International Plant Genetics Resources Institute: Rome, Italy; International Network for the Improvement of Banana and Plantain: Montpellier, France; ACP-EU Technical Centre for Agriculture and Rural Cooperation: Wageningen, The Netherlands, 1998; p. 63. [Google Scholar]
- Carlier, J.; De Waele, D.; Escalant, J.V. Global Evaluation of Musa Germplasm for Resistance to Fusarium Wilt, Mycosphaerella Leaf Spot Diseases and Nematodes: In-Depth Evaluation; The International Network for the Improvement of Banana and Plantain: Montpellier, France, 2002; p. 64. ISBN 2910810526. [Google Scholar]
- Cordeiro, Z.J.M.; Shepherd, K.; Soares Filho, W.S.; Dantas, J.L.L. Avaliação de resistência ao mal-do-Panamá em híbridos tetraploides de bananeira. Fitopatol. Bras. 1993, 18, 478–483. [Google Scholar]
- Mak, C.; Mohamed, A.A.; Liew, K.W.; Ho, Y.W. Early screening technique for Fusarium wilt resistance in banana micropropagated plants. In Banana Improvement: Cellular, Molecular Biology, and Induced Mutations; Swennen, R., Jain, M.S., Eds.; Science Publishers, Inc.: Enfield, NH, USA, 2004; pp. 219–227. ISBN 1578083400. [Google Scholar]
- Yip, M.K.; Lee, S.W.; Su, K.C.; Lin, Y.H.; Chen, T.Y.; Feng, T.Y. An easy and efficient protocol in the production of pflp transgenic banana against Fusarium wilt. Plant. Biotechnol. Rep. 2011, 5, 245–254. [Google Scholar] [CrossRef]
- Orr, R.; Pattison, A.; East, D.; Warman, N.; O’Neill, W.; Czislowski, E.; Nelson, P.N. Image-based quantification of Fusarium wilt severity in banana. Australas. Plant. Dis. Notes 2019, 14, 14. [Google Scholar] [CrossRef]
- Viljoen, A.; Mahuku, G.; Massawe, C.; Ssali, R.T.; Kimunye, J.; Mostert, G.; Ndayihanzamaso, P.; Coyne, D.L. Banana Pests and Diseases: Field Guide for Disease Diagnostics and Data Collection; International Institute of Tropical Agriculture (IITA): Ibadan, Nigeria, 2017; p. 73. [Google Scholar]
- Chen, A.; Sun, J.; Matthews, A.; Armas-Egas, L.; Chen, N.; Hamill, S.; Mintoff, S.; Tran-Nguyen, L.T.T.; Batley, J.; Aitken, E.A.B. Assessing variations in host resistance to Fusarium oxysporum f sp. cubense race 4 in Musa species, with a focus on the subtropical race 4. Front. Microbiol. 2019, 10, 1062. [Google Scholar] [CrossRef]
- Baharum, N.A.; Othman, R.Y.; Mohd-Yusuf, Y.; Tan, B.C.; Zaidi, K.; Khalid, N. The Effect of Pathogenesis-Related 10 (Pr-10) Gene on the Progression of Fusarium Wilt in Musa acuminata cv. Berangan. Sains Malays. 2018, 47, 2291–2300. [Google Scholar] [CrossRef]
- Zhang, L.; Yuan, T.; Wang, Y.; Zhang, D.; Bai, T.; Xu, S.; Wang, Y.; Tang, W.; Zheng, S.J. Identification and evaluation of resistance to Fusarium oxysporum f. sp. cubense tropical race 4 in Musa acuminata Pahang. Euphytica 2018, 106, 1–12. [Google Scholar] [CrossRef]
- Li, W.M.; Dita, M.; Wu, W.; Hu, G.B.; Xie, J.H.; Ge, X.J. Resistance sources to Fusarium oxysporum f. sp. cubense tropical race 4 in banana wild relatives. Plant. Pathol. 2015, 64, 1061–1067. [Google Scholar] [CrossRef]
- Sun, D.; Lu, X.; Hu, Y.; Li, W.; Hong, K.; Mo, Y.; Xie, J. Methyl jasmonate induced defense responses increase resistance to Fusarium oxysporum f. sp. cubense race 4 in banana. Sci Hortic. 2013, 164, 484–491. [Google Scholar] [CrossRef]
- Zuo, C.; Deng, G.; Li, B.; Huo, H.; Li, C.; Hu, C.; Kuang, R.; Yang, Q.; Dong, T.; Sheng, O.; et al. Germplasm screening of Musa spp. for resistance to Fusarium oxysporum f. sp. cubense tropical race 4 (Foc TR4). Eur. J. Plant. Pathol. 2018, 151, 723–734. [Google Scholar] [CrossRef]
- Ribeiro, L.R.; Oliveira, S.A.S.D.; Amorim, E.P.; Serejo, J.A.S.; Haddad, F. Sources of resistance to Fusarium oxysporum f. sp. cubense in banana germplasm. Rev. Bras. Frutic. 2018, 40, 1–8. [Google Scholar] [CrossRef]
- Mohamed, A.A.; Mak, C.; Liew, K.W.; Ho, Y.W. Early evaluation of banana plants at nursery stage for Fusarium wilt tolerance. In Fusarium Wilt Management: Towards Sustainable Cultivation, Proceedings of the International Workshop Banana Fusarium Wilt Diseases, Genting Highlands Resort, Malaysia, 18–20 October 1999; Molina, A., Mak, C., Liew, K.W., Ho, Y.W., Masdek, N.K., Liew, K.W., Eds.; Bioversity International: Rome, Italy, 2001. [Google Scholar]
- Wei, Y.; Hu, W.; Wang, Q.; Zeng, H.; Li, X.; Yan, Y.; Reiter, R.J.; He, C.; Shi, H. Identification, transcriptional and functional analysis of heat-shock protein 90s in banana (Musa acuminata L.) highlight their novel role in melatonin-mediated plant response to Fusarium wilt. J. Pineal Res. 2016, 62, e12367. [Google Scholar] [CrossRef]
- Garcez, M.; Martins, J.A.S.; Rodrigues, E.J.R. Evaluation of different banana genotypes for resistance to panama disease. Biosci. J. 2016, 32, 431–435. [Google Scholar] [CrossRef]
- Rodriguez, M.A.D.; Ribeiro, L.; Amorim, E.P.; Cordeiro, Z.J.M.; Silva, S.O. Metodologia Para a Caracterização de Genótipos de Bananeira Quanto à Resistência ao Mal.-do-Panamá em Casa de Vegetação; Embrapa Mandioca e Fruticultura. Comunicado Técnico, 150; Embrapa Mandioca e Fruticultura: Cruz das Almas, Brazil, 2011; 5p. [Google Scholar]
- Ghag, S.B.; Shekhawat, U.K.; Ganapathi, T.R. Host-induced post-transcriptional hairpin RNA-mediated gene silencing of vital fungal genes confers efficient resistance against Fusarium wilt in banana. Plant. Biotechnol. J. 2014, 12, 541–553. [Google Scholar] [CrossRef]
- Smith, M.K.; Langdon, P.W.; Pegg, K.G.; Daniells, J.W. Growth, yield and Fusarium wilt resistance of six FHIA tetraploid bananas (Musa spp.) grown in the Australian subtropics. Sci. Hortic. 2014, 170, 176–181. [Google Scholar] [CrossRef]
- Jones, D.R. The Improvement and Testing of Musa: A Global Partnership. Inter-National Network for the Improvement of Banana and Plantain; International Network for the Improvement of Banana and Plantain (INIBAP): Montpellier, France, 1994; p. 303. [Google Scholar]
- Mohandas, S.; Sowmya, H.D.; Saxena, A.K.; Meenakshi, S.; Rani, R.T.; Mahmood, R. Transgenic banana cv. Rasthali (AAB, Silk gp) harboring Ace-AMP1 gene imparts enhanced resistance to Fusarium oxysporum f. sp. cubense race 1. Sci. Hortic. 2013, 164, 392–399. [Google Scholar] [CrossRef]
- Nasir, N.; Pittaway, P.A.; Pegg, K.G.; Lisle, A.T. A foliar rating system for com-paring the resistance of banana cultivar grown as tissue cultured plantlets in the laboratory to Fusarium wilt. Plant. Pathol. 2003, 32, 521–526. [Google Scholar] [CrossRef]
- Ting, A.S.Y.; Mah, S.W.; Tee, C.S. Evaluating the feasibility of induced host resistance by endophytic isolate Penicillium citrinum BTF08 as a control mechanism for Fusarium wilt in banana plantlets. Biol. Control. 2012, 61, 155–159. [Google Scholar] [CrossRef]
- Paul, J.Y.; Becker, D.K.; Dickman, M.B.; Harding, R.M.; Khanna, H.K.; Dale, J.L. Apoptosis-related genes confer resistance to Fusarium wilt in transgenic ‘Lady Finger’bananas. Plant. Biotechnol. J. 2011, 9, 1141–1148. [Google Scholar] [CrossRef]
- Sun, J.B.; Peng, M.; Wang, Y.G.; Zhao, P.J.; Xia, Q.Y. Isolation and characterization of antagonistic bacteria against fusarium wilt and induction of defense related enzymes in banana. Afr. J. Microbiol. Res. 2011, 5, 509–515. [Google Scholar] [CrossRef]
- Ploetz, R.C.; Vazquez, A.; Haynes, J.L. Response of new banana accessions in South Florida to Panama disease. J. Crop. Prot. 1999, 18, 445–449. [Google Scholar] [CrossRef]
- Wu, Y.L.; Yi, G.J.; Peng, X.X. Rapid screening of Musa species for resistance to Fusarium wilt in an in vitro bioassay. Eur. J. Plant. Pathol. 2010, 128, 409–415. [Google Scholar] [CrossRef]
- Ssali, R.T.; Kiggundu, A.; Lorenzen, J.; Karamura, E.; Tushemereirwe, W.; Viljoen, A. Inheritance of resistance to Fusarium oxysporum f. sp. cubense race 1 in bananas. Euphytica 2013, 194, 425–430. [Google Scholar] [CrossRef]
- Smith, L.J.; Smith, M.K.; Tree, D.; O’Keefe, D.; Galea, V.J. Development of a small-plant bioassay to assess banana grown from tissue culture for consistent infection by Fusarium oxysporum f. sp. cubense. Australas. Plant. Path. 2008, 37, 171–179. [Google Scholar] [CrossRef]
- Li, W.; Ge, X.; Wu, W.; Wang, W.; Hu, Y.; Mo, Y.; Sun, D.; Shi, S.; Xie, J. Identification of defense-related genes in banana roots infected by Fusarium oxysporum f. sp. cubense tropical race 4. Euphytica 2015, 205, 837–849. [Google Scholar] [CrossRef]
- Ghag, S.B.; Shekhawat, U.K.S.; Ganapathi, T.R. Native cell-death genes as candidates for developing wilt resistance in transgenic banana plants. Aob Plants 2014, 6, plu037. [Google Scholar] [CrossRef]
- Saraswathi, M.S.; Kannan, G.; Uma, S.; Thangavelu, R.; Backiyarani, S. Improvement of banana cv. Rasthali (Silk, AAB) against Fusarium oxysporum f. sp. cubense (VCG 0124/5) through induced mutagenesis: Determination of LD 50 specific to mutagen, explants, toxins and in vitro and in vivo screening for Fusarium wilt resistance. Indian J. Exp. Biol. 2016, 54, 345–353. [Google Scholar] [PubMed]
- Ghag, S.B.; Shekhawat, U.K.; Ganapathi, T.R. Characterization of Fusarium wilt resistant somaclonal variants of banana cv. Rasthali by cDNA-RAPD. Mol. Biol. Rep. 2014, 41, 7929–7935. [Google Scholar] [CrossRef] [PubMed]
- Wu, Y.; Yi, G.; Peng, X.; Huang, B.; Liu, E.; Zhang, J. Systemic acquired resistance in Cavendish banana induced by infection with an incompatible strain of Fusarium oxysporum f. sp. cubense. J. Plant. Physiol. 2013, 170, 1039–1046. [Google Scholar] [CrossRef]
- Magambo, B.; Harjeet, K.; Arinaitwe, G.; Tendo, S.; Arinaitwe, I.K.; Kubiriba, J.; Tushemereirwe, W.; Dale, J. Inhibition of cell death as an approach for development of transgenic resistance against Fusarium wilt disease. Afr. J. Biotechnol. 2016, 15, 786–797. [Google Scholar] [CrossRef]
- Smith, M.K.; Daniells, J.W.; Peasley, D.; O’Neill, W.; Samuelian, S.; Wright, C.; Drenth, A. Field evaluation of six Gros Michel banana accessions (Musa spp., AAA group) for agronomic performance, resistance to Fusarium wilt race 1 and yellow Sigatoka. J. Crop. Prot. 2018, 113, 84–89. [Google Scholar] [CrossRef]
- Garcia-Bastidas, F.A.; Van der Veen, A.; Nakasato-Tagami, G.; Meijer, H.J.; Arango-Isaza, R.E.; Kema, G.H. An improved phenotyping protocol for panama disease in banana. Front. Plant. Sci. 2019, 10, 1006. [Google Scholar] [CrossRef]
- Cheng, C.; Liu, F.; Sun, X.; Tian, N.; Mensah, R.A.; Li, D.; Lai, Z. Identification of Fusarium oxysporum f. sp. cubense tropical race 4 (Foc TR4) responsive miRNAs in banana root. Sci. Rep. 2019, 9, 1–16. [Google Scholar] [CrossRef]
- Dita, M.A.; Pérez Vicente, L.; Martínez, E. Inoculation of Fusarium oxysporum f. sp. cubense causal agent of fusarium wilt in banana. In Technical Manual: Prevention and Diagnostic of Fusarium Wilt of Banana Caused by Fusarium Oxysporum f. sp. cubense Tropical Race 4 (TR4); Pérez Vicente, L., Dita, M.A., de la Parte, E.M., Eds.; Food and Agriculture Organization of the United Nations: Rome, Italy, 2014; pp. 55–58. [Google Scholar]
- Buregyeya, H.; Tumuhimbise, R.; Matovu, M.; Tumwesigye, K.S.; Kubiriba, J.; Nowankunda, K.; Tushemereirwe, W.K.; Karamura, D.; Karamura, E.; Kityo, R.M.; et al. Fusarium oxysporum Race 1 resistance and quality traits variations in apple banana germplasm. J. Plant. Breed. Crop. Sci. 2020, 12, 16–24. [Google Scholar] [CrossRef]
- Sunisha, C.; Sowmya, H.D.; Usharani, T.R.; Umesha, M.; Gopalkrishna, H.R.; Saxena, A. Deployment of Stacked Antimicrobial Genes in Banana for Stable Tolerance Against Fusarium oxysporum f. sp. cubense Through Genetic Transformation. Mol. Biotechnol. 2020, 62, 8–17. [Google Scholar] [CrossRef]
- Rocha, A.D.J.; Ferreira, M.D.S.; Rocha, L.D.S.; Oliveira, A.S.; Amorim, E.P.; Mizubuti, E.S.; Haddad, F. Interaction between Fusarium oxysporum f. sp. cubense and Radopholus similis can lead to changes in the resistance of banana cultivars to Fusarium wilt. Eur. J. Plant. Pathol. 2020, 158, 403–417. [Google Scholar] [CrossRef]
- Ahmad, F.; Martawi, N.M.; Poerba, Y.S.; Jong, H.; Schouten, H.; Kema, G.H. Genetic mapping of Fusarium wilt resistance in a wild banana Musa acuminata ssp. malaccensis accession. Appl. Genet. 2020, 133, 3409–3418. [Google Scholar] [CrossRef]
- Silva, P.R.O.; de Jesus, O.N.; Bragança, C.A.D.; Haddad, F.; Amorim, E.P.; Ferreira, C.F. Development of a thematic collection of Musa spp accessions using SCAR markers for preventive breeding against Fusarium oxysporum f. sp cubense tropical race 4. Genet. Mol. Res. 2016, 15, 15017765. [Google Scholar] [CrossRef]
- Wang, W.; Hu, Y.; Sun, D.; Staehelin, C.; Xin, D.; Xie, J. Identification and evaluation of two diagnostic markers linked to Fusarium wilt resistance (race 4) in banana (Musa spp.). Mol. Bio. Rep. 2012, 39, 451–459. [Google Scholar] [CrossRef]
- Cunha, C.M.; Hinz, R.H.; Pereira, A.; Tcacenco, F.A.; Paulino, E.C.; Stadnik, M.J.A. SCAR Marker for identifying susceptibility to Fusarium oxysporum f. sp. cubense in banana. Sci. Hortic. 2015, 191, 108–112. [Google Scholar] [CrossRef]
- Ndayihanzamaso, P.; Mostert, D.; Matthews, M.C.; Mahuku, G.; Jomanga, K.; Mpanda, H.J.; Mduma, H.; Brown, A.; Uwimana, B.; Swennen, R.; et al. Evaluation of Mchare and Matooke Bananas for Resistance to Fusarium oxysporum f. sp. cubense Race 1. Plants 2020, 9, 1082. [Google Scholar] [CrossRef]
- Bai, T.T.; Xie, W.B.; Zhou, P.P.; Wu, Z.L.; Xiao, W.C.; Zhou, L.; Sun, L.; Ruan, X.L.; Li, H.P.; Tumuhimbise, R.; et al. Transcriptome and expression profile analysis of highly resistant and susceptible banana roots challenged with Fusarium oxysporum f. sp. cubense tropical race 4. PLoS ONE 2013, 8, e73945. [Google Scholar] [CrossRef]
- Zhang, L.; Cenci, A.; Rouard, M.; Zhang, D.; Wang, Y.; Tang, W.; Zheng, S.J. Transcriptomic analysis of resistant and susceptible banana corms in response to infection by Fusarium oxysporum f. sp. cubense tropical race 4. Sci. Rep. 2019, 9, 1–14. [Google Scholar] [CrossRef]
- Wang, Y.; Xia, Q.; Wang, G.; Zhang, H.; Lu, X.; Sun, J.; Zhang, X. Differential gene expression in banana roots in response to Fusarium wilt. Can. J. Plant. Pathol. 2017, 39, 163–175. [Google Scholar] [CrossRef]
- Wang, Z.; Zhang, J.; Jia, C.; Liu, J.; Li, Y.; Yin, X.; Xu, B.; Jin, Z. De novo characterization of the banana root transcriptome and analysis of gene expression under Fusarium oxysporum f. sp. Cubense tropical race 4 infection. BMC Genom. 2012, 13, 650. [Google Scholar] [CrossRef]
- Li, C.Y.; Deng, G.M.; Yang, J.; Viljoen, A.; Jin, Y.; Kuang, R.B.; Zuo, C.W.; Lv, Z.C.; Yang, Q.S.; Sheng, O.; et al. Transcriptome profiling of resistant and susceptible Cavendish banana roots following inoculation with Fusarium oxysporum f. sp. cubense tropical race 4. BMC Genom. 2012, 13, 374. [Google Scholar] [CrossRef]
- Fei, S.; Czislowski, E.; Fletcher, S.; Peters, J.; Batley, J.; Aitken, E.; Mitter, N. Small RNA profiling of Cavendish banana roots inoculated with Fusarium oxysporum f. sp. cubense race 1 and tropical race 4. Phytopathol. Res. 2019, 1, 22. [Google Scholar] [CrossRef]
- Song, S.; Chen, X.; Huang, D.; Xu, Y.; Zeng, H.; Hu, X.; Wang, W. Identification of miRNAs differentially expressed in Fusarium wilt-resistant and susceptible banana varieties. S. Afr. J. Bot. 2016, 106, 244–249. [Google Scholar] [CrossRef]
- Li, W.; Li, C.; Li, S.; Peng, M. Long noncoding RNAs that respond to Fusarium oxysporum infection in ‘Cavendish’ banana (Musa acuminata). Sci. Rep. 2017, 7, 1–13. [Google Scholar] [CrossRef]
- Niu, Y.; Hu, B.; Li, X.; Chen, H.; Takáč, T.; Šamaj, J.; Xu, C. Comparative digital gene expression analysis of tissue-cultured plantlets of highly resistant and susceptible banana cultivars in response to Fusarium oxysporum. Int. J. Mol. Sci. 2018, 19, 350. [Google Scholar] [CrossRef]
- Subramaniam, S.; Mahmood, M.; Meon, S.; Rathinam, X. Genetic engineering for tolerance to Fusarium wilt race 1 in Musa sapientum cv. Rastali (AAB) using biolistic gun transformation system. Tree For. Sci. Biotechnol. 2010, 4, 65–75. [Google Scholar]
- Poon, N.K.; Teo, C.H. Fusarium Wilt Disease of Banana: Current Development of Fusarium Resistant Banana. J. Microbiol. Biotechnol. 2019, 4, 000134. [Google Scholar] [CrossRef]
- Ghag, S.B.; Shekhawat, U.K.S.; Ganapathi, T.R. Petunia floral defensins with unique prodomains as novel candidates for development of Fusarium wilt resistance in transgenic banana plants. PLoS ONE 2012, 7, e39557. [Google Scholar] [CrossRef]
- Hu, C.H.; Wei, Y.R.; Huang, Y.H.; Yi, G.J. An efficient protocol for the production of chit42 transgenic Furenzhi banana (Musa spp. AA group) resistant to Fusarium oxysporum. In Vitro Cell. Dev. Biol. Plant 2013, 49, 584–592. [Google Scholar] [CrossRef]
- Ghag, S.B.; Shekhawat, U.K.S.; Ganapathi, T.R. Transgenic banana plants expressing a Stellaria media defensin gene (Sm-AMP-D1) demonstrate improved resistance to Fusarium oxysporum. Plant. Cell Tissue Organ. Cult. 2014, 119, 247–255. [Google Scholar] [CrossRef]
- Dale, J.; James, A.; Paul, J.Y.; Khanna, H.; Smith, M.; Peraza-Echeverria, S.; Garcia-Bastidas, F.; Kema, G.; Waterhouse, P.; Mengersen, K.; et al. Transgenic Cavendish bananas with resistance to Fusarium wilt tropical race 4. Nat. Commun. 2017, 8, 1–8. [Google Scholar] [CrossRef]
- Dou, T.; Shao, X.; Hu, C.; Liu, S.; Sheng, O.; Bi, F.; Deng, G.; Ding, L.; Li, C.; Dong, T.; et al. Host-induced gene silencing of Foc TR 4 ERG 6/11 genes exhibits superior resistance to Fusarium wilt of banana. Plant. Biotechnol. J. 2020, 18, 11–13. [Google Scholar] [CrossRef]
- Ghag, S.B.; Shekhawat, U.K.; Hadapad, A.B.; Ganapathi, T.R. Stacking of host-induced gene silencing mediated resistance to banana bunchy top virus and fusarium wilt disease in transgenic banana plants. Curr. Trends Biotechnol. Pharm. 2015, 9, 212–221. [Google Scholar]
- Zhang, L.; Liu, L.; Cheng, P.; Shen, H.; Rong, B.; Liu, W.; Yu, G. Identification and validation of reference genes for RT-qPCR analysis in banana (Musa spp.) under Fusarium wilt resistance induction conditions. J. Phytopathol. 2017, 165, 746–754. [Google Scholar] [CrossRef]
- Chandrasekaran, M.; Raman, C.; Karthikeyan, K.; Paramasivan, M. Functional Annotation of Hypothetical Proteins Derived from Suppressive Subtraction Hybridization (SSH) Analysis Shows NPR1 (Non-Pathogenesis Related)-Like Activity. Agronomy 2019, 9, 57. [Google Scholar] [CrossRef]
- Wang, Z.; Jia, C.; Li, J.; Huang, S.; Xu, B.; Jin, Z. Activation of salicylic acid metabolism and signal transduction can enhance resistance to Fusarium wilt in banana (Musa acuminata L. AAA group, cv. Cavendish). Funct. Integr. Genomics 2015, 15, 47–62. [Google Scholar] [CrossRef]
- Cheng, Z.; Yu, X.; Li, S.; Wu, Q. Genome-wide transcriptome analysis and identification of benzothiadiazole-induced genes and pathways potentially associated with defense response in banana. BMC Genom. 2018, 19, 1–19. [Google Scholar] [CrossRef]
- Thakker, J.N.; Patel, S.; Dhandhukia, P.C. Induction of defense-related enzymes in banana plants: Effect of live and dead pathogenic strain of Fusarium oxysporum f. sp. cubense. Isrn Biotechnol. 2013, 1–6. [Google Scholar] [CrossRef]
- Nasir, N.; Dharma, A.; Habazar, T. The Chitinase Activity in Banana Seedling that Induce by Trichoderma spp as Resistance Responce to Fusarium Oxyporum f. sp. cubense. Int. J. Adv. Sci. Eng. Inf. Technol. 2016, 6, 356–360. [Google Scholar] [CrossRef]
- Magdama, F.; Monserrate-Maggi, L.; Serrano, L.; Onofre, J.G.; Jiménez-Gasco, M.D.M. Genetic Diversity of Fusarium oxysporum f. sp. cubense, the Fusarium Wilt Pathogen of Banana, in Ecuador. Plants 2020, 9, 1133. [Google Scholar] [CrossRef]
- Guo, L.; Han, L.; Yang, L.; Zeng, H.; Fan, D.; Zhu, Y.; Feng, Y.; Wang, G.; Peng, C.; Jiang, X.; et al. Genome and Transcriptome Analysis of the Fungal Pathogen Fusarium oxysporum f. sp. cubense Causing Banana Vascular Wilt Disease. PLoS ONE 2015, 10, e0117621. [Google Scholar] [CrossRef]
- Staver, C.; Pemsl, D.E.; Scheerer, L.; Vicente, L.P.; Dita, M. Ex ante assessment of returns on research investments to address the impact of Fusarium wilt tropical race 4 on global banana production. Front. Plant. Sci. 2020, 11. [Google Scholar] [CrossRef]
- Linbing, X.; Hu, Y.; Bingzhi, H.; Yuerong, W. Banana research and production in China. In Advancing Banana and Plantain R&D in Asia and the Pacific; Molina, A.B., Xu, L.B., Roa, V.N., Van den Berghand, I., Borromeo, K.H., Eds.; International Network for the Improvement of Banana and Plantain (INIBAP): Brussels, Belgium, 2004; Volume 13, p. 51. [Google Scholar]
- Simmonds, N.W. Evolution of Bananas; Longman: London, UK, 1962; pp. 5–25. [Google Scholar]
- Bancroft, J. Report of the board appointed to enquire into the cause of disease affecting livestock and plants. Votes Proc. 1876, 3, 1011–1038. [Google Scholar]
- O’Neill, W.T.; Henderson, J.; Pattemore, J.A.; O’Dwyer, C.; Perry, S.; Beasley, D.R.; Tan, Y.P.; Smyth, A.L.; Goosem, C.H.; Thomson, K.M.; et al. Detection of Fusarium oxysporum f. sp. cubense tropical race 4 strain in northern Queensland. Australas. Plant. Dis. Notes 2016, 11, 33. [Google Scholar] [CrossRef]
- Scheerer, L.; Pemsl, D.; Staver, C.; Dita, M.; Vicente, L.P. A Quantified Approach to Projecting Losses Caused by Fusarium Wilt Tropical Race 4; International Society for Horticultural Science: Brussels, Belgium, 2016; Volume 1196, pp. 211–218. [Google Scholar] [CrossRef]
- D’Hont, A.; Denoeud, F.; Aury, J.M.; Baurens, F.C.; Carreel, F.; Garsmeur, O.; Noel, B.; Bocs, S.; Droc, G.; Rouard, M.; et al. The banana (Musa acuminata) genome and the evolution of monocotyledonous plants. Nature 2012, 488, 213–217. [Google Scholar] [CrossRef]
- Raman, A.S.; White, K.I.; Ranganathan, R. Origins of allostery and evolvability in proteins: A case study. Cell 2016, 166, 468–480. [Google Scholar] [CrossRef]
- Dong, H.; Ye, Y.; Guo, Y.; Li, H. Comparison transcriptome analysis revealed resistance differences of Cavendish bananas to Fusarium oxysporum f. sp. cubense race1 and race 4. BMC Genom. 2020, 21, 122. [Google Scholar] [CrossRef]
- Li, W.M.; Dita, M.; Rouard, M.; Wu, W.; Roux, N.; Xie, J.H.; Ge, X.J. Deep RNA-seq analysis reveals key responding aspects of wild banana relative resistance to Fusarium oxysporum f. sp. cubense tropical race 4. Funct. Integr. Genom. 2020, 20, 551–562. [Google Scholar] [CrossRef]
- Chen, Y.C.; Wong, C.L.; Muzzi, F.; Vlaardingerbroek, I.; Kidd, B.N.; Schenk, P.M. Root defense analysis against Fusarium oxysporum reveals new regulators to confer resistance. Sci. Rep. 2014, 2014, 5584. [Google Scholar] [CrossRef]
- Wang, Z.; Miao, H.; Liu, J.; Xu, B.; Yao, X.; Xu, C.; Zhang, J. Musa balbisiana genome reveals subgenome evolution and functional divergence. Nat. Plants 2019, 5, 810–821. [Google Scholar] [CrossRef]
- Davey, M.W.; Gudimella, R.; Harikrishna, J.A.; Sin, L.W.; Khalid, N.; Keulemans, J. A draft Musa balbisiana genome sequence for molecular genetics in polyploid, inter-and intra-specific Musa hybrids. BMC Genom. 2013, 14, 683. [Google Scholar] [CrossRef] [PubMed]
- Wei, Y.; Liu, W.; Hu, W.; Liu, G.; Wu, C.; Liu, W.; Zeng, H.; He, Z.; Shi, H. Genome-wide analysis of autophagy-related genes in banana highlights MaATG8s in cell death and autophagy in immune response to Fusarium wilt. Plant. Cell Rep. 2017, 36, 1237–1250. [Google Scholar] [CrossRef]
- Hwang, S.C.; Ko, W.H. Cavendish banana cultivars resistant to Fusarium wilt acquired through somaclonal variation in Taiwan. Plant. Dis. 2004, 88, 580–588. [Google Scholar] [CrossRef] [PubMed]
- Ferreira, M.D.S.; Moura, É.R.D.; Lino, L.S.M.; Amorim, E.P.; Santos-Serejo, J.A.D.; Haddad, F. Selection of somaclonal variants of the cultivar ‘Prata-Anã’ for resistance to Fusarium oxysporum f. sp. cubense race 1. Rev. Bras. Frutic. 2020, 42. [Google Scholar] [CrossRef]
- Ganapathi, T.R.; Higgs, N.S.; Balint-Kurti, P.J.; Arntzen, C.J.; May, G.D.; Van Eck, J.M. Agrobacterium-mediated transformation of embryogenic cell suspensions of the banana cultivar Rasthali (AAB). Plant. Cell Rep. 2001, 20, 157–162. [Google Scholar] [CrossRef]
- Khanna, H.; Becker, D.; Kleidon, J.; Dale, J.L. Centrifugation assisted Agrobacterium tumefaciens-mediated transformation (CAAT) of embryogenic cell suspensions of banana (Musa spp. Cavendish AAA and Lady finger AAB). Mol. Breed. 2004, 14, 239–252. [Google Scholar] [CrossRef]
- Damodaran, T.; Kumar, N.; Kavino, M. Breeding and evaluation of Musa hybrids resistant to Fusarium oxysporum f. sp. cubense race 1. Fruits 2009, 64, 3–12. [Google Scholar] [CrossRef]
- Brown, A.; Tumuhimbise, R.; Amah, D.; Uwimana, B.; Nyine, M.; Mduma, H.; Talengera, D.; Karamura, D.; Kuriba, J.; Swennen, R. The genetic improvement of bananas and plantains (Musa spp.). In Genetic Improvement of Tropical Crops; Caligari, P.D.S., Ed.; Springer: Cham, Switzerland, 2017; pp. 219–240. [Google Scholar]
- Nyine, M.; Uwimana, B.; Blavet, N.; Hřibová, E.; Vanrespaille, H.; Batte, M.; Akech, V.; Brown, A.; Lorenzen, J.; Swennen, R.; et al. Genomic prediction in a multiploid crop: Genotype by environment interaction and allele dosage effects on predictive ability in Banana. Plant. Genome 2018, 11. [Google Scholar] [CrossRef]
- Dale, J.; Paul, J.Y.; Dugdale, B.; Harding, R. Modifying bananas: From transgenics to organics? Sustainability 2017, 9, 333. [Google Scholar] [CrossRef]
- Ntui, V.O.; Tripathi, J.N.; Tripathi, L. Robust CRISPR/Cas9 mediated genome editing tool for banana and plantain (Musa spp.). Curr. Plant. Biol. 2019, 21, 100128. [Google Scholar] [CrossRef]
- Tripathi, L.; Ntui, V.O.; Tripathi, J.N. Application of genetic modification and genome editing for developing climate-smart banana. Food Energy Secur. 2019, 8, e00168. [Google Scholar] [CrossRef]
- Tripathi, J.N.; Ntui, V.O.; Ron, M.; Muiruri, S.K.; Britt, A.; Tripathi, L. CRISPR/Cas9 editing of endogenous banana streak virus in the B genome of Musa spp. overcomes a major challenge in banana breeding. Commun. Biol. 2020, 2, 1–11. [Google Scholar] [CrossRef]
- Tripathi, L.; Ntui, V.O.; Tripathi, J.N. CRISPR/Cas9-based genome editing of banana for disease resistance. Curr. Opin. Plant Biol. 2020, 56, 118–126. [Google Scholar] [CrossRef]
- Arora, S.; Steuernagel, B.; Gaurav, K.; Chandramohan, S.; Long, Y.; Matny, O.; Johnson, R.; Enk, J.; Periyannan, S.; Singh, N.; et al. Resistance gene cloning from a wild crop relative by sequence capture and association genetics. Nat. Biotechnol. 2019, 37, 139–143. [Google Scholar] [CrossRef]
- Jupe, F.; Witek, K.; Verweij, W.; Sliwka, J.; Pritchard, L.; Etherington, G.J.; Maclean, D.; Cock, P.J.; Leggett, R.M.; Bryan, G.J.; et al. Resistance gene enrichment sequencing (RenSeq) enables reannotation of the NB-LRR gene family from sequenced plant genomes and rapid mapping of resistance loci in segregating populations. Plant. J. 2013, 76, 530–544. [Google Scholar] [CrossRef]

| Research Questions |
|---|
| Q1: What are the known sources of resistance (germplasm) to Fusarium oxysporum f. sp. cubense? |
| Q2: Which breeding programs work on the resistance to Fusarium wilt with respect to cultivar development? |
| Q3: Which genes are reported associated with resistance to Fusarium oxysporum f. sp. cubense in Musa spp. |
| Q4: What breeding techniques are associated with overcome Fusarium wilt? |
| Q5: Which biotechnological tools are used for assisted selection for resistance to Fusarium oxysporum f. sp. cubense? |
| Q6: Which germplasm collections have information with the potential for genetic improvement to Fusarium wilt? |
| Q7: What is the frequency of studies by country, and which programs of improvement work with crossbreeding in order to develop resistant cultivars? |
| Q8: Are there scales to assess the disease? What is the difference between them? |
| Q9: How often is the banana genome used? |
| Article | Internal Symptoms | External Symptoms | Scale Reference | |
|---|---|---|---|---|
| Rhizome Discoloration | Yellowing of the Leaf | Pseudostem Division | ||
| Degrees of Scale | ||||
| Yip et al. [69] | 0–3 | [69] * | ||
| Orr et al. [70] | 1–6 | [71] | ||
| Chen et al. [72] | 1–8 | [68] | ||
| Warman and Aitken [46] | 1–6 | [66] | ||
| Baharum et al. [73] | 1–8 | [68] | ||
| Zhang et al. [74] | 0–4 | 0–4 | [75,76] | |
| Zuo et al. [77] | 1–5 | [77] * | ||
| Ribeiro et al. [78] | 0–5 | 0–4 | [67,79] | |
| Wei et al. [80] | 0–4 | [80] * | ||
| Garcez et al. [81] | 0–5 | 0–5 | [67,82] | |
| Li et al. [75] | 0–3 | 0–3 | [75] * | |
| Ghag et al. [83] | 1–6 | [66] | ||
| Smith et al. [84] | 1–6 | [65,85] | ||
| Mohandas et al. [86] | 1–6 | 0–5 | [65,87] | |
| Ting et al. [88] | 0–5 | [88] * | ||
| Paul et al. [89] | 1–5 | 1–3 | [89] * | |
| Sun et al. [90] | 1–5 | 1–5 | [64,91] | |
| Wu et al. [92] | 1–6 | [92] ** | ||
| Ssali et al. [93] | 1–6 | [94] | ||
| Li et al. [95] | 0–4 | 0–5 | [95] * | |
| Ghag et al. [96] | 1–6 | [96] * | ||
| Saraswathi et al. [97] | 1–5 | 1–5 | [66,91] | |
| Ghag et al. [98] | 1–6 | [83] | ||
| Sun et al. [76] | 0–4 | [76] * | ||
| Wu et al. [99] | 1–6 | [99] ** | ||
| Magambo et al. [100] | 1–5 | 1–3 | [68] | |
| Smith et al. [101] | 1–6 | [65] | ||
| García-Bastidas et al. [102] | 1–6 | 1–4 | [102] * | |
| Arinaitwe et al. [31] | 1–5 | 1–6 | 1–3 | [71] |
| Cheng et al. [103] | 1–8 | [68] | ||
| Gonçalves et al. [33] | 1–5 | 1–6 | [67,104] | |
| Buregyeya et al. [105] | 1–6 | 1–3 | [94] | |
| Sunisha et al. [106] | 1–5 | 1–3 | [89] | |
| Rocha et al. [107] | 1–5 | 1–4 | [104] | |
| Ahmad et al. [108] | 1–6 | 1–4 | [102] | |
| Name | Type | Function | Citation |
|---|---|---|---|
| ScaU1001 | SCAR | Resistance to FOC TR4 | [109] |
| SuscPD | SCAR | Susceptibility to FOC 1 | [111] |
| Lipoxygenase (gene) | RAPD | Resistance to FOC 1 | [98] |
| ScaU1001 | SCAR | Resistance to FOC TR4 | [110] |
| ScaS0901 | |||
| SC1/SC2 | SCAR | Resistance to FOC TR4 | [51] |
| SC3/SC4 | |||
| SC5/SC6 | |||
| SC7/SC8 |
| Musa Germplasm | Musa Genome | Race | Level of Tolerance or Resistance to Races | Institution and Location/Country Where Germplasm Was Screened | Known Use to Mitigate Fusarium Impact | References |
|---|---|---|---|---|---|---|
| M53 | AA | Race 1 | R | Embrapa cassava and fruit growing, Brazil | In breeding programs | [33,78,107] |
| Birmanie | AA | Race 1 | R | Embrapa cassava and fruit growing, Brazil | In breeding programs | [78,107] |
| PA Songkla | AA | Race 1 | R | Embrapa cassava and fruit growing, Brazil | In breeding programs | [78] |
| Pirua | AAA | Race 1 | R | Embrapa cassava and fruit growing, Brazil | Brazil | [78] |
| Imperial | AAA | Race 1 | R | Embrapa cassava and fruit growing, Brazil | Brazil | [78] |
| Poyo | AAA | Race 1 | R | Embrapa cassava and fruit growing, Brazil [78], DAFF, Australia [84] | Brazil, Africa | [78,84] |
| BRS Vitória | AAAB | Race 1 | R | Embrapa cassava and fruit growing, Brazil | Brazil | [107] |
| Ambei | AA | Race 1 | R | Embrapa cassava and fruit growing, Brazil | In breeding programs | [78] |
| Walebo | AAA | Race 1 | R | Embrapa cassava and fruit growing, Brazil | Brazil | [78] |
| Kongo FRF 1286 | AAA | Race 1 | R | Embrapa cassava and fruit growing, Brazil | Brazil | [78] |
| Pisang Nangka | AAB | Race 1 | R | Embrapa cassava and fruit growing, Brazil | Brazil, Africa, Australia | [78] |
| Pisang Jaran | AA | Race 1 | R | Embrapa cassava and fruit growing, Brazil | In breeding programs | [78] |
| Tjau Lagada | AA | Race 1 | R | Embrapa cassava and fruit growing, Brazil [33,78] | In breeding programs | [33,78] |
| Mangana | AA | Race 1 | R | Embrapa cassava and fruit growing, Brazil | In breeding programs | [78] |
| Pisang Pipit | AAA | Race 1 | R | Embrapa cassava and fruit growing, Brazil | In breeding programs | [78] |
| Pisang Rojo Uter | AA | Race 1 | R | Embrapa cassava and fruit growing, Brazil | In breeding programs | [78] |
| 2803-01 | AA | Race 1 | R | Embrapa cassava and fruit growing, Brazil | In breeding programs | [78] |
| GN. P. Formoso | AAA | Race 1 | R | Embrapa cassava and fruit growing, Brazil | In breeding programs | [109] |
| Pisang Tongat | AA | Race 1 | R | Embrapa cassava and fruit growing, Brazil | In breeding programs | [78] |
| Mchare cultivars | AA | Race 1 | R | Stellenbosch University, South Africa (Arusha, Tanzania) | Africa | [112] |
| Mchare hybrids | AA | Race 1 | R | Stellenbosch University, South Africa (Arusha, Tanzania) | Africa | [112] |
| NARITA hybrids | AA | Race 1 | R | Stellenbosch University, South Africa (Kawanda, Uganda) | Africa | [112] |
| Figo Cinza | ABB | Race 1 | R | Embrapa cassava and fruit growing, Brazil [78] Banana Germplasm Bank of the Itajaí Research Station [111] | Brazil | [78,111] |
| M-61 | AAA | Race 1 | R | Embrapa cassava and fruit growing, Brazil | In breeding programs | [78] |
| Nanicão Magario | AAA | Race 1 | R | Embrapa cassava and fruit growing, Brazil | Brazil | [78] |
| Buitenzorg | AA | Race 1 | R | Embrapa cassava and fruit growing, Brazil | In breeding programs | [78] |
| BRS Platina | AAAB | Race 1 | R | Embrapa cassava and fruit growing, Brazil [33,78,107] Itajaí Research Station, Brazil [111] | Brazil | [33,78,107,111] |
| Nanica | AAA | Race 1 | R | Embrapa cassava and fruit growing, Brazil | Brazil | [78,107,109] |
| Pisang Ustrali | AAB | Race 1 | R | Embrapa cassava and fruit growing, Brazil | In breeding programs | [78] |
| Markatooa | AAA | Race 1 | R | Embrapa cassava and fruit growing, Brazil | In breeding programs | [78] |
| Robusta | AAA | Race 1 | R | Embrapa cassava and fruit growing, Brazil | In breeding programs | [78] |
| BRS Pacovan Ken | AAAB | Race 1 | R | Embrapa cassava and fruit growing, Brazil | Brazil | [78,107] |
| BRS Princesa | AAAB | Race 1 | R | Federal Institute of the Triangulo Mineiro, Brazil [74], Embrapa cassava and fruit growing, Brazil [33,107] | Brazil | [33,81,107] |
| BRS Japira | AAAB | Race 1 | R | Federal Institute of the Triangulo Mineiro, Brazil [81] Embrapa cassava and fruit growing, Brazil [107] | Brazil | [81,107] |
| BRS Tropical | AAAB | Race 1 | R | Federal Institute of the Triangulo Mineiro, Brazil | Brazil | [81] |
| Grand Naine | AAA | Race 1 | R | Embrapa cassava and fruit growing, Brazil [33,78,107,109], Federal University of Santa Catarina, Brazil [111] | Cavendish for export | [33,78,107,109,111] |
| Nanicão | AAA | Race 1 | R | Embrapa cassava and fruit growing, Brazil [78,111], Federal University of Santa Catarina, Brazil [111] | Brazil | [78,109,111] |
| SCS452 Corupá | AAA | Race 1 | R | Federal University of Santa Catarina, Brazil | Brazil | [111] |
| Zellig | AAA | Race 1 | R | Federal University of Santa Catarina, Brazil | Brazil | [111] |
| Figo | ABB | Race 1 | R | Embrapa cassava and fruit growing, Brazil [78], Federal University of Santa Catarina, Brazil [111] | In breeding programs | [78,111] |
| FHIA-17 | AAAA | Race 1 | R | DAFF, Australia | Honduras, Brazil | [84] |
| SH-3640.10 | AAAB | Race 1 | R | DAFF, Australia | Honduras, Brazil, Mozambique, Cameroon | [84] |
| Long Tavoy | * | Race 1 | R | University of Malaya, Kuala Lumpur, Malaysia | In breeding programs | [31] |
| Kasaska | * | Race 1 | R | University of Malaya, Kuala Lumpur, Malaysia | In breeding programs | [31] |
| Monyet | * | Race 1 | R | University of Malaya, Kuala Lumpur, Malaysia | In breeding programs | [31] |
| Mwitu Pemba | * | Race 1 | R | University of Malaya, Kuala Lumpur, Malaysia | In breeding programs | [31] |
| Hom Thong Mokho | AAA | Race 1 | R | Department of Agriculture and Fisheries (DAF), Queensland, Australia | Australia | [101] |
| Mambee Thu | AA | Race 1 | R | Embrapa cassava and fruit growing, Brazil | In breeding programs | [78] |
| PV03-79 | AAAB | Race 1 | R | Embrapa cassava and fruit growing, Brazil | In breeding programs | [78] |
| Terra Maranhão | AAB | Race 1 | R | Embrapa cassava and fruit growing, Brazil | Brazil | [107] |
| Williams | AAA | Race 1 | R | DAFF, Australia [101], Federal University of Santa Catarina, Brazil [111] | Cavendish for export | [101,111] |
| Williams | AAA | STR4 | SS | University of Queensland, Australia | Cavendish for export | [72] |
| SH-3217 | AA | STR4 | R | University of Queensland, Australia | In breeding programs | [72] |
| Ma250 | AA | STR4 | R | University of Queensland, Australia | In breeding programs | [72] |
| Pisang Bangkahulu | AA | STR4 | R | University of Queensland, Australia | In breeding programs | [72] |
| M61 Guadelope | * | STR4 | SS | University of Queensland, Australia | In breeding programs | [72] |
| CAM-020 | AAA | STR4 | S | University of Queensland, Australia | In breeding programs | [72] |
| SH-3142 | AA | TR4 | SS | (IFTR-GDAAS), Guangzhou, China | In breeding programs | [77] |
| FHIA-1 (“Gold Finger”) | AAAB | TR4 | S | GDAAS, Guangzhou, China | Australia, Brazil, Mexico, Colombia, EUA | [75] |
| GCTCV-119 | AAA | TR4 | HR | Guangdong Academy of Agricultural Sciences, Guangzhou, China | China, Taiwan, The Philippines, Mozambique. | [92] |
| M61 Guadeloupe | * | TR4 | R | University of Queensland, Australia | In breeding programs | [72] |
| CAM-020 | AAA | TR4 | R | University of Queensland, Australia | In breeding programs | [72] |
| Ibwi E | AAA | TR4 | R | (IFTR-GDAAS), Guangzhou, China | EAHBs | [77] |
| Igitsiri (Intuntu) | AAA | TR4 | R | (IFTR-GDAAS), Guangzhou, China | EAHBs | [77] |
| Ingagara | AAA | TR4 | R | (IFTR-GDAAS), Guangzhou, China | EAHBs | [77] |
| Inkira | AAA | TR4 | R | (IFTR-GDAAS), Guangzhou, China | EAHBs | [77] |
| Intokatoke | AAA | TR4 | R | (IFTR-GDAAS), Guangzhou, China | EAHBs | [77] |
| Kazirakwe | AAA | TR4 | R | (IFTR-GDAAS), Guangzhou, China | EAHBs | [77] |
| Mbwazirume | AAA | TR4 | HR | (IFTR-GDAAS), Guangzhou, China | Africa | [77] |
| Akpakpak | AAB | TR4 | HR | (IFTR-GDAAS), Guangzhou, China | Africa | [77] |
| Curaré Enano | AAB | TR4 | R | (IFTR-GDAAS), Guangzhou, China | Africa | [77] |
| Obino l’Ewai | AAB | TR4 | R | (IFTR-GDAAS), Guangzhou, China | Africa | [77] |
| Obubit Ntanga | AAB | TR4 | R | (IFTR-GDAAS), Guangzhou, China | Africa | [77] |
| Orishele False Horn | AAB | TR4 | HR | (IFTR-GDAAS), Guangzhou, China | Africa | [77] |
| Pisang Ceylan | AAB | TR4 | R | (IFTR-GDAAS), Guangzhou, China | Africa | [77] |
| Pisang Rajah | AAB | TR4 | R | (IFTR-GDAAS), Guangzhou, China | Africa | [77] |
| Musa itinerans | * | TR4 | HR | (IFTR-GDAAS), Guangzhou, China | In breeding programs | [77] |
| CIRAD930/DH Pahang | AA | TR4 | HR | (IFTR-GDAAS), Guangzhou, China | In breeding programs | [77] |
| NBA 14 | AA | TR4 | R | (IFTR-GDAAS), Guangzhou, China | In breeding programs | [77] |
| Banksii | AA | TR4 | R | (IFTR-GDAAS), Guangzhou, China | In breeding programs | [77] |
| Maia Oa | AA | TR4 | R | (IFTR-GDAAS), Guangzhou, China | In breeding programs | [77] |
| Zebrina | AA | TR4 | SS | (IFTR-GDAAS), Guangzhou, China | In breeding programs | [77] |
| Pa (Rayong) | AA | TR4 | R | (IFTR-GDAAS), Guangzhou, China | In breeding programs | [77] |
| Figue Rose | AA | TR4 | R | (IFTR-GDAAS), Guangzhou, China | In breeding programs | [77] |
| Khai (Kampengpeth) | AA | TR4 | R | (IFTR-GDAAS), Guangzhou, China | In breeding programs | [77] |
| Tani | BB | TR4 | R | (IFTR-GDAAS), Guangzhou, China | In breeding programs | [77] |
| Pisang Klutuk Wulung | BB | TR4 | R | (IFTR-GDAAS), Guangzhou, China | In breeding programs | [77] |
| Musa beccarii Callimusa | * | TR4 | R | (IFTR-GDAAS), Guangzhou, China | In breeding programs | [77] |
| Musa laterita Rhodochlamys | * | TR4 | R | (IFTR-GDAAS), Guangzhou, China | In breeding programs, | [77] |
| Musa maclayi ssp. | * | TR4 | R | (IFTR-GDAAS), Guangzhou, China | In breeding programs, | [77] |
| Khai Thong Ruang | AAA | TR4 | R | (IFTR-GDAAS), Guangzhou, China | In breeding programs | [77] |
| Kamaramasenge | AB | TR4 | R | (IFTR-GDAAS), Guangzhou, China | In breeding programs | [77] |
| Rukumamb | AAB | TR4 | R | (IFTR-GDAAS), Guangzhou, China | Australia, Papua New Guinea | [77] |
| Thap Maeo | AAB | TR4 | R | (IFTR-GDAAS), Guangzhou, China | Brazil, Honduras | [77] |
| Foconah | AAB | TR4 | R | (IFTR-GDAAS), Guangzhou, China | In breeding programs | [77] |
| Poingo | AAB | TR4 | R | (IFTR-GDAAS), Guangzhou, China | In breeding programs | [77] |
| FHIA-21 | AAAB | TR4 | R | (IFTR-GDAAS), Guangzhou, China [77], DAFF, Australia [84] | In breeding programs | [77,84] |
| Blue Java | ABB | TR4 | R | (IFTR-GDAAS), Guangzhou, China [77], Embrapa cassava and fruit growing, Brazil [107] | China, Africa, Brazil | [77,107] |
| Namwa Khom | ABB | TR4 | HR | (IFTR-GDAAS), Guangzhou, China [77], DAF, Australia [101] | China, Africa, Thailand | [77,101] |
| FHIA-02 | AAAA | TR4 | R | DAFF, Australia | Africa, Brazil, Colombia, Honduras | [72,84] |
| SH-3362 (“Pita-16”) | * | TR4 | R | DAFF, Australia | In breeding programs | [72] |
| M. yunnanensis | * | TR4 | R | South China Agricultural University | Wild germplasm | [75] |
| M. basjoo | * | TR4 | R | South China Agricultural University | Wild germplasm | [75] |
| M. nagensium | * | TR4 | R | South China Agricultural University | Wild germplasm | [75] |
| M. ruiliensis | * | TR4 | R | South China Agricultural University | Wild germplasm | [75] |
| M. velutina | * | TR4 | R | South China Agricultural University | Wild germplasm | [75] |
| Nantianqing | AAA | TR4 | MR | Dongguan Banana Vegetable Institute, China | China | [51] |
| Dongjiao 1 | AAA | TR4 | MR | Dongguan Banana Vegetable Institute, China | China | [51] |
| Kangku 1 | AAA | TR4 | R | Dongguan Banana Vegetable Institute, China | China | [51] |
| G6-2 | AAA | TR4 | R | Dongguan Banana Vegetable Institute, China | China | [51] |
| Yueke 1 | AAA | TR4 | MR | Dongguan Banana Vegetable Institute, China | China | [51] |
| Nongke 1 | AAA | TR4 | MR | Dongguan Banana Vegetable Institute, China | China | [51] |
| Kangku 5 | AAA | TR4 | HR | Dongguan Banana Vegetable Institute, China | China | [51] |
| Nantianhuang | AAA | TR4 | MR | Dongguan Banana Vegetable Institute, China | China | [51] |
| BXM51 | AAA | TR4 | MR | Dongguan Banana Vegetable Institute, China | China | [51] |
| Yueyoukang 1 | AAA | TR4 | R | South China Agricultural University | China | [113] |
| Pisang Gajih Merah | AAA | TR4 | SS | University of Queensland, Australia | Australia | [72] |
| GCTCV-218 Formosana | AAA | TR4 | R | University of Queensland, Australia and Northern Mozambique | China, Taiwan, Philippines and Mozambique. | [5,72] |
| FHIA-01 (“Goldfinger”) | AAAB | Race 1/STR4 | R | DAFF, Australia [84], FHIA, Honduras [93] | Africa, Australia, Honduras | [84,93] |
| Tuu Gia | AA | Race 1/TR4 | HR | (IFTR-GDAAS), Guangzhou, China | In breeding programs | [77] |
| Pisang Lilin | AA | Race 1/TR4 | R | (IFTR-GDAAS), Guangzhou, China | In breeding programs | [77] |
| Borneo | AA | Race 1/TR4 | R | National Agricultural Research Laboratories (NARL) [31] (IFTR-GDAAS), Guangzhou, China [80] and Wageningen University and Research, Wageningen, Netherlands [102] | In breeding programs | [31,77,102] |
| Pisang Berlin | AA | Race 1/TR4 | R | (IFTR-GDAAS), Guangzhou, China [77], Embrapa cassava and fruit growing, Brazil [78] | In breeding programs | [77,78] |
| Zebrina GF | * | Race 1/TR4 | R | University of Malaya, Kuala Lumpur, Malaysia [31], IFTR-GDAAS, Guangzhou, China [77] | In breeding programs | [31,77] |
| Pahang | AA | Race 1/STR4/TR4 | HR | University of Queensland, Australia [72], Yunnan Agricultural University, Kunming, China [74,114] and IFTR-GDAAS, Guangzhou, China [77] | In breeding programs | [72,74,77,114] |
| Calcutta-4 | AA | Race 1/STR4/TR4 | HR | University of Queensland, Australia [66] and (IFTR-GDAAS), Guangzhou, China [72] | In breeding programs | [72,77] |
| Ma851 | AA | STR4/TR4 | R | University of Queensland, Australia | In breeding programs | [72] |
| Ma852 | AA | STR4/TR4 | R | University of Queensland, Australia | In breeding programs | [72] |
| Calcutta4-IV9 | AA | STR4/TR4 | R | University of Queensland, Australia [66] and IFTR-GDAAS, Guangzhou, China [72] | In breeding programs | [72,77] |
| SH-3362 | AA | STR4/TR4 | R | University of Queensland, Australia | In breeding programs | [72] |
| SH-3142 | AA | STR4/TR4 | R | University of Queensland, Australia | In breeding programs | [72] |
| Madang Guadeloupe | AA | STR4/TR4 | R | University of Queensland, Australia | In breeding programs of | [72] |
| FHIA-1 (“Gold Finger”) | AAAB | STR4/TR4 | R | University of Queensland, Australia | Australia, Brazil, Mexico, Colombia, EUA | [72] |
| FHIA-25 | AAB | STR4/TR4 | R | University of Queensland, Australia [72], (IFTR-GDAAS), Guangzhou, China [77], Wageningen University and Research, Wageningen, Netherlands [102] | Africa, Latin America and Australia (Honduras, Colombia, Brazil, Jamaica, Mozambique) | [72,77,102] |
| GCTCV-119 | AAA | STR4/TR4 | R | University of Queensland, Australia and Northern Mozambique | China, Taiwan, The Philippines, Mozambique | [5,72] |
| Ma850 | AA | ST4/TR4 | R | University of Queensland, Australia | In breeding programs | [72] |
| Pisang Jari Buaya | AA | STR4/TR4 | R | University of Queensland, Australia [72] and (IFTR-GDAAS), Guangzhou, China [77] | In breeding programs | [72,77] |
| FHIA-18 | AAAB | STR4/TR4 | R | University of Queensland, Australia [72], IFTM Brazil [81], DAFF, Australia [84], Federal University of Santa Catarina, Brazil [111] | Africa, Latin America and Australia (Honduras, Colombia, Jamaica, Mozambique) | [72,81,84,111] |
| Article | Banana Variety | Plant Growth Stage | Race | Sampling (after Infection) | Pathways Enriched for Differentially Expressed Genes |
|---|---|---|---|---|---|
| Wang et al. [115] | Banana “Brazil” (susceptible) and “Formosana” (tolerant) | 4.5 months | FOC TR4 | 48 h | Flavonoid biosynthesis, flavone and flavonol biosynthesis, alpha-linolenic acid metabolism, starch and sucrose metabolism and phenylpropanoid biosynthesis. |
| Wang et al. [116] | Banana “Brazil” | 60 d | FOCTR4 | 0, 2, 4, 6 days | Phenylalanine metabolism, phenylpropanoid biosynthesis, drug metabolism—cytochrome P450, alpha-linolenic acid metabolism, amino sugar and nucleotide sugar metabolism. |
| Li et al. [37] | Banana “Brazil” | 50 d | FOC 1 and FOC TR4 | 3, 27, 51 h | PR proteins, phytoalexins and phenylpropanoid synthesis, cell wall modifications, biosynthesis via ethylene signaling. |
| Li et al. [117] | Banana “Brazil” and “Nongke Nº 1” (resistant) | Plants with four or five leaves | FOC TR4 | 48, 96 h | Perception of PAMP by PRRs, hormone biosynthesis and signaling, transcription factors, cell wall modification, flavonoid biosynthesis, programmed cell death, PR proteins |
| Bai et al. [113] | Banana “Brazil” and “Yueyoukang 1” resistant | 8 weeks (plants with five leaves) | FOC TR4 | 0, 5, 1, 3, 5, 10 days | PR proteins, transcription factors, cell wall modification, phenylpropanoid biosynthesis, plant hormone signal transduction. |
| Zhang et al. [114] | Musa acuminata Pahang and Brazilian | FOC STR4 | at 14 days | PR proteins, transcription factors, cell wall modification, phenylpropanoid biosynthesis, plant hormone signal transduction. | |
| Sun et al. [32] | Musa acuminata ” Guijiao 9” and Williams | 6 months | FOC TR4 | At 6 days | Membrane-bound intracellular organelle, cell wall and cytoplasm, ions, transcription factor and oxidoreductase activity, plant–pathogen interaction, plant hormone signal transduction, phenylpropanoid biosynthesis and flavonoid biosynthesis. |
| Fei et al. [118] | Cavendish banana | 3 months | FOC 1 and FOC TR4 | At 28 days | Cell components, molecular function and biological process. |
| Cheng et al. [103] | Musa acuminata cv. Tianbaojiao | 11 weeks | FOC TR4 | 5, 10, 25 h | Auxin-activated signaling pathway, cellular response, auxin stimulation, phenylpropanoid catabolic process, lignin catabolic process, lignin metabolic process, via peroxisomes. |
| Song et al. [119] | Brazilian banana and señorita banana | In the five-leaf stage | FOC 1 and FOC TR4 | In the five-leaf stage | Cellular process, metabolic process and binding of organelles and nucleic acids or proteins, regulation of biological processes and transcription factors. |
| Li et al. [120] | Cavendish banana and Brazilian (BX) | 90 days | FOC TR4 | 27 h, 51 h | Secondary metabolite biosynthesis, plant–pathogen interaction, phenylpropanoid biosynthesis and phenylalanine metabolism, fatty acid metabolism, glycerolipid and glycerophospholipid metabolism |
| Niu et al. [121] | Yueyoukang1 and Baxijiao | 2 weeks | FOC TR4 | 24 h | Cell wall biosynthesis and degradation, cell polysaccharide metabolic process, chitinase activity, pectinesterase activity and xyloglucan activity, fructose and mannose metabolism, sphingolipid metabolism, butanoate metabolism, porphyrin and chlorophyll metabolism, carotenoid and ribosome biosynthesis. |
| Hybrids | Parentage |
|---|---|
| Article | |
| Ssali et al. [93] | |
| F2 progenies | Diploid TMB2X8075 (“SH3362” (AA) × “Calcutta 4” (AA) × Sukali Ndizi (AAB) |
| Arinaitwe et al. [31] | |
| F1 progenies | Monyet (Musa acuminata ssp. Zebrina) × Kokopo (Musa acuminata ssp. Banksii) |
| Ahmad et al. [108] | |
| Musa acuminata ssp. Malaccensis (selfed) | |
| Gonçalves et al. [33] | |
| CNPMF0038 | ((M53 × Madu)) × ((Malaccensis × Tjau Lagada)) |
| CNPMF0496 | ((M61 × Pisang Lilin)) × ((Terrinha × Calcutta 4)) |
| CNPMF0513 | ((M61 × Pisang Lilin)) × ((M53 × Kumburgh)) |
| CNPMF0519 | Self-fertilization (wild diploid Tambi) |
| CNPMF0534 | ((Calcutta 4 × Madang)) × ((Borneo × Guyod)) |
| CNPMF0536 | ((Calcutta 4 × Madang)) × ((Borneo × Guyod)) |
| CNPMF0542 | ((SH3263)) × ((Malaccensis × Sinwobogi)) |
| CNPMF0557 | ((M61 × Pisang Lilin)) × ((Malaccensis × Tjau Lagada)) |
| CNPMF0565 | ((Calcutta 4 × Pahang) × (Borneo × Madang)) × Khae |
| CNPMF0572 | ((Khai × (Calcutta 4 × Madang)) × ((Calcutta 4 × Madang)) |
| CNPMF0612 | ((M53 × Madu) × Madu)) × SH3263 |
| CNPMF0731 | ((Malaccensis × Madang)) × ((Tuugia × Calcutta 4)) |
| CNPMF0767 | ((Malaccensis × Madang)) × ((Khai × (Calcutta 4 × Madang)) |
| CNPMF0811 | ((Khai × (Calcutta 4 × Madang)) × ((Calcutta 4 × Pahang) × (Borneo × Madang)) |
| CNPMF0978 | ((Calcutta 4 × Madang)) × ((Terrinha × Calcutta 4)) |
| CNPMF0993 | ((Borneo × Guyod) × (Tuugia × Calcutta 4)) × ((Khai × (Calcutta 4 × Madang)) |
| CNPMF0998 | ((Borneo × Guyod)) × ((Borneo × Guyod) × SH3263) |
| CNPMF1102 | ((Jari Buaya × (Calcutta 4 × Madang)) × ((Borneo × Guyod) × (Tuugia × Calcutta 4)) |
| CNPMF1105 | ((Borneo × Guyod) × (Calcutta 4 × Heva)) × ((Calcutta 4 × Madang)) |
| CNPMF1171 | ((Malaccensis × Madang)) × ((M53 × (Tuugia × Calcutta 4)) |
| CNPMF1272 | ((Borneo × Guyod) × (Calcutta 4 × Heva)) × ((Tuugia × Calcutta 4)) |
| CNPMF1286 | ((Calcutta 4 × Madang)) × ((Terrinha × Calcutta 4)) |
| CNPMF1323 | ((Malaccensis × Sinwobogi)) × ((Calcutta 4 × Heva)) |
| CNPMF0241 | ((Pacovan × improved diploid)) |
| CNPMF0282 | ((Pacovan × improved diploid)) |
| CNPMF0351 | ((Prata Anã × improved diploid by FHIA)) |
| CNPMF0897 | ((Prata Anã)) × ((Malaccensis × Sinwobogi) × (Zebrina × Heva)) |
| CNPMF0898 | ((Prata Anã)) × ((Malaccensis × Sinwobogi) × (Calcutta 4 × Galeo)) |
| CNPMF0906 | ((Prata Santa Maria × improved diploid)) |
| CNPMF0908 | ((Silk × improved diploid)) |
| BRS Princesa | ((Yangambi × M53)) |
| Gene | Sources | Function | Banana Cultivar | References |
|---|---|---|---|---|
| Ferredoxin (Atfd3) and ferredoxin-like protein (pflp) | Capsicum annuum | Antimicrobial peptide | cv. Pei Chiao (AAA) | [69] |
| Petunia floral defenses (PhDef1 and PhDef2) | Petunia hybrida | Antimicrobial peptide | cv. Rasthali (AAB) | [124] |
| Onion—Ace-AMP1 | Allium cepa | Antimicrobial peptide | cv. Rasthali (AAB) | [88] |
| Endochitinase (chit42) | Trichodermaharzianum | Antifungal activity | cv. Furenzhi (AA) | [125] |
| Defensin (Sm-AMP-D1) | Stellaria media | Antimicrobial peptide | cv. Rasthali (AAB) | [126] |
| Small interfering RNAs(siRNAs)/(ihpRNA) | _ | Silencing of vital fungal genes | cv. Rasthali (AAB) | [83] |
| (MusaDAD1, MusaBAG1 eMusaBI1) | Musa acuminata | Cell death is highly induced by FOC infection | cv. Rasthali (AAB) | [96] |
| Cell death (Bcl-Xl, Ced-9 e Bcl-23) | Caenorhabditiselegans | Antiapoptosis | cv. Grand Naine | [89] |
| Cell death (Ced9) | Caenorhabditiselegans | Antiapoptosis | cv. Sukali Ndizi (Musa ssp. AAB) | [100] |
| Pathogenesis-reported (MaPR-10) | Musa acuminata ssp. malaccensis | Pathogenesis (PR) | M. acuminata cv. Berangan | [73] |
| (RGA2) and (Ced9) | Musa acuminata ssp. malaccensis /Caenorhabditis elegans | Resistance analog/antiapoptosis | cv. Grand Naine | [127] |
| Chitinases and 1.3-glucanase | Oryza sativa | Disease tolerance | cv. Rasthali (AAB) | [122] |
| Synthesis of ergosterol (ERG6) | _ | Silencing of vital fungal genes | Cavendish | [128] |
| Small interfering RNAs–ihpRNA | _ | Silencing of vital fungal genes | cv. Rasthali (AAB) | [129] |
| Inductor | Application | References |
|---|---|---|
| Bacillus subtilis | Inoculation of plants with suspension in a greenhouse | [130] |
| Trichoderma asperellum | Inoculation of plants with suspension in a greenhouse | [131] |
| Abscisic acid (ABA), ethephon, methyl jasmonate (MeJA) and salicylic acid (SA) | Root treatment with inductor solutions | [132] |
| Penicillium citrinum | Inoculation of plants with suspension in a greenhouse | [88] |
| Bacillus subtilis | Treatment with in vitro fermented culture filtrate and inoculation of plants with suspension in a greenhouse | [90] |
| Benzothiadiazole (BTH) | Spraying leaves and roots | [133] |
| Interaction with dead FOC pathogen | Inoculation of plants with suspension in a greenhouse | [134] |
| Methyl jasmonate (MeJA) | Exogenous solution treatment in soil and leaves | [76] |
| A strain of FOC 1 incompatible with inducing resistance against the tropical race 4 TR4 | Systemic resistance acquired by in vitro inoculation | [99] |
| Isolates of Trichoderma spp. (T. koningii, T. viride, T. harzianum) | Biomass, liquid culture and culture filtrate | [135] |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Rocha, A.d.J.; Soares, J.M.d.S.; Nascimento, F.d.S.; Santos, A.S.; Amorim, V.B.d.O.; Ferreira, C.F.; Haddad, F.; Santos-Serejo, J.A.d.; Amorim, E.P. Improvements in the Resistance of the Banana Species to Fusarium Wilt: A Systematic Review of Methods and Perspectives. J. Fungi 2021, 7, 249. https://doi.org/10.3390/jof7040249
Rocha AdJ, Soares JMdS, Nascimento FdS, Santos AS, Amorim VBdO, Ferreira CF, Haddad F, Santos-Serejo JAd, Amorim EP. Improvements in the Resistance of the Banana Species to Fusarium Wilt: A Systematic Review of Methods and Perspectives. Journal of Fungi. 2021; 7(4):249. https://doi.org/10.3390/jof7040249
Chicago/Turabian StyleRocha, Anelita de Jesus, Julianna Matos da Silva Soares, Fernanda dos Santos Nascimento, Adriadna Souza Santos, Vanusia Batista de Oliveira Amorim, Claudia Fortes Ferreira, Fernando Haddad, Janay Almeida dos Santos-Serejo, and Edson Perito Amorim. 2021. "Improvements in the Resistance of the Banana Species to Fusarium Wilt: A Systematic Review of Methods and Perspectives" Journal of Fungi 7, no. 4: 249. https://doi.org/10.3390/jof7040249
APA StyleRocha, A. d. J., Soares, J. M. d. S., Nascimento, F. d. S., Santos, A. S., Amorim, V. B. d. O., Ferreira, C. F., Haddad, F., Santos-Serejo, J. A. d., & Amorim, E. P. (2021). Improvements in the Resistance of the Banana Species to Fusarium Wilt: A Systematic Review of Methods and Perspectives. Journal of Fungi, 7(4), 249. https://doi.org/10.3390/jof7040249








