Colletotrichum Species Causing Anthracnose of Citrus in Australia
Abstract
1. Introduction
2. Materials and Methods
2.1. Sample Collection
2.2. Isolate Preparation
2.3. Morphological and Cultural Analyses
2.4. Multigene Phylogenetic Analysis
2.4.1. DNA Extraction, PCR Amplification, and Sequencing
2.4.2. Phylogenetic Analyses
2.5. Pathogenicity Testing
2.5.1. Fruit Bioassay
2.5.2. Leaf Bioassay
2.5.3. Petal Bioassay
3. Results
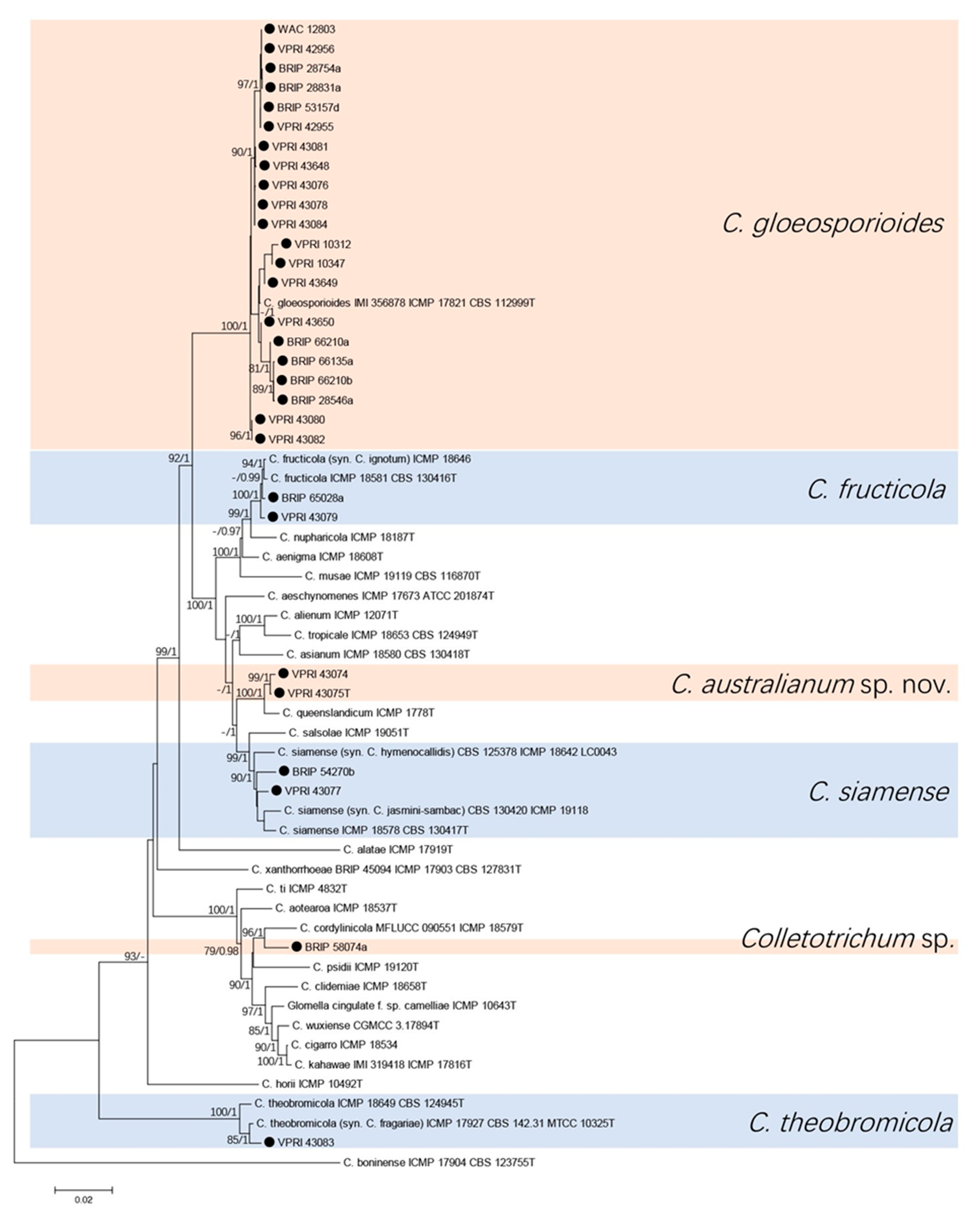
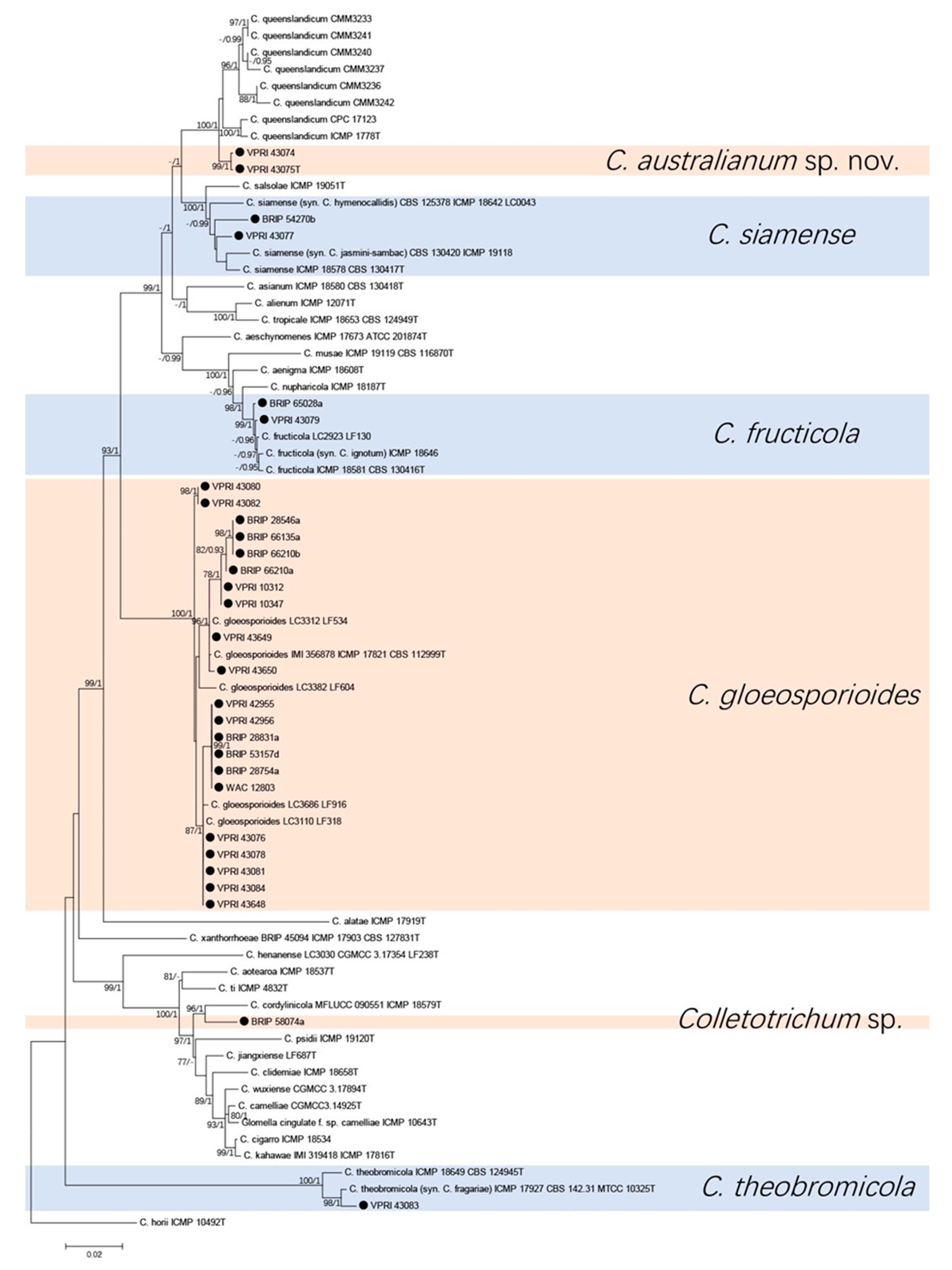
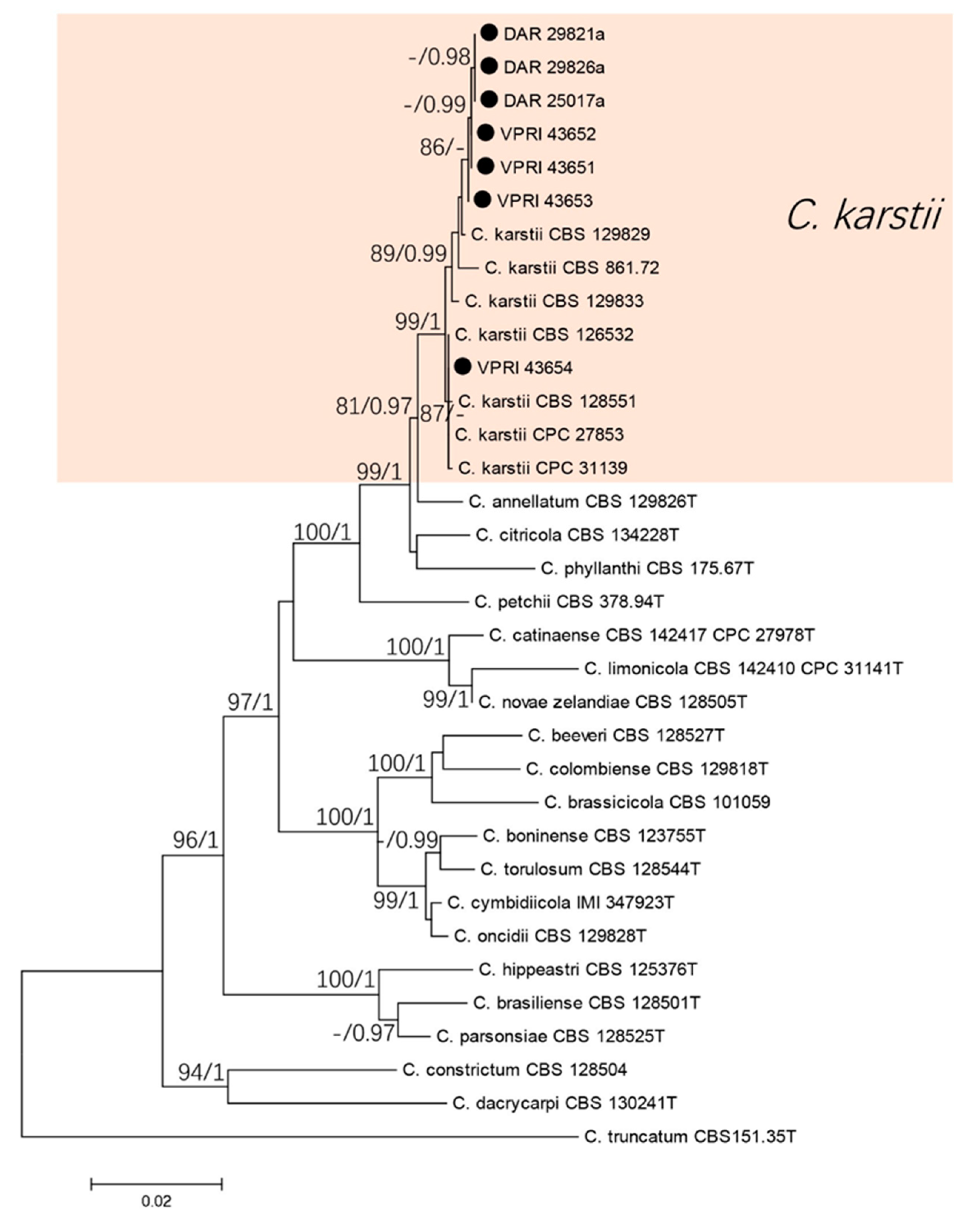
3.1. Phylogenetic Analyses
3.1.1. Gloeosporioides Species Complex
3.1.2. Boninense Species Complex
3.2. Morphological Analysis
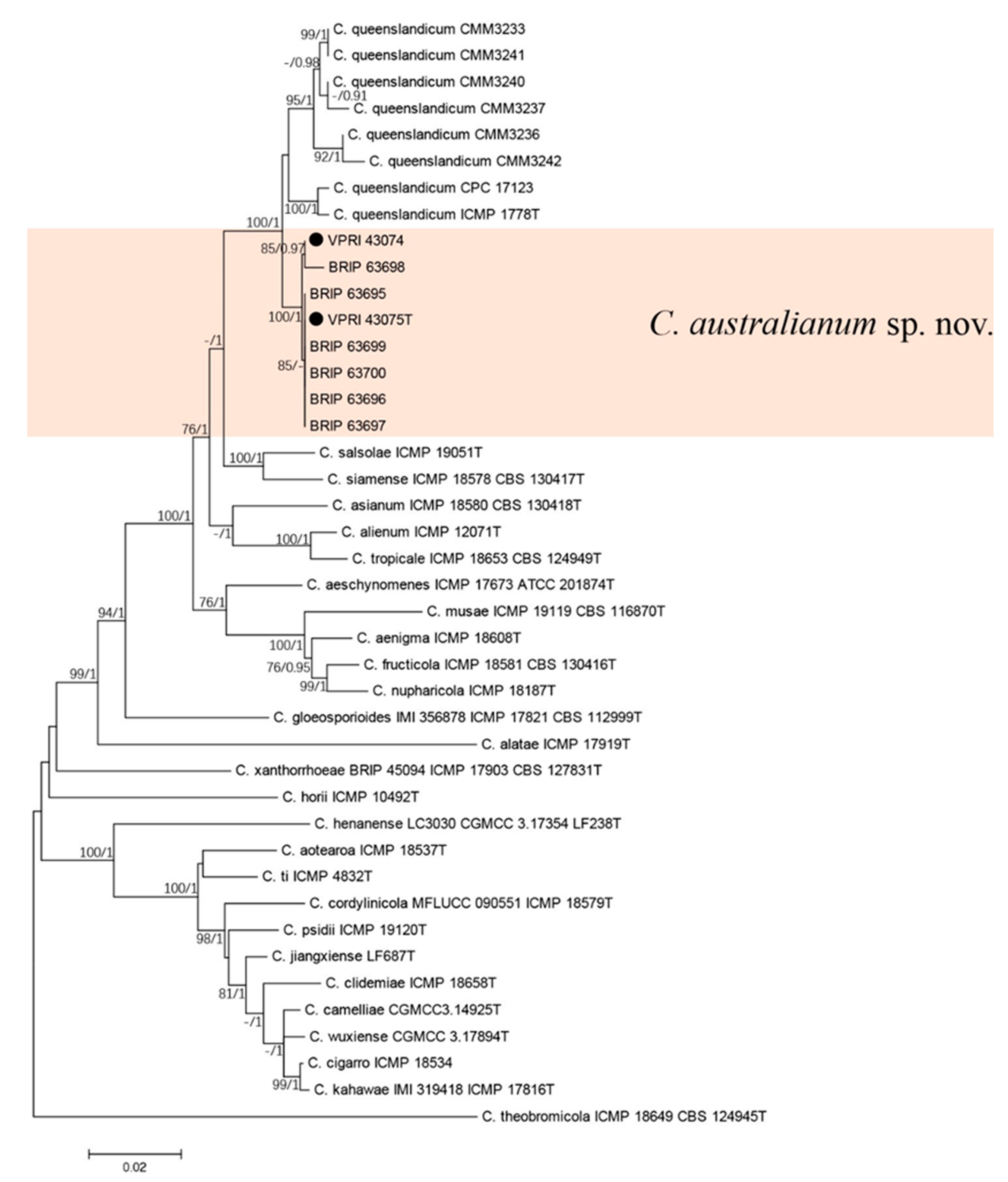
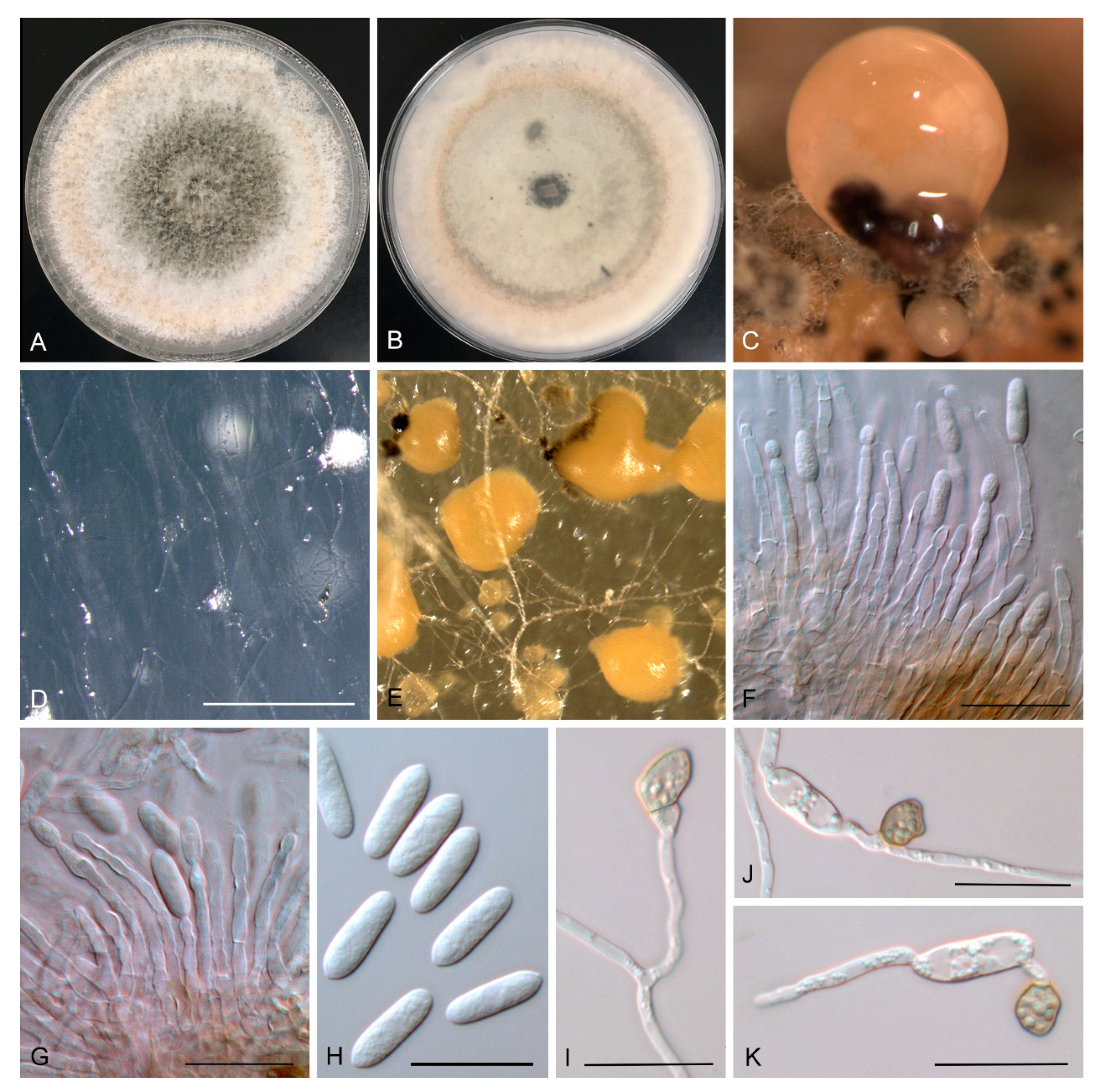
3.3. New Colletotrichum Species
3.3.1. Two-Gene Tree of New Colletotrichum Species
Taxonomy
3.4. Pathogenicity Assay
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Spiegel-Roy, P.; Goldschmidt, E.E. The Biology of Citrus; Cambridge University Press: Cambridge, UK, 2008. [Google Scholar]
- Ladaniya, M. Citrus Fruit; Academic Press: London, UK, 2008. [Google Scholar]
- Fruit and Vegetables. Available online: https://www.austrade.gov.au/search.aspx?ModuleID=8367&keywords=citrus&multiSite=False (accessed on 4 March 2020).
- Cooke, T.; Persley, D.; House, S. Diseases of Fruit Crops in Australia; CSIRO Publishing: Collingwood, ON, Canada, 2009. [Google Scholar]
- Australian Citrus Production Regions. Available online: https://citrusaustralia.com.au/growers-industry (accessed on 4 October 2020).
- The 2018 Annual Report. Available online: https://citrusaustralia.com.au/wp-content/uploads/CAL-Annual-Report-2018-FINAL.pdf (accessed on 4 October 2020).
- Phoulivong, S.; McKenzie, E.H.C.; Hyde, K.D. Cross Infection of Colletotrichum Species; A Case Study with Tropical Fruits. Curr. Res. Environ. Appl. Mycol. 2012, 2, 99–111. [Google Scholar] [CrossRef]
- Guarnaccia, V.; Groenewald, J.Z.; Polizzi, G.; Crous, P.W. High Species Diversity in Colletotrichum Associated with Citrus Diseases in Europe. Pers. Mol. Phylogeny Evol. Fungi 2017, 39, 32–50. [Google Scholar] [CrossRef]
- Marin-Felix, Y.; Groenewald, J.Z.; Cai, L.; Chen, Q.; Marincowitz, S.; Barnes, I.; Bensch, K.; Braun, U.; Camporesi, E.; Damm, U.; et al. Genera of Phytopathogenic Fungi: GOPHY 1. Stud. Mycol. 2017, 86, 99–216. [Google Scholar] [CrossRef] [PubMed]
- de Silva, D.D.; Groenewald, J.Z.; Crous, P.W.; Ades, P.K.; Nasruddin, A.; Mongkolporn, O.; Taylor, P.W.J. Identification, Prevalence and Pathogenicity of Colletotrichum Species Causing Anthracnose of Capsicum Annuum in Asia. IMA Fungus 2019, 10, 1–32. [Google Scholar] [CrossRef] [PubMed]
- Douanla-Meli, C.; Unger, J. Phylogenetic Study of The Colletotrichum Species on Imported Citrus Fruits Uncovers A Low Diversity and A New Species in The Colletotrichum gigasporum Complex. Fungal Biol. 2017, 121, 858–868. [Google Scholar] [CrossRef]
- Yin, L.; Du, S.; Cai, M.; Luo, C. Identification of Anthracnose of Wild Citrus. Southwest China J. Agric. Sci. 2017, 30, 590–594. [Google Scholar]
- Aiello, D.; Carrieri, R.; Guarnaccia, V.; Vitale, A.; Lahoz, E.; Polizzi, G. Characterization and Pathogenicity of Colletotrichum gloeosporioides and C. karstii causing Preharvest Disease on Citrus Sinensis in Italy. J. Phytopathol. 2014, 163, 168–177. [Google Scholar] [CrossRef]
- Benyahia, H.; Jrifia, A.; Smaili, C.; Afellah, M.; Lamsetef, Y.; Timmer, L.W. First Report of Colletotrichum gloeosporioides Causing Withertip on Twigs and Tear Stain on Fruit of Citrus in Morocco. Plant Pathol. 2003, 52, 798. [Google Scholar] [CrossRef]
- Cruz-Lagunas, B.; Ortega-Acosta, S.A.; Reyes-García, G.; Toribio-Jiménez, J.; Juárez-López, P.; Guillén-Sánchez, D.; Damián-Nava, A.; Romero-Ramírez, Y.; Palemón-Alberto, F. Colletotrichum gloeosporioides Causes Anthracnose on Grapefruit (Citrus paradisi) In Mexico. Australas. Plant Dis. Notes 2020, 15, 1–4. [Google Scholar] [CrossRef]
- Pérez-Mora, J.L.; Mora-Romero, G.A.; Beltrán-Peña, H.; García-León, E.; Lima, N.B.; Camacho-Tapia, M.; Tovar-Pedraza, J.M. First Report of Colletotrichum siamense and C. gloeosporioides Causing Anthracnose of Citrus Spp. In Mexico. Plant Dis. 2020. Available online: https://pubmed.ncbi.nlm.nih.gov/32910725/ (accessed on 21 December 2020).
- Fayyaz, A.; Sahi, S.T.; Nawaz-ul-Rehman, M.S.; Amrao, L. Molecular Characterization of Fungal Pathogens Associated with Citrus Withertip/Dieback from Major Citrus Growing Areas of Punjab, Pakistan. Int. J. Agric. Biol. 2020, 23, 1165–1170. [Google Scholar]
- Honger, J.O.; Offei, S.K.; Oduro, K.A.; Odamtten, G.T.; Nyaku, S.T. Identification and Species Status of The Mango Biotype of Colletotrichum gloeosporioides In Ghana. Eur. J. Plant Pathol. 2014, 140, 455–467. [Google Scholar] [CrossRef]
- Honger, J.O.; Offei, S.K.; Oduro, K.A.; Odamtten, G.T.; Nyaku, S.T. Identification and Molecular Characterisation of Colletotrichum species From Avocado, Citrus and Pawpaw in Ghana. S. Afr. J. Plant Soil 2016, 33, 177–185. [Google Scholar] [CrossRef]
- Lima, W.G.; Spósito, M.B.; Amorim, L.; Gonçalves, F.P.; de Filho, P.A.M. Colletotrichum gloeosporioides, A New Causal Agent of Citrus Post-Bloom Fruit Drop. Eur. J. Plant Pathol. 2011, 131, 157–165. [Google Scholar] [CrossRef]
- Mahiout, D.; Bendahmane, B.S.; Benkada, M.Y.; Mekouar, H.; Berrahal, N.; Rickauer, M. First Report of Colletotrichum gloeosporioides on Citrus in Algeria. Phytopathol. Mediterr. 2018, 57, 355–359. [Google Scholar]
- Ramos, A.P.; Talhinhas, P.; Sreenivasaprasad, S.; Oliveira, H. Characterization of Colletotrichum gloeosporioides, As the Main Causal Agent of Citrus Anthracnose, and C. karstii As Species Preferentially Associated with Lemon Twig Dieback in Portugal. Phytoparasitica 2016, 44, 549–561. [Google Scholar] [CrossRef]
- Daoud, H.B.H.; Baraldi, E.; Iotti, M.; Leonardi, P.; Boughalleb-M’hamdi, N. Characterization and Pathogenicity of Colletotrichum Spp. Causing Citrus Anthracnose in Tunisia. Phytopathol. Mediterr. 2019, 58, 175–185. [Google Scholar]
- Rhaiem, A.; Taylor, P.W.J. Colletotrichum gloeosporioides Associated with Anthracnose Symptoms on Citrus, A New Report for Tunisia. Eur. J. Plant Pathol. 2016, 146, 219–224. [Google Scholar] [CrossRef]
- Damm, U.; Cannon, P.F.; Woudenberg, J.H.C.; Johnston, P.R.; Weir, B.S.; Tan, Y.P.; Shivas, R.G.; Crous, P.W. The Colletotrichum boninense Species Complex. Stud. Mycol. 2012, 73, 1–36. [Google Scholar] [CrossRef]
- Peng, L.; Yang, Y.; Hyde, K.D.; Bahkali, A.H.; Liu, Z. Colletotrichum Species on Citrus Leaves in Guizhou And Yunnan Provinces, China. Cryptogam. Mycol. 2012, 33, 267–283. [Google Scholar]
- Huang, F.; Chen, G.Q.; Hou, X.; Fu, Y.S.; Cai, L.; Hyde, K.D.; Li, H.Y. Colletotrichum Species Associated with Cultivated Citrus in China. Fungal Divers. 2013, 61, 61–74. [Google Scholar] [CrossRef]
- Mayorquin, J.S.; Nouri, M.T.; Peacock, B.B.; Trouillas, F.P.; Douhan, G.W.; Kallsen, C.; Eskalen, A. Identification, Pathogenicity, And Spore Trapping of Colletotrichum karstii Associated with Twig and Shoot Dieback in California. Plant Dis. 2019, 103, 1464–1473. [Google Scholar] [CrossRef] [PubMed]
- Uysal, A.; Kurt, Ş. First Report of Colletotrichum karstii Causing Anthracnose on Citrus in The Mediterranean Region of Turkey. J. Plant Pathol. 2019, 101, 753. [Google Scholar] [CrossRef]
- Hu, W.L.; Ma, Y.Z.; Chen, J.Z. First Report of Citrus sinensis Anthracnose Caused by Colletotrichum fructicola in China. Plant Dis. 2019, 103, 1018. [Google Scholar] [CrossRef]
- Cheng, B.P.; Huang, Y.H.; Song, X.B.; Peng, A.T.; Ling, J.F.; Chen, X. First Report of Colletotrichum siamense Causing Leaf Drop and Fruit Spot of Citrus reticulata Blanco cv. Shiyue Ju in China. Plant Dis. 2013, 97, 1508. [Google Scholar] [CrossRef]
- Fayyaz, A.; Bonello, P.; Tufail, M.R.; Amrao, L.; Habib, A.; Gai, Y.; Sahi, S.T. First Report of Citrus Withertip (Tip Dieback), A Disease Complex Caused by Colletotrichum siamense And Lasiodiplodia iraniensis, On Citrus reticulata cv. Kinnow in Punjab, Pakistan. Plant Dis. 2018, 102, 2659. [Google Scholar] [CrossRef]
- Damm, U.; Cannon, P.F.; Woudenberg, J.H.C.; Crous, P.W. The Colletotrichum acutatum Species Complex. Stud. Mycol. 2012, 73, 37–113. [Google Scholar] [CrossRef]
- Kunta, M.; Park, J.W.; Vedasharan, P.; da Graça, J.V.; Terry, M.D. First Report of Colletotrichum queenslandicum on Persian Lime Causing Leaf Anthracnose in The United States. Plant Dis. 2018, 102, 677. [Google Scholar] [CrossRef]
- Perrone, G.; Magistà, D.; Ismail, A.M. First report of Colletotrichum kahawae subsp. Ciggaro on mandarin in italy. J. Plant Pathol. 2016, 98, 682. [Google Scholar]
- Weir, B.S.; Johnston, P.R.; Damm, U. The Colletotrichum gloeosporioides Species Complex. Stud. Mycol. 2012, 73, 115–180. [Google Scholar] [CrossRef]
- Shivas, R.G.; Tan, Y.P.; Edwards, J.; Dinh, Q.; Maxwell, A.; Andjic, V.; Liberato, J.R.; Anderson, C.; Beasley, D.R.; Bransgrove, K.; et al. Colletotrichum Species in Australia. Australas. Plant Pathol. 2016, 45, 447–464. [Google Scholar] [CrossRef]
- Australian Plant Pest Database|Plant Health Australia. Available online: https://www.planthealthaustralia.com.au/resources/australian-plant-pest-database/ (accessed on 22 November 2020).
- De Silva, D.D.; Ades, P.K.; Crous, P.W.; Taylor, P.W.J. Colletotrichum species Associated with Chili Anthracnose in Australia. Plant Pathol. 2016, 66, 254–267. [Google Scholar] [CrossRef]
- Rayner, R.W. A Mycological Colour Chart; Taylor & Francis, Ltd.: Abingdon, UK, 1970. [Google Scholar]
- Johnston, P.R.; Jones, D. Relationships Among Colletotrichum Isolates from Fruit-Rots Assessed Using Rdna Sequences. Mycologia 1997, 89, 420–430. [Google Scholar] [CrossRef]
- Gardes, M.; Bruns, T.D. ITS Primers with Enhanced Specificity for Basidiomycetes—Application to the Identification of Mycorrhizae and Rusts. Mol. Ecol. 1993, 2, 113–118. [Google Scholar] [CrossRef] [PubMed]
- White, T.J.; Bruns, T.D.; Lee, S.B.; Taylor, J.W. Amplification and Direct Sequencing of Fungal Ribosomal RNA Genes for Phylogenetics. PCR Protoc. Guide Methods Appl. 1990, 18, 315–322. [Google Scholar]
- Guerber, J.C.; Liu, B.; Correll, J.C.; Johnston, P.R. Characterization of Diversity in Colletotrichum acutatum sensu lato By Sequence Analysis of Two Gene Introns, Mtdna And Intron Rflps, And Mating Compatibility. Mycologia 2003, 95, 872. [Google Scholar] [CrossRef]
- Carbone, I.; Kohn, L.M. A Method for Designing Primer Sets for Speciation Studies in Filamentous Ascomycetes. Mycologia 1999, 91, 553–556. [Google Scholar] [CrossRef]
- Woudenberg, J.H.C.; Aveskamp, M.M.; de Gruyter, J.; Spiers, A.G.; Crous, P.W. Multiple Didymella Teleomorphs Are Linked to The Phoma Clematidina Morphotype. Pers. Mol. Phylogeny Evol. Fungi 2009, 22, 56–62. [Google Scholar] [CrossRef]
- Silva, D.N.; Talhinhas, P.; Várzea, V.; Cai, L.; Paulo, O.S.; Batista, D. Application of The Apn2/MAT Locus to Improve the Systematics of The Colletotrichum gloeosporioides Complex: An Example from Coffee (Coffea spp.) Hosts. Mycologia 2012, 104, 396–409. [Google Scholar] [CrossRef]
- Stephenson, S.A.; Green, J.R.; Manners, J.M.; Maclean, D.J. Cloning and Characterisation of Glutamine Synthetase From Colletotrichum gloeosporioides And Demonstration of Elevated Expression During Pathogenesis on Stylosanthes guianensis. Curr. Genet. 1997, 31, 447–454. [Google Scholar] [CrossRef]
- Crous, P.W.; Groenewald, J.Z.; Risède, J.M.; Simoneau, P.; Hywel-Jones, N.L. Calonectria Species and Their Cylindrocladium Anamorphs: Species with Sphaeropedunculate Vesicles. Stud. Mycol. 2004, 50, 415–430. [Google Scholar]
- Tamura, K.; Stecher, G.; Peterson, D.; Filipski, A.; Kumar, S. MEGA6: Molecular Evolutionary Genetics Analysis Version 6.0. Mol. Biol. Evol. 2013, 30, 2725–2729. [Google Scholar] [CrossRef]
- Liu, F.; Weir, B.S.; Damm, U.; Crous, P.W.; Wang, Y.; Liu, B.; Wang, M.; Zhang, M.; Cai, L. Unravelling Colletotrichum Species Associated with Camellia: Employing Apmat And GS Loci to Resolve Species in The C. gloeosporioides Complex. Pers. Mol. Phylogeny Evol. Fungi 2015, 35, 63–86. [Google Scholar] [CrossRef]
- Nylander, J.A.A. MrModeltest v. 2. Program Distributed by the Author; Evolutionary Biology Centre, Uppsala University: Uppsala, Sweden, 2004. [Google Scholar]
- Liu, F.; Wang, M.; Damm, U.; Crous, P.W.; Cai, L. Species Boundaries in Plant Pathogenic Fungi: A Colletotrichum Case Study. BMC Evol. Biol. 2016, 16, 81. [Google Scholar] [CrossRef] [PubMed]
- James, R.S.; Ray, J.; Tan, Y.P.; Shivas, R.G. Colletotrichum siamense, C. theobromicola and C. queenslandicum from Several Plant Species and The Identification of C. asianum in The Northern Territory, Australia. Australas. Plant Dis. Notes 2014, 9, 1–6. [Google Scholar] [CrossRef]
- Sharma, G.; Kumar, N.; Weir, B.S.; Hyde, K.D.; Shenoy, B.D. The Apmat Marker Can Resolve Colletotrichum Species: A Case Study with Mangifera Indica. Fungal Divers. 2013, 61, 117–138. [Google Scholar] [CrossRef]
- Sharma, G.; Pinnaka, A.K.; Shenoy, B.D. Resolving the Colletotrichum siamense Species Complex Using Apmat Marker. Fungal Divers. 2014, 71, 247–264. [Google Scholar] [CrossRef]
- Pitt, J.I.; Hocking, A.D. Fungi and Food Spoilage, 3rd ed.; Springer-Verlag: New York, NY, USA, 2009. [Google Scholar]
- Montri, P.; Taylor, P.W.J.; Mongkolporn, O. Pathotypes of Colletotrichum capsici, The Causal Agent of Chili Anthracnose, in Thailand. Plant Dis. 2009, 93, 17–20. [Google Scholar] [CrossRef] [PubMed]
- Talhinhas, P.; Gonçalves, E.; Sreenivasaprasad, S.; Oliveira, H. Virulence Diversity of Anthracnose Pathogens (Colletotrichum acutatum and C. gloeosporioides Species Complexes) On Eight Olive Cultivars Commonly Grown in Portugal. Eur. J. Plant Pathol. 2014, 142, 73–83. [Google Scholar] [CrossRef]
| Species | Accession Number | Host | Location | GenBank Accession Numbers | |||||||
|---|---|---|---|---|---|---|---|---|---|---|---|
| ITS | GAPDH | ACT | TUB2 | gs | ApMat | CHS-1 | HIS3 | ||||
| Gloeosporioides complex | |||||||||||
| C. aenigma | ICMP 18608 * | Persea americana | Israel | JX010244 | JX010044 | JX009443 | JX010389 | JX010078 | KM360143 | JX009774 | – |
| C. aeschynomenes | ICMP 17673; ATCC 201874 * | Aeschynomene virginica | USA | JX010176 | JX009930 | JX009483 | JX010392 | JX010081 | KM360145 | JX009799 | – |
| C. alatae | ICMP 17919 * | Dioscorea alata | India | JX010190 | JX009990 | JX009471 | JX010383 | JX010065 | KC888932 | JX009837 | – |
| C. alienum | ICMP 12071 * | Malus domestica | New Zealand | JX010251 | JX010028 | JX009572 | JX010411 | JX010101 | KM360144 | JX009882 | – |
| C. asianum | ICMP 18580; CBS 130418 * | Coffea arabica | Thailand | FJ972612 | JX010053 | JX009584 | JX010406 | JX010096 | FR718814 | JX009867 | – |
| C. aotearoa | ICMP 18537 * | Coprosma sp. | New Zealand | JX010205 | JX010005 | JX009564 | JX010420 | JX010113 | KC888930 | JX009853 | – |
| C. artocarpicola | MFLUCC 18–1167 * | Artocarpus heterophyllus | Thailand | MN415991 | MN435568 | MN435570 | MN435567 | – | – | MN435569 | – |
| C. australianum | VPRI 43074; UMC001 | Citrus reticulata | Australia, Vic | MG572137 | MG572126 | MK473452 | MG572148 | MG572159 | MG572170 | MW091986 | – |
| VPRI 43075; UMC002 * | Citrus sinensis | Australia, Vic | MG572138 | MG572127 | MN442109 | MG572149 | MG572160 | MG572171 | MW091987 | – | |
| BRIP 63695 | Capsicum annuum | Australia | KU923677 | MN442115 | MN442105 | KU923693 | KU923737 | KU923727 | MW092000 | – | |
| BRIP 63696 | Capsicum annuum | Australia | KU923678 | – | – | KU923694 | KU923738 | KU923728 | – | – | |
| BRIP 63697 | Capsicum annuum | Australia | KU923679 | – | – | KU923695 | KU923739 | KU923729 | – | – | |
| BRIP 63698 | Capsicum annuum | Australia | KU923680 | MN442116 | MN442106 | KU923696 | KU923740 | KU923730 | MW092001 | – | |
| BRIP 63699 | Capsicum annuum | Australia | KU923681 | MN442117 | MN442107 | KU923697 | KU923741 | KU923731 | MW092002 | – | |
| BRIP 63700 | Capsicum annuum | Australia | KU923682 | MN442118 | MN442108 | KU923698 | KU923742 | KU923732 | MW092003 | – | |
| C. camelliae | CGMCC 3.14925 * | Camellia sinensis | China | KJ955081 | KJ954782 | KJ954363 | KJ955230 | KJ954932 | KJ954497 | – | – |
| Glomella cingulate f. sp. camelliae | ICMP 10643 * | Camellia × williamsii | UK | JX010224 | JX009908 | JX009540 | JX010436 | JX010119 | KJ954625 | JX009891 | – |
| C. changpingense | MFLUCC 15-0022 | Fragaria ×ananassa | China | KP683152 | KP852469 | KP683093 | KP852490 | – | – | KP852449 | – |
| C. chrysophilum | CMM4268 * | Musa sp. | Brazil | KX094252 | KX094183 | KX093982 | KX094285 | KX094204 | – | KX094083 | – |
| C. conoides | CAUG17 * | Capsicum annuum | China | KP890168 | KP890162 | KP890144 | KP890174 | – | – | KP890156 | – |
| C. cordylinicola | MFLUCC 090551; ICMP 18579 * | Cordyline fruticosa | Thailand | JX010226 | JX009975 | HM470235 | JX010440 | JX010122 | JQ899274 | JX009864 | – |
| C. clidemiae | ICMP 18658 * | Clidemia hirta | USA, Hawaii | JX010265 | JX009989 | JX009537 | JX010438 | JX010129 | KC888929 | JX009877 | – |
| C. endophytica | CAUG28 | Capsicum annuum | China | KP145441 | KP145413 | KP145329 | KP145469 | – | – | KP145385 | – |
| C. fructicola | ICMP 18581; CBS 130416 * | Coffea arabica | Thailand | JX010165 | JX010033 | FJ907426 | JX010405 | JX010095 | JQ807838 | JX009866 | – |
| LC2923; LF130 | Camellia sinensis | China | KJ955083 | KJ954784 | KJ954365 | KJ955232 | KJ954934 | KJ954499 | – | – | |
| VPRI 43079; UMC006 | Citrus reticulata | Australia, Qld | MG572142 | MG572131 | MK473454 | MG572153 | MG572164 | MG572175 | MW091991 | – | |
| BRIP 65028a; VPRI 43034; B03-43034 | Citrus latifolia | Australia, Qld | MK470007 | MK470025 | MK470097 | MK470061 | MK470043 | MK470079 | MW091983 | – | |
| C. fructicola (syn. C. ignotum) | ICMP 18646 | Tetragastris panamensis | Panama | JX010173 | JX010032 | JX009581 | JX010409 | JX010099 | JQ807839 | JX009874 | – |
| C. fructivorum | CBS 133125 * | Vaccinium macrocarpon | USA | JX145145 | – | – | JX145196 | – | – | – | – |
| C. gloeosporioides | IMI 356878; ICMP 17821; CBS 112999 * | Citrus sinensis | Italy | JX010152 | JX010056 | JX009531 | JX010445 | JX010085 | JQ807843 | JX009818 | – |
| LC3110; LF318 | Camellia sinensis | China | KJ955127 | KJ954828 | KJ954407 | KJ955275 | KJ954978 | KJ954541 | – | – | |
| LC3312; LF534 | Camellia sinensis | China | KJ955158 | KJ954859 | KJ954434 | KJ955305 | KJ955009 | KJ954569 | – | – | |
| LC3382; LF604 | Camellia sinensis | China | KJ955176 | KJ954877 | KJ954450 | KJ955323 | KJ955026 | KJ954584 | – | – | |
| LC3686; LF916 | Camellia sinensis | China | KJ955226 | KJ954927 | KJ954493 | KJ955371 | KJ955076 | KJ954629 | – | – | |
| VPRI 43076; UMC003 | Citrus sinensis | Australia, Vic | MG572139 | MG572128 | MN442110 | MG572150 | MG572161 | MG572172 | MW091988 | – | |
| VPRI 43078; UMC005 | Citrus aurantifolia | Australia, Qld | MG572141 | MG572130 | MN442111 | MG572152 | MG572163 | MG572174 | MW091990 | – | |
| VPRI 43080; UMC007 | Citrus reticulata | Australia, Qld | MG572143 | MG572132 | MK473455 | MG572154 | MG572165 | MG572176 | MW091992 | – | |
| VPRI 43081; UMC008 | Citrus reticulata | Australia, Qld | MG572144 | MG572133 | MN442112 | MG572155 | MG572166 | MG572177 | MW091993 | – | |
| VPRI 43082; UMC009 | Citrus reticulata | Australia, Qld | MG572145 | MG572134 | MN442113 | MG572156 | MG572167 | MG572178 | MW091994 | – | |
| VPRI 43084; UMC011 | Citrus japonica | Australia, Vic | MG572147 | MG572136 | MN442114 | MG572158 | MG572169 | MG572180 | MW091996 | – | |
| VPRI 43648; UMC012 | Citrus sinensis | Australia, Vic | MW081160 | MW081163 | MW081166 | MW081169 | MW081175 | MW081172 | MW091997 | – | |
| VPRI 43649; UMC013 | Citrus limon | Australia, Vic | MW081161 | MW081164 | MW081167 | MW081170 | MW081176 | MW081173 | MW091998 | – | |
| VPRI 43650; UMC014 | Citrus japonica | Australia, Vic | MW081162 | MW081165 | MW081168 | MW081171 | MW081177 | MW081174 | MW091999 | – | |
| VPRI 10312; A01-10312 | Citrus sinensis | Australia, Vic | MK469996 | MK470014 | MK470086 | MK470050 | MK470032 | MK470068 | MW091972 | – | |
| VPRI 10347; A02-10347; BRIP 54771 | Citrus limon | Australia, Vic | MK469997 | MK470015 | MK470087 | MK470051; KU221374 | MK470033 | MK470069 | MW091973 | – | |
| WAC 12803; BRIP 63680a; VPRI 43024; A05-43024 | Citrus sinensis | Australia, WA | MK469998 | MK470016 | MK470088 | MK470052 | MK470034 | MK470070 | MW091974 | – | |
| BRIP 66210a; VPRI 43026; A07-43026 | Citrus reticulata | Australia, SA | MK470000 | MK470018 | MK470090 | MK470054 | MK470036 | MK470072 | MW091976 | – | |
| BRIP 66210b; VPRI 43027; A08-43027 | Citrus reticulata | Australia, SA | MK470001 | MK470019 | MK470091 | MK470055 | MK470037 | MK470073 | MW091977 | – | |
| BRIP 28546a; VPRI 43028; A09-43028 | Citrus sinensis Navel | Australia, Qld | MK470002 | MK470020 | MK470092 | MK470056 | MK470038 | MK470074 | MW091978 | – | |
| BRIP 28754a; VPRI 43030; A11-43030 | Citrus reticulata | Australia, Qld | MK470003 | MK470021 | MK470093 | MK470057 | MK470039 | MK470075 | MW091979 | – | |
| BRIP 53157d; VPRI 43031; A12-43031 | Citrus aurantifolia Tahiti | Australia, Qld | MK470004 | MK470022 | MK470094 | MK470058 | MK470040 | MK470076 | MW091980 | – | |
| BRIP 66135a; VPRI 43032; B01-43032 | Citrus reticulata Imperial Blanco | Australia, Qld | MK470005 | MK470023 | MK470095 | MK470059 | MK470041 | MK470077 | MW091981 | – | |
| BRIP 28831a; VPRI 43033; B02-43033 | Citrus sinensis | Australia, Qld | MK470006 | MK470024 | MK470096 | MK470060 | MK470042 | MK470078 | MW091982 | – | |
| VPRI 42955; G01-42955 | Citrus limon | Australia, NSW | MK470008 | MK470026 | MK470098 | MK470062 | MK470044 | MK470080 | MW091984 | – | |
| VPRI 42956; H01-42956 | Citrus sinensis | Australia, NSW | MK470009 | MK470027 | MK470099 | MK470063 | MK470045 | MK470081 | MW091985 | – | |
| C. grevilleae | CBS 132879 * | Grevillea sp. | Italy | KC297078 | KC297010 | KC296941 | KC297102 | KC297033 | – | KC296987 | – |
| C. grossum | CGMCC3.17614T; CAUG7 * | Capsicum sp. | China | KP890165 | KP890159 | KP890141 | KP890171 | – | – | KP890153 | – |
| CAU31 | Capsicum sp. | China | KP890166 | KP890160 | KP890142 | KP890172 | – | – | KP890154 | – | |
| CAUG32 | Capsicum sp. | China | KP890167 | KP890161 | KP890143 | KP890173 | – | – | KP890155 | – | |
| C. hebeiense | MFLUCC13-0726 * | Vitis vinifera cv. Cabernet Sauvignon | China | KF156863 | KF377495 | KF377532 | KF288975 | – | – | KF289008 | – |
| C. helleniense | CPC 26844; CBS 142418 * | Poncirus trifoliata | Greece | KY856446 | KY856270 | KY856019 | KY856528 | – | – | KY856186 | – |
| C. henanense | LC3030; CGMCC 3.17354; LF238 * | Camellia sinensis | China | KJ955109 | KJ954810 | KM023257 | KJ955257 | KJ954960 | KJ954524 | – | – |
| C. horii | ICMP 10492 * | Diospyros kaki | Japan | GQ329690 | GQ329681 | JX009438 | JX010450 | JX010137 | JQ807840 | JX009752 | – |
| C. hystricis | CPC 28153; CBS 142411 * | Citrus hystrix | Italy | KY856450 | KY856274 | KY856023 | KY856532 | – | – | KY856190 | – |
| C. jiangxiense | LF687 * | Camellia sinensis | China | KJ955201 | KJ954902 | KJ954471 | KJ955348 | KJ955051 | KJ954607 | – | – |
| C. cigarro | ICMP 18534 | Kunzea ericoides | New Zealand | JX010227 | JX009904 | JX009473 | JX010427 | JX010116 | HE655657 | JX009765 | – |
| C. kahawae | IMI 319418; ICMP 17816 * | Coffea arabica | Kenya | JX010231 | JX010012 | JX009452 | JX010444 | JX010130 | JQ894579 | JX009813 | – |
| C. musae | ICMP 19119; CBS 116870 * | Musa sp. | USA | JX010146 | JX010050 | JX009433 | HQ596280 | JX010103 | KC888926 | JX009896 | – |
| ICMP 17817 | Musa sapientum | Kenya | JX010142 | JX010015 | JX009432 | JX010395 | JX010084 | – | JX009815 | – | |
| C. nupharicola | ICMP 18187 * | Nuphar lutea subsp. polysepala | USA | JX010187 | JX009972 | JX009437 | JX010398 | JX010088 | JX145319 | JX009835 | – |
| C. pandanicola | MFLUCC 17-0571 | Pandanaceae | Thailand | MG646967 | MG646934 | MG646938 | MG646926 | – | – | MG646931 | – |
| C. proteae | CBS 132882 * | Protea sp. | South Africa | KC297079 | KC297009 | KC296940 | KC297101 | KC297032 | – | KC296986 | – |
| C. psidii | ICMP 19120 * | Psidium sp. | Italy | JX010219 | JX009967 | JX009515 | JX010443 | JX010133 | KC888931 | JX009901 | – |
| C. queenslandicum | ICMP 1778 * | Carica papaya | Australia | JX010276 | JX009934 | JX009447 | JX010414 | JX010104 | KC888928 | JX009899 | – |
| CPC 17123 | Syzygium australa | Australia | KP703357 | KP703282 | – | KP703439 | KP703693 | KP703778 | – | – | |
| ICMP 18705 | Coffea sp. | Fiji | JX010185 | JX010036 | JX009490 | JX010412 | JX010102 | – | JX009890 | – | |
| CMM3233 | Anacardium occidentale | Brazil, Pernambuco state | – | MF110849 | – | MF111058 | MF110996 | MF110639 | – | – | |
| CMM3241 | Anacardium occidentale | Brazil, Pernambuco state | – | MF110848 | – | MF111059 | MF111000 | MF110642 | – | – | |
| CMM3236 | Anacardium occidentale | Brazil, Pernambuco state | – | MF110850 | – | MF111060 | MF110997 | MF110640 | – | – | |
| CMM3240 | Anacardium occidentale | Brazil, Pernambuco state | – | MF110852 | – | MF111061 | MF110999 | MF110644 | – | – | |
| CMM3237 | Anacardium occidentale | Brazil, Pernambuco state | – | MF110853 | – | MF111062 | MF110998 | MF110641 | – | – | |
| CMM3242 | Anacardium occidentale | Brazil, Pernambuco state | – | – | – | MF111063 | MF111001 | MF110643 | – | – | |
| C. rhexiae | CBS 133134 * | Rhexia virginica | USA | JX145128 | – | – | JX145179 | – | – | – | – |
| C. salsolae | ICMP 19051 * | Salsola tragus | Hungary | JX010242 | JX009916 | JX009562 | JX010403 | JX010093 | KC888925 | JX009863 | – |
| C. siamense | ICMP 18578 CBS 130417 * | Citrus arabica | Thailand | JX010171 | JX009924 | FJ907423 | JX010404 | JX010094 | JQ899289 | JX009865 | – |
| VPRI 43077; UMC004 | Citrus limon | Australia, NSW | MG572140 | MG572129 | MK473453 | MG572151 | MG572162 | MG572173 | MW091989 | – | |
| BRIP 54270b; VPRI 43029; A10-43029 | Citrus australasica | Australia, Qld | MK469995 | MK470013 | MK470085 | MK470049 | MK470031 | MK470067 | MW091971 | – | |
| C. siamense (syn. C. jasmini-sambac) | CBS 130420; ICMP 19118 | Jasminum sambac | Vietnam | HM131511 | HM131497 | HM131507 | JX010415 | JX010105 | JQ807841 | JX009895 | – |
| C. siamense (syn. C. hymenocallidis) | CBS 125378; ICMP 18642; LC0043 | Hymenocallis americana | China | JX010278 | JX010019 | GQ856775 | JX010410 | JX010100 | JQ899283 | GQ856730 | – |
| C. siamense (syn. C. murrayae) | GZAAS 5.09506 | Murraya sp. | China | JQ247633 | JQ247609 | JQ247657 | JQ247644 | JQ247621 | – | – | – |
| C. syzygicola | DNCL021; MFLUCC 10-0624 * | Syzygium samarangense | Thailand | KF242094 | KF242156 | KF157801 | KF254880 | KF242125 | – | – | – |
| C. temperatum | CBS 133122 * | Vaccinium macrocarpon | USA | JX145159 | – | – | JX145211 | – | – | – | – |
| C. theobromicola | ICMP 18649; CBS 124945 * | Theobroma cacao | Panama | JX010294 | JX010006 | JX009444 | JX010447 | JX010139 | KC790726 | JX009869 | – |
| C. theobromicola (syn. C. fragariae) | ICMP 17927; CBS 142.31; MTCC 10325T | Fragaria × ananassa | USA | JX010286 | JX010024 | JX009516 | JX010373 | JX010064 | JQ807844 | JX009830 | – |
| VPRI 43083; UMC010 | Citrus aurantifolia | Australia, Qld | MG572146 | MG572135 | MK473456 | MG572157 | MG572168 | MG572179 | MW091995 | – | |
| C. ti | ICMP 4832 * | Cordyline sp. | New Zealand | JX010269 | JX009952 | JX009520 | JX010442 | JX010123 | KM360146 | JX009898 | – |
| C. tropicale | ICMP 18653; CBS 124949 * | Theobroma cacao | Panama | JX010264 | JX010007 | JX009489 | JX010407 | JX010097 | KC790728 | JX009870 | – |
| C. viniferum | GZAAS 5.08601 * | Vitis vinifera, cv. ‘Shuijing’ | China | JN412804 | JN412798 | JN412795 | JN412813 | JN412787 | – | – | – |
| C. wuxiense | CGMCC 3.17894 * | Camellia sinensis | China | KU251591 | KU252045 | KU251672 | KU252200 | KU252101 | KU251722 | KU251939 | – |
| C. xanthorrhoeae | BRIP 45094; ICMP 17903; CBS 127831 * | Xanthorrhoea preissii | Australia | JX010261 | JX009927 | JX009478 | JX010448 | JX010138 | KC790689 | JX009823 | – |
| Colletotrichum sp. | BRIP 58074a; VPRI 43025; A06-43025 | Citrus australasica | Australia, Qld | MK469999 | MK470017 | MK470089 | MK470053 | MK470035 | MK470071 | MW091975 | – |
| Boninense complex | |||||||||||
| C. annellatum | CBS 129826 * | Hevea brasiliensis | Colombia | JQ005222 | – | JQ005570 | JQ005656 | – | – | JQ005396 | JQ005483 |
| C. beeveri | CBS 128527 * | Brachyglottis repanda | New Zealand | JQ005171 | – | JQ005519 | JQ005605 | – | – | JQ005345 | JQ005432 |
| C. boninense | ICMP 17904; CBS 123755 * | Crinum asiaticum ‘Sinicum’ | Japan | JQ005153 | – | JQ005501 | JQ005588 | – | – | JQ005327 | JQ005414 |
| C. brassicicola | CBS 101059 | Brassica oleracea var. gemmifera | New Zealand | JQ005172 | – | JQ005520 | JQ005606 | – | – | JQ005346 | JQ005433 |
| C. brasiliense | CBS 128501* | Passiflora edulis | Brazil | JQ005235 | – | JQ005583 | JQ005669 | – | – | JQ005409 | JQ005496 |
| C. catinaense | CBS 142417; CPC 27978 * | Citrus reticulata | Italy, Catania | KY856400 | – | KY855971 | KY856482 | – | – | KY856136 | KY856307 |
| C. citricola | CBS 134228 * | Citrus unchiu | China | KC293576 | – | KC293616 | KC293656 | – | – | KY856140 | KY856311 |
| C. constrictum | CBS 128504 | Citrus limon | New Zealand | JQ005238 | – | JQ005586 | JQ005672 | – | – | JQ005412 | KY856313 |
| C. colombiense | CBS 129818 * | Passiflora edulis | Colombia | JQ005174 | – | JQ005522 | JQ005608 | – | – | JQ005348 | JQ005435 |
| C. cymbidiicola | IMI 347923 * | Cymbidium sp. | Australia | JQ005166 | – | JQ005514 | JQ005600 | – | – | JQ005340 | JQ005427 |
| C. dacrycarpi | CBS 130241 * | Dacrycarpus dacrydioides | New Zealand | JQ005236 | – | JQ005584 | JQ005670 | – | – | JQ005410 | JQ005497 |
| C. hippeastri | CBS 125376 * | Hippeastrum vittatum | China | JQ005231 | – | JQ005579 | JQ005665 | – | – | JQ005405 | JQ005492 |
| C. karstii | CBS 126532 | Citrus sp. | South Africa | JQ005209 | – | JQ005557 | JQ005643 | – | – | JQ005383 | JQ005470 |
| CBS 128551 | Citrus sp. | New Zealand | JQ005208 | – | JQ005556 | JQ005642 | – | – | JQ005382 | JQ005469 | |
| CBS 129829 | Gossypium hirsutum | Germany | JQ005189 | – | JQ005537 | JQ005623 | – | – | JQ005363 | JQ005450 | |
| CPC 27853 | Citrus sinensis | Italy, Catania | KY856461 | – | KY856034 | KY856543 | – | – | KY856202 | KY856377 | |
| CPC 31139 | Citrus sinensis | Italy, Catania | KY856467 | – | KY856040 | KY856549 | – | – | KY856208 | KY856383 | |
| CBS 129833 | Musa sp. | Mexico | JQ005175 | – | JQ005523 | JQ005609 | – | – | JQ005349 | JQ005436 | |
| CBS 861.72 | Bombax aquaticum | Brazil | JQ005184 | – | JQ005532 | JQ005618 | – | – | JQ005358 | JQ005445 | |
| DAR 25017a; VPRI 42941; D02-42941 | Citrus sinensis Valencia | Australia, NSW | MK470103 | – | MK470109 | MK470106 | – | – | MK470115 | MK470112 | |
| DAR 29821a; VPRI 42943; F02-42943 | Citrus sinensis Valencia | Australia, NSW | MK470104 | – | MK470110 | MK470107 | – | – | MK470116 | MK470113 | |
| DAR 29826a; VPRI 42944; G02-42944 | Citrus sinensis Valencia | Australia, NSW | MK470105 | – | MK470111 | MK470108 | – | – | MK470117 | MK470114 | |
| VPRI 43651; UMC015 | Citrus limon | Australia, Vic | MW081178 | – | MW081186 | MW081182 | – | – | MW081190 | MW081194 | |
| VPRI 43652; UMC016 | Citrus sinensis | Australia, Vic | MW081179 | – | MW081187 | MW081183 | – | – | MW081191 | MW081195 | |
| VPRI 43653; UMC017 | Citrus sinensis | Australia, Vic | MW081180 | – | MW081188 | MW081184 | – | – | MW081192 | MW081196 | |
| VPRI 43654; UMC018 | Citrus sinensis | Australia, Vic | MW081181 | – | MW081189 | MW081185 | – | – | MW081193 | MW081197 | |
| C. limonicola | CBS 142410; CPC 31141 * | Citrus limon | Malta, Gozo | KY856472 | – | KY856045 | KY856554 | – | – | KY856213 | KY856388 |
| C. novae-zelandiae | CBS 128505 * | Capsicum annuum | New Zealand | JQ005228 | – | JQ005576 | JQ005662 | – | – | JQ005402 | JQ005489 |
| C. oncidii | CBS 129828 * | Oncidium sp. | Germany | JQ005169 | – | JQ005517 | JQ005603 | – | – | JQ005343 | JQ005430 |
| C. parsonsiae | CBS 128525 * | Parsonsia capsularis | New Zealand | JQ005233 | – | JQ005581 | JQ005667 | – | – | JQ005407 | JQ005494 |
| C. petchii | CBS 378.94 * | Dracaena marginata | Italy | JQ005223 | – | JQ005571 | JQ005657 | – | – | JQ005397 | JQ005484 |
| C. phyllanthi | CBS 175.67 * | Phyllanthus acidus | India | JQ005221 | – | JQ005569 | JQ005655 | – | – | JQ005395 | JQ005482 |
| C. torulosum | CBS 128544 * | Solanum melongena | New Zealand | JQ005164 | – | JQ005512 | JQ005598 | – | – | JQ005338 | JQ005425 |
| Truncatum complex | |||||||||||
| C. truncatum | CBS 151.35 * | Phaseolus lunatus | USA | GU227862 | – | GU227960 | GU228156 | – | – | GU228352 | GU228058 |
| Dataset | Substitution Models | ||||||
|---|---|---|---|---|---|---|---|
| ITS | tub2 | act | chs-1 | his3 | |||
| boninense complex | SYM + I+G | HKY + I | HKY + G | GTR + G | HKY + I | ||
| ITS | gapdh | tub2 | act | ApMat | gs | chs-1 | |
| gloeosporioides complex | SYM + I | HKY + I | SYM + I | HKY + I | HKY + G | GTR + G | K80 + G |
| Taxon | Conidial Length (μm) | Conidial Width (μm) | Conidial Shape | Growth Rate (mm/day) 1 |
|---|---|---|---|---|
| C. gloeosporioides | (10.2–) 13.8–14.3 (–16.1) | (4.2–) 5.3–5.5 (–7.3) | Subcylindrical | 10.4–10.8 |
| C. siamense | (12.0–) 13.1–13.4 (–15.8) | (4.8–) 5.4–5.5 (–5.9) | Fusoid | 10.9–11.5 |
| C. fructicola | (12.7–) 14.2–14.6 (–17.1) | (4.6–) 5.1–5.2 (–5.7) | Cylindrical | 10.5–11.1 |
| C. theobromicola | (10.8–) 15.2–16 (–21.2) | (4.0–) 4.8–5 (–5.8) | Cylindrical | 10.5–10.7 |
| Colletotrichum sp. | (13.1–) 15.6–16 (–18.0) | (4.6–) 6.1–6.3 (–7.7) | Cylindrical | 8.9–9.7 |
| C. karstii | (11.3–) 13.2–13.6 (–14.8) | (6.4–) 7.1–7.3 (–8.3) | Cylindrical | 9.4–9.6 |
| New species | (12.7–) 14.1–14.5 (–17.2) | (3.9–) 4.5–4.7 (–5.5) | Cylindrical with one end acute | 9.7–10.3 |
| C. queenslandicum2 | (12–) 14.5–16.5 (–21.5) | (3.5–) 4.5–5 (–6) | Cylindric, straight, sometimes slightly constricted near center, ends broadly rounded | / |
| Culture | Fungus Species | Infection Incidence % | |||
|---|---|---|---|---|---|
| Fruit Bioassay (Wound) | Leaf Bioassay | Petal Bioassay | |||
| Orange Leaf | Lemon Leaf | ||||
| VPRI 43075 | C. australianum sp. nov. | 95.8 | 0 | 0 | 100 |
| VPRI 43076 | C. gloeosporioides | 0 | 0 | 0 | 100 |
| VPRI 43079 | C. fructicola | 75 | 0 | 0 | 100 |
| VPRI 43083 | C. theobromicola | 95.8 | 0 | 83.3 | 100 |
| BRIP 58074a | Colletotrichum sp. | 95.8 | 0 | 0 | 100 |
| VPRI 43654 | C. karstii | 100 | 100 | 100 | 100 |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Wang, W.; de Silva, D.D.; Moslemi, A.; Edwards, J.; Ades, P.K.; Crous, P.W.; Taylor, P.W.J. Colletotrichum Species Causing Anthracnose of Citrus in Australia. J. Fungi 2021, 7, 47. https://doi.org/10.3390/jof7010047
Wang W, de Silva DD, Moslemi A, Edwards J, Ades PK, Crous PW, Taylor PWJ. Colletotrichum Species Causing Anthracnose of Citrus in Australia. Journal of Fungi. 2021; 7(1):47. https://doi.org/10.3390/jof7010047
Chicago/Turabian StyleWang, Weixia, Dilani D. de Silva, Azin Moslemi, Jacqueline Edwards, Peter K. Ades, Pedro W. Crous, and Paul W. J. Taylor. 2021. "Colletotrichum Species Causing Anthracnose of Citrus in Australia" Journal of Fungi 7, no. 1: 47. https://doi.org/10.3390/jof7010047
APA StyleWang, W., de Silva, D. D., Moslemi, A., Edwards, J., Ades, P. K., Crous, P. W., & Taylor, P. W. J. (2021). Colletotrichum Species Causing Anthracnose of Citrus in Australia. Journal of Fungi, 7(1), 47. https://doi.org/10.3390/jof7010047





