MCC/Eisosomes Regulate Cell Wall Synthesis and Stress Responses in Fungi
Abstract
1. Introduction
2. MCC/Eisosome Domain Assembly and Structure
2.1. Discovery That MCC Domains, Eisosomes, and Plasma Membrane Furrows Correspond to the Same Membrane Compartment
2.2. Regulation of MCC/Eisosome Assembly and Disassembly
2.3. Spatial Regulation of MCC/Eisosomes
3. MCC/Eisosome Function in Cell Wall Synthesis and Morphogenesis
3.1. General Functions of MCC/Eisosomes
3.2. MCC/Eisosomes Regulate Spatial Organization of the Cell Wall and Morphogenesis
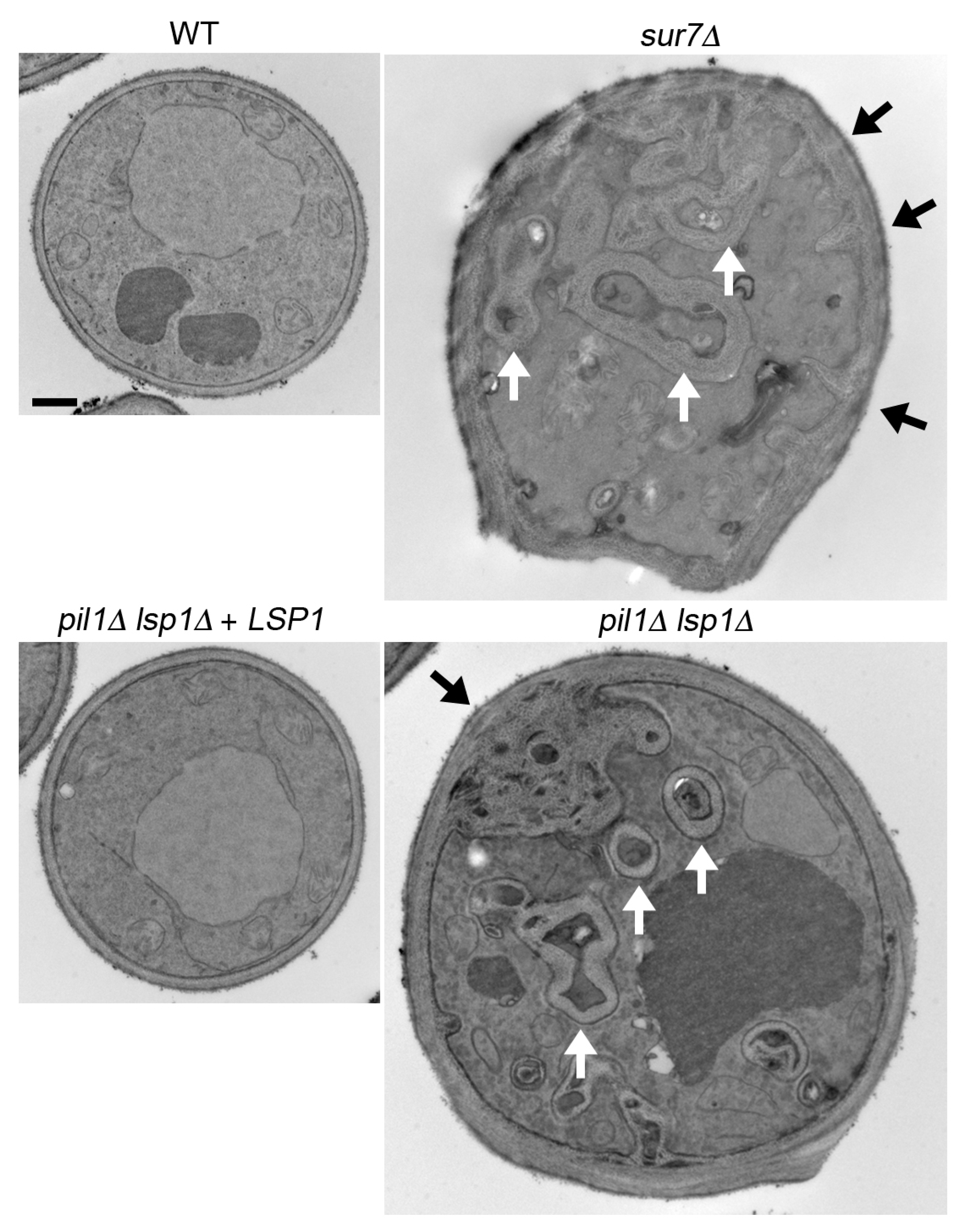
3.3. C. albicans sur7∆ Mutant Makes Thicker, but Weaker Cell Walls
3.4. Abnormal Regulation of PI4,5P2 Contributes to the Altered Cell Wall Phenotype of Eisosome Mutants
3.5. MCC/Eisosomes Contribute to Invasive Growth and Virulence of C. albicans
4. MCC/Eisosomes Affect Other Plasma Membrane Domains That Can Influence Cell Wall Synthesis
4.1. Septins Regulate Cell Wall Synthesis during Cytokinesis and Polarized Morphogenesis
4.2. Sites of Secretion
4.3. Sites of Endocytosis in the Plasma Membrane
4.4. Sites of Contact between the Endoplasmic Reticulum (ER) and Plasma Membrane
5. MCC/Eisosomes Protect Against Stress
5.1. Cell Wall Stress
5.2. Lipid Homeostasis-Related Stress
5.3. Copper/Metal Ion-Induced Stress
5.4. Oxidative Stress
6. Concluding Comments
Acknowledgments
Author Contributions
Conflicts of Interest
References
- Berchtold, D.; Walther, T.C. TORC2 plasma membrane localization is essential for cell viability and restricted to a distinct domain. Mol. Biol. Cell 2009, 20, 1565–1575. [Google Scholar] [CrossRef] [PubMed]
- Spira, F.; Mueller, N.S.; Beck, G.; von Olshausen, P.; Beig, J.; Wedlich-Soldner, R. Patchwork organization of the yeast plasma membrane into numerous coexisting domains. Nat. Cell Biol. 2012, 14, 640–648. [Google Scholar] [CrossRef] [PubMed]
- Vernay, A.; Schaub, S.; Guillas, I.; Bassilana, M.; Arkowitz, R.A. A steep phosphoinositide bis-phosphate gradient forms during fungal filamentous growth. J. Cell Biol. 2012, 198, 711–730. [Google Scholar] [CrossRef] [PubMed]
- Martin, S.W.; Konopka, J.B. Lipid raft polarization contributes to hyphal growth in Candida albicans. Eukaryot. Cell 2004, 3, 675–684. [Google Scholar] [CrossRef] [PubMed]
- Obara, K.; Yamamoto, H.; Kihara, A. Membrane protein Rim21 plays a central role in sensing ambient pH in Saccharomyces cerevisiae. J. Biol. Chem. 2012, 287, 38473–38481. [Google Scholar] [CrossRef] [PubMed]
- Sun, Y.; Carroll, S.; Kaksonen, M.; Toshima, J.Y.; Drubin, D.G. PtdIns(4,5)P2 turnover is required for multiple stages during clathrin- and actin-dependent endocytic internalization. J. Cell Biol. 2007, 177, 355–367. [Google Scholar] [CrossRef] [PubMed]
- Douglas, L.M.; Konopka, J.B. Fungal membrane organization: The eisosome concept. Annu. Rev. Microbiol. 2014, 68, 377–393. [Google Scholar] [CrossRef] [PubMed]
- Douglas, L.M.; Konopka, J.B. Plasma membrane organization promotes virulence of the human fungal pathogen Candida albicans. J. Microbiol. 2016, 54, 178–191. [Google Scholar] [CrossRef] [PubMed]
- Malinska, K.; Malinsky, J.; Opekarova, M.; Tanner, W. Visualization of protein compartmentation within the plasma membrane of living yeast cells. Mol. Biol. Cell 2003, 14, 4427–4436. [Google Scholar] [CrossRef] [PubMed]
- Malinska, K.; Malinsky, J.; Opekarova, M.; Tanner, W. Distribution of Can1p into stable domains reflects lateral protein segregation within the plasma membrane of living S. cerevisiae cells. J. Cell Sci. 2004, 117, 6031–6041. [Google Scholar] [CrossRef] [PubMed]
- Young, M.E.; Karpova, T.S.; Brugger, B.; Moschenross, D.M.; Wang, G.K.; Schneiter, R.; Wieland, F.T.; Cooper, J.A. The Sur7p family defines novel cortical domains in Saccharomyces cerevisiae, affects sphingolipid metabolism, and is involved in sporulation. Mol. Cell. Biol. 2002, 22, 927–934. [Google Scholar] [CrossRef] [PubMed]
- Walther, T.C.; Brickner, J.H.; Aguilar, P.S.; Bernales, S.; Pantoja, C.; Walter, P. Eisosomes mark static sites of endocytosis. Nature 2006, 439, 998–1003. [Google Scholar] [CrossRef] [PubMed]
- Malinsky, J.; Opekarova, M.; Grossmann, G.; Tanner, W. Membrane microdomains, rafts, and detergent-resistant membranes in plants and fungi. Annu. Rev. Plant Biol. 2013, 64, 501–529. [Google Scholar] [CrossRef] [PubMed]
- Stradalova, V.; Stahlschmidt, W.; Grossmann, G.; Blazikova, M.; Rachel, R.; Tanner, W.; Malinsky, J. Furrow-like invaginations of the yeast plasma membrane correspond to membrane compartment of Can1. J. Cell Sci. 2009, 122, 2887–2894. [Google Scholar] [CrossRef] [PubMed]
- Lee, J.H.; Heuser, J.E.; Roth, R.; Goodenough, U. Eisosome Ultrastructure and Evolution in Fungi, Microalgae and Lichens. Eukaryot. Cell 2015, 14, 1017–1042. [Google Scholar] [CrossRef] [PubMed]
- Grossmann, G.; Malinsky, J.; Stahlschmidt, W.; Loibl, M.; Weig-Meckl, I.; Frommer, W.B.; Opekarova, M.; Tanner, W. Plasma membrane microdomains regulate turnover of transport proteins in yeast. J. Cell Biol. 2008, 183, 1075–1088. [Google Scholar] [CrossRef] [PubMed]
- Buser, C.; Drubin, D.G. Ultrastructural imaging of endocytic sites in Saccharomyces cerevisiae by transmission electron microscopy and immunolabeling. Microsc. Microanal. 2013, 19, 381–392. [Google Scholar] [CrossRef] [PubMed]
- Deng, C.; Xiong, X.; Krutchinsky, A.N. Unifying fluorescence microscopy and mass spectrometry for studying protein complexes in cells. Mol. Cell. Proteom. 2009, 8, 1413–1423. [Google Scholar] [CrossRef] [PubMed]
- Wang, H.X.; Douglas, L.M.; Vesela, P.; Rachel, R.; Malinsky, J.; Konopka, J.B. Eisosomes promote the ability of Sur7 to regulate plasma membrane organization in Candida albicans. Mol. Biol. Cell 2016, 27, 1663–1675. [Google Scholar] [CrossRef] [PubMed]
- Frohlich, F.; Christiano, R.; Olson, D.K.; Alcazar-Roman, A.; DeCamilli, P.; Walther, T.C. A role for eisosomes in maintenance of plasma membrane phosphoinositide levels. Mol. Biol. Cell 2014, 25, 2797–2806. [Google Scholar] [CrossRef] [PubMed]
- Frohlich, F.; Moreira, K.; Aguilar, P.S.; Hubner, N.C.; Mann, M.; Walter, P.; Walther, T.C. A genome-wide screen for genes affecting eisosomes reveals Nce102 function in sphingolipid signaling. J. Cell Biol. 2009, 185, 1227–1242. [Google Scholar] [CrossRef] [PubMed]
- Kabeche, R.; Roguev, A.; Krogan, N.J.; Moseley, J.B. A Pil1-Sle1-Syj1-Tax4 functional pathway links eisosomes with PI(4,5)P2 regulation. J. Cell Sci. 2014, 127, 1318–1326. [Google Scholar] [CrossRef] [PubMed]
- Kabeche, R.; Howard, L.; Moseley, J.B. Eisosomes provide membrane reservoirs for rapid expansion of the yeast plasma membrane. J. Cell Sci. 2015, 128, 4057–4062. [Google Scholar] [CrossRef] [PubMed]
- Dupont, S.; Beney, L.; Ritt, J.F.; Lherminier, J.; Gervais, P. Lateral reorganization of plasma membrane is involved in the yeast resistance to severe dehydration. Biochim. Biophys. Acta 2010, 1798, 975–985. [Google Scholar] [CrossRef] [PubMed]
- Malinsky, J.; Opekarova, M.; Tanner, W. The lateral compartmentation of the yeast plasma membrane. Yeast 2010, 27, 473–478. [Google Scholar] [CrossRef] [PubMed]
- Olivera-Couto, A.; Aguilar, P.S. Eisosomes and plasma membrane organization. Mol. Genet. Genom. 2012, 287, 607–620. [Google Scholar] [CrossRef] [PubMed]
- Murphy, E.R.; Kim, K.T. Insights into eisosome assembly and organization. J. Biosci. 2012, 37, 295–500. [Google Scholar] [CrossRef]
- Grossmann, G.; Opekarova, M.; Malinsky, J.; Weig-Meckl, I.; Tanner, W. Membrane potential governs lateral segregation of plasma membrane proteins and lipids in yeast. EMBO J. 2007, 26, 1–8. [Google Scholar] [CrossRef] [PubMed]
- Brach, T.; Specht, T.; Kaksonen, M. Reassessment of the role of plasma membrane domains in the regulation of vesicular traffic in yeast. J. Cell Sci. 2011, 124, 328–337. [Google Scholar] [CrossRef] [PubMed]
- Zhang, X.; Lester, R.L.; Dickson, R.C. Pil1p and Lsp1p negatively regulate the 3-phosphoinositide-dependent protein kinase-like kinase Pkh1p and downstream signaling pathways Pkc1p and Ypk1p. J. Biol. Chem. 2004, 279, 22030–22038. [Google Scholar] [CrossRef] [PubMed]
- Walther, T.C.; Aguilar, P.S.; Frohlich, F.; Chu, F.; Moreira, K.; Burlingame, A.L.; Walter, P. Pkh-kinases control eisosome assembly and organization. EMBO J. 2007, 26, 4946–4955. [Google Scholar] [CrossRef] [PubMed]
- Fadri, M.; Daquinag, A.; Wang, S.; Xue, T.; Kunz, J. The pleckstrin homology domain proteins Slm1 and Slm2 are required for actin cytoskeleton organization in yeast and bind phosphatidylinositol-4,5-bisphosphate and TORC2. Mol. Biol. Cell 2005, 16, 1883–1900. [Google Scholar] [CrossRef] [PubMed]
- Krogan, N.J.; Cagney, G.; Yu, H.; Zhong, G.; Guo, X.; Ignatchenko, A.; Li, J.; Pu, S.; Datta, N.; Tikuisis, A.P.; et al. Global landscape of protein complexes in the yeast Saccharomyces cerevisiae. Nature 2006, 440, 637–643. [Google Scholar] [CrossRef] [PubMed]
- Moreira, K.E.; Walther, T.C.; Aguilar, P.S.; Walter, P. Pil1 controls eisosome biogenesis. Mol. Biol. Cell 2009, 20, 809–818. [Google Scholar] [CrossRef] [PubMed]
- Yu, H.; Braun, P.; Yildirim, M.A.; Lemmens, I.; Venkatesan, K.; Sahalie, J.; Hirozane-Kishikawa, T.; Gebreab, F.; Li, N.; Simonis, N.; et al. High-quality binary protein interaction map of the yeast interactome network. Science 2008, 322, 104–110. [Google Scholar] [CrossRef] [PubMed]
- Vaskovicova, K.; Awadova, T.; Vesela, P.; Balazova, M.; Opekarova, M.; Malinsky, J. mRNA decay is regulated via sequestration of the conserved 5′-3′ exoribonuclease Xrn1 at eisosome in yeast. Eur. J. Cell Biol. 2017, 96, 591–599. [Google Scholar] [CrossRef] [PubMed]
- Ghaemmaghami, S.; Huh, W.K.; Bower, K.; Howson, R.W.; Belle, A.; Dephoure, N.; O’Shea, E.K.; Weissman, J.S. Global analysis of protein expression in yeast. Nature 2003, 425, 737–741. [Google Scholar] [CrossRef] [PubMed]
- Kulak, N.A.; Pichler, G.; Paron, I.; Nagaraj, N.; Mann, M. Minimal, encapsulated proteomic-sample processing applied to copy-number estimation in eukaryotic cells. Nat. Methods 2014, 11, 319–324. [Google Scholar] [CrossRef] [PubMed]
- Karotki, L.; Huiskonen, J.T.; Stefan, C.J.; Ziolkowska, N.E.; Roth, R.; Surma, M.A.; Krogan, N.J.; Emr, S.D.; Heuser, J.; Grunewald, K.; et al. Eisosome proteins assemble into a membrane scaffold. J. Cell Biol. 2011, 195, 889–902. [Google Scholar] [CrossRef] [PubMed]
- Reijnst, P.; Walther, A.; Wendland, J. Dual-colour fluorescence microscopy using yEmCherry-/GFP-tagging of eisosome components Pil1 and Lsp1 in Candida albicans. Yeast 2011, 28, 331–338. [Google Scholar] [CrossRef] [PubMed]
- Seger, S.; Rischatsch, R.; Philippsen, P. Formation and stability of eisosomes in the filamentous fungus Ashbya gossypii. J. Cell Sci. 2011, 124, 1629–1634. [Google Scholar] [CrossRef] [PubMed]
- Kabeche, R.; Baldissard, S.; Hammond, J.; Howard, L.; Moseley, J.B. The filament-forming protein Pil1 assembles linear eisosomes in fission yeast. Mol. Biol. Cell 2011, 22, 4059–4067. [Google Scholar] [CrossRef] [PubMed]
- Vangelatos, I.; Roumelioti, K.; Gournas, C.; Suarez, T.; Scazzocchio, C.; Sophianopoulou, V. Eisosome organization in the filamentous ascomycete Aspergillus nidulans. Eukaryot. Cell 2010, 9, 1441–1454. [Google Scholar] [CrossRef] [PubMed]
- Murphy, E.R.; Boxberger, J.; Colvin, R.; Lee, S.J.; Zahn, G.; Loor, F.; Kim, K. Pil1, an eisosome organizer, plays an important role in the recruitment of synaptojanins and amphiphysins to facilitate receptor-mediated endocytosis in yeast. Eur. J. Cell Biol. 2011, 90, 825–833. [Google Scholar] [CrossRef] [PubMed]
- Ho, Y.; Gruhler, A.; Heilbut, A.; Bader, G.D.; Moore, L.; Adams, S.L.; Millar, A.; Taylor, P.; Bennett, K.; Boutilier, K.; et al. Systematic identification of protein complexes in Saccharomyces cerevisiae by mass spectrometry. Nature 2002, 415, 180–183. [Google Scholar] [CrossRef] [PubMed]
- Aguilar, P.S.; Frohlich, F.; Rehman, M.; Shales, M.; Ulitsky, I.; Olivera-Couto, A.; Braberg, H.; Shamir, R.; Walter, P.; Mann, M.; et al. A plasma-membrane E-MAP reveals links of the eisosome with sphingolipid metabolism and endosomal trafficking. Nat. Struct. Mol. Biol. 2010, 17, 901–908. [Google Scholar] [CrossRef] [PubMed]
- Moor, H.; Muhlethaler, K. Fine structure in frozen-etched yeast cells. J. Cell Biol. 1963, 17, 609–628. [Google Scholar] [CrossRef] [PubMed]
- Mulholland, J.; Preuss, D.; Moon, A.; Wong, A.; Drubin, D.; Botstein, D. Ultrastructure of the yeast actin cytoskeleton and its association with the plasma membrane. J. Cell Biol. 1994, 125, 381–391. [Google Scholar] [CrossRef] [PubMed]
- Gross, H.; Kuebler, O.; Bas, E.; Moor, H. Decoration of specific sites on freeze-fractured membranes. J. Cell Biol. 1978, 79, 646–656. [Google Scholar] [CrossRef] [PubMed]
- Barug, D.; de Groot, K. Effect of the imidazole derivative lombazole on the ultrastructure of Staphylococcus epidermidis and Candida albicans. Antimicrob. Agents Chemother. 1985, 28, 643–647. [Google Scholar] [CrossRef] [PubMed]
- Takeo, K. Lack of invaginations of the plasma membrane during budding and cell division of Saccharomyces cerevisiae and Schizosaccharomyces pombe. FEMS Microbiol. Lett. 1984, 22, 97–100. [Google Scholar] [CrossRef]
- Loibl, M.; Grossmann, G.; Stradalova, V.; Klingl, A.; Rachel, R.; Tanner, W.; Malinsky, J.; Opekarova, M. C terminus of Nce102 determines the structure and function of microdomains in the Saccharomyces cerevisiae plasma membrane. Eukaryot. Cell 2010, 9, 1184–1192. [Google Scholar] [CrossRef] [PubMed]
- Olivera-Couto, A.; Grana, M.; Harispe, L.; Aguilar, P.S. The eisosome core is composed of BAR domain proteins. Mol. Biol. Cell 2011, 22, 2360–2372. [Google Scholar] [CrossRef] [PubMed]
- Ziolkowska, N.E.; Karotki, L.; Rehman, M.; Huiskonen, J.T.; Walther, T.C. Eisosome-driven plasma membrane organization is mediated by BAR domains. Nat. Struct. Mol. Biol. 2011, 18, 854–856. [Google Scholar] [CrossRef] [PubMed]
- Zimmerberg, J.; McLaughlin, S. Membrane curvature: How BAR domains bend bilayers. Curr. Biol. 2004, 14, R250–R252. [Google Scholar] [CrossRef] [PubMed]
- Olivera-Couto, A.; Salzman, V.; Mailhos, M.; Digman, M.A.; Gratton, E.; Aguilar, P.S. Eisosomes are dynamic plasma membrane domains showing pil1-lsp1 heteroligomer binding equilibrium. Biophys. J. 2015, 108, 1633–1644. [Google Scholar] [CrossRef] [PubMed]
- Lacy, M.M.; Baddeley, D.; Berro, J. Single-molecule imaging of the BAR domain protein Pil1p reveals filament-end dynamics. Mol. Biol. Cell 2017, 28, 2251–2259. [Google Scholar] [CrossRef] [PubMed]
- Moreira, K.E.; Schuck, S.; Schrul, B.; Frohlich, F.; Moseley, J.B.; Walther, T.C.; Walter, P. Seg1 controls eisosome assembly and shape. J. Cell Biol. 2012, 198, 405–420. [Google Scholar] [CrossRef] [PubMed]
- Luo, G.; Gruhler, A.; Liu, Y.; Jensen, O.N.; Dickson, R.C. The sphingolipid long-chain base-Pkh1/2-Ypk1/2 signaling pathway regulates eisosome assembly and turnover. J. Biol. Chem. 2008, 283, 10433–10444. [Google Scholar] [CrossRef] [PubMed]
- Mascaraque, V.; Hernaez, M.L.; Jimenez-Sanchez, M.; Hansen, R.; Gil, C.; Martin, H.; Cid, V.J.; Molina, M. Phosphoproteomic analysis of protein kinase C signaling in Saccharomyces cerevisiae reveals Slt2 mitogen-activated protein kinase (MAPK)-dependent phosphorylation of eisosome core components. Mol. Cell. Proteom. 2013, 12, 557–574. [Google Scholar] [CrossRef] [PubMed]
- Zhao, H.; Michelot, A.; Koskela, E.V.; Tkach, V.; Stamou, D.; Drubin, D.G.; Lappalainen, P. Membrane-sculpting BAR domains generate stable lipid microdomains. Cell Rep. 2013, 4, 1213–1223. [Google Scholar] [CrossRef] [PubMed]
- Alvarez, F.J.; Douglas, L.M.; Rosebrock, A.; Konopka, J.B. The Sur7 protein regulates plasma membrane organization and prevents intracellular cell wall growth in Candida albicans. Mol. Biol. Cell 2008, 19, 5214–5225. [Google Scholar] [CrossRef] [PubMed]
- Grousl, T.; Opekarova, M.; Stradalova, V.; Hasek, J.; Malinsky, J. Evolutionarily conserved 5′-3′ exoribonuclease Xrn1 accumulates at plasma membrane-associated eisosomes in post-diauxic yeast. PLoS ONE 2015, 10, e0122770. [Google Scholar] [CrossRef] [PubMed]
- Thayer, N.H.; Leverich, C.K.; Fitzgibbon, M.P.; Nelson, Z.W.; Henderson, K.A.; Gafken, P.R.; Hsu, J.J.; Gottschling, D.E. Identification of long-lived proteins retained in cells undergoing repeated asymmetric divisions. Proc. Natl. Acad. Sci. USA 2014, 111, 14019–14026. [Google Scholar] [CrossRef] [PubMed]
- Badrane, H.; Nguyen, M.H.; Blankenship, J.R.; Cheng, S.; Hao, B.; Mitchell, A.P.; Clancy, C.J. Rapid redistribution of phosphatidylinositol-(4,5)-bisphosphate and septins during the Candida albicans response to caspofungin. Antimicrob. Agents Chemother. 2012, 56, 4614–4624. [Google Scholar] [CrossRef] [PubMed]
- Kabeche, R.; Madrid, M.; Cansado, J.; Moseley, J.B. Eisosomes regulate PI(4,5)P2 cortical clusters and MAP kinase signaling upon osmotic stress. J. Biol. Chem. 2015, 290, 25960–25973. [Google Scholar] [CrossRef] [PubMed]
- Douglas, L.M.; Alvarez, F.J.; McCreary, C.; Konopka, J.B. Septin function in yeast model systems and pathogenic fungi. Eukaryot. Cell 2005, 4, 1503–1512. [Google Scholar] [CrossRef] [PubMed]
- Bridges, A.A.; Gladfelter, A.S. Septin Form and Function at the Cell Cortex. J. Biol. Chem. 2015, 290, 17173–17180. [Google Scholar] [CrossRef] [PubMed]
- Bernardo, S.M.; Lee, S.A. Candida albicans SUR7 contributes to secretion, biofilm formation, and macrophage killing. BMC Microbiol. 2010, 10. [Google Scholar] [CrossRef] [PubMed]
- Wang, H.X.; Douglas, L.M.; Aimanianda, V.; Latge, J.P.; Konopka, J.B. The Candida albicans Sur7 protein is needed for proper synthesis of the fibrillar component of the cell wall that confers strength. Eukaryot. Cell 2011, 10, 72–80. [Google Scholar] [CrossRef] [PubMed]
- Drgonova, J.; Drgon, T.; Tanaka, K.; Kollar, R.; Chen, G.C.; Ford, R.A.; Chan, C.S.; Takai, Y.; Cabib, E. Rho1p, a yeast protein at the interface between cell polarization and morphogenesis. Science 1996, 272, 277–279. [Google Scholar] [CrossRef] [PubMed]
- Qadota, H.; Python, C.P.; Inoue, S.B.; Arisawa, M.; Anraku, Y.; Zheng, Y.; Watanabe, T.; Levin, D.E.; Ohya, Y. Identification of yeast Rho1p GTPase as a regulatory subunit of 1,3-β-glucan synthase. Science 1996, 272, 279–281. [Google Scholar] [CrossRef] [PubMed]
- Utsugi, T.; Minemura, M.; Hirata, A.; Abe, M.; Watanabe, D.; Ohya, Y. Movement of yeast 1,3-β-glucan synthase is essential for uniform cell wall synthesis. Genes Cells 2002, 7, 1–9. [Google Scholar] [CrossRef] [PubMed]
- Zhang, L.B.; Tang, L.; Ying, S.H.; Feng, M.G. Two eisosome proteins play opposite roles in autophagic control and sustain cell integrity, function and pathogenicity in Beauveria bassiana. Environ. Microbiol. 2017, 19, 2037–2052. [Google Scholar] [CrossRef] [PubMed]
- Berchtold, D.; Piccolis, M.; Chiaruttini, N.; Riezman, I.; Riezman, H.; Roux, A.; Walther, T.C.; Loewith, R. Plasma membrane stress induces relocalization of Slm proteins and activation of TORC2 to promote sphingolipid synthesis. Nat. Cell Biol. 2012, 14, 542–547. [Google Scholar] [CrossRef] [PubMed]
- Kamble, C.; Jain, S.; Murphy, E.; Kim, K. Requirements of Slm proteins for proper eisosome organization, endocytic trafficking and recycling in the yeast Saccharomyces cerevisiae. J. Biosci. 2011, 36, 79–96. [Google Scholar] [CrossRef] [PubMed]
- Douglas, L.M.; Wang, H.X.; Keppler-Ross, S.; Dean, N.; Konopka, J.B. Sur7 Promotes Plasma Membrane Organization and Is Needed for Resistance to Stressful Conditions and to the Invasive Growth and Virulence of Candida albicans. MBio 2012, 3, e00254-11. [Google Scholar] [CrossRef] [PubMed]
- Hopke, A.; Nicke, N.; Hidu, E.E.; Degani, G.; Popolo, L.; Wheeler, R.T. Neutrophil Attack Triggers Extracellular Trap-Dependent Candida Cell Wall Remodeling and Altered Immune Recognition. PLoS Pathog. 2016, 12, e1005644. [Google Scholar] [CrossRef] [PubMed]
- Douglas, L.M.; Wang, H.X.; Konopka, J.B. The MARVEL Domain Protein Nce102 Regulates Actin Organization and Invasive Growth of Candida albicans. MBio 2013, 4. [Google Scholar] [CrossRef] [PubMed]
- Warenda, A.J.; Konopka, J.B. Septin function in Candida albicans morphogenesis. Mol. Biol. Cell 2002, 13, 2732–2746. [Google Scholar] [CrossRef] [PubMed]
- Gladfelter, A.S.; Pringle, J.R.; Lew, D.J. The septin cortex at the yeast mother-bud neck. Curr. Opin. Microbiol. 2001, 4, 681–689. [Google Scholar] [CrossRef]
- Schmidt, M.; Varma, A.; Drgon, T.; Bowers, B.; Cabib, E. Septins, under Cla4p regulation, and the chitin ring are required for neck integrity in budding yeast. Mol. Biol. Cell 2003, 14, 2128–2141. [Google Scholar] [CrossRef] [PubMed]
- Roh, D.H.; Bowers, B.; Schmidt, M.; Cabib, E. The septation apparatus, an autonomous system in budding yeast. Mol. Biol. Cell 2002, 13, 2747–2759. [Google Scholar] [CrossRef] [PubMed]
- Khan, A.; McQuilken, M.; Gladfelter, A.S. Septins and Generation of Asymmetries in Fungal Cells. Annu. Rev. Microbiol. 2015, 69, 487–503. [Google Scholar] [CrossRef] [PubMed]
- Oh, Y.; Bi, E. Septin structure and function in yeast and beyond. Trends Cell Biol. 2011, 21, 141–148. [Google Scholar] [CrossRef] [PubMed]
- Cid, V.J.; Jimenez, J.; Molina, M.; Sanchez, M.; Nombela, C.; Thorner, J.W. Orchestrating the cell cycle in yeast: Sequential localization of key mitotic regulators at the spindle pole and the bud neck. Microbiology 2002, 148, 2647–2659. [Google Scholar] [CrossRef] [PubMed]
- Merlini, L.; Piatti, S. The mother-bud neck as a signaling platform for the coordination between spindle position and cytokinesis in budding yeast. Biol. Chem. 2011, 392, 805–812. [Google Scholar] [CrossRef] [PubMed]
- Takizawa, P.A.; DeRisi, J.L.; Wilhelm, J.E.; Vale, R.D. Plasma membrane compartmentalization in yeast by messenger RNA transport and a septin diffusion barrier. Science 2000, 290, 341–344. [Google Scholar] [CrossRef] [PubMed]
- Barral, Y.; Mermall, V.; Mooseker, M.S.; Snyder, M. Compartmentalization of the cell cortex by septins is required for maintainence of cell polarity in yeast. Mol. Cell 2000, 5, 841–851. [Google Scholar] [CrossRef]
- Chao, J.T.; Wong, A.K.; Tavassoli, S.; Young, B.P.; Chruscicki, A.; Fang, N.N.; Howe, L.J.; Mayor, T.; Foster, L.J.; Loewen, C.J. Polarization of the endoplasmic reticulum by ER-septin tethering. Cell 2014, 158, 620–632. [Google Scholar] [CrossRef] [PubMed]
- Dobbelaere, J.; Barral, Y. Spatial coordination of cytokinetic events by compartmentalization of the cell cortex. Science 2004, 305, 393–396. [Google Scholar] [CrossRef] [PubMed]
- Momany, M.; Talbot, N.J. Septins Focus Cellular Growth for Host Infection by Pathogenic Fungi. Front. Cell Dev. Biol. 2017, 5, 33. [Google Scholar] [CrossRef] [PubMed]
- Konopka, J.B. AFR1 acts in conjunction with the α-factor receptor to promote morphogenesis and adaptation. Mol. Cell. Biol. 1993, 13, 6876–6888. [Google Scholar] [CrossRef] [PubMed]
- Bharucha, J.P.; Larson, J.R.; Konopka, J.B.; Tatchell, K. Saccharomyces cerevisiae Afr1 protein is a protein phosphatase 1/Glc7-targeting subunit that regulates the septin cytoskeleton during mating. Eukaryot. Cell 2008, 7, 1246–1255. [Google Scholar] [CrossRef] [PubMed]
- Blankenship, J.R.; Cheng, S.; Woolford, C.A.; Xu, W.; Johnson, T.M.; Rogers, P.D.; Fanning, S.; Nguyen, M.H.; Clancy, C.J.; Mitchell, A.P. Mutational analysis of essential septins reveals a role for septin-mediated signaling in filamentation. Eukaryot. Cell 2014, 13, 1403–1410. [Google Scholar] [CrossRef] [PubMed]
- Sudbery, P.E. The germ tubes of Candida albicans hyphae and pseudohyphae show different patterns of septin ring localization. Mol. Microbiol. 2001, 41, 19–31. [Google Scholar] [CrossRef] [PubMed]
- Li, L.; Zhang, C.; Konopka, J.B. A Candida albicans Temperature-Sensitive cdc12–6 Mutant Identifies Roles for Septins in Selection of Sites of Germ Tube Formation and Hyphal Morphogenesis. Eukaryot. Cell 2012, 11, 1210–1218. [Google Scholar] [CrossRef] [PubMed]
- Martin, S.W.; Douglas, L.M.; Konopka, J.B. Cell cycle dynamics and quorum sensing in Candida albicans chlamydospores are distinct from budding and hyphal cells. Eukaryot. Cell 2005, 4, 1191–1202. [Google Scholar] [CrossRef] [PubMed]
- Dagdas, Y.F.; Yoshino, K.; Dagdas, G.; Ryder, L.S.; Bielska, E.; Steinberg, G.; Talbot, N.J. Septin-mediated plant cell invasion by the rice blast fungus, Magnaporthe oryzae. Science 2012, 336, 1590–1595. [Google Scholar] [CrossRef] [PubMed]
- Bridges, A.A.; Gladfelter, A.S. Fungal pathogens are platforms for discovering novel and conserved septin properties. Curr. Opin. Microbiol. 2014, 20, 42–48. [Google Scholar] [CrossRef] [PubMed]
- Vargas-Muniz, J.M.; Juvvadi, P.R.; Steinbach, W.J. Forging the ring: From fungal septins’ divergent roles in morphology, septation and virulence to factors contributing to their assembly into higher order structures. Microbiology 2016, 162, 1527–1534. [Google Scholar] [CrossRef] [PubMed]
- Munson, M.; Novick, P. The exocyst defrocked, a framework of rods revealed. Nat. Struct. Mol. Biol. 2006, 13, 577–581. [Google Scholar] [CrossRef] [PubMed]
- Heider, M.R.; Munson, M. Exorcising the exocyst complex. Traffic 2012, 13, 898–907. [Google Scholar] [CrossRef] [PubMed]
- Bonifacino, J.S. Vesicular transport earns a Nobel. Trends Cell Biol. 2014, 24, 3–5. [Google Scholar] [CrossRef] [PubMed]
- Donovan, K.W.; Bretscher, A. Tracking individual secretory vesicles during exocytosis reveals an ordered and regulated process. J. Cell Biol. 2015, 210, 181–189. [Google Scholar] [CrossRef] [PubMed]
- Luo, G.; Zhang, J.; Guo, W. The role of Sec3p in secretory vesicle targeting and exocyst complex assembly. Mol. Biol. Cell 2014, 25, 3813–3822. [Google Scholar] [CrossRef] [PubMed]
- Pleskot, R.; Cwiklik, L.; Jungwirth, P.; Zarsky, V.; Potocky, M. Membrane targeting of the yeast exocyst complex. Biochim. Biophys. Acta 2015, 1848, 1481–1489. [Google Scholar] [CrossRef] [PubMed]
- Guo, W.; Tamanoi, F.; Novick, P. Spatial regulation of the exocyst complex by Rho1 GTPase. Nat. Cell Biol. 2001, 3, 353–360. [Google Scholar] [CrossRef] [PubMed]
- Zhang, X.; Bi, E.; Novick, P.; Du, L.; Kozminski, K.G.; Lipschutz, J.H.; Guo, W. Cdc42 interacts with the exocyst and regulates polarized secretion. J. Biol. Chem. 2001, 276, 46745–46750. [Google Scholar] [CrossRef] [PubMed]
- Goode, B.L.; Eskin, J.A.; Wendland, B. Actin and endocytosis in budding yeast. Genetics 2015, 199, 315–358. [Google Scholar] [CrossRef] [PubMed]
- Weinberg, J.; Drubin, D.G. Clathrin-mediated endocytosis in budding yeast. Trends Cell Biol. 2012, 22, 1–13. [Google Scholar] [CrossRef] [PubMed]
- Kaksonen, M.; Toret, C.P.; Drubin, D.G. A modular design for the clathrin- and actin-mediated endocytosis machinery. Cell 2005, 123, 305–320. [Google Scholar] [CrossRef] [PubMed]
- Avinoam, O.; Schorb, M.; Beese, C.J.; Briggs, J.A.; Kaksonen, M. Endocytic sites mature by continuous bending and remodeling of the clathrin coat. Science 2015, 348, 1369–1372. [Google Scholar] [CrossRef] [PubMed]
- Skruzny, M.; Brach, T.; Ciuffa, R.; Rybina, S.; Wachsmuth, M.; Kaksonen, M. Molecular basis for coupling the plasma membrane to the actin cytoskeleton during clathrin-mediated endocytosis. Proc. Natl. Acad. Sci. USA 2012, 109, E2533–E2542. [Google Scholar] [CrossRef] [PubMed]
- Sivadon, P.; Peypouquet, M.F.; Doignon, F.; Aigle, M.; Crouzet, M. Cloning of the multicopy suppressor gene SUR7: Evidence for a functional relationship between the yeast actin-binding protein Rvs167 and a putative membranous protein. Yeast 1997, 13, 747–761. [Google Scholar] [CrossRef]
- Wong, A.K.; Chao, J.T.; Loewen, C.J. Barriers to uniformity within the endoplasmic reticulum. Curr. Opin. Cell Biol. 2014, 29, 31–38. [Google Scholar] [CrossRef] [PubMed]
- Henne, W.M.; Liou, J.; Emr, S.D. Molecular mechanisms of inter-organelle ER-PM contact sites. Curr. Opin. Cell Biol. 2015, 35, 123–130. [Google Scholar] [CrossRef] [PubMed]
- Stradalova, V.; Blazikova, M.; Grossmann, G.; Opekarova, M.; Tanner, W.; Malinsky, J. Distribution of cortical endoplasmic reticulum determines positioning of endocytic events in yeast plasma membrane. PLoS ONE 2012, 7, e35132. [Google Scholar] [CrossRef] [PubMed]
- Schuck, S.; Prinz, W.A.; Thorn, K.S.; Voss, C.; Walter, P. Membrane expansion alleviates endoplasmic reticulum stress independently of the unfolded protein response. J. Cell Biol. 2009, 187, 525–536. [Google Scholar] [CrossRef] [PubMed]
- Pichler, H.; Gaigg, B.; Hrastnik, C.; Achleitner, G.; Kohlwein, S.D.; Zellnig, G.; Perktold, A.; Daum, G. A subfraction of the yeast endoplasmic reticulum associates with the plasma membrane and has a high capacity to synthesize lipids. Eur. J. Biochem. 2001, 268, 2351–2361. [Google Scholar] [CrossRef] [PubMed]
- Tavassoli, S.; Chao, J.T.; Young, B.P.; Cox, R.C.; Prinz, W.A.; de Kroon, A.I.; Loewen, C.J. Plasma membrane—Endoplasmic reticulum contact sites regulate phosphatidylcholine synthesis. EMBO Rep. 2013, 14, 434–440. [Google Scholar] [CrossRef] [PubMed]
- Manford, A.G.; Stefan, C.J.; Yuan, H.L.; Macgurn, J.A.; Emr, S.D. ER-to-plasma membrane tethering proteins regulate cell signaling and ER morphology. Dev. Cell 2012, 23, 1129–1140. [Google Scholar] [CrossRef] [PubMed]
- Maeda, K.; Anand, K.; Chiapparino, A.; Kumar, A.; Poletto, M.; Kaksonen, M.; Gavin, A.C. Interactome map uncovers phosphatidylserine transport by oxysterol-binding proteins. Nature 2013, 501, 257–261. [Google Scholar] [CrossRef] [PubMed]
- Moser von Filseck, J.; Copic, A.; Delfosse, V.; Vanni, S.; Jackson, C.L.; Bourguet, W.; Drin, G. Phosphatidylserine transport by ORP/Osh proteins is driven by phosphatidylinositol 4-phosphate. Science 2015, 349, 432–436. [Google Scholar] [CrossRef] [PubMed]
- Yoshikawa, K.; Tanaka, T.; Furusawa, C.; Nagahisa, K.; Hirasawa, T.; Shimizu, H. Comprehensive phenotypic analysis for identification of genes affecting growth under ethanol stress in Saccharomyces cerevisiae. FEMS Yeast Res. 2009, 9, 32–44. [Google Scholar] [CrossRef] [PubMed]
- Hosiner, D.; Sponder, G.; Graschopf, A.; Reipert, S.; Schweyen, R.J.; Schuller, C.; Aleschko, M. Pun1p is a metal ion-inducible, calcineurin/Crz1p-regulated plasma membrane protein required for cell wall integrity. Biochim. Biophys. Acta 2011, 1808, 1108–1119. [Google Scholar] [CrossRef] [PubMed]
- Platta, H.W.; Stenmark, H. Endocytosis and signaling. Curr. Opin. Cell Biol. 2011, 23, 393–403. [Google Scholar] [CrossRef] [PubMed]
- Divito, C.B.; Amara, S.G. Close encounters of the oily kind: Regulation of transporters by lipids. Mol. Interv. 2009, 9, 252–262. [Google Scholar] [CrossRef] [PubMed]
- Groves, J.T.; Kuriyan, J. Molecular mechanisms in signal transduction at the membrane. Nat. Struct. Mol. Biol. 2010, 17, 659–665. [Google Scholar] [CrossRef] [PubMed]
- Garcia-Marques, S.; Randez-Gil, F.; Dupont, S.; Garre, E.; Prieto, J.A. Sng1 associates with Nce102 to regulate the yeast Pkh-Ypk signalling module in response to sphingolipid status. Biochim. Biophys. Acta 2016, 1863, 1319–1333. [Google Scholar] [CrossRef] [PubMed]
- Breslow, D.K.; Collins, S.R.; Bodenmiller, B.; Aebersold, R.; Simons, K.; Shevchenko, A.; Ejsing, C.S.; Weissman, J.S. Orm family proteins mediate sphingolipid homeostasis. Nature 2010, 463, 1048–1053. [Google Scholar] [CrossRef] [PubMed]
- Han, S.; Lone, M.A.; Schneiter, R.; Chang, A. Orm1 and Orm2 are conserved endoplasmic reticulum membrane proteins regulating lipid homeostasis and protein quality control. Proc. Natl. Acad. Sci. USA 2010, 107, 5851–5856. [Google Scholar] [CrossRef] [PubMed]
- Roelants, F.M.; Breslow, D.K.; Muir, A.; Weissman, J.S.; Thorner, J. Protein kinase Ypk1 phosphorylates regulatory proteins Orm1 and Orm2 to control sphingolipid homeostasis in Saccharomyces cerevisiae. Proc. Natl. Acad. Sci. USA 2011, 108, 19222–19227. [Google Scholar] [CrossRef] [PubMed]
- Niles, B.J.; Mogri, H.; Hill, A.; Vlahakis, A.; Powers, T. Plasma membrane recruitment and activation of the AGC kinase Ypk1 is mediated by target of rapamycin complex 2 (TORC2) and its effector proteins Slm1 and Slm2. Proc. Natl. Acad. Sci. USA 2012, 109, 1536–1541. [Google Scholar] [CrossRef] [PubMed]
- Sun, Y.; Miao, Y.; Yamane, Y.; Zhang, C.; Shokat, K.M.; Takematsu, H.; Kozutsumi, Y.; Drubin, D.G. Orm protein phosphoregulation mediates transient sphingolipid biosynthesis response to heat stress via the Pkh-Ypk and Cdc55-PP2A pathways. Mol. Biol. Cell 2012, 23, 2388–2398. [Google Scholar] [CrossRef] [PubMed]
- Hodgkinson, V.; Petris, M.J. Copper homeostasis at the host-pathogen interface. J. Biol. Chem. 2012, 287, 13549–13555. [Google Scholar] [CrossRef] [PubMed]
- Li, L.; Naseem, S.; Sharma, S.; Konopka, J.B. Flavodoxin-Like Proteins Protect Candida albicans from Oxidative Stress and Promote Virulence. PLoS Pathog. 2015, 11, e1005147. [Google Scholar] [CrossRef] [PubMed]
- Brown, A.J.; Haynes, K.; Quinn, J. Nitrosative and oxidative stress responses in fungal pathogenicity. Curr. Opin. Microbiol. 2009, 12, 384–391. [Google Scholar] [CrossRef] [PubMed]
- Brown, A.J.; Budge, S.; Kaloriti, D.; Tillmann, A.; Jacobsen, M.D.; Yin, Z.; Ene, I.V.; Bohovych, I.; Sandai, D.; Kastora, S.; et al. Stress adaptation in a pathogenic fungus. J. Exp. Biol. 2014, 217, 144–155. [Google Scholar] [CrossRef] [PubMed]
- Chauhan, N.; Latge, J.P.; Calderone, R. Signalling and oxidant adaptation in Candida albicans and Aspergillus fumigatus. Nat. Rev. Microbiol. 2006, 4, 435–444. [Google Scholar] [CrossRef] [PubMed]
- Dantas Ada, S.; Day, A.; Ikeh, M.; Kos, I.; Achan, B.; Quinn, J. Oxidative stress responses in the human fungal pathogen, Candida albicans. Biomolecules 2015, 5, 142–165. [Google Scholar] [CrossRef] [PubMed]
- Pedroso, N.; Gomes-Alves, P.; Marinho, H.S.; Brito, V.B.; Boada, C.; Antunes, F.; Herrero, E.; Penque, D.; Cyrne, L. The plasma membrane-enriched fraction proteome response during adaptation to hydrogen peroxide in Saccharomyces cerevisiae. Free Radic. Res. 2012, 46, 1267–1279. [Google Scholar] [CrossRef] [PubMed]
- North, M.; Tandon, V.J.; Thomas, R.; Loguinov, A.; Gerlovina, I.; Hubbard, A.E.; Zhang, L.; Smith, M.T.; Vulpe, C.D. Genome-wide functional profiling reveals genes required for tolerance to benzene metabolites in yeast. PLoS ONE 2011, 6, e24205. [Google Scholar] [CrossRef] [PubMed]
- Kim, Y.; Chay, K.O.; Kim, I.; Song, Y.B.; Kim, T.Y.; Han, S.J.; Ahn, Y.; Cho, S.H.; Hoe, K.L.; Ahn, B.W.; et al. Redox regulation of the tumor suppressor PTEN by glutaredoxin 5 and Ycp4. Biochem. Biophys. Res. Commun. 2011, 407, 175–180. [Google Scholar] [CrossRef] [PubMed]
- Koch, K.; Hromic, A.; Sorokina, M.; Strandback, E.; Reisinger, M.; Gruber, K.; Macheroux, P. Structure, biochemical and kinetic properties of recombinant Pst2p from Saccharomyces cerevisiae, a FMN-dependent NAD(P)H:quinone oxidoreductase. Biochim. Biophys. Acta 2017, 1865, 1046–1056. [Google Scholar] [CrossRef] [PubMed]
| Protein | Location | Function | Localization Reference | Copies/cell 1 |
|---|---|---|---|---|
| Sur7 | MCC | Sur7 family tetraspan | [9,11] | 17,000 |
| Fmp45 | MCC | Sur7 family tetraspan | [11] | 329 |
| Pun1 | MCC | Sur7 family tetraspan | [16] | 1660 |
| Ynl194c | MCC | Sur7 family tetraspan | [11] | ND |
| Nce102 | MCC | Nce102 family tetraspan | [16] | 1824 |
| Fhn1 | MCC | Nce102 family tetraspan | [16] | ND |
| Can1 | MCC | H+-driven Arg permease | [9] | ND |
| Fur4 | MCC | H+-driven uracil permease | [10] | 3 |
| Tat2 | MCC | H+-driven Trp and Tyr permease | [28] | 752 |
| Pil1 | eisosome | BAR domain | [12] | 115,000 |
| Lsp1 | eisosome | BAR domain | [12] | 104,000 |
| Pkh1 | eisosome | Ser/Thr protein kinase | [30,31] | 221 |
| Pkh2 | eisosome | Ser/Thr protein kinase | [30,31] | 229 |
| Eis1 | eisosome | Unknown | [16] | 5570 |
| Slm1 | eisosome | BAR domain and PH domain | [16,32] | 5190 |
| Slm2 | eisosome | BAR domain and PH domain | [16,32] | 2610 |
| Seg1 | eisosome | Unknown | [18,33] | 982 |
| Mdg1 | eisosome | Unknown | [16] | 1240 |
| Ygr130c | eisosome | Unknown | [16,18] | 10,300 |
| Pst2 | eisosome | Similar to flavodoxin-like proteins | [16] | 2330 |
| Rfs1 | eisosome | Similar to flavodoxin-like proteins | [16] | 7060 |
| Ycp4 | eisosome | Similar to flavodoxin-like proteins | [16] | 14,600 |
| Msc3 | eisosome | Protein of unknown function | [34,35] | 131 |
| Xrn1 | eisosome | Exonuclease | [36] | 11,700 |
© 2017 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Foderaro, J.E.; Douglas, L.M.; Konopka, J.B. MCC/Eisosomes Regulate Cell Wall Synthesis and Stress Responses in Fungi. J. Fungi 2017, 3, 61. https://doi.org/10.3390/jof3040061
Foderaro JE, Douglas LM, Konopka JB. MCC/Eisosomes Regulate Cell Wall Synthesis and Stress Responses in Fungi. Journal of Fungi. 2017; 3(4):61. https://doi.org/10.3390/jof3040061
Chicago/Turabian StyleFoderaro, Jenna E., Lois M. Douglas, and James B. Konopka. 2017. "MCC/Eisosomes Regulate Cell Wall Synthesis and Stress Responses in Fungi" Journal of Fungi 3, no. 4: 61. https://doi.org/10.3390/jof3040061
APA StyleFoderaro, J. E., Douglas, L. M., & Konopka, J. B. (2017). MCC/Eisosomes Regulate Cell Wall Synthesis and Stress Responses in Fungi. Journal of Fungi, 3(4), 61. https://doi.org/10.3390/jof3040061





