Biological and Genomic Insights into Fusarium acuminatum Causing Needle Blight in Pinus tabuliformis
Abstract
1. Introduction
2. Materials and Methods
2.1. Sample Collection and Fungal Isolation
2.2. Pathogenicity Tests on Host Needles
2.3. Molecular Identification and Phylogenetic Analysis of Fungal Isolate
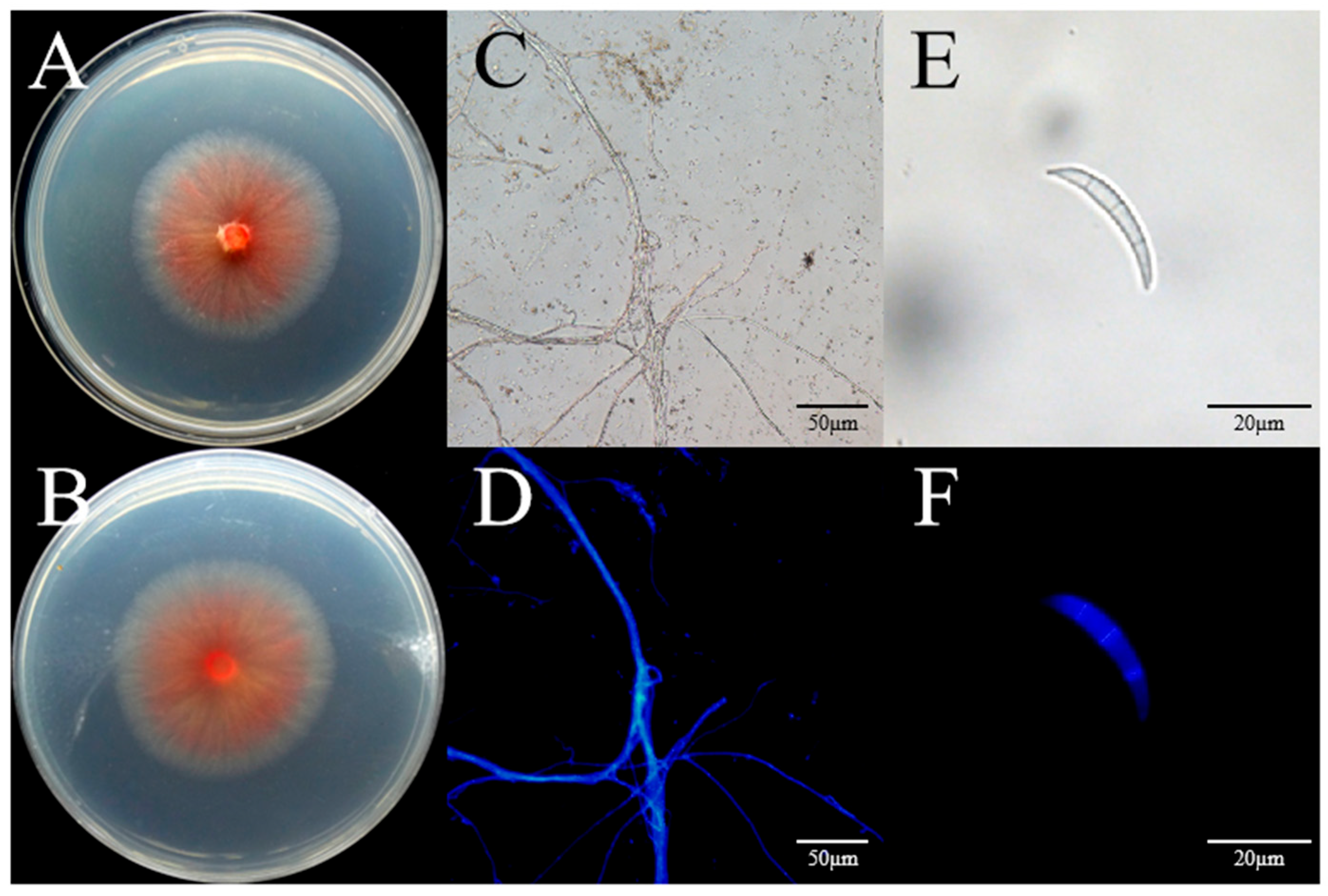
2.4. Morphological Study
2.5. Culture Characteristics of the Representative Strain
2.5.1. Analysis of Growth Characteristics at Different pH Values
2.5.2. Evaluation of Optimum Temperature
2.5.3. Analysis of Preferred Carbon and Nitrogen Sources
2.6. Whole-Genome Sequencing and Assembly
2.6.1. DNA Extraction
2.6.2. Genome Sequencing and Assembly
2.6.3. Gene Prediction and Functional Annotation
2.6.4. Statistical Analysis
3. Results
3.1. Natural Symptoms and Pathogen Isolation
3.2. AP-3 Isolate Is Pathogenic on P.tabuliformis
3.3. Identification of the AP-3 Isolate
3.4. Effect of pH on the Pathogen Growth
3.5. Effect of Temperature on the Pathogen Growth
3.6. Effect of Carbon Sources on the Growth of the AP-3 Isolate
3.7. Effect of Different Nitrogen Sources on the Growth of AP-3
3.8. Genome Sequencing and Assembly of F. acuminatum (AP-3)
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Wang, X.; Zhao, L.; Yang, X.X.; Liu, Z.L. Complete chloroplast genome sequence of Pinus tabuliformis var. henryi (Mast.) C.T.Kuan 1983 (Pinaceae). Mitochondrial DNA B Resour. 2024, 9, 83–87. [Google Scholar] [CrossRef] [PubMed]
- Niu, S.; Li, W.; Li, Y. Chinese pine (Pinus tabuliformis Carr.). Trends Genet. 2022, 38, 409–411. [Google Scholar] [CrossRef] [PubMed]
- Wang, M.B.; Gao, F.Q. Genetic variation in Chinese pine (Pinus tabulaeformis), a woody species endemic to China. Biochem. Genet. 2009, 47, 154–164. [Google Scholar] [CrossRef] [PubMed]
- Chen, K.; Abbott, R.J.; Milne, R.I.; Tian, X.M.; Liu, J. Phylogeography of Pinus tabulaeformis Carr. (Pinaceae), a dominant species of coniferous forest in northern China. Mol. Ecol. 2008, 17, 4276–4288. [Google Scholar] [CrossRef]
- Zhang, Y.; Han, J.; Wang, L.; Jing, X.; Wang, Y.; Liu, P. Response of Pinus tabuliformis saplings to continuous autumn fertilization treatments in the mountains of Eastern Liaoning Province, China. PeerJ 2022, 10, e12853. [Google Scholar] [CrossRef]
- Jin, W.T.; Gernandt, D.S.; Wehenkel, C.; Xia, X.M.; Wei, X.X.; Wang, X.Q. Phylogenomic and ecological analyses reveal the spatiotemporal evolution of global pines. Proc. Natl. Acad. Sci. USA 2021, 118, e2022302118. [Google Scholar] [CrossRef]
- Ning, P.; Zhang, M.; Bai, T.; Zhang, B.; Yang, L.; Dang, S.; Yang, X.; Gao, R. Dendroclimatic response of Pinus tabuliformis Carr. along an altitudinal gradient in the warm temperate region of China. Front. Plant Sci. 2023, 14, 1147229. [Google Scholar] [CrossRef]
- Mesanza, N.; García-García, D.; Raposo, E.R.; Raposo, R.; Iturbide, M.; Pascual, M.T.; Barrena, I.; Urkola, A.; Berano, N.; Sáez de Zerain, A.; et al. Weather variables associated with spore dispersal of Lecanosticta acicola causing Pine needle blight in Northern Spain. Plants 2021, 10, 2788. [Google Scholar] [CrossRef]
- Capron, A.; Feau, N.; Heinzelmann, R.; Barnes, I.; Benowicz, A.; Bradshaw, R.E.; Dale, A.; Lewis, K.J.; Owen, T.J.; Reich, R.; et al. Signatures of post-glacial genetic isolation and human-driven migration in the Dothistroma needle blight pathogen in Western Canada. Phytopathology 2021, 111, 116–127. [Google Scholar] [CrossRef]
- Qiao, C.; Zhao, R.; Li, D.; Ding, X. A new species of biscogniauxia associated with pine needle blight on Pinus thunbergii in China. Forests 2024, 15, 956. [Google Scholar] [CrossRef]
- Qi, M.; Xie, C.X.; Chen, Q.W.; Yu, Z.D. Pestalotiopsis trachicarpicola, a novel pathogen causes twig blight of Pinus bungeana (Pinaceae: Pinoideae) in China. Antonie Leeuwenhoek 2021, 114, 1–9. [Google Scholar] [CrossRef]
- Ding, X.L.; Lin, S.X.; Zhao, R.W.; Ye, J.R. First report of needle blight on Pinus thunbergii caused by Fusarium proliferatum in China. Plant Dis. 2022, 106, 2989. [Google Scholar] [CrossRef]
- Seo, S.T.; Park, M.J.; Park, J.H.; Shin, H.D. First report of brown spot needle blight on Pinus thunbergii caused by Lecanosticta acicola in Korea. Plant Dis. 2012, 96, 914. [Google Scholar] [CrossRef]
- Fabre, B.; Ioos, R.; Piou, D.; Marçais, B. Is the emergence of Dothistroma needle blight of pine in France caused by the cryptic species Dothistroma pini? Phytopathology 2012, 102, 47–54. [Google Scholar] [CrossRef]
- Chen, W.; Fan, X.; Tian, C.; Li, F.; Chou, G. Identification and characterization of Neocosmospora silvicola causing canker disease on Pinus armandii in China. Plant Dis. 2023, 107, 3026–3036. [Google Scholar] [CrossRef]
- Ma, L.J.; Geiser, D.M.; Proctor, R.H.; Rooney, A.P.; O’Donnell, K.; Trail, F.; Gardiner, D.M.; Manners, J.M.; Kazan, K. Fusarium pathogenomics. Annu. Rev. Microbiol. 2013, 67, 399–416. [Google Scholar] [CrossRef] [PubMed]
- Sánchez, C. Fusarium as a promising fungal genus with potential application in bioremediation for pollutants mitigation: A review. Biotechnol. Adv. 2024, 77, 108476. [Google Scholar] [CrossRef] [PubMed]
- Munkvold, G.P. Epidemiology of Fusarium diseases and their mycotoxins in maize ears. Eur. J. Plant Pathol. 2003, 109, 705–713. [Google Scholar] [CrossRef]
- Chakroun, Y.; Oueslati, S.; Pinson-Gadais, L.; Abderrabba, M.; Savoie, J.M. Characterization of Fusarium acuminatum: A potential enniatins producer in Tunisian Wheat. J. Fungi 2022, 8, 458. [Google Scholar] [CrossRef]
- Xu, J.; Jiao, B.; Xia, H.; Dai, T. First report of Fusarium acuminatum causing Dianthus chinensis root rot and foliage blight in China. Plant Dis. 2023, 107, 2254. [Google Scholar] [CrossRef]
- Ignjatov, M.; Bjelic, D.; Nikolic, Z.; Milosevic, D.; Gvozdanovic-Varga, J.; Marinkovic, J.; Ivanovic, Z. First report of Fusarium acuminatum causing garlic bulb rot in Serbia. Plant Dis. 2017, 101, 1047–1048. [Google Scholar] [CrossRef]
- Xie, Y.J.; Sun, Q.; Zhang, S.-L.; Shi, X.C.; Herrera-Balandrano, D.D.; Wang, S.Y.; Laborda, P. First report of Fusarium acuminatum causing leaf blight on garlic in China. Plant Dis. 2023, 107, 213. [Google Scholar] [CrossRef]
- Soufiani, M.G.; Abrinbana, M.; Pouralibaba, H.R. Identification and pathogenicity assessment of Fusarium species associated with grass pea yellowing and wilting in Iran. Physiol. Mol. Plant Pathol. 2025, 136, 102587. [Google Scholar] [CrossRef]
- Parkunan, V.; Ji, P. First report of leaf spot on onion caused by Fusarium acuminatum in Georgia. Plant Dis. 2013, 97, 1384. [Google Scholar] [CrossRef] [PubMed]
- Sun, X.; Yang, R.; Tang, H.; Ma, M.; Chen, H.; Chang, X.; Zhang, M.; Gong, G. Diversity and pathogenicity of Fusarium species associated with Fusarium head blight in wheat and maize cropping systems in Sichuan Province. Sci. Rep. 2025, 15, 5984. [Google Scholar] [CrossRef] [PubMed]
- Jiang, Y.; Zhang, L.; Wang, S.; Li, Y.; Bai, Y.; Cao, L. Evaluation and control of Fusarium acuminatum causing leaf spot in Saposhnikovia divaricata in Northeast China. Plant Dis. 2024, 108, 1139–1145. [Google Scholar] [CrossRef]
- Wang, C.; Hao, D.; Jiao, W.; Li, J.; Yuan, J.; Ma, Y.; Wang, X.; Xu, A.; Wang, M.; Wang, Y. Identification and fungicide sensitivity of Fusarium spp. associated with root rot of Scutellaria baicalensis in Shanxi Province, China. Phytopathology 2024, 114, 1533–1541. [Google Scholar] [CrossRef]
- Vetrova, S.; Alyokhina, K.; Engalycheva, I.; Kozar, E.; Mukhina, K.; Sletova, M.; Krivenkov, L.; Tikhonova, T.; Kameneva, A.; Frolova, S.; et al. Identification and pathogenicity of Fusarium species associated with onion basal rot in the Moscow region of Russian Federation. J. Fungi 2024, 10, 331. [Google Scholar] [CrossRef]
- Yu, X.; Zhang, J.; Zhang, X.; Yang, X.; Xu, X.; Lin, J.; Bing, H.; Wang, X.; Zhao, J.; Xiang, W. Identification and pathogenicity of fungi associated with leaf spot of muskmelon in Eastern Shandong Province, China. Plant Dis. 2022, 106, 872–890. [Google Scholar] [CrossRef]
- Lazreg, F.; Belabid, L.; Sanchez, J.; Gallego, E.; Garrido-Cardenas, J.A.; Elhaitoum, A. First report of Fusarium acuminatum causing damping-off disease on Aleppo pine in Algeria. Plant Dis. 2013, 97, 557. [Google Scholar] [CrossRef]
- Fang, Z.D. Research Methods for Plant Pathology, 3rd ed.; China Agriculture Press: Beijing, China, 1998; pp. 156–168. [Google Scholar]
- Xu, X.; Zhang, L.; Yang, X.; Cao, H.; Li, J.; Cao, P.; Guo, L.; Wang, X.; Zhao, J.; Xiang, W. Alternaria spp. associated with leaf blight of maize in Heilongjiang Province, China. Plant Dis. 2022, 106, 572–584. [Google Scholar] [CrossRef]
- White, T.J.; Bruns, T.D.; Lee, S.B.; Taylor, J.W. Amplification and direct sequencing of fungal ribosomal RNA genes for phylogenetics. In PCR Protocols: A Guide to Methods and Applications; Innis, M.A., Gelfand, D.H., Sninsky, J.J., White, T.J., Eds.; Academic Press: New York, NY, USA, 1990; pp. 315–322. [Google Scholar]
- Lee, S.B.; Taylor, J.W. Isolation of DNA from fungal mycelia and single spores. In PCR Protocols: A Guide to Methods and Applications; Innis, M.A., Gelfand, D.H., Sninsky, J.J., Eds.; Academic Press Inc.: New York, NY, USA, 1990; pp. 282–287. [Google Scholar]
- Kumar, S.; Stecher, G.; Li, M.; Knyaz, C.; Tamura, K. MEGA X: Molecular evolutionary genetics analysis across computing platforms. Mol. Biol. Evol. 2018, 35, 1547–1549. [Google Scholar] [CrossRef]
- O’Donnell, K.; Kistler, H.C.; Cigelnik, E.; Ploetz, R.C. Multiple evolutionary origins of the fungus causing Panama disease of banana: Concordant evidence from nuclear and mitochondrial gene genealogies. Proc. Natl. Acad. Sci. USA 1998, 95, 2044–2049. [Google Scholar] [CrossRef]
- Schoch, C.L.; Robbertse, B.; Robert, V.; Vu, D.; Cardinali, G.; Irinyi, L.; Meyer, W.; Nilsson, R.H.; Hughes, K.; Miller, A.N.; et al. Finding needles in Haystacks: Linking scientific names, reference specimens and molecular data for fungi. Database 2014, 2014, bau061. [Google Scholar] [CrossRef] [PubMed]
- Brankovics, B.; van Diepeningen, A.D.; de Hoog, G.S.; van der Lee, T.A.J.; Waalwijk, C. Detecting introgression between members of the Fusarium fujikuroi and F. oxysporum species complexes by comparative mitogenomics. Front. Microbiol. 2020, 11, 1092. [Google Scholar] [CrossRef] [PubMed]
- Kondo, T. Stem and root rot of gentian caused by Fusarium avenaceum. J. Gen. Plant Pathol. 2019, 85, 440–443. [Google Scholar] [CrossRef]
- Hassan, O.; Chang, T. Morphological and molecular characteristics of fungal species associated with crown rot of strawberry in South Korea. Mol. Biol. Rep. 2022, 49, 51–62. [Google Scholar] [CrossRef]
- Chen, S.Z.; Chen, T.X.; Ji, X.X.; Yang, C.D.; Chen, X.R. Identification of the pathogen of the potato dry rot disease and screening of fungicides in the Minxian District of Gansu Province. Pratac. Sci. 2017, 11, 2218–2225. [Google Scholar]
- Fraga Meizoso, M.; Aguín, O.; López-Burgos, M.; Salinero, C.; Mansilla, J.P. The complexity of mycobiota associated with chestnut galls induced by Dryocosmus kuriphilus in Galicia (Northwestern Spain). iForest 2024, 17, 378–385. [Google Scholar] [CrossRef]
- Kim, H.S.; Lohmar, J.M.; Busman, M.; Brown, D.W.; Naumann, T.A.; Divon, H.H.; Lysøe, E.; Uhlig, S.; Proctor, R.H. Correction to: Identification and distribution of gene clusters required for synthesis of sphingolipid metabolism inhibitors in diverse species of the filamentous fungus Fusarium. BMC Genom. 2020, 21, 712. [Google Scholar] [CrossRef]
- Vu, D.; Groenewald, M.; de Vries, M.; Gehrmann, T.; Stielow, B.; Eberhardt, U.; Al-Hatmi, A.; Groenewald, J.Z.; Cardinali, G.; Houbraken, J.; et al. Large-scale generation and analysis of filamentous fungal DNA barcodes boosts coverage for kingdom fungi and reveals thresholds for fungal species and higher taxon delimitation. Stud. Mycol. 2019, 92, 135–154. [Google Scholar] [CrossRef]
- Stenglein, S.A.; Rodriguero, M.S.; Chandler, E.; Jennings, P.; Salerno, G.L.; Nicholson, P. Phylogenetic relationships of Fusarium poae based on EF-1α and mtSSU sequences. Fungal Biol. 2010, 114, 96–106. [Google Scholar] [CrossRef]
- Abbas, M.F.; Naz, F.; Rauf, C.A.; Mehmood, N.; Zhang, X.; Rosli, B.H.; Gleason, M.L. First report of Fusarium solani causing fruit rot of loquat (Eriobotrya japonica) in Pakistan. Plant Dis. 2017, 101, 839. [Google Scholar] [CrossRef]
- Mahajan, A.; Kaur, L.; Singh, G.; Dhawan, R.K.; Singh, L. Multipotentiality of Luliconazole against various fungal strains: Novel topical formulations and patent review. Recent Adv. Anti-Infect. Drug Discov. 2021, 16, 182–195. [Google Scholar] [CrossRef]
- Laraba, I.; Busman, M.; Geiser, D.M.; O’Donnell, K. Phylogenetic diversity and mycotoxin potential of emergent phytopathogens within the Fusarium tricinctum species complex. Phytopathology 2022, 112, 1284–1298. [Google Scholar] [CrossRef] [PubMed]
- Zhang, N.; O’Donnell, K.; Sutton, D.A.; Nalim, F.A.; Summerbell, R.C.; Padhye, A.A.; Geiser, D.M. Members of the Fusarium solani species complex that cause infections in both humans and plants are common in the environment. J. Clin. Microbiol. 2006, 44, 2186–2190. [Google Scholar] [CrossRef] [PubMed]
- Montville, T.J.; Goldstein, P.K. Sodium bicarbonate reduces viability and alters aflatoxin distribution of Aspergillus parasiticus in Czapek’s agar. Appl. Environ. Microbiol. 1987, 53, 2303–2307. [Google Scholar] [CrossRef] [PubMed]
- Ranallo-Benavidez, T.R.; Jaron, K.S.; Schatz, M.C. GenomeScope 2.0 and Smudgeplot for reference-free profiling of polyploid genomes. Nat. Commun. 2020, 11, 1432. [Google Scholar] [CrossRef]
- Hu, J.; Fan, J.; Sun, Z.; Liu, S. NextPolish:a fast and efficient genome polishing tool for long-read assembly. Bioinformatics 2020, 36, 2253–2255. [Google Scholar] [CrossRef]
- Seppey, M.; Manni, M.; Zdobnov, E. MBUSCO: Assessing genome assembly and annotation completeness. Methods Mol. Biol. 2019, 1962, 227–245. [Google Scholar]
- Li, H.; Durbin, R. Fast and accurate short read alignment with Burrows-Wheeler transform. Bioinformatics 2009, 25, 1754–1760. [Google Scholar] [CrossRef]
- Bruna, T.; Hoff, K.J.; Lomsadze, A.; Stanke, M.; Borodovsky, M. BRAKER2: Automatic eukaryotic genome annotation with GeneMark-EP+ and AUGUSTUS supported by a protein database. NAR Genom. Bioinform. 2021, 3, lqaa108. [Google Scholar] [CrossRef]
- Stanke, M.; Diekhans, M.; Baertsch, R.; Haussler, D. Using native and syntenically mapped cDNA alignments to improve de novo gene finding. Bioinformatics 2008, 24, 637–644. [Google Scholar] [CrossRef] [PubMed]
- Bruna, T.; Lomsadze, A.; Borodovsky, M. GeneMark-EP: Eukaryotic gene prediction with self-training in the space of genes and proteins. NAR Genom. Bioinform. 2020, 2, lqaa026. [Google Scholar] [CrossRef] [PubMed]
- Jones, P.; Binns, D.; Chang, H.Y.; Fraser, M.; Li, W.; McAnulla, C.; McWilliam, H.; Maslen, J.; Mitchell, A.; Nuka, G.; et al. InterProScan 5: Genome-scale protein function classification. Bioinformatics 2014, 30, 1236–1240. [Google Scholar] [CrossRef] [PubMed]
- Huerta-Cepas, J.; Szklarczyk, D.; Heller, D.; Hernández-Plaza, A.; Forslund, S.K.; Cook, H.; Mende, D.R.; Letunic, I.; Rattei, T.; Jensen, L.J.; et al. EggNOG 5.0: A hierarchical, functionally and phylogenetically annotated orthology resource based on 5090 organisms and 2502 viruses. Nucleic Acids Res. 2019, 47, D309–D314. [Google Scholar] [CrossRef]
- Lombard, V.; Golaconda Ramulu, H.; Drula, E.; Coutinho, P.M.; Henrissat, B. The carbohydrate-active enzymes database (CAZy) in 2013. Nucleic Acids Res. 2014, 42, D490–D495. [Google Scholar] [CrossRef]
- Zhang, M.J.; Zheng, X.R.; Li, H.; Chen, F.M. Alternaria alternata, the causal agent of a new needle blight disease on Pinus bungeana. J. Fungi 2023, 9, 71. [Google Scholar] [CrossRef]
- van der Nest, A.; Wingfield, M.J.; Ortiz, P.C.; Barnes, I. Biodiversity of Lecanosticta pine-needle blight pathogens suggests a Mesoamerican Centre of origin. IMA Fungus 2019, 10, 2. [Google Scholar] [CrossRef]
- Wang, Y.H.; Dai, Y.; Kong, W.L.; Zhu, M.L.; Wu, X.Q. Improvement of Sphaeropsis shoot blight disease resistance by applying the ectomycorrhizal fungus Hymenochaete sp. Rl and mycorrhizal helper bacterium Bacillus pumilus HR10 to Pinus thunbergii. Phytopathology 2022, 112, 1226–1234. [Google Scholar] [CrossRef]
- Solheim, H.; Vuorinen, M. First report of Mycosphaerella pini causing red band needle blight on scots pine in Norway. Plant Dis. 2011, 95, 875. [Google Scholar] [CrossRef]
- Li, H.; Peng, B.Y.; Xie, J.Y.; Bai, Y.Q.; Li, D.W.; Zhu, L.H. Pestalotiopsis jiangsuensis sp. nov. causing needle blight on Pinus massoniana in China. J. Fungi 2024, 10, 230. [Google Scholar] [CrossRef] [PubMed]
- Barnes, I.; Kirisits, T.; Wingfield, M.J.; Wingfield, B.D. Needle blight of pine caused by two species of Dothistroma in Hungary. Forest Patholog. 2011, 41, 361–369. [Google Scholar] [CrossRef]
- Lin, R.; Kong, W.L.; Ye, J.R.; Wu, X.Q. First report of Pestalotiopsis clavata, P. chamaeropis and P. lushanensis causing shoot blight on Cedrus deodara in China. Crop Prot. 2025, 190, 107119. [Google Scholar]
- Yang, J.; Wang, S.; Zhang, Y.; Chen, Y.; Zhou, H.; Zhang, G. Identification, culture characteristics and whole-genome analysis of Pestalotiopsis neglecta causing black spot blight of Pinus sylvestris var. mongolica. J. Fungi 2023, 9, 564. [Google Scholar] [CrossRef]
- Shen, S.; Li, W.; Wang, J. Inhibitory activity of Halobacillus trueperi S61 and its active extracts on potato dry rot. Bioengineered 2022, 13, 3852–3867. [Google Scholar] [CrossRef]
- Leslie, J.F.; Summerell, B.A. The Fusarium Laboratory Manual; Blackwell Publishing: Ames, IA, USA, 2006. [Google Scholar]
- O’Donnell, K.; Ward, T.J.; Robert, V.; Crous, P.W.; Geiser, D.M.; Kang, S. DNA sequence-based identification of Fusarium: Current status and future directions. Phytoparasitica 2015, 43, 583–595. [Google Scholar] [CrossRef]
- Srivastava, S.; Kadooka, C.; Uchida, J.Y. Fusarium species as pathogen on orchids. Microbiol. Res. 2018, 207, 188–195. [Google Scholar] [CrossRef]
- Poveda, J.; Martín-García, J.; Zamora-Brauweiler, P.; Pastor, M.; Díez, J.J. Biological control of damping-off by Fusarium oxysporum and F. verticillioides on pine and oak seedlings using edible ectomycorrhizal fungi. Pedobiologia 2024, 105, 150973. [Google Scholar] [CrossRef]
- Fulton, J.C.; Yu, P.L.; Smith, K.E.; Huguet-Tapia, J.C.; Hudson, O.; Meeks, A.; Quesada, T.; McKeever, K.; Brawner, J.T. Comparative genomics of Fusarium circinatum isolates used to screen southern Pines for pitch canker resistance. Mol. Plant Microbe Interact. 2022, 35, 477–487. [Google Scholar] [CrossRef]
- Lazreg, F.; Belabid, L.; Sanchez, J.; Gallego, E.; Garrido-Cardenas, J.A.; Elhaitoum, A. First report of Fusarium equiseti causing damping-off disease on Aleppo Pine in Algeria. Plant Dis. 2014, 98, 1268. [Google Scholar] [CrossRef] [PubMed]
- Lazreg, F.; Belabid, L.; Sanchez, J.; Gallego, E.; Garrido-Cardenas, J.A.; Elhaitoum, A. First report of Fusarium chlamydosporum causing damping-Off disease on Aleppo Pine in Algeria. Plant Dis. 2013, 97, 1506. [Google Scholar] [CrossRef] [PubMed]
- Ju, F.; Qi, Z.; Tan, J.; Dai, T. Development of green fluorescent protein-tagged strains of Fusarium acuminatum via PEG-mediated genetic transformation. Microorganisms 2024, 12, 2427. [Google Scholar] [CrossRef] [PubMed]
- Ding, D.; Shao, Y.; Zhao, J.; Lin, J.; Zhang, X.; Wang, X.; Xu, X.; Xu, C. Identification and pathogenicity of Alternaria and Fusarium species associated with bagged apple black spot disease in Shaanxi, China. Front. Microbiol. 2024, 15, 1457315. [Google Scholar] [CrossRef]
- Cai, Y.B.; Dou, T.; Gao, F.T.; Du, J.F.; Wang, F.Q.; Nian, W.K.; Wang, M.; Zhang, Z.Y.; Liu, X.Y. Identification and biological characteristics of Fusarium acuminatum causing leaf blight of Rehmannia glutinosa. China J. Chin. Mater. Med. 2022, 47, 1824–1830. [Google Scholar]
- Wu, Y.; Xie, L.; Jiang, Y.; Li, T. Prediction of effector proteins and their implications in pathogenicity of phytopathogenic filamentous fungi: A review. Int. J. Biol. Macromol. 2022, 206, 188–202. [Google Scholar] [CrossRef]
- Blin, K.; Shaw, S.; Steinke, K.; Villebro, R.; Ziemert, N.; Lee, S.Y.; Medema, M.H.; Weber, T. AntiSMASH 5.0: Updates to the secondary metabolite genome mining pipeline. Nucleic Acids Res. 2019, 47, W81–W87. [Google Scholar] [CrossRef]
- Marín, P.; Moretti, A.; Ritieni, A.; Jurado, M.; Vázquez, C.; González-Jaén, M.T. Phylogenetic analyses and toxigenic profiles of Fusarium equiseti and F. acuminatum isolated from cereals from Southern Europe. Food Microbiol. 2012, 31, 229–237. [Google Scholar] [CrossRef]
- Schütt, F.; Nirenberg, H.I.; Demi, G. Moniliformin production in the genus Fusarium. Mycotoxin Res. 1998, 14, 35–40. [Google Scholar] [CrossRef]






| Locus a | Primer | Primer Sequence | PCR Conditions | Reference |
|---|---|---|---|---|
| ITS | ITS1 ITS4 | TCCGTAGGTGAACCTGCGG | 94 °C for 5 min (94 °C for 40 s, 58 °C for 40 s, and 72 °C for 60 s) × 35 cycles, 72 °C for 7 min | [33] |
| TCCTCCGCTTATTGATATGC | ||||
| TEF-1α | TEF1 | ATGGGTAAGGAAGACAAGAC | 94 °C for 3 min (94 °C for 60 s, 60 °C for 60 s, and 72 °C for 3 min) × 32 cycles, 72 °C for 5 min | [36] |
| TEF2 | GGAAGTACCAGTGATCATGTT |
| Original Name | Culture Accession Number(s) | Type Status | Accession No. ITS | Accession No. TEF-1α | Reference |
|---|---|---|---|---|---|
| Fusarium acutatum | CBS 402.97 | Type of Fusarium avenaceum | NR111142.1 | MT010989.1 | [37,38] |
| Fusarium avenaceum | NRRL 54939 | Type of Fusarium avenaceum | PP336534.1 | MH582391.1 | [39] |
| Plectosphaerella cucumerina | SJB163 | Type of Plectosphaerella cucumerina | LC633900.1 | LC633926.1 | [40] |
| Fusarium avenaceum | GSLZA-A-2 | Type of Fusarium avenaceum | KX029335.1 | KX029338.1 | [41] |
| Fusarium avenaceum | FU-EFA 1092.12 | Type of Fusarium avenaceum | PP660998.1 | PP726199.1 | [42] |
| Fusarium acuminatum | F829 | Type of Fusarium acuminatum | JABEEU010000081.1(genome) | [43] | |
| Fusarium tricinctum | CBS 393.93 | Type of Fusarium tricinctum | MH862424.1 | LC468080.1 | [44,45] |
| Fusarium fujikuroi | FUS53 | Type of Fusarium fujikuroi | MK630074.1 | LC468051.1 | [46,47] |
| Fusarium iranicum | FRC R8875 | Type of Fusarium iranicum | OL832291.1 | OL772863.1 | [48] |
| Fusarium solani | NRRL 32810 | Type of Fusarium solani | DQ094577.1 | DQ247118.1 | [49] |
| Features | AP-3 |
|---|---|
| ONT * long reads | 5.18 Gb (~125×) |
| Illunima short reads | 3.69 Gb (~89×) |
| Assembly size (bp) | 41,498,617 |
| Contig number | 14 |
| Contig N50 (bp) | 4,642,861 |
| Contig L50 | 4 |
| Average contig length (bp) | 2,964,186 |
| Maximum contig length (bp) | 6,454,297 |
| GC content | 47.97% |
| Repeat sequence | 3.04% |
| Illumina reads mapping rate | 99.10% |
| Protein-coding genes | 13,408 |
| Genes annotated by Pfam # | 9765 |
| Genes annotated by GO # | 4321 |
| Genes annotated by KEGG # | 4788 |
| Genes annotated by KOG # | 10,166 |
| Genes annotated by CAZy # | 421 |
| Pathogen-host interaction genes | 3185 |
| Cytochrome P450 enzymes | 120 |
| Putative secreted proteins | 694 |
| SMBGCs $ | 51 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2025 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Song, L.; Xu, Y.; Liu, T.; Wang, H.; Wang, X.; Fu, C.; Xie, X.; Abubakar, Y.S.; Felix, A.; Yang, R.; et al. Biological and Genomic Insights into Fusarium acuminatum Causing Needle Blight in Pinus tabuliformis. J. Fungi 2025, 11, 636. https://doi.org/10.3390/jof11090636
Song L, Xu Y, Liu T, Wang H, Wang X, Fu C, Xie X, Abubakar YS, Felix A, Yang R, et al. Biological and Genomic Insights into Fusarium acuminatum Causing Needle Blight in Pinus tabuliformis. Journal of Fungi. 2025; 11(9):636. https://doi.org/10.3390/jof11090636
Chicago/Turabian StyleSong, Linin, Yuying Xu, Tianjin Liu, He Wang, Xinyue Wang, Changxiao Fu, Xiaoling Xie, Yakubu Saddeeq Abubakar, Abah Felix, Ruixian Yang, and et al. 2025. "Biological and Genomic Insights into Fusarium acuminatum Causing Needle Blight in Pinus tabuliformis" Journal of Fungi 11, no. 9: 636. https://doi.org/10.3390/jof11090636
APA StyleSong, L., Xu, Y., Liu, T., Wang, H., Wang, X., Fu, C., Xie, X., Abubakar, Y. S., Felix, A., Yang, R., Jing, X., Lu, G., Bao, J., & Ye, W. (2025). Biological and Genomic Insights into Fusarium acuminatum Causing Needle Blight in Pinus tabuliformis. Journal of Fungi, 11(9), 636. https://doi.org/10.3390/jof11090636








