Evaluation of the Potential Risk Posed by Emerging Yr5-Virulent and Predominant Races of Puccinia striiformis f. sp. tritici on Bread Wheat (Triticum aestivum L.) Varieties Grown in Türkiye
Abstract
1. Introduction
2. Materials and Methods
2.1. Genetic Materials
2.2. Multiplication of Pst Urediniospores and Reaction Tests of the Varieties
2.3. Extraction of Genomic DNA
2.4. Molecular Detection of Yr Genes
2.5. Data Analysis
3. Results
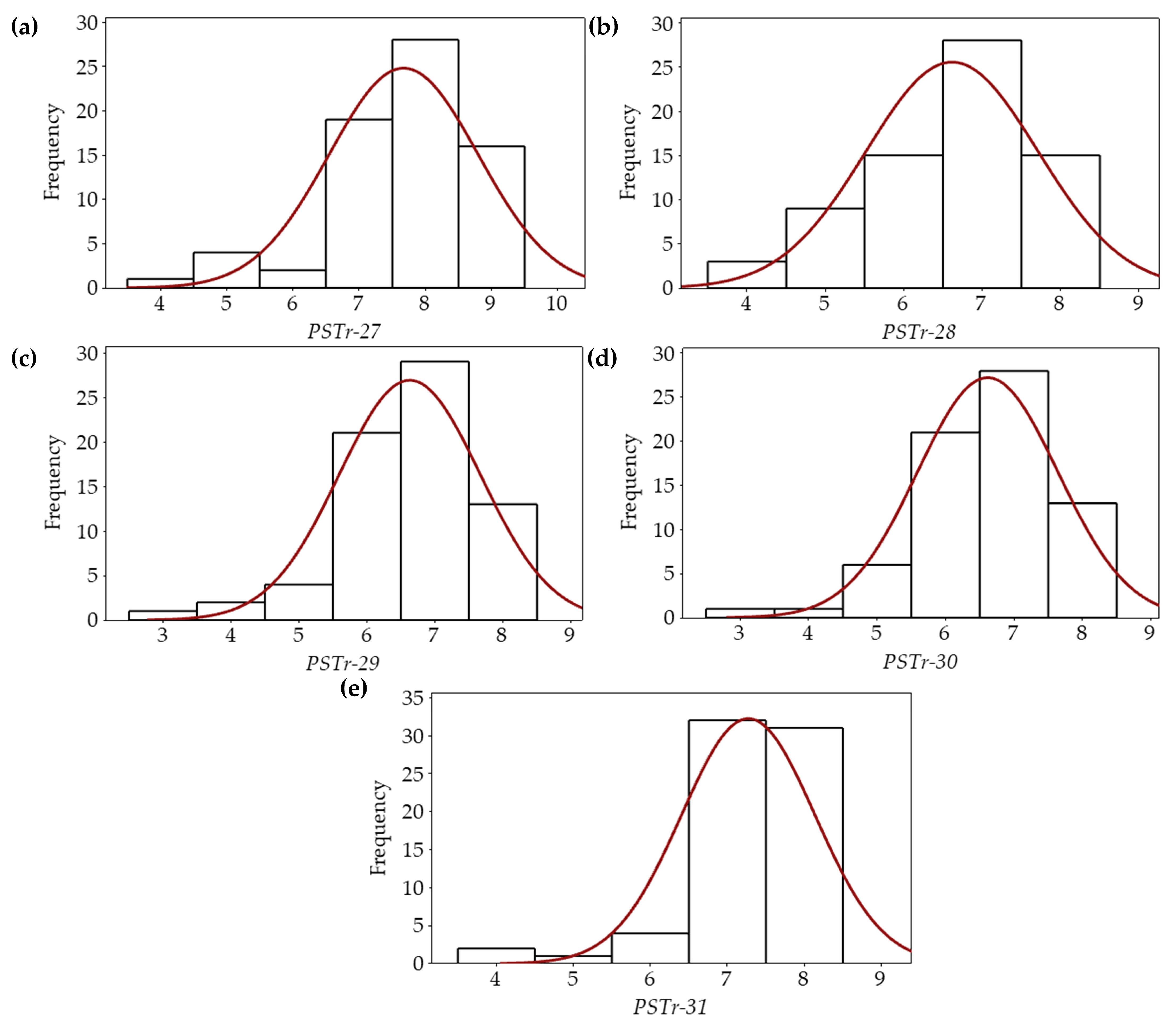
3.1. Reaction Tests of the Bread Wheat Varieties to Pst Races
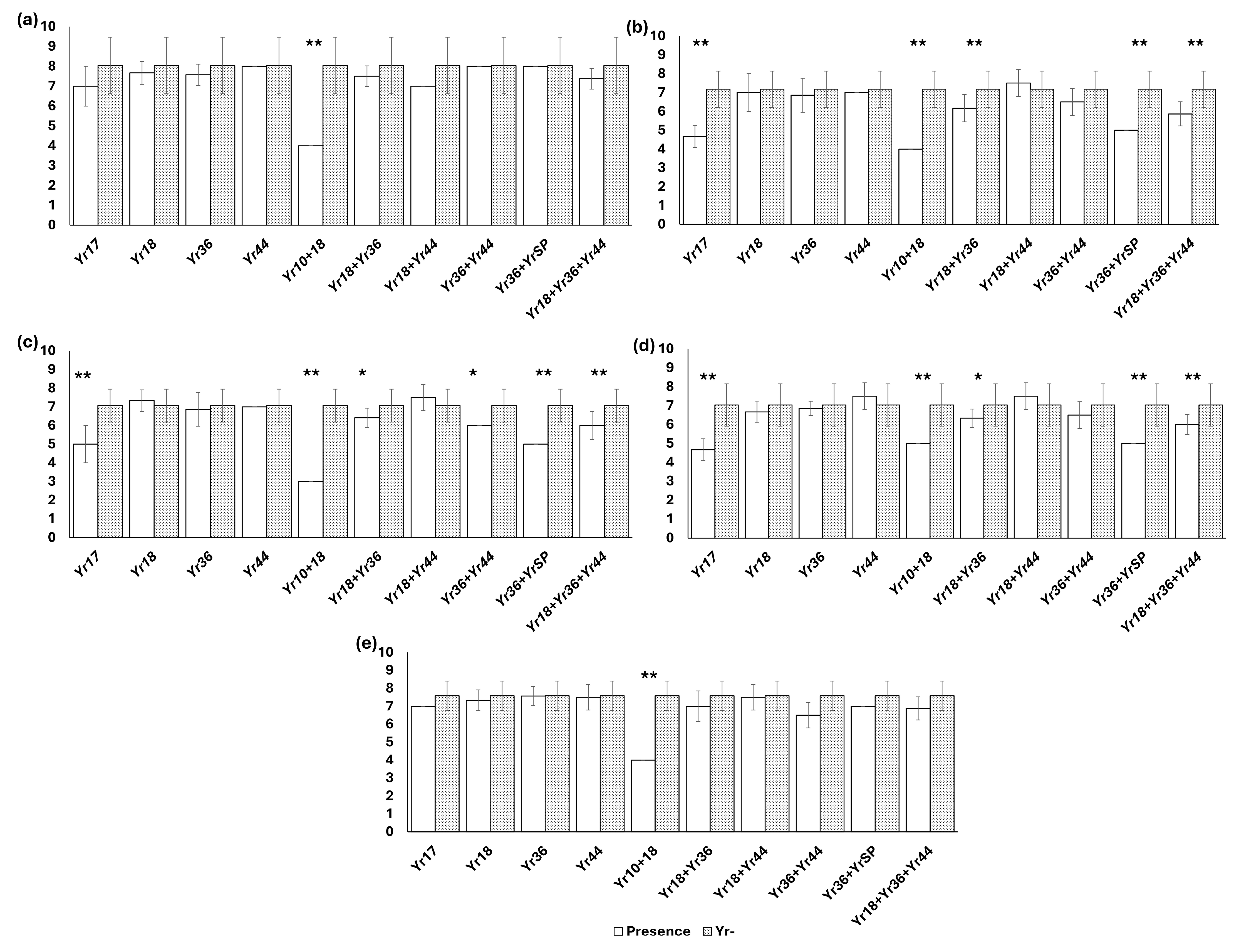
3.2. Molecular Detection of Yr Genes
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- FAOSTAT. Crops and Livestock Products. Available online: https://www.fao.org/faostat/en/#data/QCL (accessed on 24 February 2025).
- Schwessinger, B. Fundamental wheat stripe rust research in the 21st century. New Phytol. 2017, 213, 1625–1631. [Google Scholar] [CrossRef]
- Chen, X.M. Pathogens which threaten food security: Puccinia striiformis, the wheat stripe rust pathogen. Food Secur. 2020, 12, 239–251. [Google Scholar] [CrossRef]
- Braun, H.; Saari, E.E. An assesment of the potential of Puccinia striiformis f. sp. tritici to cause yield losses in wheat on the Anatolian Plateau of Turkey. In Proceedings of the Eight European and Mediterranean Cereal Rusts and Mildews Conference, Weihenstephan, Germany, 8–11 September 1992. [Google Scholar]
- Mamluk, O.F.; Cetin, L.; Braun, H.J.; Bolat, N.; Bertschinger, L.; Makkouk, K.M.; Yildirim, A.F.; Saari, E.E.; Zencirci, N.; Albustan, S.; et al. Current status of wheat and barley diseases in the Central Anatolian Plateau of Turkey. Phytopathol. Mediterr. 1997, 36, 167–181. [Google Scholar]
- Cat, A.; Tekin, M.; Akan, K.; Akar, T.; Catal, M. Virulence characterization of the wheat stripe rust pathogen, Puccinia striiformis f. sp. tritici, in Turkey from 2018 to 2020. Can. J. Plant Pathol. 2023, 45, 158–167. [Google Scholar] [CrossRef]
- Akan, K. Evaluation of some lines selected from bread wheat landraces for their reaction to yellow (stripe) Rust. Fresen Environ. Bull. 2019, 28, 6339–6351. [Google Scholar]
- Cat, A.; Tekin, M.; Catal, M.; Akan, K.; Akar, T. Wheat stripe rust and breeding studies for resistance to the disease. Mediterr. Agric. Sci. 2017, 30, 97–105. [Google Scholar]
- Louwers, J.M.; van Silfhout, C.H.; Stubbs, R.W. Race Analysis in Wheat in Developing Countries; Report 1992-11; Research Institute for Plant Protection (IPO-DLO): Wageningen, The Netherlands, 1992. [Google Scholar]
- Mert, Z.; Dusunceli, F.; Akan, K.; Cetin, L.; Yazar, S.; Bolat, N.; Yorgancilar, A.; Unsal, R.; Ercan, B.; Ozseven, I.; et al. An overview of the network for important cereal diseases management research in Turkey between 2003 and 2011. In Proceedings of the 13th International Cereal Rusts and Powdery Mildews Conference, Beijing, China, 28 August–1 September 2012; pp. 208–209. [Google Scholar]
- Wellings, C.R.; Singh, R.P.; McIntosch, R.A.; Pretorius, Z.A. The development and application of near isogenic lines for the stripe (yellow) rust pathosystem. In Proceedings of the 11th International Cereal Rusts and Powdery Mildew Conference, Norwich, UK, 22–27 August 2004; p. Abstr. A1.39. [Google Scholar]
- Cheng, P.; Chen, X.M. Molecular mapping of a gene for stripe rust resistance in spring wheat cultivar IDO377s. Theor. Appl. Genet. 2010, 121, 195–204. [Google Scholar] [CrossRef]
- Wan, A.M.; Chen, X.M. Virulence characterization of Puccinia striiformis f. sp. tritici using a new set of Yr single-gene line differentials in the United States in 2010. Plant Dis. 2014, 98, 1534–1542. [Google Scholar] [CrossRef]
- Cat, A.; Tekin, M.; Akan, K.; Akar, T.; Catal, M. Races of Puccinia striiformis f. sp. tritici identified from the coastal areas of Turkey. Can. J. Plant Pathol. 2021, 43, S323–S332. [Google Scholar] [CrossRef]
- Chen, X.M.; Wang, M.N.; Wan, A.M.; Bai, Q.; Li, M.J.; Lopez, P.F.; Maccaferri, M.; Mastrangelo, A.M.; Barnes, C.W.; Cruz, D.F.C.; et al. Virulence characterization of Puccinia striiformis f. sp. tritici collections from six countries in 2013 to 2020. Can. J. Plant Pathol. 2021, 43, S308–S322. [Google Scholar] [CrossRef]
- McIntosh, R.A.; Dubcovsky, J.; Rogers, W.J.; Xia, X.C.; Raupp, W.J. Catalogue of gene symbols for wheat: 2020 supplement. GrainGenes 2020, 66, 109–128. [Google Scholar]
- Tekin, M.; Cat, A.; Akan, K.; Catal, M.; Akar, T. A new virulent race of wheat stripe rust pathogen (Puccinia striiformis f. sp. tritici) on the resistance gene Yr5 in Turkey. Plant Dis. 2021, 105, 3292. [Google Scholar] [CrossRef]
- Nagarajan, S.; Nayar, S.K.; Bahadur, P. Race-13 (67s8) of Puccinia striiformis virulent on Triticum spelta var album in India. Plant Dis. 1986, 70, 173. [Google Scholar] [CrossRef]
- Wellings, C.R.; McIntosch, R.A. Puccinia striiformis f.sp. tritici in Australasia: Pathogenic changes during the first 10 years. Plant Pathol. 1990, 39, 316–325. [Google Scholar] [CrossRef]
- Zhang, G.S.; Zhao, Y.Y.; Kang, Z.S.; Zhao, J. First report of a Puccinia striiformis f. sp. tritici race virulent to wheat stripe rust resistance gene Yr5 in China. Plant Dis. 2020, 104, 284–285. [Google Scholar] [CrossRef]
- Kharouf, S.; Hamzeh, S.; Azmeh, S.F. Races identification of wheat rusts in Syria during the 2019 growing season. Arab. J. Plant Prot. 2021, 39, 1–13. [Google Scholar] [CrossRef]
- El Amil, R.; Ali, S.; Bahri, B.; Leconte, M.; de Vallavieille-Pope, C.; Nazari, K. Pathotype diversification in the invasive PstS2 clonal lineage of Puccinia striiformis f. sp. tritici causing yellow rust on durum and bread wheat in Lebanon and Syria in 2010-2011. Plant Pathol. 2020, 69, 618–630. [Google Scholar] [CrossRef]
- Akci, N.; Karakaya, A. Puccinia graminis f. sp. tritici races identified on wheat and Berberis spp. in northern Turkey. Indian Phytopathol. 2021, 74, 1105–1109. [Google Scholar] [CrossRef]
- Tekin, M.; Cat, A.; Akar, T.; Catal, M. Super spreaders of wheat stripe rust pathogen (Puccinia striiformis f. sp. tritici) in Turkey. In Proceedings of the 3rd International Eurasian Conference on Biological and Chemical Sciences, Ankara, Türkiye, 19–20 March 2020; p. 568. [Google Scholar]
- Singh, R.P.; William, H.M.; Huerta-Espino, J.; Rosewarne, G. Wheat rust in Asia: Meeting the challenges with old and new technologies. In Proceedings of the 4th International Crop Science Congress, Brisbane, Australia, 26 September–1 October 2004; pp. 1–13. [Google Scholar]
- Zhang, G.S.; Liu, W.; Wang, L.; Cheng, X.R.; Tian, X.X.; Du, Z.M.; Kang, Z.S.; Zhao, J. Evaluation of the potential risk of the emerging virulent races of Puccinia striiformis f. sp. to 165 Chinese wheat cultivars. Plant Dis. 2022, 106, 1867–1874. [Google Scholar] [CrossRef]
- Sorensen, C.K.; Thach, T.; Hovmoller, M.S. Evaluation of spray and point inoculation methods for the phenotyping of Puccinia striiformis on wheat. Plant Dis. 2016, 100, 1064–1070. [Google Scholar] [CrossRef]
- McNeal, F.H.; Konzak, C.F.; Smith, E.P.; Tate, W.S.; Russell, T.S. A Uniform System for Recording and Processing Cereal Research Data; REP-10904; CIMMYT: Mexico City, Mexico, 1971. [Google Scholar]
- Chen, X.M.; Soria, M.A.; Yan, G.P.; Sun, J.; Dubcovsky, J. Development of sequence tagged site and cleaved amplified polymorphic sequence markers for wheat stripe rust resistance gene Yr5. Crop Sci. 2003, 43, 2058–2064. [Google Scholar] [CrossRef]
- Singh, R.; Datta, D.; Priyamvada; Singh, S.; Tiwari, R. A diagnostic PCR based assay for stripe rust resistance gene Yr10 in wheat. Acta Phytopathol. Entomol. Hung. 2009, 44, 11–18. [Google Scholar] [CrossRef]
- Murphy, L.R.; Santra, D.; Kidwell, K.; Yan, G.P.; Chen, X.M.; Campbell, K.G. Linkage maps of wheat stripe rust resistance genes Yr5 and Yr15 for use in marker-assisted selection. Crop Sci. 2009, 49, 1786–1790. [Google Scholar] [CrossRef]
- Helguera, M.; Khan, I.A.; Kolmer, J.; Lijavetzky, D.; Zhong-qi, L.; Dubcovsky, J. PCR assays for the Lr37-Yr17-Sr38 cluster of rust resistance genes and their use to develop isogenic hard red spring wheat lines. Crop Sci. 2003, 43, 1839–1847. [Google Scholar] [CrossRef]
- Krattinger, S.G.; Lagudah, E.S.; Spielmeyer, W.; Singh, R.P.; Huerta-Espino, J.; McFadden, H.; Bossolini, E.; Selter, L.L.; Keller, B. A putative ABC transporter confers durable resistance to multiple fungal pathogens in wheat. Science 2009, 323, 1360–1363. [Google Scholar] [CrossRef] [PubMed]
- Zhang, X.J.; Han, D.J.; Zeng, Q.D.; Duan, Y.H.; Yuan, F.P.; Shi, J.D.; Wang, Q.L.; Wu, J.H.; Huang, L.L.; Kang, Z.S. Fine mapping of wheat stripe rust resistance gene Yr26 based on collinearity of wheat with Brachypodium distachyon and rice. PLoS ONE 2013, 8, e57885. [Google Scholar] [CrossRef] [PubMed]
- Fu, D.L.; Uauy, C.; Distelfeld, A.; Blechl, A.; Epstein, L.; Chen, X.M.; Sela, H.A.; Fahima, T.; Dubcovsky, J. A Kinase-START gene confers temperature-dependent resistance to wheat stripe rust. Science 2009, 323, 1357–1360. [Google Scholar] [CrossRef] [PubMed]
- Sui, X.X.; Wang, M.N.; Chen, X.M. Molecular mapping of a stripe rust resistance gene in spring wheat cultivar Zak. Phytopathology 2009, 99, 1209–1215. [Google Scholar] [CrossRef]
- Feng, J.Y.; Wang, M.N.; Chen, X.M.; See, D.R.; Zheng, Y.L.; Chao, S.M.; Wan, A.M. Molecular mapping of YrSP and its relationship with other genes for stripe rust resistance in wheat chromosome 2BL. Phytopathology 2015, 105, 1206–1213. [Google Scholar] [CrossRef]
- Jeger, M.J.; Fielder, H.; Beale, T.; Szyniszewska, A.M.; Parnell, S.; Cunniffe, N.J. What can be learned by a synoptic review of plant disease epidemics and outbreaks published in 2021? Phytopathology 2023, 113, 1141–1158. [Google Scholar] [CrossRef]
- Fielder, H.; Beale, T.; Jeger, M.J.; Oliver, G.; Parnell, S.; Szyniszewska, A.M.; Taylor, P.; Cunniffe, N.J. A synoptic review of plant disease epidemics and outbreaks published in 2022. Phytopathology 2024, 114, 1717–1732. [Google Scholar] [CrossRef] [PubMed]
- Singh, R.P.; Hodson, D.P.; Huerta-Espino, J.; Jin, Y.; Bhavani, S.; Njau, P.; Herrera-Foessel, S.; Singh, P.K.; Singh, S.; Govindan, V. The emergence of Ug99 races of the stem rust fungus is a threat to world wheat production. Annu. Rev. Phytopathol. 2011, 49, 465–481. [Google Scholar] [CrossRef] [PubMed]
- Zhang, G.S.; Sun, M.D.; Ma, X.Y.; Liu, W.; Du, Z.M.; Kang, Z.S.; Zhao, J. Yr5-virulent races of Puccinia striiformis f. sp. tritici possess relative parasitic fitness higher than current main predominant races and potential risk. J. Integr. Agric. 2024, 23, 2674–2685. [Google Scholar] [CrossRef]
- Coriton, O.; Jahier, J.; Leconte, M.; Huteau, V.; Trotoux, G.; Dedryver, F.; de-Vallavieille-Pope, C. Double dose efficiency of the yellow rust resistance gene Yr17 in bread wheat lines. Plant Breed. 2020, 139, 263–271. [Google Scholar] [CrossRef]
- Liu, R.; Lu, J.; Zhou, M.; Zheng, S.G.; Liu, Z.H.; Zhang, C.H.; Du, M.; Wang, M.X.; Li, Y.F.; Wu, Y.; et al. Developing stripe rust resistant wheat (Triticum aestivum L.) lines with gene pyramiding strategy and marker-assisted selection. Genet. Resour. Crop Evol. 2020, 67, 381–391. [Google Scholar] [CrossRef]
- Wang, L.C.; Tang, X.R.; Wu, J.H.; Shen, C.; Dai, M.F.; Wang, Q.L.; Zeng, Q.D.; Kang, Z.S.; Wu, Y.F.; Han, D.J. Stripe rust resistance to a burgeoning Puccinia striiformis f. sp. tritici race CYR34 in current Chinese wheat cultivars for breeding and research. Euphytica 2019, 215, 68. [Google Scholar] [CrossRef]
- Yazdani, M.; Yassaie, M.; Rezaee, S.; Zakeri, A.K.; Patpour, M. Exploring Iranian wheat landraces for stripe rust resistance genes and validation of selected genes using molecular markers. Can. J. Plant Pathol. 2023, 45, 451–462. [Google Scholar] [CrossRef]
- Eagles, H.A.; Bariana, H.S.; Ogbonnaya, F.C.; Rebetzke, G.J.; Hollamby, G.J.; Henry, R.J.; Henschke, P.H.; Carter, M. Implementation of markers in Australian wheat breeding. Aust. J. Agric. Res. 2001, 52, 1349–1356. [Google Scholar] [CrossRef]
- Baloch, F.S.; Ali, A.; Tajibayev, D.; Nadeem, M.A.; Ölmez, F.; Aktas, H.; Alsaleh, A.; Coemertpay, G.; Imren, M.; Mustafa, Z.; et al. Stripe rust resistance gene in Turkish and Kazakhstan wheat germplasms and the potential of Turkish wild emmer for stripe rust breeding. Genet. Resour. Crop Evol. 2024, 71, 2699–2719. [Google Scholar] [CrossRef]
- Abbas, S.; Li, Y.F.; Lu, J.; Hu, J.M.; Zhang, X.N.; Lv, X.; Shahzad, A.; Ao, D.H.; Abbas, M.; Wu, Y.; et al. Evaluation of stripe rust resistance and analysis of resistance genes in wheat genotypes from Pakistan and Southwest China. Front. Plant Sci. 2024, 15, 1494566. [Google Scholar] [CrossRef] [PubMed]
- Ul Islam, B.; Mir, S.; Dar, M.S.; Khan, G.H.; Shikari, A.B.; Sofi, N.U.; Mohiddin, F.; Ahangar, M.A.; Jehangir, I.A.; Kumar, S.; et al. Characterization of pre-breeding wheat (Triticum aestivum L.) germplasm for stripe rust resistance using field phenotyping and genotyping. Plants 2023, 12, 3239. [Google Scholar] [CrossRef] [PubMed]
- Sucur, R.; Ali, A.; Mortazavi, P.; Mladenov, V.; Jockovic, B.; Gou, J.Y.; Nadeem, M.A.; Baloch, F.S.; Chung, Y.S. Molecular screening of stripe rust and powdery mildew resistance genes in European bread wheat using the validated gene-specific SSR markers. Physiol. Mol. Plant Pathol. 2025, 139, 102743. [Google Scholar] [CrossRef]
- Leharwan, M.; Singh, A.K.; Kumar, A.; Kashyap, P.L.; Kumar, S.; Singh, R.; Gangwar, O.P. Phenotyping and deciphering genetic resistance to yellow rust in wheat through marker-assisted analysis. Physiol. Mol. Plant Pathol. 2025, 139, 102757. [Google Scholar] [CrossRef]
- Kokhmetova, A.; Rsaliyev, A.; Malysheva, A.; Atishova, M.; Kumarbayeva, M.; Keishilov, Z. Identification of stripe rust resistance genes in common wheat cultivars and breeding lines from Kazakhstan. Plants 2021, 10, 2303. [Google Scholar] [CrossRef] [PubMed]
| Gene | Marker Type | Marker | Annealing Temp (°C) | Sequence (5′→3′) | Expected Fragment (bp) | Reference |
|---|---|---|---|---|---|---|
| Yr5 | STS/CAPS | STS7 | 45 | GTACAATTCACCTAGAGT | 289 (+) | [29] |
| STS8 | GCAAGTTTTCTCCCTAT | 182 (−) | ||||
| Yr10 | Gene-specific | Yr 10 F | 64 | TCAAAGACATCAAGAGCCGC | 543 (+) | [30] |
| Yr 10 R | TGGCCTACATGAACTCTGGAT | |||||
| Yr15 | SSR | Xbarc8 F | 50 | GCGGGGGCGAAACATACACATAAAAACA | 200 (+) | [31] |
| Xbarc8 R | GCGGGAATCATGCATAGGAAAACAGAA | 280 (−) | ||||
| Yr17 | SCAR | VENTRIUP | 65 | AGGGGCTACTGACCAAGGCT | 262 (+) | [32] |
| LN2 | TGCAGCTACAGCAGTATGTACACAAAA | |||||
| Yr18 | Gene-specific | L34DINT9F | 51 | TTGATGAAACCAGTTTTTTTTCTA | 517 (+) | [33] |
| L34PLUSR | GCCATTTAACATAATCATGATGGA | |||||
| Yr26 | STS | we173 | 57 | GGGACAAGGGGAGTTGAAGC | 451/500 (+) | [34] |
| GAGAGTTCCAAGCAGAACAC | 730 (−) | |||||
| Yr36 | Gene-specific | Yr36START | 52 | GGCCACACTGCAATACTATACC | 871 (+) | [35] |
| CACAAATCCTGGCTGTGGAC | ||||||
| Yr44 | STS | pWB5 | 49 | GGTGCAATTTGAGTTTGGAGT | 380 (+) | [36] |
| pWN1R1 | GGTGTTGACTGGAGAATCCG | |||||
| YrSP | STS | dp269 | 55 | CTGCTGTCACCGCTCTCC | 190 (+) | [37] |
| AGTCACACGCCCTACTCTCC | 201 (−) |
| Variety | Pst Races | ||||
|---|---|---|---|---|---|
| PSTr-27 | PSTr-28 | PSTr-29 | PSTr-30 | PSTr-31 | |
| Bezostaja-1 | S | MR | MR | MR | S |
| Kıraç 66 | MR | MR | R | MR | MR |
| Cumhuriyet 75 | S | S | S | S | S |
| Gerek 79 | S | S | S | S | S |
| Atay-85 | S | S | S | S | S |
| Kate A-1 | S | MR | MR | MR | S |
| Dağdaş 94 | S | MR | MR | MR | MR |
| Sultan 95 | S | S | S | MR | S |
| Kaşifbey 95 | S | MR | S | MR | S |
| İkizce 96 | MR | S | S | S | S |
| Pamukova 97 | S | S | S | S | S |
| Pehlivan | S | S | MR | S | S |
| Karacadağ 98 | S | S | S | S | S |
| Gönen 98 | S | MR | MR | MR | MR |
| Ziyabey 98 | MR | MR | MR | MR | S |
| Karahan-99 | S | MR | S | MR | S |
| Ceyhan-99 | S | MR | MR | MR | S |
| Flamura 85 | S | MR | S | S | S |
| Bayraktar 2000 | MR | MR | MR | MR | S |
| Demir 2000 | S | S | S | S | S |
| Sönmez 2001 | S | MR | MR | MR | S |
| Alparslan | S | MR | MR | S | S |
| Pandas | S | S | S | S | S |
| Sagittario | S | MR | MR | MR | S |
| Canik 2003 | S | S | S | S | S |
| Tosunbey | S | S | MR | MR | S |
| Ahmetağa | S | S | S | S | S |
| Krasunia odes’ka | S | S | S | S | S |
| Kenanbey | S | S | MR | S | S |
| Aldane | S | S | S | S | S |
| Selimiye | S | MR | S | MR | S |
| ES 26 | S | S | MR | S | S |
| Esperia | S | S | MR | S | MR |
| Cömert | S | S | S | S | S |
| Altındane | MR | S | S | S | S |
| Vittorio | S | MR | MR | MR | S |
| Quality | S | S | S | S | S |
| Rumeli | S | MR | S | MR | MR |
| Aglika | S | MR | MR | MR | S |
| Dinç | MR | MR | MR | R | MR |
| Gökkan | S | S | S | S | S |
| Segor | S | S | S | S | S |
| Adelaide | S | MR | MR | MR | S |
| Avorio | S | MR | MR | MR | S |
| Tekin | S | S | S | S | S |
| Nevzatbey | S | S | MR | S | S |
| Yakamoz | S | S | S | S | S |
| Bora | S | MR | MR | MR | S |
| Genesi | S | MR | MR | MR | MR |
| Glosa | S | S | S | S | S |
| Masaccio | S | S | S | S | S |
| Efe | S | S | MR | MR | S |
| Kale | S | S | S | MR | S |
| Yüksel | S | S | S | S | S |
| Leuta | S | S | S | S | S |
| Duru | S | S | S | S | S |
| Hüseyinbey | S | MR | MR | MR | S |
| Albachiara | S | S | S | S | S |
| Damla | S | S | S | S | S |
| Koç | S | MR | S | S | S |
| İzvor | S | S | S | S | S |
| Anafarta | S | S | S | S | S |
| Abide | S | S | S | S | S |
| Eylül | S | S | S | S | S |
| Albaşak | S | S | S | S | S |
| Boldane | S | MR | MR | MR | S |
| Beyaz-I | S | S | S | S | S |
| Shiro | MR | MR | MR | MR | S |
| Destra | S | S | S | S | S |
| Alba | S | S | S | MR | S |
| No of Variety | Pst Races | ||||
|---|---|---|---|---|---|
| PSTr-27 | PSTr-28 | PSTr-29 | PSTr-30 | PSTr-31 | |
| 2 | + | − | − | − | − |
| 2 | − | + | − | − | − |
| 4 | − | − | + | − | − |
| 3 | − | − | − | + | − |
| 1 | − | + | + | − | − |
| 3 | − | + | − | + | − |
| 2 | − | − | + | + | − |
| 1 | − | − | + | − | + |
| 12 | − | + | + | + | − |
| 1 | − | + | − | + | + |
| 3 | + | + | + | + | − |
| 3 | − | + | + | + | + |
| 2 | + | + | + | + | + |
| 31 | − | − | − | − | − |
| Total | 7 | 27 | 28 | 29 | 7 |
| Race | PSTr-27 | PSTr-28 | PSTr-29 | PSTr-30 | PSTr-31 |
|---|---|---|---|---|---|
| PSTr-27 | 1 | ||||
| PSTr-28 | 0.519 ** | 1 | |||
| PSTr-29 | 0.527 ** | 0.727 ** | 1 | ||
| PSTr-30 | 0.516 ** | 0.770 ** | 0.721 ** | 1 | |
| PSTr-31 | 0.464 ** | 0.601 ** | 0.581 ** | 0.543 ** | 1 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2025 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Akan, K.; Cat, A.; Yurduseven, M.; Tekin, Y.S.; Yeken, M.Z.; Tekin, M. Evaluation of the Potential Risk Posed by Emerging Yr5-Virulent and Predominant Races of Puccinia striiformis f. sp. tritici on Bread Wheat (Triticum aestivum L.) Varieties Grown in Türkiye. J. Fungi 2025, 11, 635. https://doi.org/10.3390/jof11090635
Akan K, Cat A, Yurduseven M, Tekin YS, Yeken MZ, Tekin M. Evaluation of the Potential Risk Posed by Emerging Yr5-Virulent and Predominant Races of Puccinia striiformis f. sp. tritici on Bread Wheat (Triticum aestivum L.) Varieties Grown in Türkiye. Journal of Fungi. 2025; 11(9):635. https://doi.org/10.3390/jof11090635
Chicago/Turabian StyleAkan, Kadir, Ahmet Cat, Medine Yurduseven, Yesim Sila Tekin, Mehmet Zahit Yeken, and Mehmet Tekin. 2025. "Evaluation of the Potential Risk Posed by Emerging Yr5-Virulent and Predominant Races of Puccinia striiformis f. sp. tritici on Bread Wheat (Triticum aestivum L.) Varieties Grown in Türkiye" Journal of Fungi 11, no. 9: 635. https://doi.org/10.3390/jof11090635
APA StyleAkan, K., Cat, A., Yurduseven, M., Tekin, Y. S., Yeken, M. Z., & Tekin, M. (2025). Evaluation of the Potential Risk Posed by Emerging Yr5-Virulent and Predominant Races of Puccinia striiformis f. sp. tritici on Bread Wheat (Triticum aestivum L.) Varieties Grown in Türkiye. Journal of Fungi, 11(9), 635. https://doi.org/10.3390/jof11090635






