Zebrafish as a New Tool in Heart Preservation Research
Abstract
1. Introduction
2. Current Status in Heart Preservation Research
2.1. Gold Standard in Heart Preservation
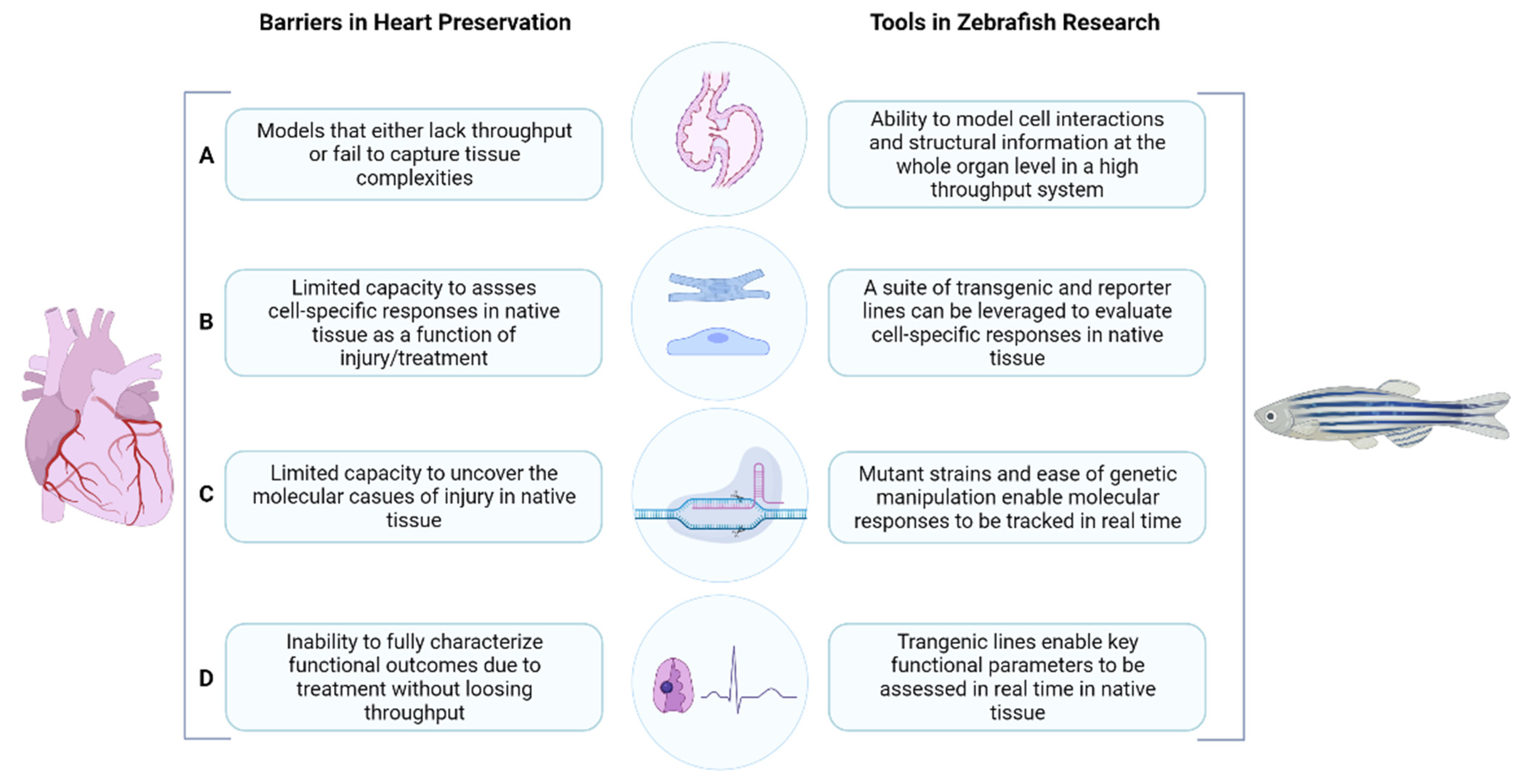
2.2. Barriers and New Technologies in Heart Preservation
3. The Zebrafish: Our Solution to Overcome Barriers in Heart Preservation Research
4. Tools in Zebrafish Research
4.1. Isolated Organ and Transplantation Assays
4.2. Cell-Specific Assays
4.3. Molecular Assays
4.4. Functional Assays
5. Conclusions
Author Contributions
Funding
Conflicts of Interest
References
- Alraies, M.C.; Eckman, P. Adult heart transplant: Indications and outcomes. J. Thorac. Dis. 2014, 6, 1120–1128. [Google Scholar]
- Bhagra, S.K.; Pettit, S.; Parameshwar, J. Cardiac transplantation: Indications, eligibility and current outcomes. Heart 2019, 105, 252. [Google Scholar] [CrossRef]
- Giwa, S.; Lewis, J.K.; Alvarez, L.; Langer, R.; Roth, A.E.; Church, G.M.; Markmann, J.F.; Sachs, D.H.; Chandraker, A.; Wertheim, J.A.; et al. The promise of organ and tissue preservation to transform medicine. Nat. Biotechnol. 2017, 35, 530–542. [Google Scholar] [CrossRef] [PubMed]
- Farr, M.; Truby, L.K.; Lindower, J.; Jorde, U.; Taylor, S.; Chen, L.; Gass, A.; Stevens, G.; Reyentovich, A.; Mancini, D.; et al. Potential for donation after circulatory death heart transplantation in the United States: Retrospective analysis of a limited UNOS dataset. Am. J. Transplant. 2020, 20, 525–529. [Google Scholar] [CrossRef] [PubMed]
- Khush, K.K.; Menza, R.; Nguyen, J.; Zaroff, J.G.; Goldstein, B.A. Donor predictors of allograft use and recipient outcomes after heart transplantation. Circ. Heart Fail. 2013, 6, 300–309. [Google Scholar] [CrossRef]
- Fahy, G.M.; Wowk, B.; Wu, J. Cryopreservation of complex systems: The missing link in the regenerative medicine supply chain. Rejuvenation Res. 2006, 9, 279–291. [Google Scholar] [CrossRef] [PubMed]
- Lewis, J.K.; Bischof, J.C.; Braslavsky, I.; Brockbank, K.G.; Fahy, G.M.; Fuller, B.J.; Rabin, Y.; Tocchio, A.; Woods, E.J.; Wowk, B.G.; et al. The Grand Challenges of Organ Banking: Proceedings from the first global summit on complex tissue cryopreservation. Cryobiology 2016, 72, 169–182. [Google Scholar] [CrossRef] [PubMed]
- Ishine, N.; Rubinsky, B.; Lee, C.Y. Transplantation of mammalian livers following freezing: Vascular damage and functional recovery. Cryobiology 2000, 40, 84–89. [Google Scholar] [CrossRef] [PubMed]
- Berendsen, T.A.; Bruinsma, B.G.; Puts, C.F.; Saeidi, N.; Usta, O.B.; Uygun, B.E.; Izamis, M.L.; Toner, M.; Yarmush, M.L.; Uygun, K.; et al. Supercooling enables long-term transplantation survival following 4 days of liver preservation. Nat. Med. 2014, 20, 790–793. [Google Scholar] [CrossRef] [PubMed]
- Seguchi, R.; Watanabe, G.; Kato, H.; Yamaguchi, S. Subzero 12-hour nonfreezing cryopreservation of porcine heart in a variable magnetic field. Transplant. Direct. 2015, 1, e33. [Google Scholar] [CrossRef] [PubMed]
- Ehrlich, L.E.; Fahy, G.M.; Wowk, B.G.; Malen, J.A.; Rabin, Y. Thermal analyses of a human kidney and a rabbit kidney during cryopreservation by vitrification. J. Biomech. Eng. 2018, 140, 0110051–0110058. [Google Scholar] [CrossRef] [PubMed]
- Tessier, S.N.; Weng, L.; Moyo, W.D.; Au, S.H.; Wong, K.H.K.; Angpraseuth, C.; Stoddard, A.E.; Lu, C.; Nieman, L.T.; Sandlin, R.D.; et al. Effect of ice nucleation and cryoprotectants during high subzero-preservation in endothelialized microchannels. ACS Biomater. Sci. Eng. 2018, 4, 3006–3015. [Google Scholar] [CrossRef] [PubMed]
- de Vries, R.J.; Tessier, S.N.; Banik, P.D.; Nagpal, S.; Cronin, S.E.J.; Ozer, S.; Hafiz, E.O.A.; van Gulik, T.M.; Yarmush, M.L.; Markmann, J.F.; et al. Supercooling extends preservation time of human livers. Nat. Biotechnol. 2019, 37, 1131–1136. [Google Scholar] [CrossRef]
- Que, W.; Hu, X.; Fujino, M.; Terayama, H.; Sakabe, K.; Fukunishi, N.; Zhu, P.; Yi, S.Q.; Yamada, Y.; Zhong, L.; et al. Prolonged cold ischemia time in mouse heart transplantation using supercooling preservation. Transplantation 2020, 104, 1879–1889. [Google Scholar] [CrossRef]
- Abt, P.L.; Desai, N.M.; Crawford, M.D.; Forman, L.M.; Markmann, J.W.; Olthoff, K.M.; Markmann, J.F. Survival following liver transplantation from non-heart-beating donors. Ann. Surg. 2004, 239, 87–92. [Google Scholar] [CrossRef]
- Sade, R.M. Brain death, cardiac death, and the dead donor rule. J. South Carol. Med. Assoc. 2011, 107, 146–149. [Google Scholar]
- Manara, A.R.; Murphy, P.G.; O’Callaghan, G. Donation after circulatory death. Br. J. Anaesth. 2012, 108, 108–121. [Google Scholar] [CrossRef] [PubMed]
- Hessheimer, A.J.; Riquelme, F.; Fundora-Suárez, Y.; García Pérez, R.; Fondevila, C. Normothermic perfusion and outcomes after liver transplantation. Transplant. Rev. 2019, 33, 200–208. [Google Scholar] [CrossRef]
- Osaki, S.; Anderson, J.E.; Johnson, M.R.; Edwards, N.M.; Kohmoto, T. The potential of cardiac allografts from donors after cardiac death at the University of Wisconsin Organ Procurement Organization. Eur. J. Cardiothorac. Surg. 2010, 37, 74–79. [Google Scholar] [CrossRef] [PubMed]
- Taylor, M.J.; Baicu, S.C. Current state of hypothermic machine perfusion preservation of organs: The clinical perspective. Cryobiology 2010, 60, S20–S35. [Google Scholar] [CrossRef]
- Beuth, J.; Falter, F.; Ribeiro, R.V.P.; Badiwala, M.; Meineri, M. New strategies to expand and optimize heart donor pool: Ex vivo heart perfusion and donation after circulatory death: A review of current research and future trends. Anesth. Analg. 2019, 128, 406–413. [Google Scholar] [CrossRef] [PubMed]
- Goldsmith, J.R.; Jobin, C. Think small: Zebrafish as a model system of human pathology. J. Biomed. Biotechnol. 2012, 2012, 817341. [Google Scholar] [CrossRef] [PubMed]
- Rafferty, S.A.; Quinn, T.A. A beginner’s guide to understanding and implementing the genetic modification of zebrafish. Prog. Biophys. Mol. Biol. 2018, 138, 3–19. [Google Scholar] [CrossRef]
- Udvadia, A.J.; Linney, E. Windows into development: Historic, current, and future perspectives on transgenic zebrafish. Dev. Biol. 2003, 256, 1–17. [Google Scholar] [CrossRef]
- Kupperman, E.; An, S.; Osborne, N.; Stainier, D.Y.R. A sphingosine-1-phosphate receptor regulates cell migration during vertebrate heart development. Nature 2000, 406, 192–195. [Google Scholar] [CrossRef]
- Poss, K.D.; Wilson, L.G.; Keating, M.T. Heart regeneration in zebrafish. Science 2002, 298, 2188–2190. [Google Scholar] [CrossRef]
- Xu, X.; Meiler, S.E.; Zhong, T.P.; Mohideen, M.; Crossley, D.A.; Burggren, W.W.; Fishman, M.C. Cardiomyopathy in zebrafish due to mutation in an alternatively spliced exon of titin. Nat. Genet. 2002, 30, 205–209. [Google Scholar] [CrossRef]
- Huang, C.C.; Monte, A.; Cook, J.M.; Kabir, M.S.; Peterson, K.P. Zebrafish heart failure models for the evaluation of chemical probes and drugs. Assay Drug. Dev. Technol. 2013, 11, 561–572. [Google Scholar] [CrossRef]
- Baker, K.; Warren, K.S.; Yellen, G.; Fishman, M.C. Defective “pacemaker” current (Ih) in a zebrafish mutant with a slow heart rate. Proc. Natl. Acad. Sci. USA 1997, 94, 4554–4559. [Google Scholar] [CrossRef]
- Kopp, R.; Schwerte, T.; Pelster, B. Cardiac performance in the zebrafish breakdance mutant. J. Exp. Biol. 2005, 208, 2123–2134. [Google Scholar] [CrossRef]
- Barnard, C.N. The operation. A human cardiac transplant: An interim report of a successful operation performed at Groote Schuur Hospital, Cape Town. S. Afr. Med. J. 1967, 41, 1271–1274. [Google Scholar]
- Brink, J.G.; Hassoulas, J. The first human heart transplant and further advances in cardiac transplantation at Groote Schuur Hospital and the University of Cape Town-with reference to: The operation. A human cardiac transplant: An interim report of a successful operation performed at Groote Schuur Hospital, Cape Town. Cardiovasc. J. Afr. 2009, 20, 31–35. [Google Scholar]
- Sliwa, K.; Zilla, P. 50th Anniversary of the first human heart transplant—How is it seen today? Eur. Heart J. 2017, 38, 3402–3404. [Google Scholar] [CrossRef]
- Thomas, F.T.; Szentpetery, S.S.; Mammana, R.E.; Wolfgang, T.C.; Lower, R.R. Long-distance transportation of human hearts for transplantation. Ann. Thorac. Surg. 1978, 26, 344–350. [Google Scholar] [CrossRef]
- Rajab, T.K.; Singh, S.K. Donation after cardiac death heart transplantation in America is clinically necessary and ethically justified. Circ. Heart Fail. 2018, 11, e004884. [Google Scholar] [CrossRef] [PubMed]
- Salehi, S.; Tran, K.; Grayson, W.L. Advances in perfusion systems for solid organ preservation. Yale J. Biol. Med. 2018, 91, 301–312. [Google Scholar] [PubMed]
- Institute of Medicine (US) Committee on Organ Procurement and Transplantation Policy. Organ Procurement and Transplantation: Assessing Current Policies and the Potential Impact of the DHHS Final Rule; National Academies Press: Washington, DC, USA, 1999. [Google Scholar]
- Goldsmith, K.A.; Demiris, N.; Gooi, J.H.; Sharples, L.D.; Jenkins, D.P.; Dhital, K.K.; Tsui, S.S. Life-years gained by reducing donor heart ischemic times. Transplantation 2009, 87, 243–248. [Google Scholar] [CrossRef] [PubMed]
- Rylski, B.; Berchtold-Herz, M.; Olschewski, M.; Zeh, W.; Schlensak, C.; Siepe, M.; Beyersdorf, F. Reducing the ischemic time of donor hearts will decrease morbidity and costs of cardiac transplantations. Interact. Cardiov. Thorac. Surg. 2010, 10, 945–947. [Google Scholar] [CrossRef] [PubMed]
- Valero-Masa, M.J.; González-Vílchez, F.; Almenar-Bonet, L.; Crespo-Leiro, M.G.; Manito-Lorite, N.; Sobrino-Márquez, J.M.; Gómez-Bueno, M.; Delgado-Jiménez, J.F.; Pérez-Villa, F.; Brossa Loidi, V.; et al. Cold ischemia > 4 hours increases heart transplantation mortality. An analysis of the Spanish heart transplantation registry. Int. J. Cardiol. 2020, 319, 14–19. [Google Scholar] [CrossRef]
- Monteagudo Vela, M.; García Sáez, D.; Simon, A.R. Current approaches in retrieval and heart preservation. Ann. Cardiothorac. Surg. 2018, 7, 67–74. [Google Scholar] [CrossRef]
- Latchana, N.; Peck, J.R.; Whitson, B.; Black, S.M. Preservation solutions for cardiac and pulmonary donor grafts: A review of the current literature. J. Thorac. Dis. 2014, 6, 1143–1149. [Google Scholar]
- Ferng, A.S.; Schipper, D.; Connell, A.M.; Marsh, K.M.; Knapp, S.; Khalpey, Z. Novel vs clinical organ preservation solutions: Improved cardiac mitochondrial protection. J. Cardiothorac. Surg. 2017, 12, 7. [Google Scholar] [CrossRef]
- Wagner, F.M. Donor heart preservation and perfusion. Appl. Cardiopulm. Pathophysiol. 2011, 15, 198–206. [Google Scholar]
- Wicomb, W.N.; Cooper, D.K.; Novitzky, D.; Barnard, C.N. Cardiac transplantation following storage of the donor heart by a portable hypothermic perfusion system. Ann. Thorac. Surg. 1984, 37, 243–248. [Google Scholar] [CrossRef]
- Bruinsma, B.G.; Uygun, K. Subzero organ preservation: The dawn of a new ice age? Curr. Opin. Organ Transplant. 2017, 22, 281–286. [Google Scholar] [CrossRef]
- Luu, B.E.; Storey, K.B. Solving donor organ shortage with insights from freeze tolerance in nature: Activating endogenous antioxidant systems with non-coding RNA to precondition donor organs. Bioessays 2018, 40, e1800092. [Google Scholar] [CrossRef]
- Taylor, M.J.; Weegman, B.P.; Baicu, S.C.; Giwa, S.E. New approaches to cryopreservation of cells, tissues, and organs. Transfus. Med. Hemother. 2019, 46, 197–215. [Google Scholar] [CrossRef] [PubMed]
- Fahy, G.M.; Wowk, B.; Wu, J.; Phan, J.; Rasch, C.; Chang, A.; Zendejas, E. Cryopreservation of organs by vitrification: Perspectives and recent advances. Cryobiology 2004, 48, 157–178. [Google Scholar] [CrossRef] [PubMed]
- Finger, E.B.; Bischof, J.C. Cryopreservation by vitrification: A promising approach for transplant organ banking. Curr. Opin. Organ Transplant. 2018, 23, 353–360. [Google Scholar] [CrossRef] [PubMed]
- Dhital, K.K.; Iyer, A.; Connellan, M.; Chew, H.C.; Gao, L.; Doyle, A.; Hicks, M.; Kumarasinghe, G.; Soto, C.; Dinale, A.; et al. Adult heart transplantation with distant procurement and ex-vivo preservation of donor hearts after circulatory death: A case series. Lancet 2015, 385, 2585–2591. [Google Scholar] [CrossRef]
- Senior, M. Beating the organ clock. Nat. Biotechnol. 2018, 36, 488–492. [Google Scholar] [CrossRef] [PubMed]
- Kirk, R.; Dipchand, A.I. Continuous donor perfusion for heart preservation. Progr. Pediatr. Cardiol. 2017, 46, 15–18. [Google Scholar] [CrossRef]
- De Vries, R.J.; Yarmush, M.; Uygun, K. Systems engineering the organ preservation process for transplantation. Curr. Opin. Biotechnol. 2019, 58, 192–201. [Google Scholar] [CrossRef] [PubMed]
- Eshmuminov, D.; Becker, D.; Bautista Borrego, L.; Hefti, M.; Schuler, M.J.; Hagedorn, C.; Muller, X.; Mueller, M.; Onder, C.; Graf, R.; et al. An integrated perfusion machine preserves injured human livers for 1 week. Nat. Biotechnol. 2020, 38, 189–198. [Google Scholar] [CrossRef]
- Cypel, M.; Yeung, J.C.; Liu, M.; Anraku, M.; Chen, F.; Karolak, W.; Sato, M.; Laratta, J.; Azad, S.; Madonik, M.; et al. Normothermic ex vivo lung perfusion in clinical lung transplantation. N. Engl. J. Med. 2011, 364, 1431–1440. [Google Scholar] [CrossRef]
- Cornet, C.; Calzolari, S.; Miñana-Prieto, R.; Dyballa, S.; van Doornmalen, E.; Rutjes, H.; Savy, T.; D’Amico, D.; Terriente, J. ZeGlobalTox: An innovative approach to address organ drug toxicity using zebrafish. Int. J. Mol. Sci. 2017, 18, 864. [Google Scholar] [CrossRef] [PubMed]
- Poon, K.L.; Brand, T. The zebrafish model system in cardiovascular research: A tiny fish with mighty prospects. Glob. Cardiol. Sci. Pract. 2013, 2013, 9–28. [Google Scholar] [CrossRef]
- Cavalcante, L.D.S.; Toner, M.; Uygun, K.; Tessier, S.N. Leveraging the zebrafish to model organ transplantation. Curr. Opin. Organ Transplant. 2019, 24, 613–619. [Google Scholar] [CrossRef]
- Pieperhoff, S.; Wilson, K.S.; Baily, J.; de Mora, K.; Maqsood, S.; Vass, S.; Taylor, J.; Del-Pozo, J.; MacRae, C.A.; Mullins, J.J.; et al. Heart on a plate: Histological and functional assessment of isolated adult zebrafish hearts maintained in culture. PLoS ONE 2014, 9, e96771. [Google Scholar] [CrossRef]
- González-Rosa, J.M.; Peralta, M.; Mercader, N. Pan-epicardial lineage tracing reveals that epicardium derived cells give rise to myofibroblasts and perivascular cells during zebrafish heart regeneration. Dev. Biol. 2012, 370, 173–186. [Google Scholar] [CrossRef]
- Patra, C.; Kontarakis, Z.; Kaur, H.; Rayrikar, A.; Mukherjee, D.; Stainier, D.Y.R. The zebrafish ventricle: A hub of cardiac endothelial cells for in vitro cell behavior studies. Sci. Rep. 2017, 7, 2687. [Google Scholar] [CrossRef]
- Kugler, E.; Plant, K.; Chico, T.; Armitage, P. Enhancement and Segmentation Workflow for the Developing Zebrafish Vasculature. J. Imaging 2019, 5, 14. [Google Scholar] [CrossRef]
- Itou, J.; Oishi, I.; Kawakami, H.; Glass, T.J.; Richter, J.; Johnson, A.; Lund, T.C.; Kawakami, Y. Migration of cardiomyocytes is essential for heart regeneration in zebrafish. Development 2012, 139, 4133–4142. [Google Scholar] [CrossRef] [PubMed]
- Kao, R.M.; Rurik, J.G.; Farr, G.H., 3rd; Dong, X.R.; Majesky, M.W.; Maves, L. Pbx4 is required for the temporal onset of zebrafish myocardial differentiation. J. Dev. Biol. 2015, 3, 93–111. [Google Scholar] [CrossRef] [PubMed]
- Sallin, P.; Jaźwińska, A. Acute stress is detrimental to heart regeneration in zebrafish. Open Biol. 2016, 6, 160012. [Google Scholar] [CrossRef]
- Yue, M.S.; Plavicki, J.S.; Li, X.Y.; Peterson, R.E.; Heideman, W. A co-culture assay of embryonic zebrafish hearts to assess migration of epicardial cells in vitro. BMC Dev. Biol. 2015, 15, 50. [Google Scholar] [CrossRef]
- Sánchez-Iranzo, H.; Galardi-Castilla, M.; Sanz-Morejón, A.; González-Rosa, J.M.; Costa, R.; Ernst, A.; Sainz de Aja, J.; Langa, X.; Mercader, N. Transient fibrosis resolves via fibroblast inactivation in the regenerating zebrafish heart. Proc. Natl. Acad. Sci. USA 2018, 115, 4188–4193. [Google Scholar] [CrossRef] [PubMed]
- Chi, N.C.; Shaw, R.M.; Jungblut, B.; Huisken, J.; Ferrer, T.; Arnaout, R.; Scott, I.; Beis, D.; Xiao, T.; Baier, H.; et al. Genetic and physiologic dissection of the vertebrate cardiac conduction system. PLoS Biol. 2008, 6, e109. [Google Scholar] [CrossRef] [PubMed]
- Reischauer, S.; Arnaout, R.; Ramadass, R.; Stainier, D.Y. Actin binding GFP allows 4D In Vivo imaging of myofilament dynamics in the zebrafish heart and the identification of Erbb2 signaling as a remodeling factor of myofibril architecture. Circ. Res. 2014, 115, 845–856. [Google Scholar] [CrossRef]
- Whitesell, T.R.; Kennedy, R.M.; Carter, A.D.; Rollins, E.L.; Georgijevic, S.; Santoro, M.M.; Childs, S.J. An α-smooth muscle actin (acta2/αsma) zebrafish transgenic line marking vascular mural cells and visceral smooth muscle cells. PLoS ONE 2014, 9, e90590. [Google Scholar]
- Lenard, A.; Xie, H.M.; Pastuer, T.; Shank, T.; Libbrecht, C.; Kingsley, M.; Riedel, S.S.; Yuan, Z.F.; Zhu, N.; Neff, T.; et al. Using zebrafish to model erythroid lineage toxicity and regeneration. Haematologica 2016, 101, e164–e167. [Google Scholar] [CrossRef]
- Parente, V.; Balasso, S.; Pompilio, G.; Verduci, L.; Colombo, G.I.; Milano, G.; Guerrini, U.; Squadroni, L.; Cotelli, F.; Pozzoli, O.; et al. Hypoxia/reoxygenation cardiac injury and regeneration in zebrafish adult heart. PLoS ONE 2013, 8, e53748. [Google Scholar] [CrossRef]
- Wittamer, V.; Bertrand, J.Y.; Gutschow, P.W.; Traver, D. Characterization of the mononuclear phagocyte system in zebrafish. Blood 2011, 117, 7126–7135. [Google Scholar] [CrossRef]
- Hui, S.P.; Sheng, D.Z.; Sugimoto, K.; Gonzalez-Rajal, A.; Nakagawa, S.; Hesselson, D.; Kikuchi, K. Zebrafish regulatory T cells mediate organ-specific regenerative programs. Dev. Cell 2017, 43, 659–672.e5. [Google Scholar] [CrossRef] [PubMed]
- Ando, K.; Fukuhara, S.; Izumi, N.; Nakajima, H.; Fukui, H.; Kelsh, R.N.; Mochizuki, N. Clarification of mural cell coverage of vascular endothelial cells by live imaging of zebrafish. Development 2016, 143, 1328–1339. [Google Scholar] [CrossRef] [PubMed]
- Santhakumar, K.; Judson, E.C.; Elks, P.M.; McKee, S.; Elworthy, S.; van Rooijen, E.; Walmsley, S.S.; Renshaw, S.A.; Cross, S.S.; van Eeden, F.J. A zebrafish model to study and therapeutically manipulate hypoxia signaling in tumorigenesis. Cancer Res. 2012, 72, 4017–4027. [Google Scholar] [CrossRef] [PubMed]
- Boyle, E.M., Jr.; Pohlman, T.H.; Cornejo, C.J.; Verrier, E.D. Endothelial cell injury in cardiovascular surgery: Ischemia-reperfusion. Ann. Thorac. Surg. 1996, 62, 1868–1875. [Google Scholar] [CrossRef]
- Gao, W.; Bentley, R.C.; Madden, J.F.; Clavien, P.A. Apoptosis of sinusoidal endothelial cells is a critical mechanism of preservation injury in rat liver transplantation. Hepatology 1998, 27, 1652–1660. [Google Scholar] [CrossRef] [PubMed]
- Hortells, L.; Johansen, A.K.Z.; Yutzey, K.E. Cardiac fibroblasts and the extracellular matrix in regenerative and nonregenerative hearts. J. Cardiovasc. Dev. Dis. 2019, 6, 29. [Google Scholar] [CrossRef]
- Kanisicak, O.; Khalil, H.; Ivey, M.J.; Karch, J.; Maliken, B.D.; Correll, R.N.; Brody, M.J.; Lin, S.C.J.; Aronow, B.J.; Tallquist, M.D.; et al. Genetic lineage tracing defines myofibroblast origin and function in the injured heart. Nat. Commun. 2016, 7, 12260. [Google Scholar] [CrossRef] [PubMed]
- González-Rosa, J.M.; Mercader, N. Cryoinjury as a myocardial infarction model for the study of cardiac regeneration in the zebrafish. Nat. Protoc. 2012, 7, 782–788. [Google Scholar] [CrossRef]
- Baeten, J.T.; de Jong, J.L.O. Genetic Models of Leukemia in Zebrafish. Front. Cell Dev. Biol. 2018, 6, 115. [Google Scholar] [CrossRef]
- Ong, S.G.; Lee, W.H.; Theodorou, L.; Kodo, K.; Lim, S.Y.; Shukla, D.H.; Briston, T.; Kiriakidis, S.; Ashcroft, M.; Davidson, S.M.; et al. Hif-1 reduces ischaemia-reperfusion injury in the heart by targeting the mitochondrial permeability transition pore. Cardiovasc. Res. 2014, 104, 24–36. [Google Scholar] [CrossRef]
- Mandic, M.; Tzaneva, V.; Careau, V.; Perry, S.F. Hif-1α paralogs play a role in the hypoxic ventilatory response of larval and adult zebrafish (Danio rerio). J. Exp. Biol. 2019, 222, jeb195198. [Google Scholar] [CrossRef] [PubMed]
- Jopling, C.; Suñé, G.; Faucherre, A.; Fabregat, C.; Izpisua Belmonte, J.C. Hypoxia induces myocardial regeneration in zebrafish. Circulation 2012, 126, 3017–3027. [Google Scholar] [CrossRef]
- Pfeiffer, J.; Tarbashevich, K.; Bandemer, J.; Palm, T.; Raz, E. Rapid progression through the cell cycle ensures efficient migration of primordial germ cells-the role of Hsp90. Dev. Biol. 2018, 436, 84–93. [Google Scholar] [CrossRef] [PubMed]
- Bill, B.R.; Petzold, A.M.; Clark, K.J.; Schimmenti, L.A.; Ekker, S.C. A primer for morpholino use in zebrafish. Zebrafish 2009, 6, 69–77. [Google Scholar] [CrossRef] [PubMed]
- Blechinger, S.R.; Warren, J.T., Jr.; Kuwada, J.Y.; Krone, P.H. Developmental toxicology of cadmium in living embryos of a stable transgenic zebrafish line. Environ. Health Perspect. 2002, 110, 1041–1046. [Google Scholar] [CrossRef]
- Babayigit, A.; Duy Thanh, D.; Ethirajan, A.; Manca, J.; Muller, M.; Boyen, H.-G.; Conings, B. Assessing the toxicity of Pb- and Sn-based perovskite solar cells in model organism Danio rerio. Sci. Rep. 2016, 6, 18721. [Google Scholar] [CrossRef]
- Liu, C.C.; Li, L.; Lam, Y.W.; Siu, C.W.; Cheng, S.H. Improvement of surface ECG recording in adult zebrafish reveals that the value of this model exceeds our expectation. Sci. Rep. 2016, 6, 25073. [Google Scholar] [CrossRef]
- Lenning, M.; Fortunato, J.; Le, T.; Clark, I.; Sherpa, A.; Yi, S.; Hofsteen, P.; Thamilarasu, G.; Yang, J.; Xu, X.; et al. Real-time monitoring and analysis of zebrafish electrocardiogram with anomaly detection. Sensors 2017, 18, 61. [Google Scholar] [CrossRef] [PubMed]
- Zhao, Y.; Yun, M.; Nguyen, S.A.; Tran, M.; Nguyen, T.P. In Vivo surface electrocardiography for adult zebrafish. J. Vis. Exp. 2019, 150. [Google Scholar] [CrossRef]
- Garcia-Puig, A.; Mosquera, J.L.; Jiménez-Delgado, S.; García-Pastor, C.; Jorba, I.; Navajas, D.; Canals, F.; Raya, A. Proteomics analysis of extracellular matrix remodeling during zebrafish heart regeneration. Mol. Cell Proteom. 2019, 18, 1745–1755. [Google Scholar] [CrossRef] [PubMed]
- Machackova, J.; Barta, J.; Dhalla, N.S. Myofibrillar remodeling in cardiac hypertrophy, heart failure and cardiomyopathies. Can. J. Cardiol. 2006, 22, 953–968. [Google Scholar] [CrossRef]
- Pauty, J.; Usuba, R.; Takahashi, H.; Suehiro, J.; Fujisawa, K.; Yano, K.; Nishizawa, T.; Matsunaga, Y.T. A vascular permeability assay using an in vitro human microvessel model mimicking the inflammtory condition. Nanotheranostics 2017, 1, 103–113. [Google Scholar] [CrossRef] [PubMed]
| Solutions | University of Wisconsin Solution | Celsior® | Custodiol® HTK Solution |
|---|---|---|---|
| Type | Intracellular | Extracellular | Extracellular |
| Buffer | Phosphate | Histidine | Histidine |
| Antioxidant | Glutathione, Allopurinol | Glutathione, Mannitol | Mannitol |
| Colloid | Hydroxyethyl starch, Lactobionic acid, Raffinose | Mannitol, Lactobionic acid | Mannitol |
| Intended use | Flushing/sold storage | Flushing/cold storage | Flushing/cold storage |
| Other | High viscosity and tendency of particle formation (might require filtration) | Combines elements of UW and HTK | Lower viscosity and potassium compared to UW |
| References | [42,44] | [42,43,44] | [42,44] |
| Regulatory Region | Cardiovascular Structure | Construct | References |
|---|---|---|---|
| kdrl | Cardiac endothelial cells/endocardium | Tg(kdrl:EGFP) (cytosolic) Tg(kdrl:Hsa.HRAS-mCherry) (plasma membrane) | [62,63] |
| myl7 | Cardiomyocytes | Tg(myl7:GFP) | [62] |
| myl7 | Cardiomyocytes | Tg3(myl7:Kaede) | [64] |
| myl7 | Nucleus of cardiomyocytes | Tg(myl7:h2afva-mCherry)sd12 | [65] |
| myl7 | Endocardial cells | Tg(Tie2:EGFP) | [66] |
| tcf21 | Epicardial cells | Tg(tcf21:DsRed2) | [67] |
| wt1a | Cardiac fibroblasts | Tg(−6.8kbwt1a:GFP) | [68] |
| col1a2 | Collagen producing cells | Tg(col1a2:LOXP-mCherry-NTR) | [68] |
| postn | Perivascular and epicardial cells (uninjured hearts), activated fibroblasts (injured hearts) | Tg(postnb:citrine) | [68] |
| myh6 | atrial cells | Tg(myh6:EGFP)s958 | [65] |
| myl7 (previous name cmlc2) | Cardiac specific calcium indicator | Tg(myl7:GCaMP) | [69] |
| myl7 (previous name cmlc2) | Cardiac filamentous actin (F-actin) | Tg(myl7:LifeAct-GFP) | [70] |
| fli1a | Endothelial filamentous actin (F-actin) | Tg(fli1a:LifeAct-mClover) | [62] |
| acta2 | Mural cell α-smooth muscle actin | Tg(acta2:EGFP) Tg(acta2:mCherry) | [71] |
| gata1a | Blood cells (erythroid) | Tg(gata1:DsRed) (progenitor cells) Tg(gata1:DsRed;globin:GFP) (mature erythrocytes) | [72] |
| mpo/lyz (previous name lysC) | Blood cells (leukocytes I) Double transgene (neutrophils and macrophages) | Tg(MPO:EGFP) × Tg(LysC:DsRed) | [73] |
| mhc2dab | Blood cells (leukocytes II) B lymphocytes, eosinophils, Macrophages and dendritic cells | Tg(mhc2dab:GFP) | [74] |
| cd45 | Blood cells (leukocytes III) Myeloid cells and T Lymphocytes | Tg(cd45:DsRed) | [74] |
| foxp3a | Regulatory T cells | TgBAC(foxp3a:TagRFP)vcc3 | [75] |
| fli1a | Vasculature/endothelial cells | Tg(fli1a:Myr-mCherry) | [76] |
| pdgfrb | Vasculature/mural cells | TgBAC(pdgfrb:EGFP), TgBAC(pdgfrb:mCherry) | [76] |
| egln3 (previous name phd3 ) | Hypoxia | TgBAC(egln3:EGFP) | [77] |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Da Silveira Cavalcante, L.; Tessier, S.N. Zebrafish as a New Tool in Heart Preservation Research. J. Cardiovasc. Dev. Dis. 2021, 8, 39. https://doi.org/10.3390/jcdd8040039
Da Silveira Cavalcante L, Tessier SN. Zebrafish as a New Tool in Heart Preservation Research. Journal of Cardiovascular Development and Disease. 2021; 8(4):39. https://doi.org/10.3390/jcdd8040039
Chicago/Turabian StyleDa Silveira Cavalcante, Luciana, and Shannon N. Tessier. 2021. "Zebrafish as a New Tool in Heart Preservation Research" Journal of Cardiovascular Development and Disease 8, no. 4: 39. https://doi.org/10.3390/jcdd8040039
APA StyleDa Silveira Cavalcante, L., & Tessier, S. N. (2021). Zebrafish as a New Tool in Heart Preservation Research. Journal of Cardiovascular Development and Disease, 8(4), 39. https://doi.org/10.3390/jcdd8040039






