1. Introduction
Coronary Artery disease (CAD) is the most common type of heart disease which leads to morbidity and mortality among men and women, irrespective of race and ethnicity [
1,
2]. CAD is an intricate disease that is influenced by many environmental and familial risk factors [
3,
4]. In fact, many of the novel and traditional risk factors, such as lipoprotein-cholesterol, elevated low density lipoprotein, cholesterol, high blood pressure, smoking, obesity and diabetes are inadequate to fully understand who is at risk for CAD [
5]. A complex series of events involving multiple biological pathways and genes are responsible for the progression of CAD [
6,
7]. It has been suggested by epidemiologic and genetic studies that certain genetic variants, which include polymorphisms in several genes, are associated with an inflated prevalence of CAD among high or low risk subjects. Study of the genetic elements would improve risk evaluation and offer better measures for the prevention and treatment of the CAD disease.
LDLR, the low-density lipoprotein receptor, is a cell surface glycoprotein, which has the ability of binding and up taking of plasma LDL particles and plays a significant role in maintaining the cellular cholesterol homeostasis [
8]. Mutations in the
LDLR gene can lead to a significant rise in plasma LDL levels, which in turn may result in an increased risk for atherosclerosis and coronary heart disease [
9]. Various mutations of the
LDLR have been described so far, influencing exons, splicing sites and the promoter regions. Mutations in genes of other proteins involved in LDL receptor adaptor protein (LDLRAP1) and LDL uptake and metabolism (ApoB) and in LDLR intracellular recycling (propoprotein convertase subtilisin/kexin type 9 serine protease, PCSK9) have also been shown to be involved in genetic hypercholesterolemia [
10]. A single nucleotide polymorphism (SNP) in
LDLR exon 12, rs688 is linked with low-density lipoprotein cholesterol (LDL-C) and coronary heart disease (CAD) in a gender-independent way [
11,
12].
LDLR rs688 acts as a modulator of alterative exon splicing, which can lead to a shift in the reading frame and an altered gene transcript [
13,
14]. The frequency of genetic polymorphism of LDLR (rs688) across the globe is summarized in
Table 1. The non-coding SNPs in
LDLR have also been found to have functions; for instance, rs17248720 in the intergenic region [
15] and in the promoter region c.-139C > G [
16], c.-101T > C, c.-121T > C [
17], and c.-49C > T [
18], are involved in regulation of gene expression and have been reported to cause familial hypercholesterolemia (FH). [
19].
Notably, the allele frequencies and association analyses can differ widely amongst African, Americans, European Caucasians, Hispanics, Asians, and other ethnic groups. It is also noteworthy that associations found in one ethnic group may not translate to the same associations in other ethnic groups. The association of the single nucleotide polymorphisms with the severity of coronary arteries has been demonstrated [
20,
21]. Therefore, this study was pursued to examine the frequency of LDLR (rs688C > T) gene polymorphism in coronary heart diseases in an Indian population.
2. Materials and Methods
This study was a hospital-based case control study. Subjects were collected from different hospitals. Informed written consent was obtained for all study subjects. The research study was approved by the Ethical Committee of Punjabi University.
2.1. Inclusion Criteria
The case control study included clinically-confirmed coronary artery disease cases. The diagnosis was confirmed using electrocardiogram (ECG), echocardiogram stress test, cardiac catheterization and angiogram heart scan. Out of 200 CAD patients, 180 were males and 20 were females.
2.2. Exclusion Criteria
The exclusion criteria involved patients with who had previously had coronary bypass surgery or percutaneous transluminal coronary angioplasty (PTCA) because of their treated coronary status.
2.3. Collection of Clinical History
Informed written consent was obtained from all CAD patients, as well as healthy controls. Both CAD patients as well as healthy controls were interviewed using a structured questionnaire regarding epidemiological/demographic data, past history, history of addiction, particularly smoking, family history of MI or CAD. The detailed laboratory and clinical data were collected to determine relevant clinical history.
2.4. Coronary Artery Disease and Its Risk Factors
The traditional risk factors for coronary artery disease are high LDL cholesterol, low HDL cholesterol, high blood pressure, family history of a patient , diabetes, smoking, being post-menopausal for women and being older than 45 for men. The family history was the presence of a first degree relative with coronary artery disease at the age of <55 years for men and <60 years for women.
2.5. Coronary Angiography
Coronary angiography is a procedure that uses contrast dye, usually containing iodine, and X-ray pictures to detect blockages in the coronary arteries that are caused by plaque buildup. All 200 CAR cases underwent coronary angiography. This procedure is used to diagnose coronary heart disease and coronary microvascular disease after chest pain, sudden cardiac arrest, or abnormal results from tests, such as an electrocardiogram of the heart or an exercise stress test. It is important to detect blockages because, over time, they can cause chest pain, especially with physical activity or stress, or a heart attack. Coronary angiography detects the severity of coronary artery disease that can be expressed by the number of affected vessels (one, two, or three vessel disease).
2.6. DNA Extraction
DNA was extracted by using DNeasy Blood Kit (cat 69506) from Qiagen (Hilden, Germany) as per the manufactures instructions. The extracted DNA was dissolved in nuclease-free water and stored at 4 °C until use. The quality and integrity of DNA was checked using NanoDrop™ (Thermo Fisher Scientific 168 Third Avenue, Waltham, MA, USA).
2.7. LDLR (rs688) Genotyping
LDLR-rs688C > T gene polymorphism was detected using an allele-specific PCR. Allele-specific PCR is based on the use of sequence-specific PCR primers that allow amplification of test DNA, only when the target allele is contained within the sample. The LDLR-rs688C/T genotyping primers were designed using primer3 software [
22], as depicted in
Table 1.
AS-PCR was performed in a reaction volume of 25 μL containing template DNA (50 ng), F1 −0.35 μL, F2 −0.35 μL, RI −0.30 μL, and 25 pmol of each primer, and 10 μL of Green Master Mix (Qiagen). A final volume of 25 μL was adjusted by adding nuclease free ddH
2O. The PCR cocktail was prepared as depicted in
Table 2. Finally 2 μL of DNA in each PCR tube from each patient as well as control.
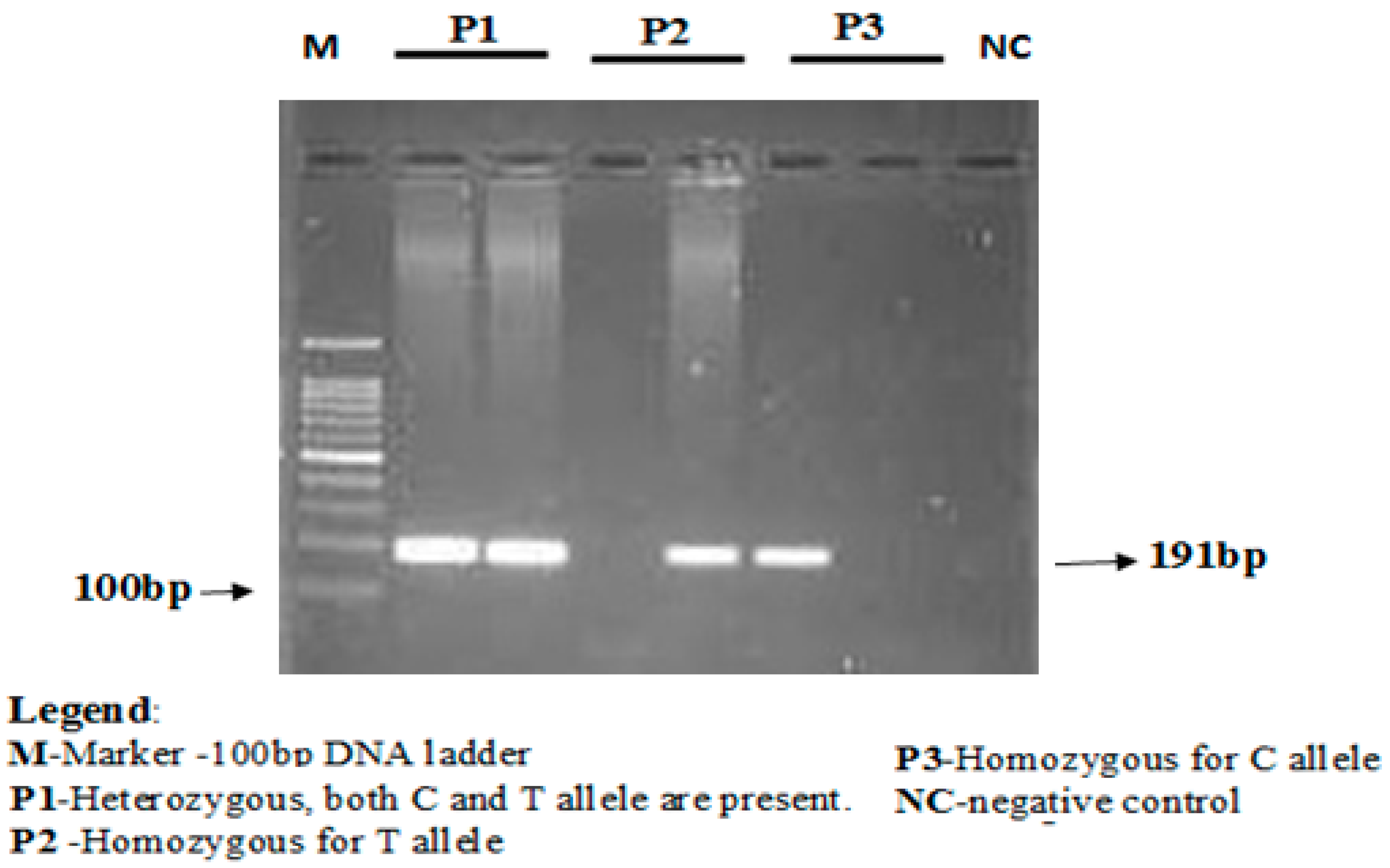
The concentration of the mutant-specific primer was raised, and concentrations of common reverse primer were lowered to favor amplification from the mutant allele. The annealing temperature was lowered from 67 to 68 °C to favor the binding of both forward primers that contained mismatches with the templates. PCR conditions used were as follows, initial denaturation for 3 min at 94 °C followed by 35 cycles of 30 s at 94 °C (denaturation), 30 s at 67 °C (annealing), 1 min at 72 °C (elongation), and finally followed by 10 min at 72 °C (final elongation). PCR products were stored at 4 °C till further analyses. The amplification products were separated using electrophoresis via 2% agarose gel stained with ethidium bromide. Lengths of PCR products formed were 191 bp for both rs688C allele and rs688T allele, as depicted in
Figure 1.
The number of cycles was increased from 30 to 40 cycles, significantly enhancing the yields of all three PCR products. Together, these changes resulted in a more robust amplification of the mutant allele and a less competing reaction from the control, as evidenced by the relative intensities of the corresponding bands on agarose gel electrophoresis.
2.8. Statistical Analysis
Deviations from Hardy-Weinberg disequilibrium (HWD) were calculated using the Chi-square (χ2) goodness-of-fit test. Group differences were compared using Student’s two-sample t-test or one-way analysis of variance (ANOVA) for continuous variables and Chi-squared for categorical variables. Differences of the LDLR-rs688C/T allele and genotype frequencies between groups were evaluated using the Chi-square test. The associations between LDLR-rs688C/T genotypes and risk of CVD were estimated by computing the odds ratios (ORs), risk ratios (RRs) and risk differences (RDs) with 95% confidence intervals (CIs). Allele frequencies among cases, as well as controls, were evaluated using the Chi-square Hardy-Weinberg equilibrium test. A p value < 0.05 was considered significant. All statistical analyses were performed using SPSS 16.0 (IBM, Chicago, IL, USA)
4. Discussion
In India, more than 10.5 million deaths occur annually, and it has been reported that CAD led deaths occurred in 20% in men and 16.9% in women. According to 2010–2013 RGI data (27), proportionate mortality from CAD increased to 23% of total and 32% of adult deaths in the years 2010–2013 [
23]. Coronary artery disease (CAD) is a leading cause of morbidity and mortality worldwide and has become a major public health burden in India [
24].
Traditional risk factors, such as lipid-rich diet, advanced age, smoking, hypertension, diabetes mellitus and dyslipidemia, are associated with an increased risk of CAD. RNA made containing the rs688 (T) SNP, a variant near exon 12 of the low-density lipoprotein receptor, which is a receptor for ApOE proteins, is spliced at a lower efficiency in males. Recent genome-wide association studies have highlighted that common variations at the
LDLR locus are strongly associated with proatherogenic lipid profile and with CAD [
25].
In our study, we reported a significant difference in genotype distribution among the CAD cases and matched healthy controls (
p = 0.0031). The frequencies of all three genotypes, CC, CT, and TT, reported in patients were 14%, 65% and 21% and in healthy controls 18%, 73% and 9% respectively. The
LDLR TT genotype as well as LDLR T allele was associated with increased susceptibility to CAD in Indian patients. The frequencies of LDLR rs688 genotypes CC, CT, and TT were obtained from different studies performed in different countries, as shown in
Table 7. Teslovich et al. [
26], through genome-wide association studies reported single nucleotide polymorphisms at the
LDLR locus that contribute to inter-individual variation in serum lipid concentrations. In the same study it was reported that several SNPs within
LDLR, such as rs12983082, rs2738446, rs1799898, rs9789302, and rs5925, were in linkage disequilibrium with
LDLR rs688C/T
r2 > 0.8, but none were associated with plasma lipids, suggesting that rs688 is the causative underlying polymorphism. Our study observed that a higher percentage of TT (21%) genotype was found in patients than in healthy control TT (9%). Similar results were in obtained in Italy (21%) and the USA (21%), whereas a lesser percentage of the TT genotype (2% and 4%) was reported in two studies from Taiwan, as depicted in
Table 7.
It was reported that the minor variant of LDLR rs688 (Asn
591 ACC→ACT) gene has been reported to be associated with a 4–10% increase in plasma cholesterol levels in several independent populations [
29]. Mutations in the
LDLR gene can lead to elevated plasma LDL levels, resulting in an increased risk for atherosclerosis and coronary heart disease [
30]. Several studies have established that the rs688 SNP is associated with significantly decreased LDLR exon 12 splicing efficiency in women in vivo. The rs688 C/T in exon 12 has been recently shown to alter splicing efficiency, with the T allele being associated with increased total and LDL-cholesterol levels in premenopausal women. These effects on splicing may be physiologically relevant because of the presence of rs688 minor allele being associated with increased total and LDL-cholesterol in female members of the Framingham Offspring Study. It was reported that an LDLR SNP present in approximately 60% of Caucasians is associated with significant (10%) increases in total and LDL-cholesterol in pre-menopausal women [
31]. There is clear epidemiological evidence that increased levels of LDL lead to cardiovascular disease (mostly coronary disease), and it is estimated that elevated cholesterol contributes to 4.4 million deaths per year worldwide [
32]. Samani et al. [
33], reported that the minor allele at the LDLR rs688 SNP has recently been associated with increased risk of coronary artery disease in a combined genome-wide analysis of British and German cohorts. Similar results were documented in our study, in which LDLR rs688 SNP is associated with an increased risk of developing coronary artery disease.