Molecular Surveillance and Whole Genomic Characterization of Bovine Rotavirus A G6P[1] Reveals Interspecies Reassortment with Human and Feline Strains in China
Simple Summary
Abstract
1. Introduction
2. Materials and Methods
2.1. Clinical Sample Collection and Preparation
2.2. Genomic Screening for Rotavirus Group A
2.3. Virus Isolation Workflow
2.4. Viral Plaque Purification and Growth Curve Determination
2.5. Whole-Genome Sequencing
2.6. Viral Segment Genotyping and Phylogenetic Analysis
3. Results
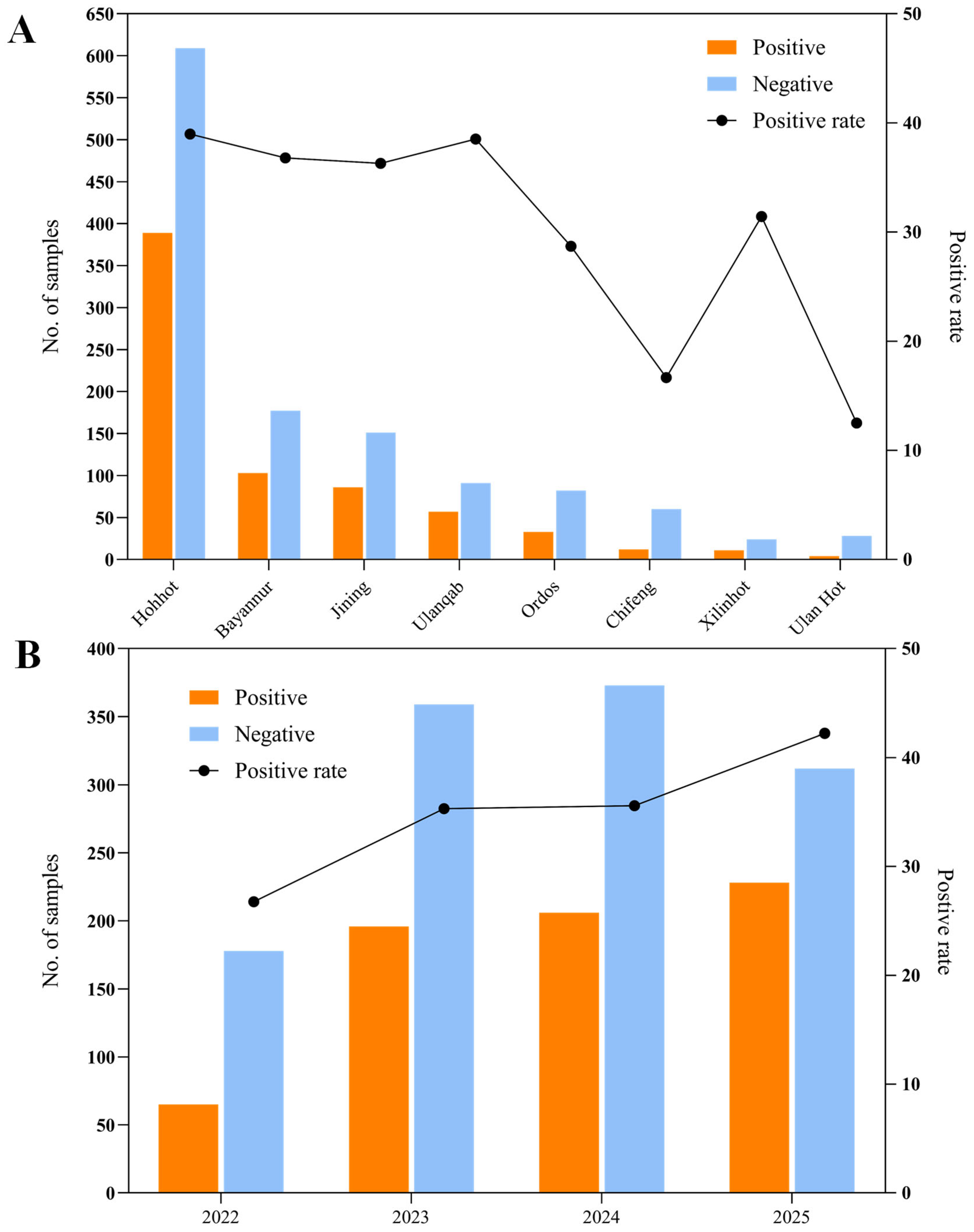
3.1. Positive Rate of Bovine Rotavirus A
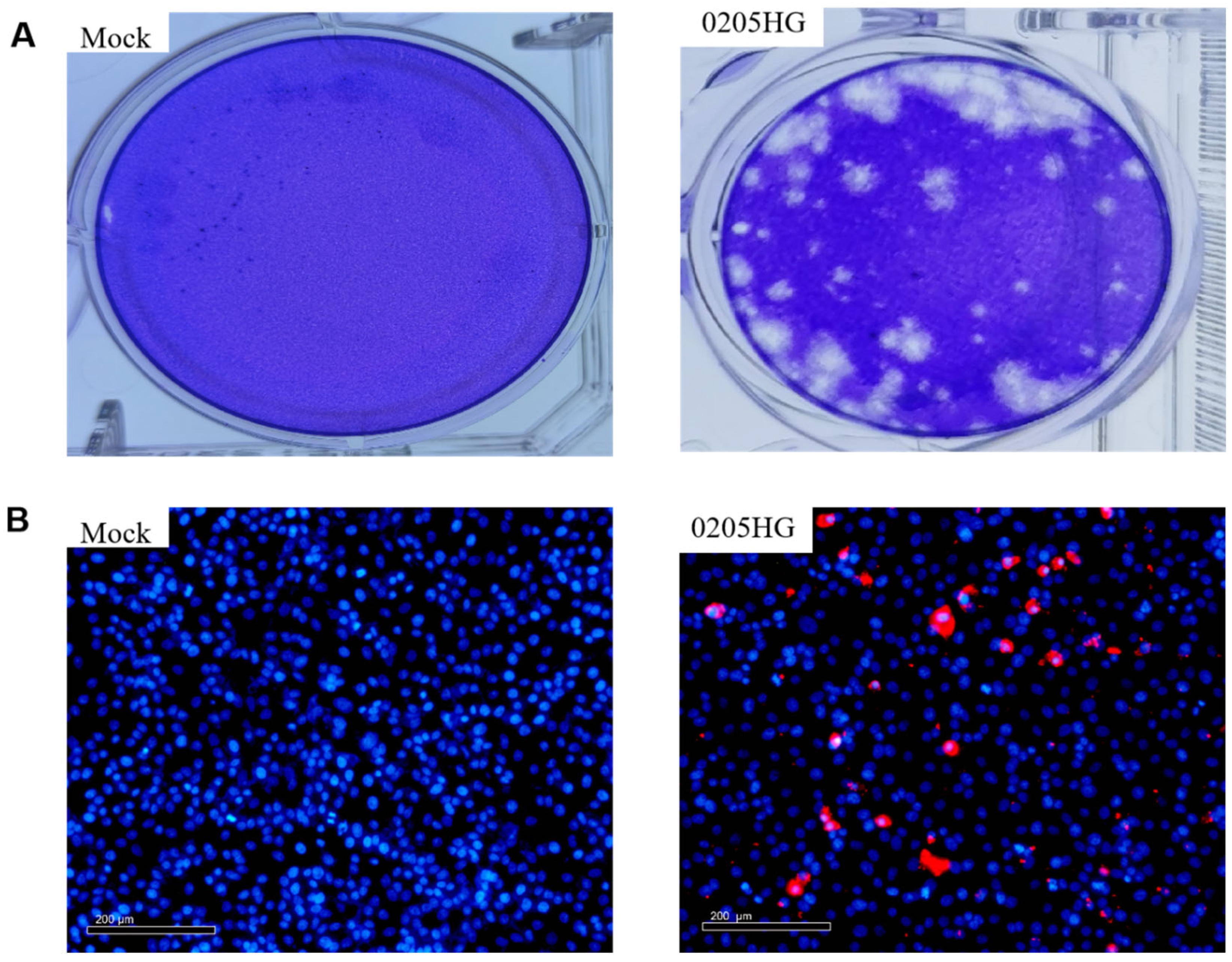
3.2. Viral Isolation and Identification
3.3. Indirect Immunofluorescence Assay
3.4. TCID50 Measurements of the Isolated Strain
3.5. Whole-Genome Sequence of the BRV Strain 0205HG
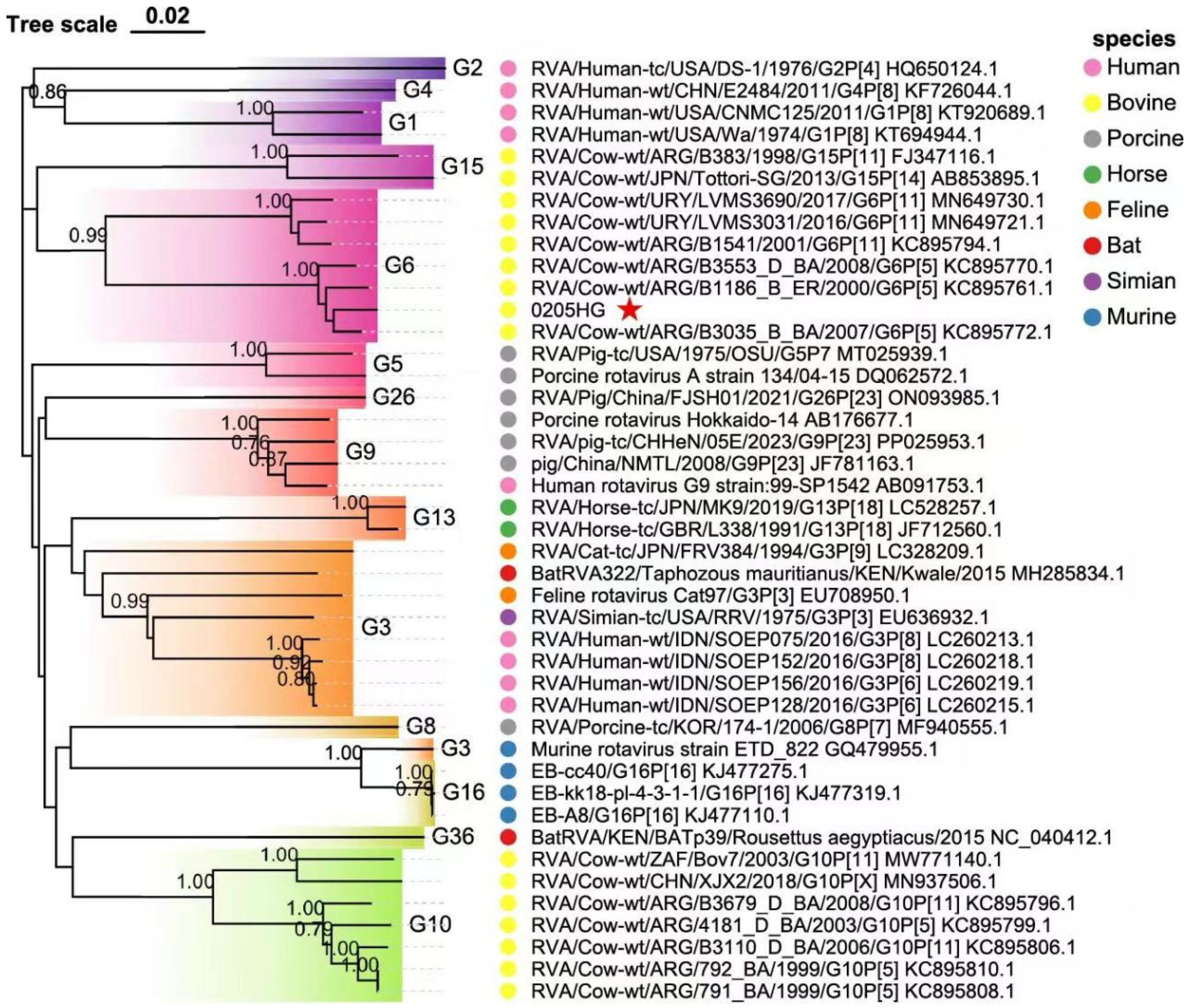
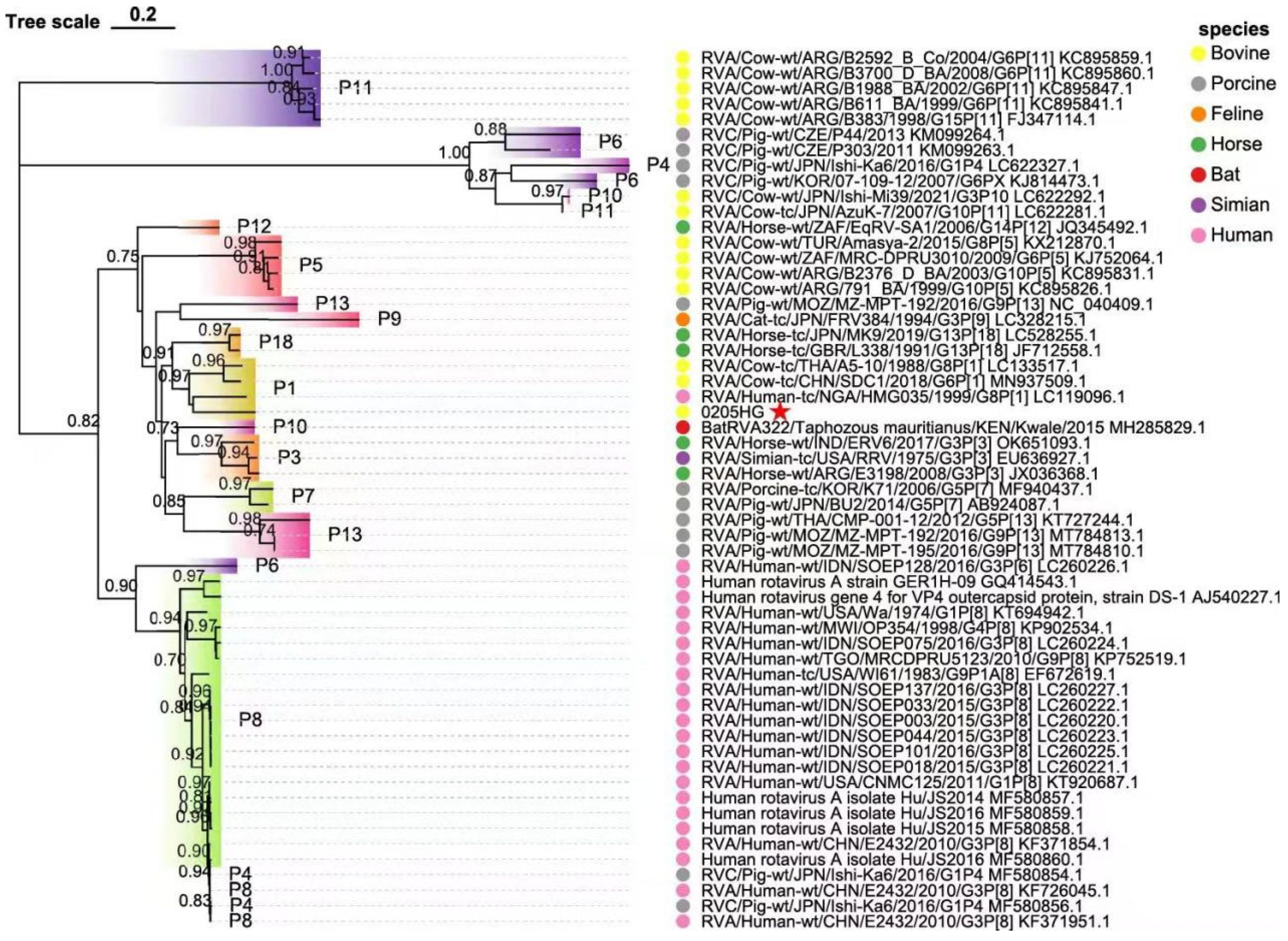
3.6. Molecular Phylogeny of Structural Protein Genes
3.7. Molecular Phylogeny of Nonstructural Protein Genes
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Jessop, E.; Li, L.; Renaud, D.L.; Verbrugghe, A.; Macnicol, J.; Gamsjäger, L.; Gomez, D.E. Neonatal Calf Diarrhea and Gastrointestinal Microbiota: Etiologic Agents and Microbiota Manipulation for Treatment and Prevention of Diarrhea. Vet. Sci. 2024, 11, 108. [Google Scholar] [CrossRef] [PubMed]
- Ghonaim, A.H.; Hopo, M.G.; Ghonaim, N.H.; Jiang, Y.; He, Q.; Li, W. The Epidemiology of Circulating Rotavirus Associated with Diarrhea in Egyptian Kids and Calves: A Review. Zoonoses 2023, 3, 985. [Google Scholar] [CrossRef]
- Hosein, H.I.; Rouby, S.R.; Menshawy, A.M.S.; Abolwafaa, M.; Ghonaim, A.; Hongmei, Z.; Qigai, H.; Wentao, L. Molecular Epidemiology and Associated Risk Factors for Bovine Rotavirus Infection in Cattle and Buffalo Calves in Different Governorates in Egypt. Egypt. J. Vet. Sci. 2025, 1–12. [Google Scholar] [CrossRef]
- Hosein, H.I.; Rouby, S.R.; Abdel, S.; Abdel, A.; Abo-elwafa, M.; Menshawy, A.M.S.; Abdel-ra, A.M.; Madkour, B.S. Epidemiologic Features and Evidence of New Subtypes of Cryptosporidium parvum in Diarrheic Calves in Egypt. Zoonoses 2024, 4, 967. [Google Scholar] [CrossRef]
- Maier, G.U.; Breitenbuecher, J.; Gomez, J.P.; Samah, F.; Fausak, E.; Van Noord, M. Vaccination for the Prevention of Neonatal Calf Diarrhea in Cow-Calf Operations: A Scoping Review. Vet. Anim. Sci. 2022, 15, 100238. [Google Scholar] [CrossRef]
- Ghonaim, A.H.; Rouby, S.R.; Nageeb, W.M.; Elgendy, A.A.; Xu, R.; Jiang, C.; Ghonaim, N.H.; He, Q.; Li, W. Insights into Recent Advancements in Human and Animal Rotavirus Vaccines: Exploring New Frontiers. Virol. Sin. 2024, 40, 1–14. [Google Scholar] [CrossRef]
- Chen, Y.; Wu, R.; Mi, W.; Ghonaim, A.H.; Ren, W.; Yang, L.; Ruan, S.; He, Q.; Chen, H.; Jiang, Y. Molecular Evolution and Genetic Characteristics of G3P[3] Group A Canine Rotavirus Isolated in Wuhan, China. J. Gen. Virol. 2022, 103, 001784. [Google Scholar] [CrossRef]
- Park, G.N.; Choe, S.E.; Cha, R.M.; Shin, J.; Kim, K.S.; An, B.H.; Kim, S.Y.; Hyun, B.H.; An, D.J. Genetic Diversity of Bovine Group A Rotavirus Strains Circulating in Korean Calves during 2014 and 2018. Animals 2022, 12, 3555. [Google Scholar] [CrossRef]
- Dharmarajan, G.; Li, R.; Chanda, E.; Dean, K.R.; Dirzo, R.; Jakobsen, K.S.; Khan, I.; Leirs, H.; Shi, Z.; Wolfe, N.D.; et al. The Animal Origin of Major Human Infectious Diseases: What Can Past Epidemics Teach Us About Preventing the Next Pandemic ? Zoonoses 2022, 2, 989. [Google Scholar] [CrossRef]
- Erkyihun, G.A.; Alemayehu, M.B. One Health Approach for the Control of Zoonotic Diseases. Zoonoses 2022, 2, 963. [Google Scholar] [CrossRef]
- Li, W.; Yi, X.; Wu, B.; Li, X.; Ye, B.; Deng, Z.; Runa, A.; Hu, S.; Li, D.; Wu, H.; et al. Neonatal Calf Diarrhea Is Associated with Decreased Bacterial Diversity and Altered Gut Microbiome Profiles. Fermentation 2023, 9, 827. [Google Scholar] [CrossRef]
- Sun, M.; Pratama, A.C.; Qiu, H.; Liu, Z.; He, F. Toward Innovative Veterinary Nanoparticle Vaccines. Anim. Dis. 2024, 4, 14. [Google Scholar] [CrossRef]
- Bazzi, R.; Alaboudi, A.; Rácz, G. The Role of Veterinarians in the One Health Approach to Antimicrobial Resistance Perspectives in Jordan. Anim. Dis. 2022, 2, 1. [Google Scholar] [CrossRef]
- Li, C.; Wang, X.; Zhu, Q.; Sun, D. Isolation and Identification of BRV G6P[1] Strain in Heilongjiang Province, Northeast China. Front. Vet. Sci. 2024, 11, 1416465. [Google Scholar] [CrossRef]
- Zhao, C.; Hu, X.; Zhang, N.; Fu, Y. Emerging Role of Ruminal Microbiota in the Development of Perinatal Bovine Diseases. Anim. Zoonoses 2025, 1, 86–98. [Google Scholar] [CrossRef]
- Vlasova, A.N.; Amimo, J.O.; Saif, L.J. Porcine Rotaviruses: Epidemiology, Immune Responses and Control Strategies. Viruses 2017, 9, 48. [Google Scholar] [CrossRef] [PubMed]
- Oluwaseun, S.; Yang, C.; Jian, S.; Tu, S.; Yin, J.; Song, Y.; Sun, Q. Health Impact of Rotavirus Vaccination in China ABSTRACT. Hum. Vaccines Immunother. 2024, 20, 2386750. [Google Scholar] [CrossRef] [PubMed]
- Wu, L.; Jing, Z.; Pan, Y.; Guo, L.; Li, Z.; Feng, L.; Tian, J. Emergence of a Novel Pathogenic Porcine G1P[7] Rotavirus in China. Virology 2024, 598, 110185. [Google Scholar] [CrossRef] [PubMed]
- Mohamed, F.F.; Mansour, S.M.G.; El-Araby, I.E.; Mor, S.K.; Goyal, S.M. Molecular Detection of Enteric Viruses from Diarrheic Calves in Egypt. Arch. Virol. 2017, 162, 129–137. [Google Scholar] [CrossRef]
- Cheng, X.; Wu, W.; Teng, F.; Yan, Y.; Li, G.; Wang, L.; Wang, X.; Wang, R.; Zhou, H.; Jiang, Y.; et al. Isolation and Characterization of Bovine RVA from Northeast China, 2017–2020. Life 2021, 11, 1389. [Google Scholar] [CrossRef]
- Aksoy, E.; Azkur, A.K. Genome Characterization of a Turkish Bovine Rotavirus Field Isolate by Shotgun Metagenomics. Arch. Virol. 2023, 168, 159. [Google Scholar] [CrossRef]
- Liu, X.; Tang, C.; Yan, N.; Wang, Y.; Zhang, B. Detection and Molecular Characteristics of Bovine Rotavirus A in Dairy Calves in China Specimen Collection. J. Vet. Sci. 2021, 22, e69. [Google Scholar] [CrossRef]
- Yamada, M.; Kubota, K.; Takahashi, S.; Toyofuku, C.; Hakim, H.; Alam, M.S.; Hasan, M.A.; Shoham, D.; Takehara, K. Longitudinal and Cross-Sectional Detection of Four Bovine Enteric Viruses by Multiplex-Reverse Transcription Polymerase Chain Reaction: Identification of Possible Indicator Viruses to Assess Biosecurity Level at Bovine Farms. J. Vet. Med. Sci. 2020, 82, 314–319. [Google Scholar] [CrossRef]
- Elkady, G.; Zhu, J.; Peng, Q.; Chen, M.; Liu, X.; Chen, Y.; Hu, C.; Chen, H.; Guo, A. Isolation and Whole Protein Characterization of Species A and B Bovine Rotaviruses from Chinese Calves. Infect. Genet. Evol. 2021, 89, 104715. [Google Scholar] [CrossRef] [PubMed]
- Zhang, G.; Hao, J.; Li, L.; Jiang, Y.; Hao, P.; Yu, J.; Ren, L.; Li, C.; Wang, M. Construction and Immunogenicity of Recombinant Lactiplantibacillus Plantarum Vaccine Candidate Expressing SARS-CoV-2 Omicron S1 Protein. Anim. Zoonoses 2025, 1, 13–23. [Google Scholar] [CrossRef]
- Bolger, A.M.; Lohse, M.; Usadel, B. Trimmomatic: A Flexible Trimmer for Illumina Sequence Data. Bioinformatics 2014, 30, 2114–2120. [Google Scholar] [CrossRef] [PubMed]
- Langmead, B.; Salzberg, S.L. Fast Gapped-Read Alignment with Bowtie 2. Nat. Methods 2012, 9, 357–359. [Google Scholar] [CrossRef]
- Bankevich, A.; Nurk, S.; Antipov, D.; Gurevich, A.A.; Dvorkin, M.; Kulikov, A.S.; Lesin, V.M.; Nikolenko, S.I.; Pham, S.; Prjibelski, A.D.; et al. SPAdes: A New Genome Assembly Algorithm and Its Applications to Single-Cell Sequencing. J. Comput. Biol. 2012, 19, 455–477. [Google Scholar] [CrossRef]
- Pickett, B.E.; Sadat, E.L.; Zhang, Y.; Noronha, J.M.; Squires, R.B.; Hunt, V.; Liu, M.; Kumar, S.; Zaremba, S.; Gu, Z.; et al. ViPR: An Open Bioinformatics Database and Analysis Resource for Virology Research. Nucleic Acids Res. 2012, 40, D593. [Google Scholar] [CrossRef]
- Kumar, S.; Stecher, G.; Li, M.; Knyaz, C.; Tamura, K. MEGA X: Molecular Evolutionary Genetics Analysis across Computing Platforms. Mol. Biol. Evol. 2018, 35, 1547–1549. [Google Scholar] [CrossRef]
- Xie, J.; Chen, Y.; Cai, G.; Cai, R.; Hu, Z.; Wang, H. Tree Visualization By One Table (TvBOT): A Web Application for Visualizing, Modifying and Annotating Phylogenetic Trees. Nucleic Acids Res. 2023, 51, W587–W592. [Google Scholar] [CrossRef]
- Díaz Alarcón, R.G.; Liotta, D.J.; Miño, S. Zoonotic RVA: State of the Art and Distribution in the Animal World. Viruses 2022, 14, 1554. [Google Scholar] [CrossRef]
- Wang, D.; Wang, L.; Liu, Q.; Zhao, Y. Virulence Factors in Biofilm Formation and Therapeutic Strategies for Staphylococcus Aureus: A Review. Anim. Zoonoses 2025, 1, 188–202. [Google Scholar] [CrossRef]
- Yan, N.; Li, R.; Wang, Y.; Zhang, B.; Yue, H.; Tang, C. High Prevalence and Genomic Characteristics of G6P[1] Bovine Rotavirus A in Yak in China. J. Gen. Virol. 2020, 101, 701–711. [Google Scholar] [CrossRef] [PubMed]
- Fritzen, J.T.T.; Lorenzetti, E.; Oliveira, M.V.; Bon, V.R.; Ayres, H.; Alfieri, A.F.; Alfieri, A.A. Cross-Sectional Study of the G and P Genotypes of Rotavirus A Field Strains Circulating in Regularly Vaccinated Dairy Cattle Herds. Trop. Anim. Health Prod. 2019, 51, 887–892. [Google Scholar] [CrossRef] [PubMed]
- Singh, S.; Singh, R.; Singh, K.P.; Singh, V.; Malik, Y.P.S.; Kamdi, B.; Singh, R.; Kashyap, G. Immunohistochemical and Molecular Detection of Natural Cases of Bovine Rotavirus and Coronavirus Infection Causing Enteritis in Dairy Calves. Microb. Pathog. 2020, 138, 103814. [Google Scholar] [CrossRef]
- Bertoni, E.; Aduriz, M.; Bok, M.; Vega, C.; Saif, L.; Aguirre, D.; Cimino, R.O.; Miño, S.; Parreño, V. First Report of Group A Rotavirus and Bovine Coronavirus Associated with Neonatal Calf Diarrhea in the Northwest of Argentina. Trop. Anim. Health Prod. 2020, 52, 2761–2768. [Google Scholar] [CrossRef]
- Qin, Y.-F.; Gong, Q.-L.; Zhang, M.; Sun, Z.-Y.; Wang, W.; Wei, X.-Y.; Chen, Y.; Zhang, Y.; Zhao, Q.; Jiang, J. Prevalence of Bovine Rotavirus among Bovidae in China during 1984–2021: A Systematic Review and Meta-Analysis. Microb. Pathog. 2022, 169, 105661. [Google Scholar] [CrossRef]
- Elkady, G.; Chen, Y.; Hu, C.; Chen, J.; Chen, X.; Guo, A. MicroRNA Profile of MA-104 Cell Line Associated With the Pathogenesis of Bovine Rotavirus Strain Circulated in Chinese Calves. Front. Microbiol. 2022, 13, 854348. [Google Scholar] [CrossRef]
- Nikam, A.P.; Meshram, P.D.; Vanjari, A.V.; Mundada, S. V Comparative Study of Spray-Drying and Freeze-Drying Techniques for Increasing Fenofibrate’s Solubility and Dissolution Rate. BIO Integr. 2024, 5, 985. [Google Scholar] [CrossRef]
- Ghonaim, A.H.; Ghonaim, H.A.; Maria, C.; Li, W. Equine Rotavirus: A Hidden Threat to Public Health and Equine Welfare. In BT-Veterinary Virology of Domestic and Pet Animals; Wang, L., Ed.; Springer Nature Switzerland: Cham, Switzerland, 2025; pp. 1–20. ISBN 978-3-031-54690-7. [Google Scholar]
- Nagai, M.; Shimada, S.; Fujii, Y.; Moriyama, H.; Oba, M.; Katayama, Y.; Tsuchiaka, S.; Okazaki, S.; Omatsu, T.; Furuya, T.; et al. H2 Genotypes of G4P[6], G5P[7], and G9[23] Porcine Rotaviruses Show Super-Short RNA Electropherotypes. Vet. Microbiol. 2015, 176, 250–256. [Google Scholar] [CrossRef]
- Bwogi, J.; Jere, K.C.; Karamagi, C.; Byarugaba, D.K.; Namuwulya, P.; Baliraine, F.N.; Desselberger, U.; Iturriza-Gomara, M. Whole Genome Analysis of Selected Human and Animal Rotaviruses Identified in Uganda from 2012 to 2014 Reveals Complex Genome Reassortment Events between Human, Bovine, Caprine and Porcine Strains. PLoS ONE 2017, 12, e0178855. [Google Scholar] [CrossRef]




| Gene | Genotype | Closest Strain | Nucleotide Sequence Identity (%) | Amino Acid Sequence Identity (%) | Host Origin | Accession Number |
|---|---|---|---|---|---|---|
| VP1 | R2 | RVA/Human-wt/HUN/BP1062/2004/G8P[14] | 93.4% | 99.36% | Human | FN665688 |
| VP2 | C2 | RVA/Yak-tc/CHN/HY-1/2018/G6P[11] | 96.13% | 99.42% | Bovine | QDJ94338 |
| VP3 | M2 | RVA/Cat-wt/ITA/BA222/2005/G3P[9] | 90.51% | 95.21% | Feline | GU827408 |
| VP4 | P[1] | RVA/Human-tc/NGA/HMG035/1999/G8P[1] | 91.80% | 90.65% | Human | LC119096 |
| VP6 | I2 | RVA/Cow-tc/USA/NCDV/1967/G9P[16] | 99.63% | 98.89% | Bovine | DQ870496 |
| VP7 | G6 | RVA/Cow-wt/ARG/B3035_B_BA/2007/G6P[5] | 96.62% | 96.72% | Bovine | OM643804 |
| NSP1 | A3 | RVA/Cow-tc/THA/A44/1989/G10P[11] | 94.48% | 94.97% | Bovine | LC133575.1 |
| NSP2 | N2 | RVA/Cow-tc/JPN/AzuK-7/2007/G10P[11] | 95.49% | 96.85% | Bovine | LC553635 |
| NSP3 | T6 | RVA/Cow-tc/USA/B223/1983/G10P[11] | 98.84% | 99.42% | Bovine | LC133555 |
| NSP4 | E2 | RVA/Cow-tc/JPN/BRV106/1983/G6P[1] | 99.55% | 98.63% | Bovine | AB748604.1 |
| NSP5 | H3 | RVA/Human-wt/USA/3000015004/2014/G6P[14] | 97.82% | 95.92% | Human | OP680466.1 |
| Strain | G | P | I | R | C | M | A | N | T | E | H |
|---|---|---|---|---|---|---|---|---|---|---|---|
| 0205HG | 6 | 1 | 2 | 2 | 2 | 2 | 3 | 2 | 6 | 2 | 3 |
| RVA/Human-tc/USA/DS-1/1976/G2P1B[4] | 2 | 4 | 2 | 2 | 2 | 2 | 2 | 2 | 2 | 2 | 2 |
| RVA/Human-tc/JPN/AU-1/1982/G3P3[9] | 3 | 9 | 3 | 3 | 3 | 3 | 3 | 3 | 3 | 3 | 3 |
| RVA/Cow-tc/ARG/B3700/2008/G6P[11] | 6 | 11 | 2 | 5 | 2 | 2 | 3 | 2 | 6 | 12 | 3 |
| RVA/Hu-wt/SVN/SI-R56/07/2007/G6P[11] | 6 | 11 | 2 | 2 | 2 | 2 | 13 | 2 | 6 | 2 | 3 |
| RVA/Cow-wt/IRL/CIT-A21/2006/G6P[11] | 6 | 11 | 2 | 2 | 2 | 2 | 3 | 2 | 6 | 2 | 3 |
| RVA/Cow-wt/SVN/SI-B17/2004/G6P[11] | 6 | 11 | 2 | 2 | 2 | 2 | 3 | 2 | 6 | 2 | 3 |
| RVA/Rabbit-tc/NLD/K1130027/2011/G6P[11] | 6 | 11 | 2 | 2 | 2 | 2 | 13 | 2 | 6 | 2 | 3 |
| RVA/Cow-wt/ZAF/MRC-456/2009/G6P[11] | 6 | 11 | 2 | 2 | 2 | 2 | 13 | 2 | 6 | 2 | 3 |
| RVA/Cow-wt/ZAF/MRC-3005/2009/G6P[11] | 6 | 11 | 2 | 2 | 2 | 2 | 13 | 2 | 6 | 2 | 3 |
| RVA/Cow-tc/UK/GBR/1973/G6P[5] | 6 | 5 | 2 | 2 | 2 | 2 | 3 | 2 | 7 | 2 | 3 |
| RVA/Cow-tc/ARG/B3035/2007/G6P[5] | 6 | 5 | 2 | 5 | 2 | 2 | 3 | 2 | 6 | 12 | 3 |
| RVA/Cow-tc/JPN/OH-4/1982/G6P[5] | 6 | 5 | 2 | 2 | 2 | 2 | 13 | 2 | 6 | 2 | 3 |
| RVA/Cow-wt/ZAF/1603/2007/G6P[5] | 6 | 5 | 2 | 2 | 2 | 2 | 3 | 2 | 6 | 2 | 3 |
| RVA/Cow-tc/FRN/RF/1982/G6P[1] | 6 | 1 | 2 | 2 | 2 | 2 | 3 | 2 | 6 | 2 | 3 |
| RVA/Cow-tc/USA/WC3/1981/G6P[1] | 6 | 1 | 2 | 2 | 2 | 2 | 3 | 2 | 6 | 2 | 3 |
| RVA/Cow-tc/USA/NCDV/1967/G6P[1] | 6 | 1 | 2 | 2 | 2 | 2 | 3 | 2 | 6 | 2 | 3 |
| RVA/Goat-tc/UGA/BUW-14/2014/G6P[1] | 6 | 1 | 2 | 2 | 2 | 2 | 11 | 2 | 6 | 2 | 3 |
| RVA/Goat-tc/BGD/GO34/1999/G6P[1] | 6 | 1 | 2 | 2 | 2 | 2 | 11 | 2 | 6 | 2 | 3 |
| RVA/Human-wt/GHA/PML1965/2012/G6P[6] | 6 | 6 | 2 | 2 | 2 | 2 | 2 | 2 | 2 | 2 | 2 |
| RVA/Human-wt/BEL/B1711/2002/G6P[6] | 6 | 6 | 2 | 2 | 2 | 2 | 2 | 2 | 2 | 2 | 2 |
| RVA/Human-wt/GNB/MRC-DPRU5625/2011/G6P[6] | 6 | 6 | 2 | 2 | 2 | 2 | 2 | 2 | 2 | 2 | 2 |
| RVA/Human-wt/CMR/MA202/2011/G6P[6] | 6 | 6 | 2 | 2 | 2 | 2 | 2 | 2 | 2 | 2 | 2 |
| RVA/Human-wt/BGD/KH2288/2011/G6P[8] | 6 | 8 | 2 | 2 | 2 | 2 | 11 | 2 | 6 | 2 | 3 |
| RVA/Human-wt/JPN/KF17/2010/G6P[9] | 6 | 9 | 2 | 2 | 2 | 2 | 3 | 2 | 3 | 3 | 3 |
| RVA/Human-tc/ITA/PA169/1988/G6P[14] | 6 | 14 | 2 | 2 | 2 | 2 | 3 | 2 | 6 | 2 | 3 |
| RVA/Human-wt/HUN/Hun5/1997/G6P[14] | 6 | 14 | 2 | 2 | 2 | 2 | 11 | 2 | 6 | 2 | 3 |
| RVA/Human-wt/EGY/EGY3399/2004/G6P[14] | 6 | 14 | 2 | 2 | 2 | 2 | 11 | 2 | 6 | 2 | 3 |
| RVA/Human-wt/IND/N-1/2009/G6P[14] | 6 | 14 | 2 | 2 | 2 | 2 | 11 | 2 | 6 | 2 | 3 |
| RVA/Human-tc/AUS/MG6/1993/G6P[14] | 6 | 14 | 2 | 2 | 2 | 2 | 11 | 2 | 6 | 2 | 3 |
| RVA/Human-wt/BEL/B10925/1997/G6P[14] | 6 | 14 | 2 | 2 | 2 | 2 | 3 | 2 | 6 | 2 | 3 |
| RVA/Human-wt/HUN/BP1879/2003/G6P[14] | 6 | 14 | 2 | 2 | 2 | 2 | 11 | 2 | 6 | 2 | 3 |
| RVA/Camel–wt/SDN/MRC–DPRU447/2002/G8P[11] | 8 | 11 | 2 | 2 | 2 | 2 | 18 | 2 | 6 | 2 | 1 |
| RVA/Cow-tc/THA/A44/1989/G10P[11] | 10 | 11 | 2 | 2 | 2 | 2 | 3 | 2 | 6 | 2 | 3 |
| RVA/Cow-tc/USA/B223/1983/G10P[11] | 10 | 11 | 2 | 2 | 2 | 2 | 13 | 2 | 6 | 2 | 3 |
| RVA/Giraff e-wt/IRL/GirRV/2008/G10P[11] | 10 | 11 | 2 | 2 | 2 | 2 | 3 | 2 | 6 | 2 | 3 |
| RVA/Cow-tc/JPN/KK3/1983/G10P[11] | 10 | 11 | 2 | 2 | 2 | 2 | 13 | 2 | 6 | 2 | 3 |
| RVA/Human-wt/IND/N155/200 3/G10P[11] | 10 | 11 | 2 | 2 | 2 | 2 | 1 | 1 | 1 | 2 | 3 |
| RVA/Cow-tc/CHN/DQ-75/2008/G10P[11] | 10 | 11 | 2 | 2 | 2 | 2 | 3 | 2 | 6 | 2 | 3 |
| RVA/Cow-tc/ARG/B383/1998/G15P[11] | 15 | 11 | 2 | 5 | 2 | 2 | 13 | 2 | 6 | 12 | 3 |
| RVA/Human-tc/USA/DS-1/1976/G2P1B[4] | 2 | 4 | 2 | 2 | 2 | 2 | 2 | 2 | 2 | 2 | 2 |
| RVA/Human-re/GBR/Re/1987/G10P[14] | 10 | 14 | 2 | 2 | 2 | 1 | 3 | 2 | 6 | 2 | 3 |
| RVA/Human-wt/IND/BCH 5836/2015/G3P[4] | 3 | 4 | 1 | 1 | 1 | 1 | 1 | 1 | 2 | 1 | 1 |
| RVA/Cow-wt/ZAF/1604/2007/G8P[1] | 8 | 1 | 2 | 2 | 2 | 2 | 3 | 2 | 6 | 2 | 3 |
| RVA/Human-wt/IND/BCH 7221/2015/G3P[8] | 3 | 8 | 1 | 1 | 1 | 1 | 1 | 1 | 1 | 1 | 1 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2025 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Ghonaim, A.H.; Lei, M.; Zeng, Y.; Xu, Q.; Hong, B.; Li, D.; Yang, Z.; Zhou, J.; Liu, C.; He, Q.; et al. Molecular Surveillance and Whole Genomic Characterization of Bovine Rotavirus A G6P[1] Reveals Interspecies Reassortment with Human and Feline Strains in China. Vet. Sci. 2025, 12, 742. https://doi.org/10.3390/vetsci12080742
Ghonaim AH, Lei M, Zeng Y, Xu Q, Hong B, Li D, Yang Z, Zhou J, Liu C, He Q, et al. Molecular Surveillance and Whole Genomic Characterization of Bovine Rotavirus A G6P[1] Reveals Interspecies Reassortment with Human and Feline Strains in China. Veterinary Sciences. 2025; 12(8):742. https://doi.org/10.3390/vetsci12080742
Chicago/Turabian StyleGhonaim, Ahmed H., Mingkai Lei, Yang Zeng, Qian Xu, Bo Hong, Dongfan Li, Zhengxin Yang, Jiaru Zhou, Changcheng Liu, Qigai He, and et al. 2025. "Molecular Surveillance and Whole Genomic Characterization of Bovine Rotavirus A G6P[1] Reveals Interspecies Reassortment with Human and Feline Strains in China" Veterinary Sciences 12, no. 8: 742. https://doi.org/10.3390/vetsci12080742
APA StyleGhonaim, A. H., Lei, M., Zeng, Y., Xu, Q., Hong, B., Li, D., Yang, Z., Zhou, J., Liu, C., He, Q., Zhang, Y., & Li, W. (2025). Molecular Surveillance and Whole Genomic Characterization of Bovine Rotavirus A G6P[1] Reveals Interspecies Reassortment with Human and Feline Strains in China. Veterinary Sciences, 12(8), 742. https://doi.org/10.3390/vetsci12080742









