An Environmental Data Collection for COVID-19 Pandemic Research
Abstract
:1. Summary
2. Data Description
2.1. Raw Measurements and Data Sources
2.1.1. MERRA-2 Temperature and Humidity Reanalysis
2.1.2. IMERG Precipitation Estimation
2.1.3. NPP/VIIRS Nighttime Light radiance
2.1.4. Aura-OMI Air Pollution Observation
2.1.5. Ground-Based Air Quality Data
2.2. Derived Product and Metadata
2.2.1. Daily/Monthly Global Environmental Factors Reprocessing
2.2.2. Environmental Factors of Multiple Administration Levels
3. Methods
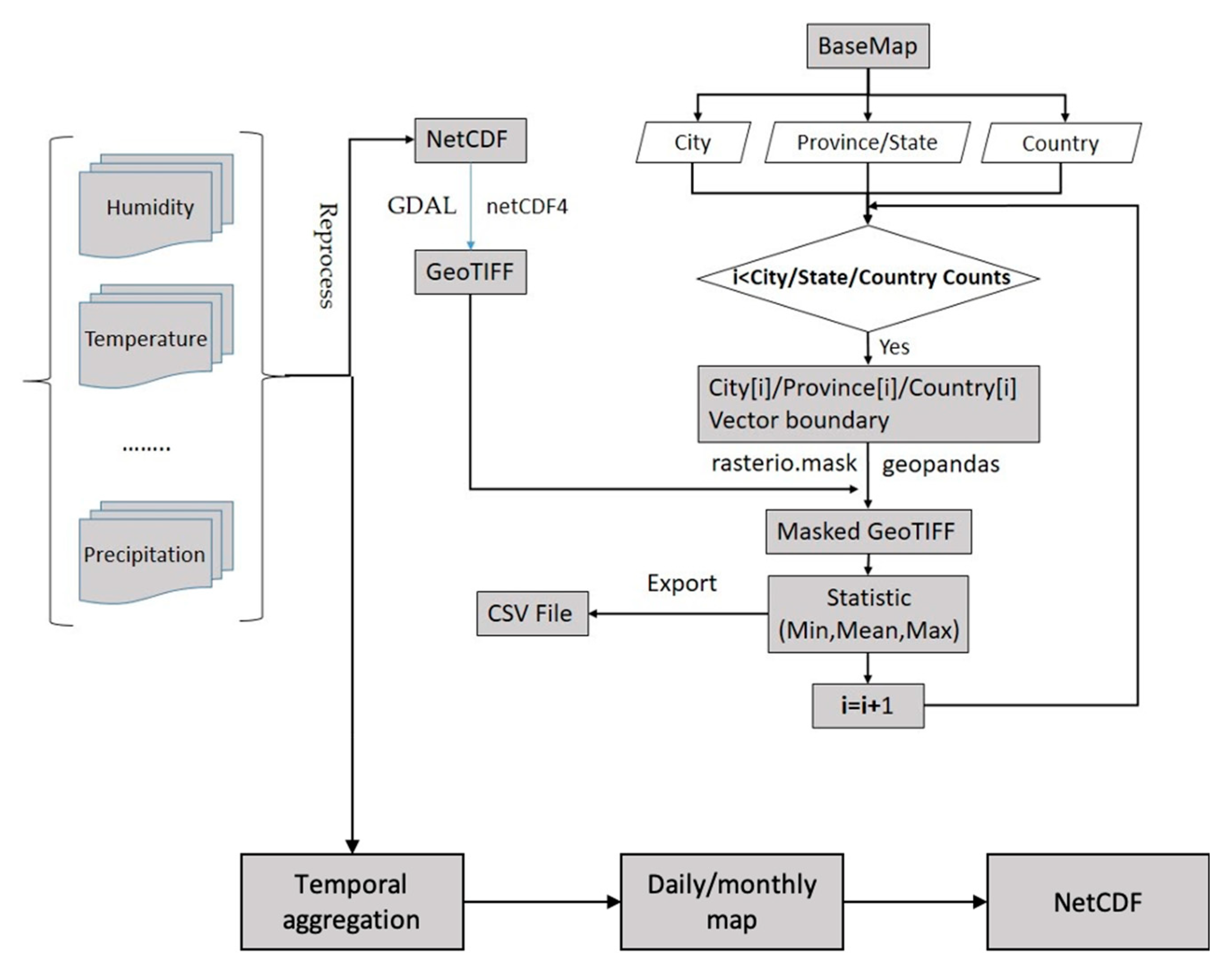
3.1. Spatiotemporal Aggregation and Collocation
3.2. Collocating Environmental Factors with COVID-19 Case Data
3.3. Data Computing and Storage on AWS Cloud Platform
4. Data Sharing
5. Quality Control
6. Conclusions
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Kagan, A.R.; Levi, L. Health and environment—Psychosocial stimuli: A review. Soc. Sci. Med. 1974, 8, 225–241. [Google Scholar] [CrossRef]
- Chan, K.H.; Peiris, J.S.M.; Lam, S.Y.; Poon, L.L.; Yuen, K.-Y.; Seto, W.H. The Effects of Temperature and Relative Humidity on the Viability of the SARS Coronavirus. Adv. Virol. 2011, 2011, 1–7. [Google Scholar] [CrossRef] [PubMed]
- Chen, B.; Liang, H.; Yuan, X.; Hu, Y.; Xu, M.; Zhao, Y.; Zhang, B.; Tian, F.; Zhu, X. Roles of meteorological conditions in COVID-19 transmission on a worldwide scale. MedRxiv 2020. [Google Scholar] [CrossRef] [Green Version]
- Bashir, M.F.; Ma, B.; Komal, B.; Bashir, M.A.; Tan, D.; Bashir, M. Correlation between climate indicators and COVID-19 pandemic in New York, USA. Sci. Total Environ. 2020, 728, 138835. [Google Scholar] [CrossRef] [PubMed]
- Liu, Q.; Sha, D.; Liu, W.; Houser, P.; Zhang, L.; Hou, R.; Lan, H.; Flynn, C.; Lu, M.; Hu, T.; et al. Spatiotemporal Patterns of COVID-19 Impact on Human Activities and Environment in Mainland China Using Nighttime Light and Air Quality Data. Remote Sens. 2020, 12, 1576. [Google Scholar] [CrossRef]
- Prata, D.N.; Rodrigues, W.; Bermejo, P.H. Temperature significantly changes COVID-19 transmission in (sub) tropical cities of Brazil. Sci. Total Environ. 2020, 729, 138862. [Google Scholar] [CrossRef]
- Shereen, M.A.; Khan, S.; Kazmi, A.; Bashir, N.; Siddique, R. COVID-19 infection: Origin, transmission, and characteristics of human coronaviruses. J. Adv. Res. 2020, 24, 91–98. [Google Scholar] [CrossRef]
- Bashir, M.F.; Bilal, B.J.M.A.; Komal, B. Correlation between environmental pollution indicators and COVID-19 pandemic: A brief study in Californian context. Environ. Res. 2020, 187, 109652. [Google Scholar] [CrossRef]
- Murgante, B.; Borruso, G.; Balletto, G.; Castiglia, P.; Dettori, M. Why Italy First? Health, Geographical and Planning Aspects of the COVID-19 Outbreak. Sustainability 2020, 12, 5064. [Google Scholar] [CrossRef]
- You, H.; Wu, X.; Guo, X. Distribution of COVID-19 Morbidity Rate in Association with Social and Economic Factors in Wuhan, China: Implications for Urban Development. Int. J. Environ. Res. Public Health 2020, 17, 3417. [Google Scholar] [CrossRef]
- Pirouz, B.; Haghshenas, S.S.; Piro, P.; Haghshenas, S.S. Investigating a Serious Challenge in the Sustainable Development Process: Analysis of Confirmed cases of COVID-19 (New Type of Coronavirus) Through a Binary Classification Using Artificial Intelligence and Regression Analysis. Sustainability 2020, 12, 2427. [Google Scholar] [CrossRef] [Green Version]
- Peng, Z.; Wang, R.; Liu, L.; Wu, H. Exploring Urban Spatial Features of COVID-19 Transmission in Wuhan Based on Social Media Data. ISPRS Int. J. Geo-Inf. 2020, 9, 402. [Google Scholar] [CrossRef]
- What Is a Data Platform? Available online: https://looker.com/definitions/data-platform#:~:text=A%20data%20platform%20is%20an,technologies%20for%20strategic%20business%20purposes (accessed on 14 June 2020).
- World Health Organization. Coronavirus Disease (COVID-2019) Situation Reports. Available online: https://www.who.int/emergencies/diseases/novel-coronavirus-2019/situation-reports (accessed on 14 June 2020).
- Center for Disease Control and Prevention. Get the Facts About Coronavirus. Available online: https://www.cdc.gov/coronavirus/2019-ncov/index.html (accessed on 14 June 2020).
- Virginia Department of Health. COVID-19 in Virginia. Available online: https://www.vdh.virginia.gov/coronavirus/ (accessed on 14 June 2020).
- Maryland Department of Health. COVID-19 Outbreak. Available online: https://phpa.health.maryland.gov/Pages/Novel-coronavirus.aspx (accessed on 14 June 2020).
- Government of the District of Columbia. Coronavirus Data. Available online: https://coronavirus.dc.gov/page/coronavirus-data (accessed on 14 June 2020).
- The New York Times. The Coronavirus Outbreak. Available online: https://www.nytimes.com/news-event/coronavirus (accessed on 14 June 2020).
- Dong, E.; Du, H.; Gardner, L.M. An interactive web-based dashboard to track COVID-19 in real time. Lancet Infect. Dis. 2020, 20, 533–534. [Google Scholar] [CrossRef]
- 1 Point 3 Acres. Global COVID-19 Tracker & Interactive Chart. Available online: https://coronavirus.1point3acres.com/en/ (accessed on 14 June 2020).
- Yang, C.; Sha, D.; Liu, Q.; Li, Y.; Lan, H.; Guan, W.W.; Hu, T.; Li, Z.; Zhang, Z.; Thompson, J.H.; et al. Taking the pulse of COVID-19: A spatiotemporal perspective. arXiv 2020, arXiv:2005.04224v1. [Google Scholar]
- Wang, J.; Tang, K.; Feng, K.; Lv, W. High Temperature and High Humidity Reduce the Transmission of COVID-19. SSRN Electron. J. 2020. [Google Scholar] [CrossRef] [Green Version]
- Sajadi, M.M.; Habibzadeh, P.; Vintzileos, A.; Shokouhi, S.; Miralles-Wilhelm, F.; Amoroso, A. Temperature and Latitude Analysis to Predict Potential Spread and Seasonality for COVID-19. SSRN Electron. J. 2020. [Google Scholar] [CrossRef]
- Gelaro, R.; Mccarty, W.; Suárez, M.J.; Todling, R.; Molod, A.; Takacs, L.; Randles, C.; Darmenov, A.; Bosilovich, M.G.; Reichle, R.H.; et al. The Modern-Era Retrospective Analysis for Research and Applications, Version 2 (MERRA-2). J. Clim. 2017, 30, 5419–5454. [Google Scholar] [CrossRef]
- Ruiz, M.O.; Chaves, L.F.; Hamer, G.L.; Sun, T.; Brown, W.M.; Walker, E.D.; Haramis, L.; Goldberg, T.L.; Kitron, U. Local impact of temperature and precipitation on West Nile virus infection in Culex species mosquitoes in northeast Illinois, USA. Parasites Vectors 2010, 3, 19. [Google Scholar] [CrossRef] [Green Version]
- Huffman, G.J.; Bolvin, D.T.; Nelkin, E.J. Integrated Multi-satellitE Retrievals for GPM (IMERG) technical documentation. NASA/GSFC Code 2015, 612, 2019. [Google Scholar]
- Liu, Q.; Li, Y.; Yu, M.; Chiu, L.S.; Hao, X.; Duffy, D.Q.; Yang, C. Daytime Rainy Cloud Detection and Convective Precipitation Delineation Based on a Deep Neural Network Method Using GOES-16 ABI Images. Remote Sens. 2019, 11, 2555. [Google Scholar] [CrossRef] [Green Version]
- Román, M.O.; Wang, Z.; Sun, Q.; Kalb, V.; Miller, S.D.; Molthan, A.; Schultz, L.; Bell, J.; Stokes, E.C.; Pandey, B.; et al. Nasa’s black marble nighttime lights product suite. Remote Sens. Environ. 2018, 210, 113–143. [Google Scholar] [CrossRef]
- Russell, A.R.; Valin, L.C.; Cohen, R.C. Trends in OMI NO2 observations over the United States: Effects of emission control technology and the economic recession. Atmos. Chem. Phys. Discuss. 2012, 12, 12197–12209. [Google Scholar] [CrossRef] [Green Version]
- Levelt, P.; Oord, G.V.D.; Dobber, M.; Malkki, A.; Visser, H.; De Vries, J.; Stammes, P.; Lundell, J.; Saari, H. The ozone monitoring instrument. IEEE Trans. Geosci. Remote Sens. 2006, 44, 1093–1101. [Google Scholar] [CrossRef]
- Isaifan, R. The dramatic impact of Coronavirus outbreak on air quality: Has it saved as much as it has killed so far? Glob. J. Environ. Sci. Manag. 2020, 6, 275–288. [Google Scholar]
- Yang, C.; Clarke, K.; Shekhar, S.; Tao, C.V. Big Spatiotemporal Data Analytics: A research and innovation frontier. Int. J. Geogr. Inf. Sci. 2019, 34, 1075–1088. [Google Scholar] [CrossRef] [Green Version]
- Yang, C.; Yu, M.; Li, Y.; Hu, F.; Jiang, Y.; Liu, Q.; Sha, D.; Xu, M.; Gu, J. Big Earth data analytics: A survey. Big Earth Data 2019, 3, 83–107. [Google Scholar] [CrossRef] [Green Version]
| Data Sources | Features and Information Offered | Roles in COVID-19 Related Studies | Download Address |
|---|---|---|---|
| MERRA-2 | Temperature, humidity, environmental condition | Suitability of virus spread, spread range and rate | https://disc.gsfc.nasa.gov/datasets/M2T1NXSLV_5.12.4/summary?keywords=MERRA2_400.tavg1_2d_slv_Nx |
| IMERG | Precipitation rate | Spread range and rate | https://disc.gsfc.nasa.gov/datasets/GPM_3IMERGHHE_06/summary?keywords=IMERG |
| NPP VIIRS/DNB | Nighttime light radiance, human activities, community distributions, human-gathering levels | Human activities impact | https://ladsweb.modaps.eosdis.nasa.gov/ |
| Aura-OMI | Concentration of air pollutants | Mortality rate, human activities impact | https://disc.gsfc.nasa.gov/datasets/OMNO2d_003/summary?keywords=omi |
| Ground-based air pollution observations | Air quality index, air pollution concentration | Mortality rate, human activities impact | http://data.cma.cn/ |
| Dataset | Attribute Name | Dimension | Description | Units |
|---|---|---|---|---|
| Daily Reanalyzed Specific Humidity | daily_QV2M | xy | 2-m specific humidity | kg kg−1 |
| nlat | y | latitude | degree north | |
| nlon | x | longitude | degree east | |
| Daily Reanalyzed Near Surface Temperature | daily_T2M | xy | 2-m temperature | K |
| nlat | y | latitude | degree north | |
| nlon | x | longitude | degree east | |
| Daily Precipitation | daily_precipitation | xy | daily precipitation | mm/hour |
| nlat | y | latitude | degree north | |
| nlon | x | longitude | degree east | |
| Monthly Nighttime Light Radiance | monthly_mean_radiance | number of pixels | monthly mean radiance | nW/(cm2 sr) |
| nlat | number of pixels | latitude | degree north | |
| nlon | number of pixels | longitude | degree east |
| Attribute Name | Description | Format | Units | |
|---|---|---|---|---|
| Environmental Factors of City-level | GID_2 | Used to uniquely identify the city | String | None |
| Max | Maximum Temperature/Humidity/ Precipitation/ TVCD | Float | K/(kg kg−1)/(mm/hour)/ (molec/cm2) | |
| Mean | Average Temperature/Humidity/ Precipitation/ TVCD | Float | K/(kg kg−1)/(mm/hour)/ (molec/cm2) | |
| Min | Minimum Temperature/Humidity/ Precipitation/ TVCD | Float | K/(kg kg−1)/(mm/hour)/ (molec/cm2) | |
| Environmental Factors of Province-level/State-level | GID_1 | Used to uniquely identify the province/state | String | None |
| Max | Maximum Temperature/Humidity/ Precipitation/ TVCD | Float | K/(kg kg−1)/(mm/hour)/ (molec/cm2) | |
| Mean | Average Temperature/Humidity/ Precipitation/ TVCD | Float | K/(kg kg−1)/(mm/hour)/ (molec/cm2) | |
| Min | Minimum Temperature/Humidity/ Precipitation/ TVCD | Float | K/(kg kg−1)/(mm/hour)/ (molec/cm2) | |
| Environmental Factors of Country-levels | GID_0 | Used to uniquely identify the country | String | None |
| Max | Maximum Temperature/Humidity/ Precipitation/ TVCD | Float | K/(kg kg−1)/(mm/hour)/ (molec/cm2) | |
| Mean | Average Temperature/Humidity/ Precipitation/ TVCD | Float | K/(kg kg−1)/(mm/hour)/ (molec/cm2) | |
| Min | Minimum Temperature/Humidity/ Precipitation/ TVCD | Float | K/(kg kg−1)/(mm/hour)/ (molec/cm2) |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Liu, Q.; Liu, W.; Sha, D.; Kumar, S.; Chang, E.; Arora, V.; Lan, H.; Li, Y.; Wang, Z.; Zhang, Y.; et al. An Environmental Data Collection for COVID-19 Pandemic Research. Data 2020, 5, 68. https://doi.org/10.3390/data5030068
Liu Q, Liu W, Sha D, Kumar S, Chang E, Arora V, Lan H, Li Y, Wang Z, Zhang Y, et al. An Environmental Data Collection for COVID-19 Pandemic Research. Data. 2020; 5(3):68. https://doi.org/10.3390/data5030068
Chicago/Turabian StyleLiu, Qian, Wei Liu, Dexuan Sha, Shubham Kumar, Emily Chang, Vishakh Arora, Hai Lan, Yun Li, Zifu Wang, Yadong Zhang, and et al. 2020. "An Environmental Data Collection for COVID-19 Pandemic Research" Data 5, no. 3: 68. https://doi.org/10.3390/data5030068
APA StyleLiu, Q., Liu, W., Sha, D., Kumar, S., Chang, E., Arora, V., Lan, H., Li, Y., Wang, Z., Zhang, Y., Zhang, Z., Harris, J. T., Chinala, S., & Yang, C. (2020). An Environmental Data Collection for COVID-19 Pandemic Research. Data, 5(3), 68. https://doi.org/10.3390/data5030068







