TIRF Microscope Image Sequences of Fluorescent IgE-FcεRI Receptor Complexes inside a FcεRI-Centric Synapse in RBL-2H3 Cells
Abstract
1. Summary
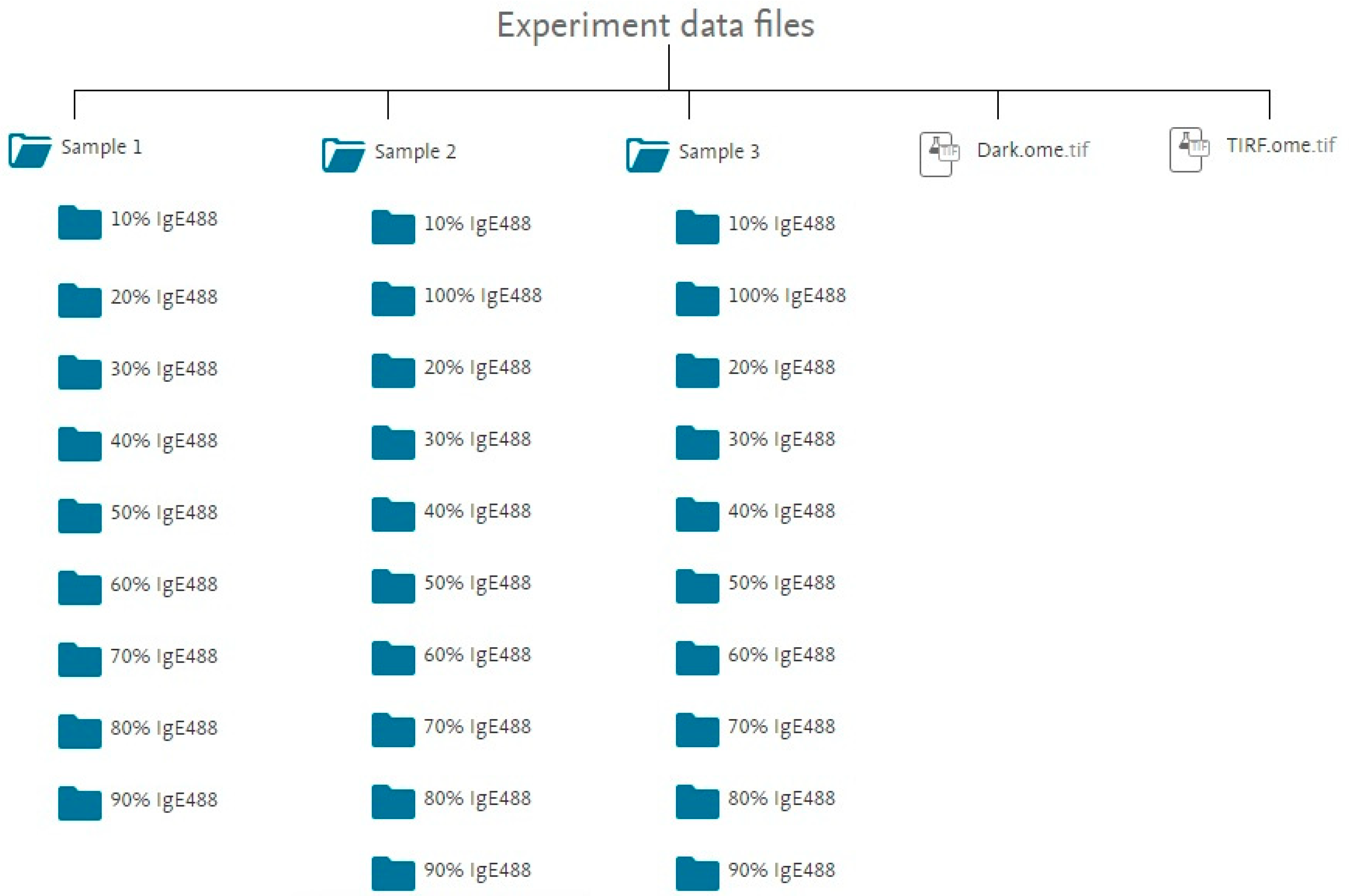
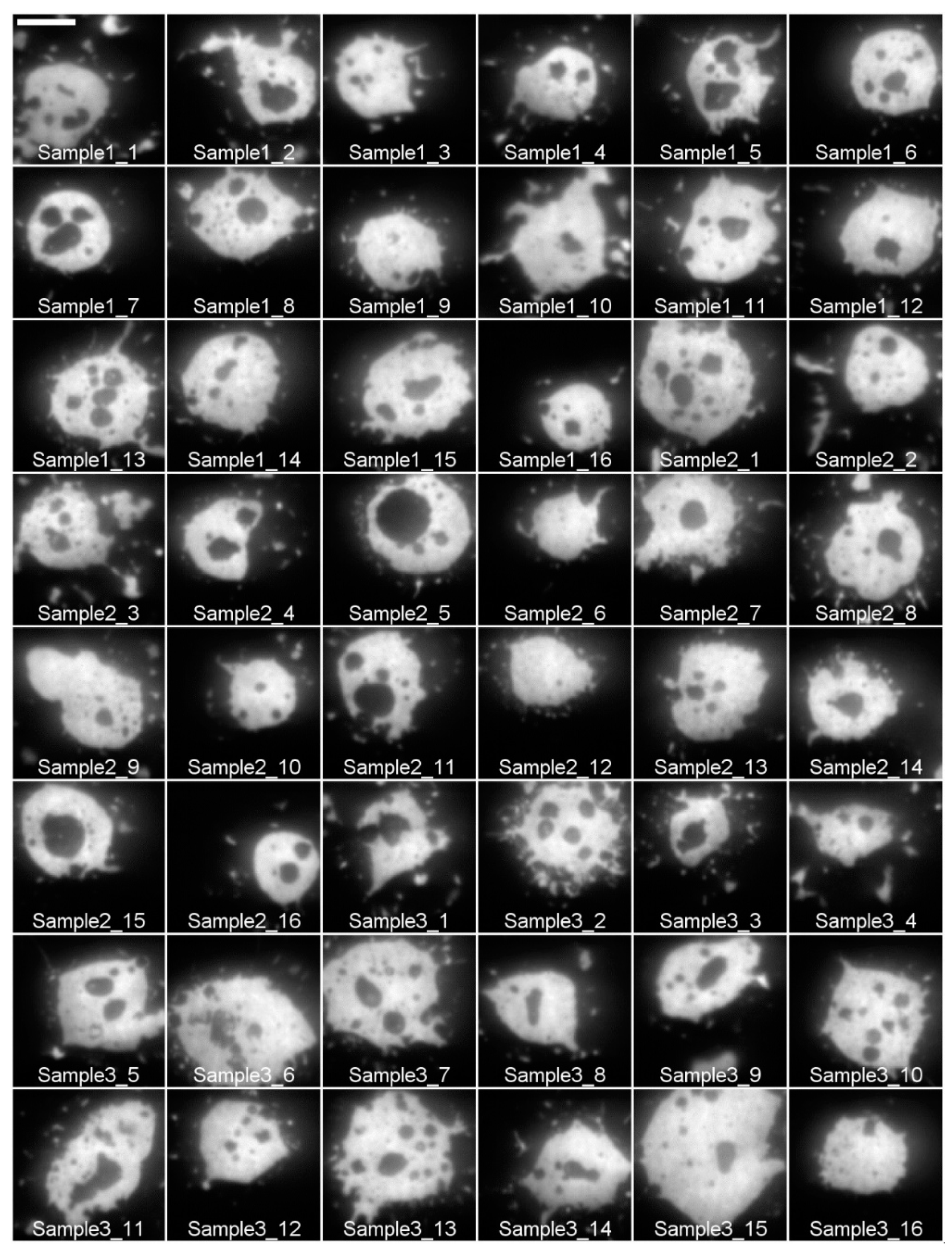
2. Data Description
- [%] refers to the percentage of IgE488 and 100% - [%] gives the corresponding percentage of IgEdark for a given image stack.
- [#] is the sample number. [#] can be 1, 2, or 3.
- [cell #] is an integer number ranging from 1 to 26, identifying a particular cell for a given IgE488 percentage and sample number.
3. Methods
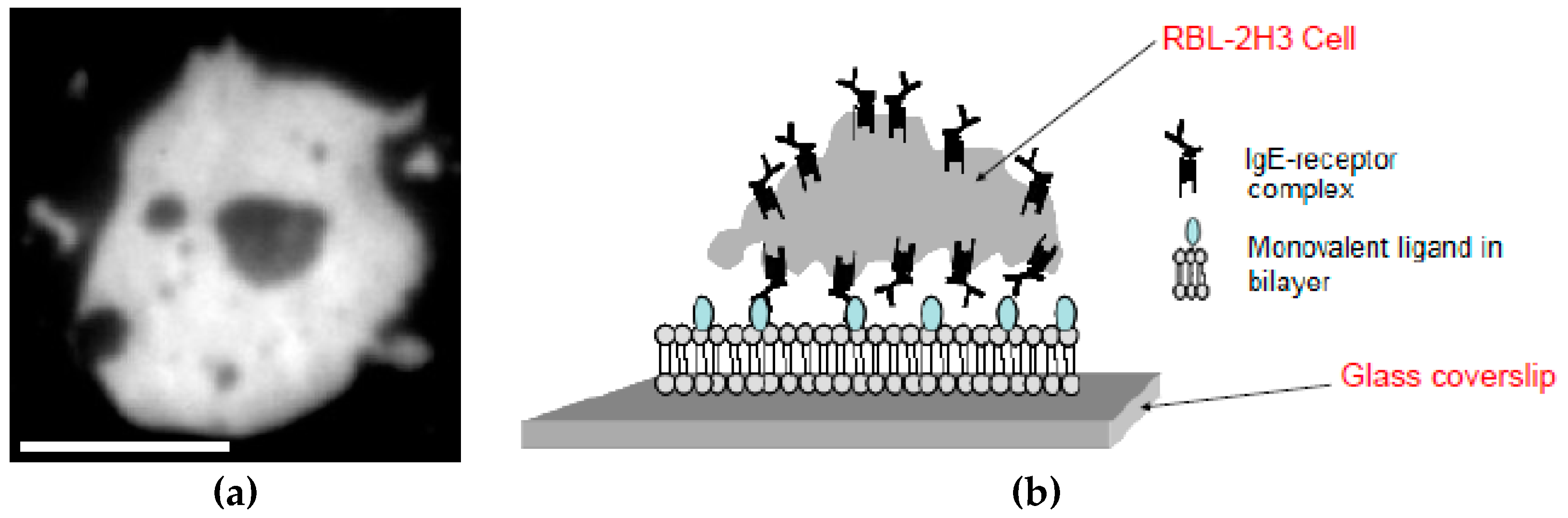
3.1. Labeling of RBL-2H3 Cells at Varying Fluorescent and Dark Anti-DNP IgE Concentrations
3.2. Supported Lipid Bilayer
3.3. TIRF Microscopy
3.4. Calibration and Image Correction
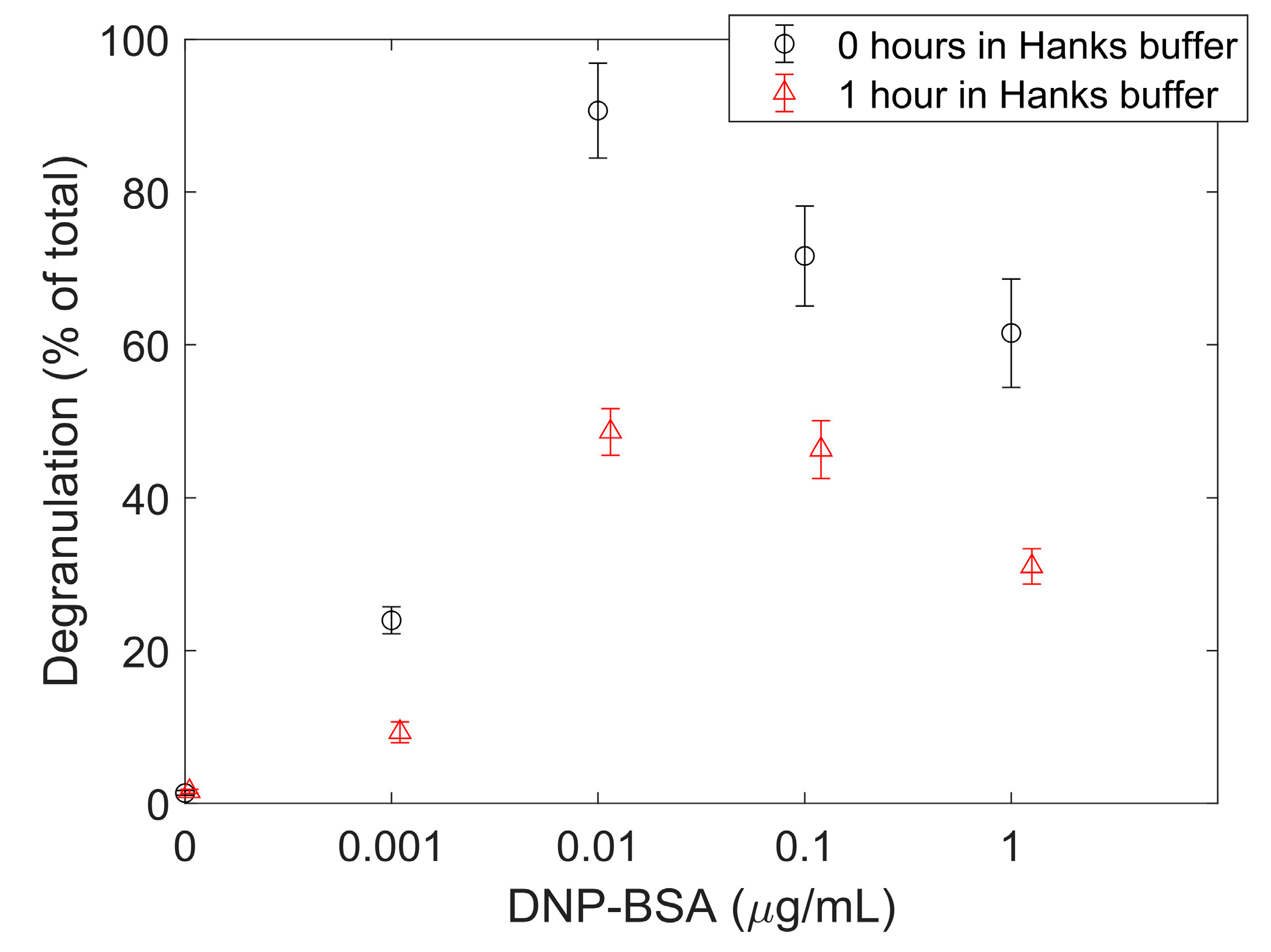
3.5. Degranulation Assay
4. User Notes
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Radhakrishnan, K.; Halász, A.; Vlachos, D.; Edwards, J.S. Quantitative understanding of cell signaling: The importance of membrane organization. Curr. Opin. Biotechnol. 2010, 21, 677–682. [Google Scholar] [CrossRef] [PubMed]
- Metzger, H. Transmembrane signaling: The joy of aggregation. J. Immunol. 1992, 149, 1477–1487. [Google Scholar] [PubMed]
- Sever, R.; Brugge, J.S. Signal transduction in cancer. Cold Spring Harb. Perspect. Med. 2015, 5, a006098. [Google Scholar] [CrossRef] [PubMed]
- Sigalov, A.B. Transmembrane signaling: From understanding to action. Self. Nonself. 2010, 1, 341–342. [Google Scholar] [CrossRef][Green Version]
- Colgan, J.D.; Hankel, I.L. Signaling pathways critical for allergic airway inflammation. Curr. Opin. Allergy Clin. Immunol. 2010, 10, 42–47. [Google Scholar] [CrossRef] [PubMed]
- Galli, S.J.; Tsai, M.; Piliponsky, A.M. The development of allergic inflammation. Nature 2008, 454, 445–454. [Google Scholar] [CrossRef]
- Thomas, J.L.; Feder, T.J.; Webb, W.W. Effects of protein concentration on IgE receptor mobility in rat basophilic leukemia cell plasma membranes. Biophys. J. 1992, 61, 1402–1412. [Google Scholar] [CrossRef][Green Version]
- Thomas, J.L.; Holowka, D.; Baird, B.; Webb, W.W. Large-scale co-aggregation of fluorescent lipid probes with cell surface proteins. J. Cell Biol. 1994, 125, 795–802. [Google Scholar] [CrossRef]
- Posner, R.G.; Subramanian, K.; Goldstein, B.; Thomas, J.; Feder, T.; Holowka, D.; Baird, B. Simultaneous cross-linking by two nontriggering bivalent ligands causes synergistic signaling of IgE Fc epsilon RI complexes. J. Immunol. 1995, 155, 3601–3609. [Google Scholar]
- Passante, E.; Ehrhardt, C.; Sheridan, H.; Frankish, N. RBL-2H3 cells are an imprecise model for mast cell mediator release. Inflamm. Res. 2009, 58, 611–618. [Google Scholar] [CrossRef]
- Carroll-Portillo, A.; Spendier, K.; Pfeiffer, J.; Griffiths, G.; Li, H.; Lidke, K.A.; Oliver, J.M.; Lidke, D.S.; Thomas, J.L.; Wilson, B.S.; et al. Formation of a mast cell synapse: Fc epsilon RI membrane dynamics upon binding mobile or immobilized ligands on surfaces. J. Immunol. 2010, 184, 1328–1338. [Google Scholar] [CrossRef]
- Spendier, K.; Thomas, J.L. Mast Cells: Phenotypic Features, Biological Functions, and Role in Immunity; Nova Science Publishers: Hauppauge, NY, USA, 2013; Chapter 2- Spatial Patterns in Mast Cell Activation; ISBN 978-162-618-166-3. [Google Scholar]
- Spendier, K.; Lidke, K.A.; Lidke, D.S.; Thomas, J.L. Single-particle tracking of immunoglobulin e receptors (FcRI) in micron-sized clusters and receptor patches. FEBS Lett. 2012, 586, 416–421. [Google Scholar] [CrossRef]
- Spendier, K.; Carroll-Portillo, A.; Lidke, K.A.; Wilson, B.S.; Timlin, J.A.; Thomas, J.L. Distribution and dynamics of rat basophilic leukemia immunoglobulin E receptors (FcepsilonRI) on planar ligand-presenting surfaces. Biophys. J. 2010, 99, 388–397. [Google Scholar] [CrossRef]
- Machado, R.; Bendesky, J.; Brown, M.; Spendier, K.; Hagen, G.M. Imaging Membrane Curvature inside a FcεRI-Centric Synapse in RBL-2H3 Cells Using TIRF Microscopy with Polarized Excitation. J. Imaging 2019, 5, 63. [Google Scholar] [CrossRef]
- Jung, S.-R.; Hille, B. Optical approaches for visualization of arrestin binding to muscarinic receptor. Methods Cell Biol. 2019, 149, 1–18. [Google Scholar]
- Kalkur, R.S.; Ballast, A.C.; Triplett, A.R.; Spendier, K. Effects of deuterium oxide on cell growth and vesicle speed in RBL-2H3 cells. PeerJ 2014, 2, e553. [Google Scholar] [CrossRef]
- Yi, J.; Balagopalan, L.; Nguyen, T.; McIntire, K.M.; Samelson, L.E. TCR microclusters form spatially segregated domains and sequentially assemble in calcium-dependent kinetic steps. Nat. Commun. 2019, 10, 277. [Google Scholar] [CrossRef] [PubMed]
- Tolar, P.; Hanna, J.; Krueger, P.D.; Pierce, S.K. The Constant Region of the Membrane Immunoglobulin Mediates B Cell-Receptor Clustering and Signaling in Response to Membrane Antigens. Immunity 2009, 30, 44–55. [Google Scholar] [CrossRef]
- Andrews, N.L.; Lidke, K.A.; Pfeiffer, J.R.; Burns, A.R.; Wilson, B.S.; Oliver, J.M.; Lidke, D.S. Actin restricts FcɛRI diffusion and facilitates antigen-induced receptor immobilization. Nat. Cell Biol. 2008, 10, 955–963. [Google Scholar] [CrossRef]
- Su, X.; Ditlev, J.A.; Hui, E.; Xing, W.; Banjade, S.; Okrut, J.; King, D.S.; Taunton, J.; Rosen, M.K.; Vale, R.D. Phase separation of signaling molecules promotes T cell receptor signal transduction. Science 2016, 352, 595–599. [Google Scholar] [CrossRef]
- Oppong, E.; Hedde, P.N.; Sekula-Neuner, S.; Yang, L.; Brinkmann, F.; Dörlich, R.M.; Hirtz, M.; Fuchs, H.; Nienhaus, G.U.; Cato, A.C.B. Localization and Dynamics of Glucocorticoid Receptor at the Plasma Membrane of Activated Mast Cells. Small 2014, 10, 1991–1998. [Google Scholar] [CrossRef]
- Varma, R.; Campi, G.; Yokosuka, T.; Saito, T.; Dustin, M.L. T Cell Receptor-Proximal Signals Are Sustained in Peripheral Microclusters and Terminated in the Central Supramolecular Activation Cluster. Immunity 2006, 25, 117–127. [Google Scholar] [CrossRef]
- Davis, R.E.; Ngo, V.N.; Lenz, G.; Tolar, P.; Young, R.M.; Romesser, P.B.; Kohlhammer, H.; Lamy, L.; Zhao, H.; Yang, Y.; et al. Chronic active B-cell-receptor signalling in diffuse large B-cell lymphoma. Nature 2010, 463, 88–92. [Google Scholar] [CrossRef]
- Axelrod, D. Cell-substrate contacts illuminated by total internal reflection fluorescence. J. Cell Biol. 1981, 89, 141–145. [Google Scholar] [CrossRef]
- Monks, C.R.F.; Freiberg, B.A.; Kupfer, H.; Sciaky, N.; Kupfer, A. Three-dimensional segregation of supramolecular activation clusters in T cells. Nature 1998, 395, 82–86. [Google Scholar] [CrossRef]
- Kaizuka, Y.; Douglass, A.D.; Varma, R.; Dustin, M.L.; Vale, R.D. Mechanisms for segregating T cell receptor and adhesion molecules during immunological synapse formation in Jurkat T cells. Proc. Natl. Acad. Sci. USA 2007, 104, 20296–20301. [Google Scholar] [CrossRef]
- Carroll-Portillo, A.; Cannon, J.L.; Te Riet, J.; Holmes, A.; Kawakami, Y.; Kawakami, T.; Cambi, A.; Lidke, D.S. Mast cells and dendritic cells form synapses that facilitate antigen transfer for T cell activation. J. Cell Biol. 2015, 210, 851–864. [Google Scholar] [CrossRef]
- Mantri, C.K.; John, A.L. St. Immune synapses between mast cells and γδ T cells limit viral infection. J. Clin. Investig. 2019, 129, 1094–1108. [Google Scholar] [CrossRef]
- Drawbond, R.; Thomas, J.; Spendier, K. Suppressed concentration fluctuations in rat basophilic leukemia cell synapse: (indirect) evidence for large signaling complexes. Bull. Am. Phys. Soc. 2014, 59, 11. [Google Scholar]
- The OME-TIFF Format—OME Data Model and File Formats 5.6.3 Documentation. Available online: https://docs.openmicroscopy.org/ome-model/5.6.3/ome-tiff/ (accessed on 19 May 2019).
- Edelstein, A.; Amodaj, N.; Hoover, K.; Vale, R.; Stuurman, N. Computer Control of Microscopes Using µManager. In Current Protocols in Molecular Biology; John Wiley & Sons, Inc.: Hoboken, NJ, USA, 2010; Volume 92, pp. 14.20.1–14.20.17. [Google Scholar]
- Drawbond, R.; Spendier, K. TIRF Microscopy Image Sequences of Fluorescent IgE-FcεRI inside Immunological Model Synapse in RBL-2H3 Cells Dataset. Mendeley Data 2019, v2. [Google Scholar]
- Hank’s Balanced Salt Solution (HBSS) without Phenol Red. Cold Spring Harb. Protoc. 2006, 2006, pdb.rec548.
- Taylor, J.R.; John, R. An Introduction to Error Analysis: The Study of Uncertainties in Physical Measurements; University Science Books: Sausalito, CA, USA, 1997; ISBN 978-093-570-275-0. [Google Scholar]
- Lidke, K.A.; Rieger, B.; Lidke, D.S.; Jovin, T.M. The role of photon statistics in fluorescence anisotropy imaging. IEEE Trans. Image Process. 2005, 14, 1237–1245. [Google Scholar] [CrossRef]
- Smith, A.J.; Pfeiffer, J.R.; Zhang, J.; Martinez, A.M.; Griffiths, G.M.; Wilson, B.S. Microtubule-dependent transport of secretory vesicles in RBL-2H3 cells. Traffic 2003, 4, 302–312. [Google Scholar] [CrossRef]
- Fukuishi, N.; Murakami, S.; Ohno, A.; Yamanaka, N.; Matsui, N.; Fukutsuji, K.; Yamada, S.; Itoh, K.; Akagi, M. Does β-hexosaminidase function only as a degranulation indicator in mast cells? The primary role of β-hexosaminidase in mast cell granules. J. Immunol. 2014, 193, 1886–1894. [Google Scholar] [CrossRef]
- DIPimage & DIPlib. Available online: http://www.diplib.org/ (accessed on 1 February 2019).
- ImageJ. Image processing and analysis in Java. Available online: https://imagej.nih.gov/ij/index.html (accessed on 1 February 2019).
- Model, M.A.; Burkhardt, J.K. A standard for calibration and shading correction of a fluorescence microscope. Cytometry 2001, 44, 309–316. [Google Scholar] [CrossRef]
| % IgE488 (%) | % IgEdark (%) | Volume of IgE488 Added to Cell Suspension (µL) 1 | Volume of IgEdark Added to Cell Suspension (µL) 1 | Concentration of IgE488 in Cell Suspension (μg/mL) 2 | Concentration of IgEdark in Cell Suspension (μg/mL) 2 |
|---|---|---|---|---|---|
| 10 | 90 | 0.5 ± 0.1 | 3.8 ± 0.1 | 0.06 ± 0.1 | 0.53 ± 0.3 |
| 20 | 80 | 1.0 ± 0.1 | 3.4 ± 0.1 | 0.12 ± 0.2 | 0.47 ± 0.2 |
| 30 | 70 | 1.5 ± 0.1 | 3.0 ± 0.1 | 0.18 ± 0.2 | 0.41 ± 0.2 |
| 40 | 60 | 2.0 ± 0.1 | 2.6 ± 0.1 | 0.24 ± 0.2 | 0.36 ± 0.2 |
| 50 | 50 | 2.5 ± 0.1 | 2.1 ± 0.1 | 0.30 ± 0.3 | 0.30 ± 0.2 |
| 60 | 40 | 3.0 ± 0.1 | 1.7 ± 0.1 | 0.36 ± 0.3 | 0.24 ± 0.2 |
| 70 | 30 | 3.5 ± 0.1 | 1.3 ± 0.1 | 0.42 ± 0.4 | 0.18 ± 0.2 |
| 80 | 20 | 4.0 ± 0.1 | 0.9 ± 0.1 | 0.48 ± 0.4 | 0.12 ± 0.1 |
| 90 | 10 | 4.5 ± 0.1 | 0.4 ± 0.1 | 0.54 ± 0.5 | 0.06 ± 0.1 |
| 100 | 0 | 5.0 ± 0.1 | 0.0 | 0.60 ± 0.5 | 0.00 |
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Drawbond, R.; Spendier, K. TIRF Microscope Image Sequences of Fluorescent IgE-FcεRI Receptor Complexes inside a FcεRI-Centric Synapse in RBL-2H3 Cells. Data 2019, 4, 111. https://doi.org/10.3390/data4030111
Drawbond R, Spendier K. TIRF Microscope Image Sequences of Fluorescent IgE-FcεRI Receptor Complexes inside a FcεRI-Centric Synapse in RBL-2H3 Cells. Data. 2019; 4(3):111. https://doi.org/10.3390/data4030111
Chicago/Turabian StyleDrawbond, Rachel, and Kathrin Spendier. 2019. "TIRF Microscope Image Sequences of Fluorescent IgE-FcεRI Receptor Complexes inside a FcεRI-Centric Synapse in RBL-2H3 Cells" Data 4, no. 3: 111. https://doi.org/10.3390/data4030111
APA StyleDrawbond, R., & Spendier, K. (2019). TIRF Microscope Image Sequences of Fluorescent IgE-FcεRI Receptor Complexes inside a FcεRI-Centric Synapse in RBL-2H3 Cells. Data, 4(3), 111. https://doi.org/10.3390/data4030111





