Transcriptome Dataset of Leaf Tissue in Agave H11648
Abstract
:1. Introduction
2. Results
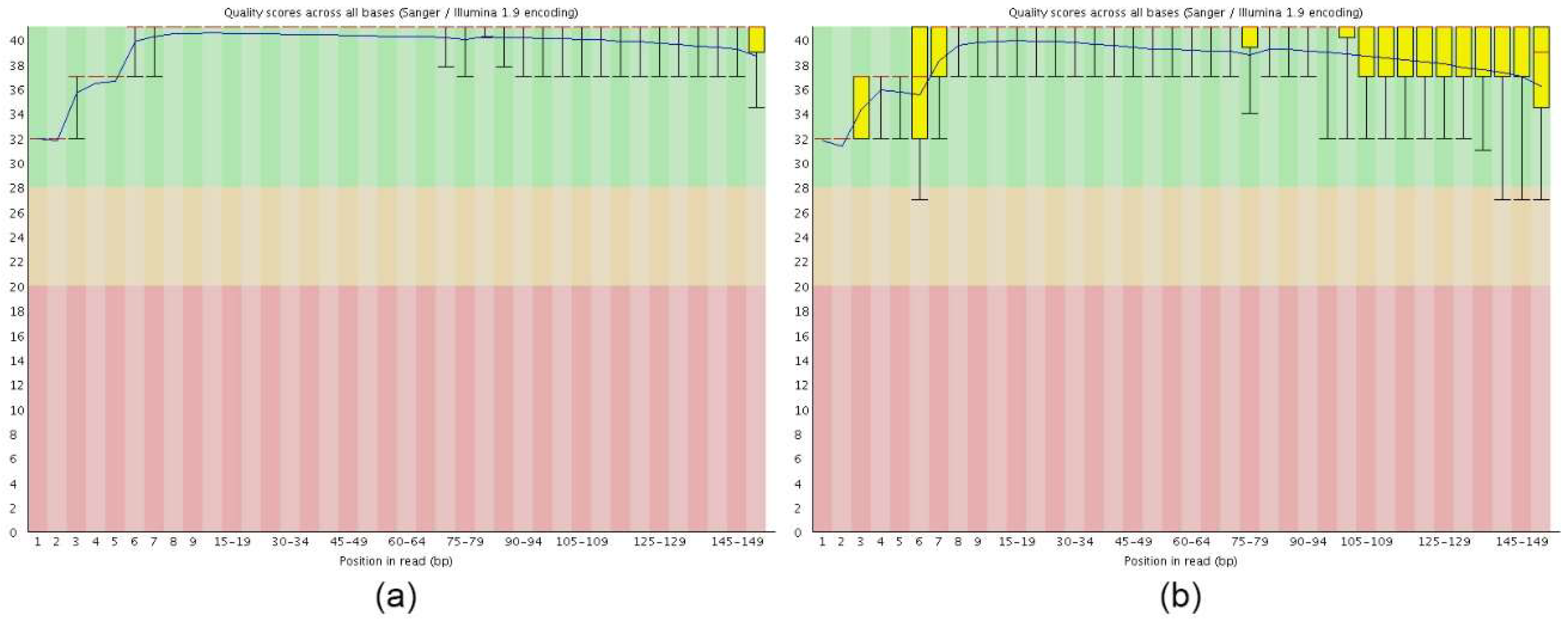
2.1. Illumina Sequencing and De Novo Assembly
2.2. Value of the Data
2.3. Data Records
3. Materials and Methods
3.1. Plant Material and RNA Isolation
3.2. Library Construction and Illumina Sequencing
3.3. Data Processing
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Li, Y.; Mai, Y.W.; Ye, L. Sisal fibre and its composites: A review of recent developments. Compos. Sci. Tech. 2000, 60, 2037–2055. [Google Scholar] [CrossRef]
- Food and Agriculture Organization of the United Nations (FAO). Available online: http://www.fao.org/faostat (accessed on 30 March 2019).
- Gao, J.; Yang, F.; Zhang, S.; Li, J.; Chen, H.; Liu, Q.; Zheng, J.; Xi, J.; Yi, K. Expression of a hevein-like gene in transgenic Agave hybrid No. 11648 enhances tolerance against zebra stripe disease. Plant Cell Tissue Organ Culture 2014, 119, 579–585. [Google Scholar] [CrossRef]
- Nava-Cruz, N.Y.; Medina-Morales, M.A.; Martinez, J.L.; Rodriguez, R.; Aguilar, C.N. Agave biotechnology: An overview. Crit. Rev. Biotechnol. 2015, 35, 546–559. [Google Scholar] [CrossRef] [PubMed]
- Robert, M.L.; Lim, K.Y.; Hanson, L.; Sanchez-Teyer, F.; Bennett, M.D.; Leitch, A.R.; Leitch, I.J. Wild and agronomically important Agave species (Asparagaceae) show proportional increases in chromosome number, genome size, and genetic markers with increasing ploidy. Bot. J. Linn. Soc. 2010, 158, 215–222. [Google Scholar] [CrossRef]
- Schuster, S.C. Next-generation sequencing transforms today’s biology. Nat. Methods 2008, 5, 16–18. [Google Scholar] [CrossRef] [PubMed]
- Gross, S.M.; Martin, J.A.; Simpson, J.; Abraham-Juarez, M.J.; Wang, Z.; Visel, A. De novo transcriptome assembly of drought tolerant CAM plants, Agave deserti and Agave tequilana. BMC Genom. 2013, 14, 563. [Google Scholar] [CrossRef]
- Abraham, P.E.; Yin, H.; Borland, A.M.; Weighill, D.; Lim, S.D.; De Paoli, H.C.; Engle, N.; Jones, P.C.; Agh, R.; Weston, D.J.; et al. Transcript, protein and metabolite temporal dynamics in the CAM plant Agave. Nat. Plants 2016, 2, 16178. [Google Scholar] [CrossRef] [PubMed]
- Cervantes-Pérez, S.A.; Espinal-Centeno, A.; Oropeza-Aburto, A.; Caballero-Pérez, J.; Falcon, F.; Aragón-Raygoza, A.; Sánchez-Segura, L.; Herrera-Estrella, L.; Cruz-Hernández, A.; Cruz-Ramírez, A. Transcriptional profiling of the CAM plant Agave salmiana reveals conservation of a genetic program for regeneration. Dev. Biol. 2018, 442, 28–39. [Google Scholar] [CrossRef]
- Sarwar, M.B.; Ahmad, Z.; Rashid, B.; Hassan, S.; Gregersen, P.L.; Leyva, M.O.; Nagy, I.; Asp, T.; Husnain, T. De novo assembly of Agave sisalana transcriptome in response to drought stress provides insight into the tolerance mechanisms. Sci. Rep. 2019, 9, 396. [Google Scholar]
- Huang, X.; Xiao, M.; Xi, J.; He, C.; Zheng, J.; Chen, H.; Gao, J.; Zhang, S.; Wu, W.; Liang, Y.; et al. De novo transcriptome assembly of Agave H11648 by Illumina sequencing and identification of cellulose synthase genes in Agave species. Genes 2019, 10, 103. [Google Scholar] [CrossRef]
- Huang, X.; Wang, B.; Xi, J.; Zhang, Y.; He, C.; Zheng, J.; Gao, J.; Chen, H.; Zhang, S.; Wu, W.; et al. Transcriptome comparison reveals distinct selection patterns in domesticated and wild Agave species, the important CAM plants. Int. J. Genom. 2018, 2018, 5716518. [Google Scholar] [CrossRef] [PubMed]
- Smith, M.L.; Glass, G.V. Meta-analysis of psychotherapy outcome studies. Am. Psychol. 1977, 32, 752–760. [Google Scholar] [CrossRef] [PubMed]
- Luo, X.; Xu, L.; Liang, D.; Wang, Y.; Zhang, W.; Zhu, X.; Zhu, Y.; Jiang, H.; Tang, M.; Liu, L. Comparative transcriptomics uncovers alternative splicing and molecular marker development in radish (Raphanus sativus L.). BMC Genom. 2017, 18, 505. [Google Scholar] [CrossRef] [PubMed]
- Harkess, A.; Zhou, J.; Xu, C.; Bowers, J.E.; Van der Hulst, R.; Ayyampalayam, S.; Mercati, F.; Riccardi, P.; McKain, M.R.; Kakrana, A.; et al. The asparagus genome sheds light on the origin and evolution of a young Y chromosome. Nat. Commun. 2017, 8, 1279. [Google Scholar] [CrossRef] [Green Version]
- Verta, J.P.; Landry, C.R.; MacKay, J. Dissection of expression-quantitative trait locus and allele specificity using a haploid/diploid plant system—insights into compensatory evolution of transcriptional regulation within populations. New Phytol. 2016, 211, 159–171. [Google Scholar] [CrossRef] [PubMed]
- Singh, N.K. miRNAs target databases: Developmental methods and target identification techniques with functional annotations. Cell Mol. Life Sci. 2017, 74, 2239–2261. [Google Scholar] [CrossRef] [PubMed]
- Huang, X.; Chen, J.; Bao, Y.; Liu, L.; Jiang, H.; An, X.; Dai, L.; Wang, B.; Peng, D. Transcript profiling reveals auxin and cytokinin signaling pathways and transcription regulation during in vitro organogenesis of ramie (Boehmeria nivea L. Gaud). PLoS ONE 2014, 9, e113768. [Google Scholar] [CrossRef] [PubMed]
- FastQC. Available online: http://www.bioinformatics.babraham.ac.uk/projects/fastqc/ (accessed on 30 March 2019).
- Martin, M. Cutadapt removes adapter sequences from high-throughput sequencing reads. EMBnet J. 2011, 17, 10–12. [Google Scholar] [CrossRef]
- Bolger, A.M.; Lohse, M.; Usadel, B. Trimmomatic: A flexible trimmer for Illumina sequence data. Bioinformatics 2014, 30, 2114–2120. [Google Scholar] [CrossRef] [PubMed]
- Grabherr, M.G.; Haas, B.J.; Yassour, M.; Levin, J.Z.; Thompson, D.A.; Amit, I.; Adiconis, X.; Fan, L.; Raychowdhury, R.; Zeng, Q.; et al. Full-length transcriptome assembly from RNA-Seq data without a reference genome. Nat. Biotechnol. 2011, 29, 644–652. [Google Scholar] [CrossRef]
- Li, B.; Dewey, C.N. RSEM: Accurate transcript quantification from RNA-Seq data with or without a reference genome. BMC Bioinform. 2011, 12, 323. [Google Scholar] [CrossRef] [PubMed]
- Mortazavi, A.; Williams, B.A.; Mccue, K.; Schaeffer, L.; Wold, B. Mapping and quantifying mammalian transcriptomes by RNA-Seq. Nat. Methods 2008, 5, 621–628. [Google Scholar] [CrossRef] [PubMed]


© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Huang, X.; Xie, L.; Gbokie, T., Jr.; Xi, J.; Yi, K. Transcriptome Dataset of Leaf Tissue in Agave H11648. Data 2019, 4, 62. https://doi.org/10.3390/data4020062
Huang X, Xie L, Gbokie T Jr., Xi J, Yi K. Transcriptome Dataset of Leaf Tissue in Agave H11648. Data. 2019; 4(2):62. https://doi.org/10.3390/data4020062
Chicago/Turabian StyleHuang, Xing, Li Xie, Thomas Gbokie, Jr., Jingen Xi, and Kexian Yi. 2019. "Transcriptome Dataset of Leaf Tissue in Agave H11648" Data 4, no. 2: 62. https://doi.org/10.3390/data4020062
APA StyleHuang, X., Xie, L., Gbokie, T., Jr., Xi, J., & Yi, K. (2019). Transcriptome Dataset of Leaf Tissue in Agave H11648. Data, 4(2), 62. https://doi.org/10.3390/data4020062





