Review on Facial-Recognition-Based Applications in Disease Diagnosis
Abstract
:1. Introduction
2. The Facial Recognition System: Approaches and Algorithms
2.1. Image Capture
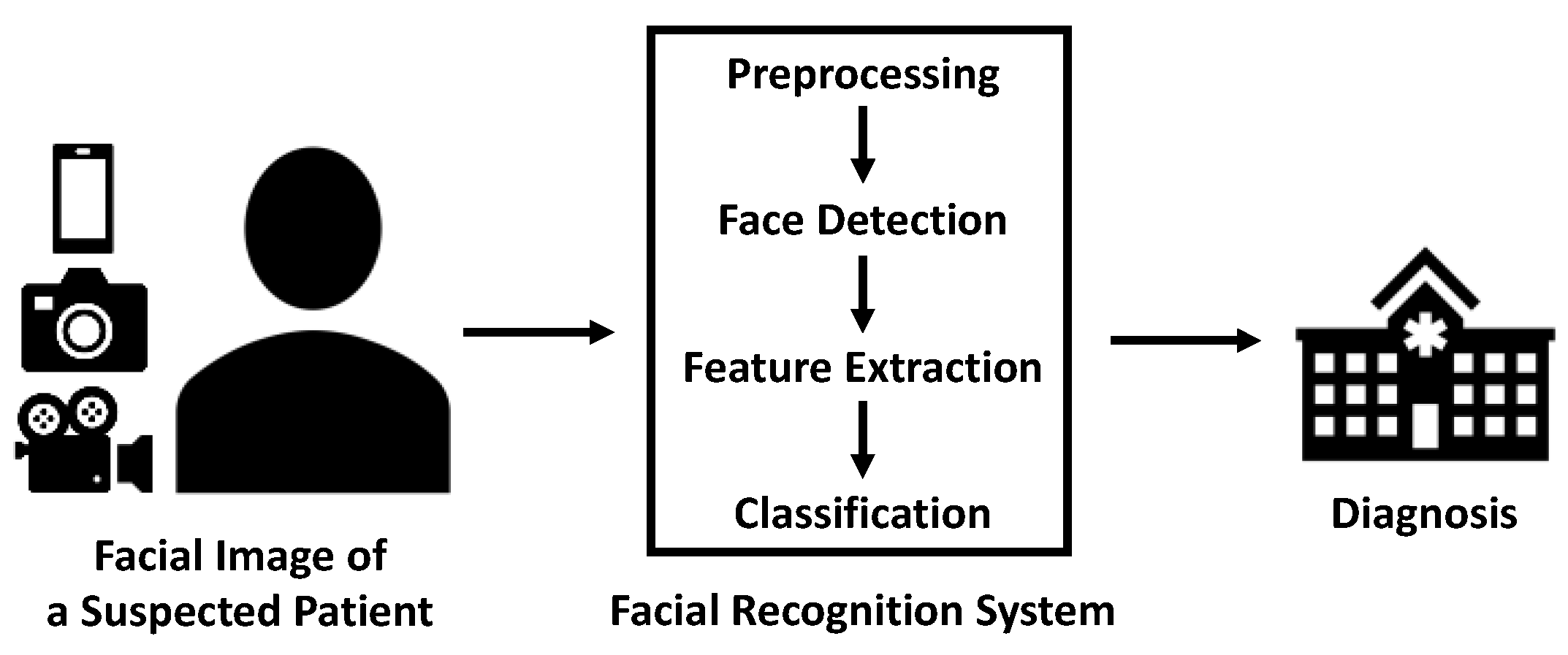
2.2. Workflow of Facial Recognition Technology
2.3. Facial Analysis Algorithms
2.3.1. Traditional Methods
2.3.2. Deep Learning Methods
2.3.3. Mature Software
3. The Facial Recognition System: Applications and Advantages
3.1. Performance in Varieties of Disease
3.1.1. Endocrine and Metabolic Diseases
3.1.2. Genetic and Chromosome Abnormalities
3.1.3. Neuromuscular Diseases
3.1.4. Other Types of Disease
3.2. Clinical Applications
3.3. Advantages over Traditional Methods
3.3.1. Accurate and Objective
3.3.2. Comprehensive and Informative
3.3.3. Improvement of Healthcare System
4. Future Outlook
4.1. Expansion of Database Volume
4.2. Factors Affecting Diagnostic Accuracy
4.3. Integration of Novel Technology
4.4. Applications beyond Diagnosis
4.5. From Research to Products
4.6. Privacy and Security
5. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Kaur, P.; Krishan, K.; Sharma, S.K.; Kanchan, T. Facial-Recognition Algorithms: A Literature Review. Med. Sci. Law 2020, 60, 131–139. [Google Scholar] [CrossRef]
- Fontaine, X.; Achanta, R.; Süsstrunk, S. Face Recognition in Real-World Images. In Proceedings of the 2017 IEEE International Conference on Acoustics, Speech and Signal Processing (ICASSP), New Orleans, LA, USA, 5–9 March 2017; pp. 1482–1486. [Google Scholar] [CrossRef] [Green Version]
- Kosilek, R.P.; Frohner, R.; Würtz, R.P.; Berr, C.M.; Schopohl, J.; Reincke, M.; Schneider, H.J. Diagnostic Use of Facial Image Analysis Software in Endocrine and Genetic Disorders: Review, Current Results and Future Perspectives. Eur. J. Endocrinol. 2015, 173, M39–M44. [Google Scholar] [CrossRef] [Green Version]
- Gurovich, Y.; Hanani, Y.; Bar, O.; Nadav, G.; Fleischer, N.; Gelbman, D.; Basel-Salmon, L.; Krawitz, P.M.; Kamphausen, S.B.; Zenker, M.; et al. Identifying Facial Phenotypes of Genetic Disorders Using Deep Learning. Nat. Med. 2019, 25, 60–64. [Google Scholar] [CrossRef]
- Ali, M.R.; Myers, T.; Wagner, E.; Ratnu, H.; Dorsey, E.R.; Hoque, E. Facial Expressions Can Detect Parkinson’s Disease: Preliminary Evidence from Videos Collected Online. NPJ Digit. Med. 2021, 4, 1–4. [Google Scholar] [CrossRef]
- Loos, H.S.; Wieczorek, D.; Würtz, R.P.; von der Malsburg, C.; Horsthemke, B. Computer-Based Recognition of Dysmorphic Faces. Eur. J. Hum. Genet. 2003, 11, 555–560. [Google Scholar] [CrossRef] [Green Version]
- Meintjes, E.M.; Douglas, T.S.; Martinez, F.; Vaughan, C.L.; Adams, L.P.; Stekhoven, A.; Viljoen, D. A Stereo-Photogrammetric Method to Measure the Facial Dysmorphology of Children in the Diagnosis of Fetal Alcohol Syndrome. Med. Eng. Phys. 2002, 24, 683–689. [Google Scholar] [CrossRef]
- Wachtman, G.S.; Cohn, J.F.; VanSwearingen, J.M.; Manders, E.K. Automated Tracking of Facial Features in Patients with Facial Neuromuscular Dysfunction. Plast. Reconstr. Surg. 2001, 107, 1124–1133. [Google Scholar] [CrossRef]
- Rajpurkar, P.; Chen, E.; Banerjee, O.; Topol, E.J. AI in Health and Medicine. Nat. Med. 2022, 28, 31–38. [Google Scholar] [CrossRef]
- Yu, K.-H.; Beam, A.L.; Kohane, I.S. Artificial Intelligence in Healthcare. Nat. Biomed. Eng. 2018, 2, 719–731. [Google Scholar] [CrossRef]
- Hallgrímsson, B.; Aponte, J.D.; Katz, D.C.; Bannister, J.J.; Riccardi, S.L.; Mahasuwan, N.; McInnes, B.L.; Ferrara, T.M.; Lipman, D.M.; Neves, A.B.; et al. Automated Syndrome Diagnosis by Three-Dimensional Facial Imaging. Genet. Med. 2020, 22, 1682–1693. [Google Scholar] [CrossRef]
- Meng, T.; Guo, X.; Lian, W.; Deng, K.; Gao, L.; Wang, Z.; Huang, J.; Wang, X.; Long, X.; Xing, B. Identifying Facial Features and Predicting Patients of Acromegaly Using Three-Dimensional Imaging Techniques and Machine Learning. Front. Endocrinol. 2020, 11, 492. [Google Scholar] [CrossRef]
- Ridha, A.M.; Shehieb, W.; Yacoub, P.; Al-Balawneh, K.; Arshad, K. Smart Prediction System for Facial Paralysis. In Proceedings of the 2020 7th International Conference on Electrical and Electronics Engineering (ICEEE), Antalya, Turkey, 14–16 April 2020; pp. 321–327. [Google Scholar]
- Anwarul, S.; Dahiya, S. A Comprehensive Review on Face Recognition Methods and Factors Affecting Facial Recognition Accuracy. In Proceedings of the ICRIC 2019, Recent Innovations in Computing, Jammu, India, 8–9 March 2019; Springer: Berlin/Heidelberg, Germany, 2020; pp. 495–514. [Google Scholar] [CrossRef]
- Gao, X.; Li, S.Z.; Liu, R.; Zhang, P. Standardization of Face Image Sample Quality. In Proceedings of the International Conference on Biometrics, Seoul, Korea, 27–29 August 2007; pp. 242–251. [Google Scholar] [CrossRef] [Green Version]
- Song, W.; Lei, Y.; Chen, S.; Pan, Z.; Yang, J.-J.; Pan, H.; Du, X.; Cai, W.; Wang, Q. Multiple Facial Image Features-Based Recognition for the Automatic Diagnosis of Turner Syndrome. Comput. Ind. 2018, 100, 85–95. [Google Scholar] [CrossRef]
- Hong, D.; Zheng, Y.-Y.; Xin, Y.; Sun, L.; Yang, H.; Lin, M.-Y.; Liu, C.; Li, B.-N.; Zhang, Z.-W.; Zhuang, J.; et al. Genetic Syndromes Screening by Facial Recognition Technology: VGG-16 Screening Model Construction and Evaluation. Orphanet. J. Rare Dis. 2021, 16, 344. [Google Scholar] [CrossRef]
- Bandini, A.; Rezaei, S.; Guarin, D.L.; Kulkarni, M.; Lim, D.; Boulos, M.I.; Zinman, L.; Yunusova, Y.; Taati, B. A New Dataset for Facial Motion Analysis in Individuals With Neurological Disorders. IEEE J. Biomed. Health Inform. 2021, 25, 1111–1119. [Google Scholar] [CrossRef]
- Jayaraman, U.; Gupta, P.; Gupta, S.; Arora, G.; Tiwari, K. Recent Development in Face Recognition. Neurocomputing 2020, 408, 231–245. [Google Scholar] [CrossRef]
- Banerjee, D.; Yu, K. Robotic Arm-Based Face Recognition Software Test Automation. IEEE Access 2018, 6, 37858–37868. [Google Scholar] [CrossRef]
- AbdAlmageed, W.; Mirzaalian, H.; Guo, X.; Randolph, L.M.; Tanawattanacharoen, V.K.; Geffner, M.E.; Ross, H.M.; Kim, M.S. Assessment of Facial Morphologic Features in Patients With Congenital Adrenal Hyperplasia Using Deep Learning. JAMA Netw. Open 2020, 3, e2022199. [Google Scholar] [CrossRef]
- Ji, S.; Xu, W.; Yang, M.; Yu, K. 3D Convolutional Neural Networks for Human Action Recognition. IEEE Trans. Pattern Anal. Mach. Intell. 2013, 35, 221–231. [Google Scholar] [CrossRef] [Green Version]
- Jin, B.; Qu, Y.; Zhang, L.; Gao, Z. Diagnosing Parkinson Disease Through Facial Expression Recognition: Video Analysis. J. Med. Internet Res. 2020, 22, e18697. [Google Scholar] [CrossRef]
- Baltrušaitis, T.; Zadeh, A.; Lim, Y.; Morency, L.-P. OpenFace 2.0: Facial Behavior Analysis Toolkit. In Proceedings of the 13th IEEE International Conference on Automatic Face & Gesture Recognition (FG 2018), Xi’an, China, 15–19 May 2018. [Google Scholar] [CrossRef]
- Nam, U.; Lee, K.; Ko, H.; Lee, J.-Y.; Lee, E.C. Analyzing Facial and Eye Movements to Screen for Alzheimer’s Disease. Sensors 2020, 20, 5349. [Google Scholar] [CrossRef]
- Colao, A.; Grasso, L.F.S.; Giustina, A.; Melmed, S.; Chanson, P.; Pereira, A.M.; Pivonello, R. Acromegaly. Nat. Rev. Dis Primers 2019, 5, 20. [Google Scholar] [CrossRef] [PubMed]
- Loriaux, D.L. Diagnosis and Differential Diagnosis of Cushing’s Syndrome. N. Engl. J. Med. 2017, 376, 1451–1459. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Learned-Miller, E.; Lu, Q.; Paisley, A.; Trainer, P.; Blanz, V.; Dedden, K.; Miller, R. Detecting Acromegaly: Screening for Disease with a Morphable Model. Med. Image Comput. Comput. Assist. Interv. 2006, 9, 495–503. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Frohner, R.; Würtz, R.P.; Kosilek, R.; Schneider, H.J. Optimierung Der Gesichtsklassifikation Bei Der Erkennung von Akromegalie. J. Klin. Endokrinol. Stoffw. 2013, 6, 20–24. [Google Scholar]
- Gencturk, B.; Nabiyev, V.V.; Ustubioglu, A.; Ketenci, S. Automated Pre-Diagnosis of Acromegaly Disease Using Local Binary Patterns and Its Variants. In Proceedings of the 36th International Conference on Telecommunications and Signal Processing (TSP), Rome, Italy, 2–4 July 2013; pp. 817–821. [Google Scholar] [CrossRef]
- Kong, X.; Gong, S.; Su, L.; Howard, N.; Kong, Y. Automatic Detection of Acromegaly From Facial Photographs Using Machine Learning Methods. EBioMedicine 2018, 27, 94–102. [Google Scholar] [CrossRef] [Green Version]
- Wei, R.; Jiang, C.; Gao, J.; Xu, P.; Zhang, D.; Sun, Z.; Liu, X.; Deng, K.; Bao, X.; Sun, G.; et al. Deep-Learning Approach to Automatic Identification of Facial Anomalies in Endocrine Disorders. Neuroendocrinology 2020, 110, 328–337. [Google Scholar] [CrossRef]
- Popp, K.H.; Kosilek, R.P.; Frohner, R.; Stalla, G.K.; Athanasoulia-Kaspar, A.; Berr, C.; Zopp, S.; Reincke, M.; Witt, M.; Würtz, R.P.; et al. Computer Vision Technology in the Differential Diagnosis of Cushing’s Syndrome. Exp. Clin. Endocrinol. Diabetes 2019, 127, 685–690. [Google Scholar] [CrossRef] [Green Version]
- Kosilek, R.P.; Schopohl, J.; Grunke, M.; Reincke, M.; Dimopoulou, C.; Stalla, G.K.; Würtz, R.P.; Lammert, A.; Günther, M.; Schneider, H.J. Automatic Face Classification of Cushing’s Syndrome in Women—A Novel Screening Approach. Exp. Clin. Endocrinol. Diabetes 2013, 121, 561–564. [Google Scholar] [CrossRef] [Green Version]
- Bull, M.J. Down Syndrome. N. Engl. J. Med. 2020, 382, 2344–2352. [Google Scholar] [CrossRef]
- Zhao, Q.; Okada, K.; Rosenbaum, K.; Kehoe, L.; Zand, D.J.; Sze, R.; Summar, M.; Linguraru, M.G. Digital Facial Dysmorphology for Genetic Screening: Hierarchical Constrained Local Model Using ICA. Med. Image Anal. 2014, 18, 699–710. [Google Scholar] [CrossRef]
- Burçin, K.; Vasif, N.V. Down Syndrome Recognition Using Local Binary Patterns and Statistical Evaluation of the System. Expert Syst. Appl. 2011, 38, 8690–8695. [Google Scholar] [CrossRef]
- Saraydemir, S.; Taşpınar, N.; Eroğul, O.; Kayserili, H.; Dinçkan, N. Down Syndrome Diagnosis Based on Gabor Wavelet Transform. J. Med. Syst. 2012, 36, 3205–3213. [Google Scholar] [CrossRef] [PubMed]
- Zhao, Q.; Rosenbaum, K.; Okada, K.; Zand, D.J.; Sze, R.; Summar, M.; Linguraru, M.G. Automated Down Syndrome Detection Using Facial Photographs. Annu Int Conf IEEE Eng. Med. Biol Soc. 2013, 2013, 3670–3673. [Google Scholar] [CrossRef]
- Qin, B.; Liang, L.; Wu, J.; Quan, Q.; Wang, Z.; Li, D. Automatic Identification of Down Syndrome Using Facial Images with Deep Convolutional Neural Network. Diagnostics 2020, 10, 487. [Google Scholar] [CrossRef]
- Srisraluang, W.; Rojnueangnit, K. Facial Recognition Accuracy in Photographs of Thai Neonates with Down Syndrome among Physicians and the Face2Gene Application. Am. J. Med. Genet. A 2021, 185, 3701–3705. [Google Scholar] [CrossRef] [PubMed]
- Vorravanpreecha, N.; Lertboonnum, T.; Rodjanadit, R.; Sriplienchan, P.; Rojnueangnit, K. Studying Down Syndrome Recognition Probabilities in Thai Children with De-Identified Computer-Aided Facial Analysis. Am. J. Med. Genet. A 2018, 176, 1935–1940. [Google Scholar] [CrossRef]
- Porras, A.R.; Bramble, M.S.; Mosema Be Amoti, K.; Spencer, D.; Dakande, C.; Manya, H.; Vashist, N.; Likuba, E.; Ebwel, J.M.; Musasa, C.; et al. Facial Analysis Technology for the Detection of Down Syndrome in the Democratic Republic of the Congo. Eur. J. Med. Genet. 2021, 64, 104267. [Google Scholar] [CrossRef]
- Gravholt, C.H.; Viuff, M.H.; Brun, S.; Stochholm, K.; Andersen, N.H. Turner Syndrome: Mechanisms and Management. Nat. Rev. Endocrinol. 2019, 15, 601–614. [Google Scholar] [CrossRef]
- Pan, Z.; Shen, Z.; Zhu, H.; Bao, Y.; Liang, S.; Wang, S.; Li, X.; Niu, L.; Dong, X.; Shang, X.; et al. Clinical Application of an Automatic Facial Recognition System Based on Deep Learning for Diagnosis of Turner Syndrome. Endocrine 2021, 72, 865–873. [Google Scholar] [CrossRef]
- Basel-Vanagaite, L.; Wolf, L.; Orin, M.; Larizza, L.; Gervasini, C.; Krantz, I.; Deardoff, M. Recognition of the Cornelia de Lange Syndrome Phenotype with Facial Dysmorphology Novel Analysis. Clin. Genet. 2016, 89, 557–563. [Google Scholar] [CrossRef]
- Latorre-Pellicer, A.; Ascaso, Á.; Trujillano, L.; Gil-Salvador, M.; Arnedo, M.; Lucia-Campos, C.; Antoñanzas-Pérez, R.; Marcos-Alcalde, I.; Parenti, I.; Bueno-Lozano, G.; et al. Evaluating Face2Gene as a Tool to Identify Cornelia de Lange Syndrome by Facial Phenotypes. Int. J. Mol. Sci. 2020, 21, 1042. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Hadj-Rabia, S.; Schneider, H.; Navarro, E.; Klein, O.; Kirby, N.; Huttner, K.; Wolf, L.; Orin, M.; Wohlfart, S.; Bodemer, C.; et al. Automatic Recognition of the XLHED Phenotype from Facial Images. Am. J. Med. Genet. A 2017, 173, 2408–2414. [Google Scholar] [CrossRef] [PubMed]
- Liehr, T.; Acquarola, N.; Pyle, K.; St-Pierre, S.; Rinholm, M.; Bar, O.; Wilhelm, K.; Schreyer, I. Next Generation Phenotyping in Emanuel and Pallister-Killian Syndrome Using Computer-Aided Facial Dysmorphology Analysis of 2D Photos. Clin. Genet. 2018, 93, 378–381. [Google Scholar] [CrossRef] [PubMed]
- Amudhavalli, S.M.; Hanson, R.; Angle, B.; Bontempo, K.; Gripp, K.W. Further Delineation of Aymé-Gripp Syndrome and Use of Automated Facial Analysis Tool. Am. J. Med. Genet. A 2018, 176, 1648–1656. [Google Scholar] [CrossRef]
- Pode-Shakked, B.; Finezilber, Y.; Levi, Y.; Liber, S.; Fleischer, N.; Greenbaum, L.; Raas-Rothschild, A. Shared Facial Phenotype of Patients with Mucolipidosis Type IV: A Clinical Observation Reaffirmed by next Generation Phenotyping. Eur. J. Med. Genet. 2020, 63, 103927. [Google Scholar] [CrossRef]
- Wang, Y.; Li, N.; Su, Z.; Xu, Y.; Liu, S.; Chen, Y.; Li, X.; Shen, Y.; Hung, C.; Wang, J.; et al. The Phenotypic Spectrum of Kabuki Syndrome in Patients of Chinese Descent: A Case Series. Am. J. Med. Genet. A 2020, 182, 640–651. [Google Scholar] [CrossRef]
- Porras, A.R.; Summar, M.; Linguraru, M.G. Objective Differential Diagnosis of Noonan and Williams-Beuren Syndromes in Diverse Populations Using Quantitative Facial Phenotyping. Mol. Genet. Genom. Med. 2021, 9, e1636. [Google Scholar] [CrossRef]
- Gautam, R.; Sharma, M. Prevalence and Diagnosis of Neurological Disorders Using Different Deep Learning Techniques: A Meta-Analysis. J. Med. Syst. 2020, 44, 49. [Google Scholar] [CrossRef]
- Wang, B.; Wu, X. Application of Artificial Intelligence in Recognition and Evaluation of Facial Paralysis. Chin. J. Med. Instrum. 2022, 46, 57–62. [Google Scholar] [CrossRef]
- Ngo, T.H.; Seo, M.; Matsushiro, N.; Chen, Y.-W. Quantitative Analysis of Facial Paralysis Based on Filters of Concentric Modulation. In Proceedings of the 2015 12th International Conference on Fuzzy Systems and Knowledge Discovery (FSKD), Zhangjiajie, China, 15–17 August 2015; pp. 1758–1763. [Google Scholar]
- Guo, Z.; Shen, M.; Duan, L.; Zhou, Y.; Xiang, J.; Ding, H.; Chen, S.; Deussen, O.; Dan, G. Deep Assessment Process: Objective Assessment Process for Unilateral Peripheral Facial Paralysis via Deep Convolutional Neural Network. In Proceedings of the 2017 IEEE 14th International Symposium on Biomedical Imaging (ISBI 2017), Melbourne, Australia, 18–21 April 2017; pp. 135–138. [Google Scholar]
- Storey, G.; Jiang, R.; Keogh, S.; Bouridane, A.; Li, C.-T. 3DPalsyNet: A Facial Palsy Grading and Motion Recognition Framework Using Fully 3D Convolutional Neural Networks. arXiv 2019, arXiv:1905.13607. [Google Scholar] [CrossRef]
- Tăuţan, A.-M.; Ionescu, B.; Santarnecchi, E. Artificial Intelligence in Neurodegenerative Diseases: A Review of Available Tools with a Focus on Machine Learning Techniques. Artif. Intell. Med. 2021, 117, 102081. [Google Scholar] [CrossRef] [PubMed]
- Bandini, A.; Orlandi, S.; Escalante, H.J.; Giovannelli, F.; Cincotta, M.; Reyes-Garcia, C.A.; Vanni, P.; Zaccara, G.; Manfredi, C. Analysis of Facial Expressions in Parkinson’s Disease through Video-Based Automatic Methods. J. Neurosci. Methods 2017, 281, 7–20. [Google Scholar] [CrossRef] [PubMed]
- Rajnoha, M.; Mekyska, J.; Burget, R.; Eliasova, I.; Kostalova, M.; Rektorova, I. Towards Identification of Hypomimia in Parkinson’s Disease Based on Face Recognition Methods. In Proceedings of the 2018 10th International Congress on Ultra Modern Telecommunications and Control Systems and.d Workshops (ICUMT), Moscow, Russia, 5–9 November 2018; pp. 1–4. [Google Scholar]
- Jung, S.-G.; An, J.; Kwak, H.; Salminen, J.; Jansen, B. Inferring Social Media Users’ Demographics from Profile Pictures: A Face++ Analysis on Twitter Users. In Proceedings of the International Conference on Electronic Business (ICEB), Dubai, United Arab Emirates, 4–8 December 2017. [Google Scholar]
- Hou, X.; Zhang, Y.; Wang, Y.; Wang, X.; Zhao, J.; Zhu, X.; Su, J. A Markerless 2D Video, Facial Feature Recognition-Based, Artificial Intelligence Model to Assist With Screening for Parkinson Disease: Development and Usability Study. J. Med. Internet Res. 2021, 23, e29554. [Google Scholar] [CrossRef] [PubMed]
- Umeda-Kameyama, Y.; Kameyama, M.; Tanaka, T.; Son, B.-K.; Kojima, T.; Fukasawa, M.; Iizuka, T.; Ogawa, S.; Iijima, K.; Akishita, M. Screening of Alzheimer’s Disease by Facial Complexion Using Artificial Intelligence. Aging 2021, 13, 1765–1772. [Google Scholar] [CrossRef] [PubMed]
- Zhuang, Y.; McDonald, M.; Uribe, O.; Yin, X.; Parikh, D.; Southerland, A.M.; Rohde, G.K. Facial Weakness Analysis and Quantification of Static Images. IEEE J. Biomed. Health Inform. 2020, 24, 2260–2267. [Google Scholar] [CrossRef]
- Forte, C.; Voinea, A.; Chichirau, M.; Yeshmagambetova, G.; Albrecht, L.M.; Erfurt, C.; Freundt, L.A.; Carmo, L.O.E.; Henning, R.H.; van der Horst, I.C.C.; et al. Deep Learning for Identification of Acute Illness and Facial Cues of Illness. Front. Med. 2021, 8, 661309. [Google Scholar] [CrossRef]
- Lin, S.; Li, Z.; Fu, B.; Chen, S.; Li, X.; Wang, Y.; Wang, X.; Lv, B.; Xu, B.; Song, X.; et al. Feasibility of Using Deep Learning to Detect Coronary Artery Disease Based on Facial Photo. Eur. Heart J. 2020, 41, 4400–4411. [Google Scholar] [CrossRef]
- Valentine, M.; Bihm, D.C.J.; Wolf, L.; Hoyme, H.E.; May, P.A.; Buckley, D.; Kalberg, W.; Abdul-Rahman, O.A. Computer-Aided Recognition of Facial Attributes for Fetal Alcohol Spectrum Disorders. Pediatrics 2017, 140, e20162028. [Google Scholar] [CrossRef]
- Chen, Y.; Liu, W.; Zhang, L.; Yan, M.; Zeng, Y. Hybrid Facial Image Feature Extraction and Recognition for Non-Invasive Chronic Fatigue Syndrome Diagnosis. Comput. Biol. Med. 2015, 64, 30–39. [Google Scholar] [CrossRef]
- Miller, M.Q.; Hadlock, T.A.; Fortier, E.; Guarin, D.L. The Auto-eFACE: Machine Learning-Enhanced Program Yields Automated Facial Palsy Assessment Tool. Plast. Reconstr. Surg. 2021, 147, 467–474. [Google Scholar] [CrossRef]
- Kruszka, P.; Addissie, Y.A.; McGinn, D.E.; Porras, A.R.; Biggs, E.; Share, M.; Crowley, T.B.; Chung, B.H.Y.; Mok, G.T.K.; Mak, C.C.Y.; et al. 22q11.2 Deletion Syndrome in Diverse Populations. Am. J. Med. Genet. A 2017, 173, 879–888. [Google Scholar] [CrossRef] [PubMed]
- Miller, R.E.; Learned-Miller, E.G.; Trainer, P.; Paisley, A.; Blanz, V. Early Diagnosis of Acromegaly: Computers vs Clinicians. Clin. Endocrinol. 2011, 75, 226–231. [Google Scholar] [CrossRef] [PubMed]
- Schneider, H.J.; Kosilek, R.P.; Günther, M.; Roemmler, J.; Stalla, G.K.; Sievers, C.; Reincke, M.; Schopohl, J.; Würtz, R.P. A Novel Approach to the Detection of Acromegaly: Accuracy of Diagnosis by Automatic Face Classification. J. Clin. Endocrinol. Metab. 2011, 96, 2074–2080. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Chen, S.; Pan, Z.-X.; Zhu, H.-J.; Wang, Q.; Yang, J.-J.; Lei, Y.; Li, J.-Q.; Pan, H. Development of a Computer-Aided Tool for the Pattern Recognition of Facial Features in Diagnosing Turner Syndrome: Comparison of Diagnostic Accuracy with Clinical Workers. Sci. Rep. 2018, 8, 9317. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Mishima, H.; Suzuki, H.; Doi, M.; Miyazaki, M.; Watanabe, S.; Matsumoto, T.; Morifuji, K.; Moriuchi, H.; Yoshiura, K.-I.; Kondoh, T.; et al. Evaluation of Face2Gene Using Facial Images of Patients with Congenital Dysmorphic Syndromes Recruited in Japan. J. Hum. Genet. 2019, 64, 789–794. [Google Scholar] [CrossRef]
- Porras, A.R.; Rosenbaum, K.; Tor-Diez, C.; Summar, M.; Linguraru, M.G. Development and Evaluation of a Machine Learning-Based Point-of-Care Screening Tool for Genetic Syndromes in Children: A Multinational Retrospective Study. Lancet Digit. Health 2021, 3, e635–e643. [Google Scholar] [CrossRef]
- Guo, X.; Meng, T.; Huang, J.; Wang, X.; Lian, W.; Deng, K.; Gao, L.; Wang, Z.; Xing, B.; Long, X. 3D Facial Analysis in Acromegaly: Gender-Specific Features and Clinical Correlations. Front. Endocrinol. 2018, 9, 722. [Google Scholar] [CrossRef]
- Fan, Y.; Li, Y.; Li, Y.; Feng, S.; Bao, X.; Feng, M.; Wang, R. Development and Assessment of Machine Learning Algorithms for Predicting Remission after Transsphenoidal Surgery among Patients with Acromegaly. Endocrine 2020, 67, 412–422. [Google Scholar] [CrossRef]
- Topol, E.J. High-Performance Medicine: The Convergence of Human and Artificial Intelligence. Nat. Med. 2019, 25, 44–56. [Google Scholar] [CrossRef]
- Klare, B.F.; Burge, M.J.; Klontz, J.C.; Vorder Bruegge, R.W.; Jain, A.K. Face Recognition Performance: Role of Demographic Information. IEEE Trans. Inf. Forensics Secur. 2012, 7, 1789–1801. [Google Scholar] [CrossRef] [Green Version]
- Langevin, R.; Ali, M.R.; Sen, T.; Snyder, C.; Myers, T.; Dorsey, E.R.; Hoque, M.E. The PARK Framework for Automated Analysis of Parkinson’s Disease Characteristics. Proc. ACM Interact. Mob. Wearable Ubiquitous Technol. 2019, 3, 1–22. [Google Scholar] [CrossRef]
- Pantel, J.T.; Zhao, M.; Mensah, M.A.; Hajjir, N.; Hsieh, T.-C.; Hanani, Y.; Fleischer, N.; Kamphans, T.; Mundlos, S.; Gurovich, Y.; et al. Advances in Computer-Assisted Syndrome Recognition and Differentiation in a Set of Metabolic Disorders. bioRxiv 2017, 219394. [Google Scholar]
- Wu, D.; Chen, S.; Zhang, Y.; Zhang, H.; Wang, Q.; Li, J.; Fu, Y.; Wang, S.; Yang, H.; Du, H.; et al. Facial Recognition Intensity in Disease Diagnosis Using Automatic Facial Recognition. J. Pers. Med. 2021, 11, 1172. [Google Scholar] [CrossRef] [PubMed]
- Face Live Detection Method Based on Physiological Motion Analysis. Tsinghua Sci. Technol. 2009, 14, 685–690. [CrossRef]
- Guanjie Liu, Y.W.; Guanjie Liu, Y.W. A Computer-Aided System for Ocular Myasthenia Gravis Diagnosis. Tsinghua Sci. Technol. 2021, 26, 749–758. [Google Scholar] [CrossRef]
- Arac, A. Machine Learning for 3D Kinematic Analysis of Movements in Neurorehabilitation. Curr. Neurol. Neurosci. Rep. 2020, 20, 29. [Google Scholar] [CrossRef]
- Khashman, A. Application of an Emotional Neural Network to Facial Recognition. Neural Comput. Appl. 2009, 18, 309–320. [Google Scholar] [CrossRef]
- Hassan, T.; Seus, D.; Wollenberg, J.; Weitz, K.; Kunz, M.; Lautenbacher, S.; Garbas, J.-U.; Schmid, U. Automatic Detection of Pain from Facial Expressions: A Survey. IEEE Trans. Pattern. Anal. Mach. Intell. 2021, 43, 1815–1831. [Google Scholar] [CrossRef] [Green Version]
- Zhao, X.; Zou, J.; Li, H.; Dellandrea, E.; Kakadiaris, I.A.; Chen, L. Automatic 2.5-D Facial Landmarking and Emotion Annotation for Social Interaction Assistance. IEEE Trans. Cybern 2016, 46, 2042–2055. [Google Scholar] [CrossRef]
- Boonipat, T.; Asaad, M.; Lin, J.; Glass, G.E.; Mardini, S.; Stotland, M. Using Artificial Intelligence to Measure Facial Expression Following Facial Reanimation Surgery. Plast. Reconstr. Surg. 2020, 146, 1147–1150. [Google Scholar] [CrossRef]
- Vabalas, A.; Gowen, E.; Poliakoff, E.; Casson, A.J. Machine Learning Algorithm Validation with a Limited Sample Size. PLoS ONE 2019, 14, e0224365. [Google Scholar] [CrossRef] [PubMed]
- Niazi, M.K.K.; Parwani, A.V.; Gurcan, M.N. Digital Pathology and Artificial Intelligence. Lancet Oncol. 2019, 20, e253–e261. [Google Scholar] [CrossRef]
- Hosny, A.; Parmar, C.; Quackenbush, J.; Schwartz, L.H.; Aerts, H.J.W.L. Artificial Intelligence in Radiology. Nat. Rev. Cancer 2018, 18, 500–510. [Google Scholar] [CrossRef] [PubMed]
- U.S. Food and Drug Administration Artificial Intelligence/Machine Learning (AI/ML)-Based Software as a Medical Device (SaMD) Action Plan. 2021. Available online: https://www.fda.gov (accessed on 20 May 2022).
- Katsanis, S.H.; Claes, P.; Doerr, M.; Cook-Deegan, R.; Tenenbaum, J.D.; Evans, B.J.; Lee, M.K.; Anderton, J.; Weinberg, S.M.; Wagner, J.K. A Survey of U.S. Public Perspectives on Facial Recognition Technology and Facial Imaging Data Practices in Health and Research Contexts. PLoS ONE 2021, 16, e0257923. [Google Scholar] [CrossRef]
- Smith, M.; Miller, S. The Ethical Application of Biometric Facial Recognition Technology. AI Soc. 2022, 37, 167–175. [Google Scholar] [CrossRef]
- Almeida, D.; Shmarko, K.; Lomas, E. The Ethics of Facial Recognition Technologies, Surveillance, and Accountability in an Age of Artificial Intelligence: A Comparative Analysis of US, EU, and UK Regulatory Frameworks. AI Ethics 2021, 1–11. [Google Scholar] [CrossRef]
- Roundtree, A.K. Ethics and Facial Recognition Technology: An Integrative Review. In Proceedings of the 2021 3rd World Symp. Artif. (WSAI), Guangzhou, China, 18–20 June 2021; pp. 10–19. [Google Scholar] [CrossRef]
- Hongsong Chen, Y.Z.; Hongsong Chen, Y.Z. Security Issues and Defensive Approaches in Deep Learning Frameworks. Tsinghua Sci. Technol. 2021, 26, 894–905. [Google Scholar] [CrossRef]
- Silva, S.H.; Najafirad, P. Opportunities and Challenges in Deep Learning Adversarial Robustness: A Survey. arXiv 2020, arXiv:2007.00753. [Google Scholar]
- Yang, X.; Yang, D.; Dong, Y.; Yu, W.; Su, H.; Zhu, J. Delving into the Adversarial Robustness on Face Recognition. arXiv 2020, arXiv:2007.04118. [Google Scholar]
- Rozsa, A.; Günther, M.; Rudd, E.M.; Boult, T.E. Facial Attributes: Accuracy and Adversarial Robustness. Pattern Recognit. Lett. 2019, 124, 100–108. [Google Scholar] [CrossRef] [Green Version]
| Category | Algorithm |
|---|---|
| Appearance-based | Principal Component Analysis (PCA) Eigenface-Based Methods PCA Algorithm Kernel Principal Component Analysis (Kernel PCA) 2D-Image Principal Component Analysis (2D Image IPCA) Linear Discriminant Analysis (LDA) Discriminant Common Vectors (DCV) Independent Component Analysis (ICA) IPCA-ICA Super Vector Machine (SVM) |
| Feature-based | Geometric Features Local Binary Patterns (LBP) Elastic Bunch Graph Matching (EBGM) Histogram of Oriented Gradients (HoG) Elastic Bunch Graph (EBG) Hidden Markov Model (HMM) |
| Deep learning | Probabilistic-Decision-Based Neural Networks (PDBNN) Radial Basis Function (RBF) Convolutional Neural Network (CNN) |
| Study | Disease | Method | Sample Size | Efficacy |
|---|---|---|---|---|
| Basel-Vanagaite et al. [46] | Cornelia de Lange syndrome | FDNA | 31 cases in training set, 17 cases in testing set | Accuracy = 87% (training), accuracy = 94% (testing) |
| Latorre-Pellicer et al. [47] | Cornelia de Lange syndrome | Face2Gene | 49 cases | Accuracy = 83.7% |
| Hadj-Rabia et al. [48] | X-linked hypohidrotic ectodermal dysplasia | Face2Gene | 136 cases, 717 controls | AUC ≥ 0.98 |
| Liehr et al. [49] | Emanuel syndrome (ES) Pallister-Killian syndrome (PKS) | Face2Gene | 59 ES, 70 PKS, 973 controls, 973 others | AUC ≥ 0.98 |
| Amudhavalli et al. [50] | Aymé-Gripp syndrome | Face2Gene | 13 cases, 20 controls, 20 DS | AUC = 0.994 (controls), AUC = 0.994 (DS) |
| Pode-Shakked et al. [51] | Mucolipidosis type IV | Face2Gene | 26 cases, 98 controls, 99 others | AUC = 0.822 (controls), AUC = 0.885 (others) |
| Wang et al. [52] | Kabuki syndrome | Face2Gene | 14 cases | Accuracy = 93% |
| AbdAlmageed et al. [21] | Congenital adrenal hyperplasia | DNN | 102 cases, 144 controls | AUC = 92% |
| Porras et al. [53] | Noonan syndrome (NS) Williams-Beuren syndrome (WBS) | LBP, SVM | 286 NS, 161 WBS | Accuracy = 85.68% |
| Study | Disease | Data | Sample Size | Method | Efficacy |
|---|---|---|---|---|---|
| Bandini et al. [60] | PD | Video | 17 PD, 17 HC | Intraface tracking algorithm, Euclidean distance, SVM | Difference (p < 0.05) between PD and HC |
| Rajnoha et al. [61] | PD | Image | 50 PD, 50 HC | Random Forests, XGBoost | Accuracy = 67.33% |
| Jin et al. [23] | PD | Video | 33 PD, 31 HC | Face++ [62], tremor extraction, LSTM neural network | Precision = 86% |
| Ali et al. [5] | PD | Video | 61 PD, 543 HC | OpenFace 2.0 [24], SVM | Accuracy = 95.6% |
| Hou et al. [63] | PD | Video | 70 PD, 70 HC | HOG, LBP, SVM, k-NN, Random Forests | F1 = 88% |
| Nam et al. [25] | AD | Video | 17 AD, 17 HC | OpenFace 2.0 [24], extract movement coordinates to calculate Spearman’s correlation coefficient | Difference (p < 0.05) between AD and HC |
| Umeda et al. [64] | AD | Image | 121 AD, 117 HC | Xception, SENet50, ResNet50, VGG16, and simple CNN with SGD and Adam optimizer | Xception with Adam showed the best accuracy = 94% |
| Bandini et al. [18] | ALS | Video | 11 ALS, 11 HC | AAM, CLM, ERT, SDM, FAN | Accuracy = 88.9% |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Qiang, J.; Wu, D.; Du, H.; Zhu, H.; Chen, S.; Pan, H. Review on Facial-Recognition-Based Applications in Disease Diagnosis. Bioengineering 2022, 9, 273. https://doi.org/10.3390/bioengineering9070273
Qiang J, Wu D, Du H, Zhu H, Chen S, Pan H. Review on Facial-Recognition-Based Applications in Disease Diagnosis. Bioengineering. 2022; 9(7):273. https://doi.org/10.3390/bioengineering9070273
Chicago/Turabian StyleQiang, Jiaqi, Danning Wu, Hanze Du, Huijuan Zhu, Shi Chen, and Hui Pan. 2022. "Review on Facial-Recognition-Based Applications in Disease Diagnosis" Bioengineering 9, no. 7: 273. https://doi.org/10.3390/bioengineering9070273
APA StyleQiang, J., Wu, D., Du, H., Zhu, H., Chen, S., & Pan, H. (2022). Review on Facial-Recognition-Based Applications in Disease Diagnosis. Bioengineering, 9(7), 273. https://doi.org/10.3390/bioengineering9070273






