Recent Advances in Biological Recycling of Polyethylene Terephthalate (PET) Plastic Wastes
Abstract
:1. Introduction
2. PET Properties and Applications
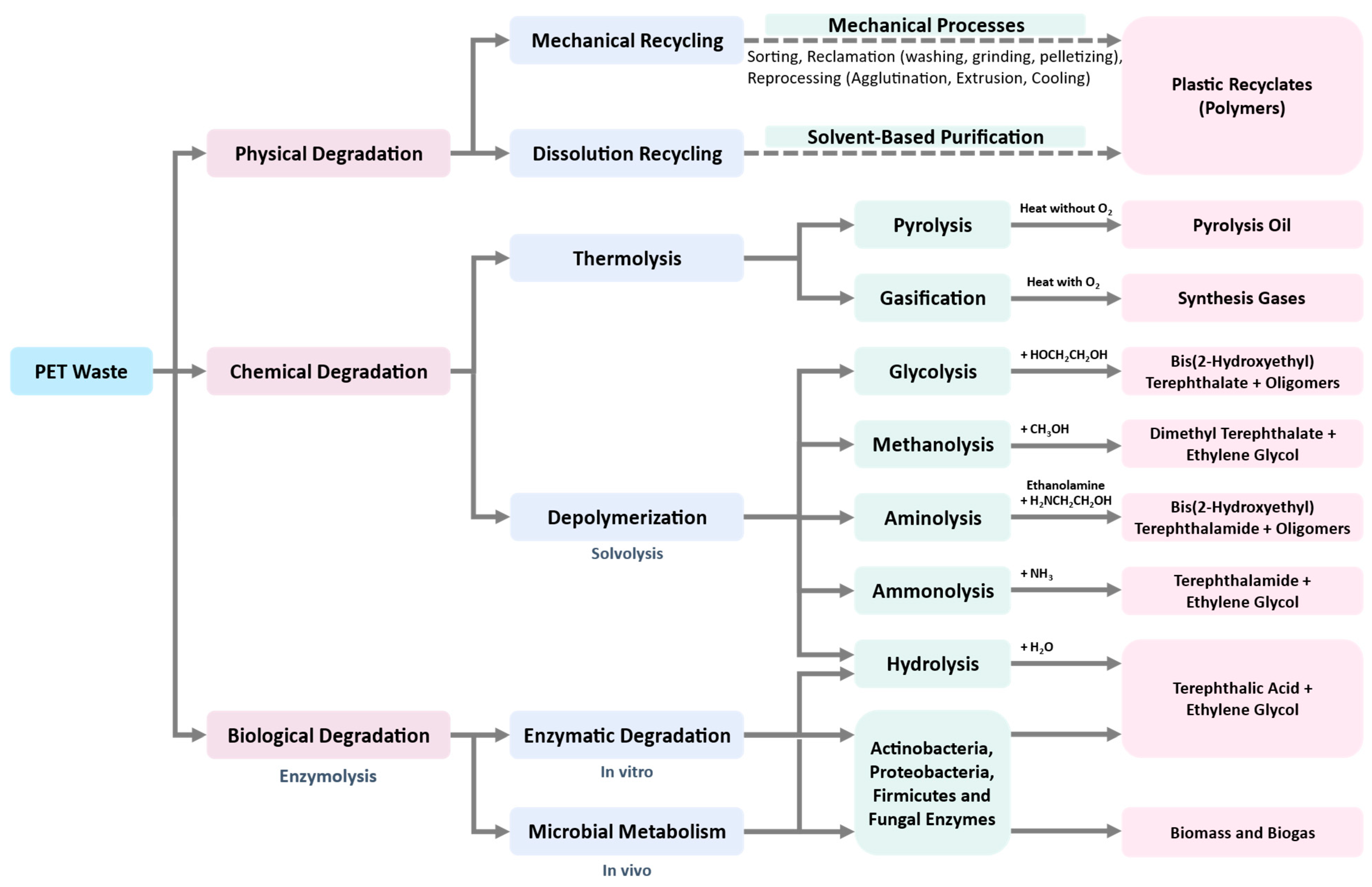
3. Conventional Approaches for Recycling of PET and Other Plastics
3.1. Landfilling
3.2. Energy Recovery and Incineration
3.3. Downgauging and Reuse of Packaging Plastic Materials
3.4. Four Categories of Plastic Recycling
3.4.1. Primary Recycling
3.4.2. Secondary Recycling (Mechanical Recycling)
3.4.3. Tertiary Recycling (Chemical or Feedstock Recycling)
3.4.4. Quaternary Recycling (Energy Recovery)
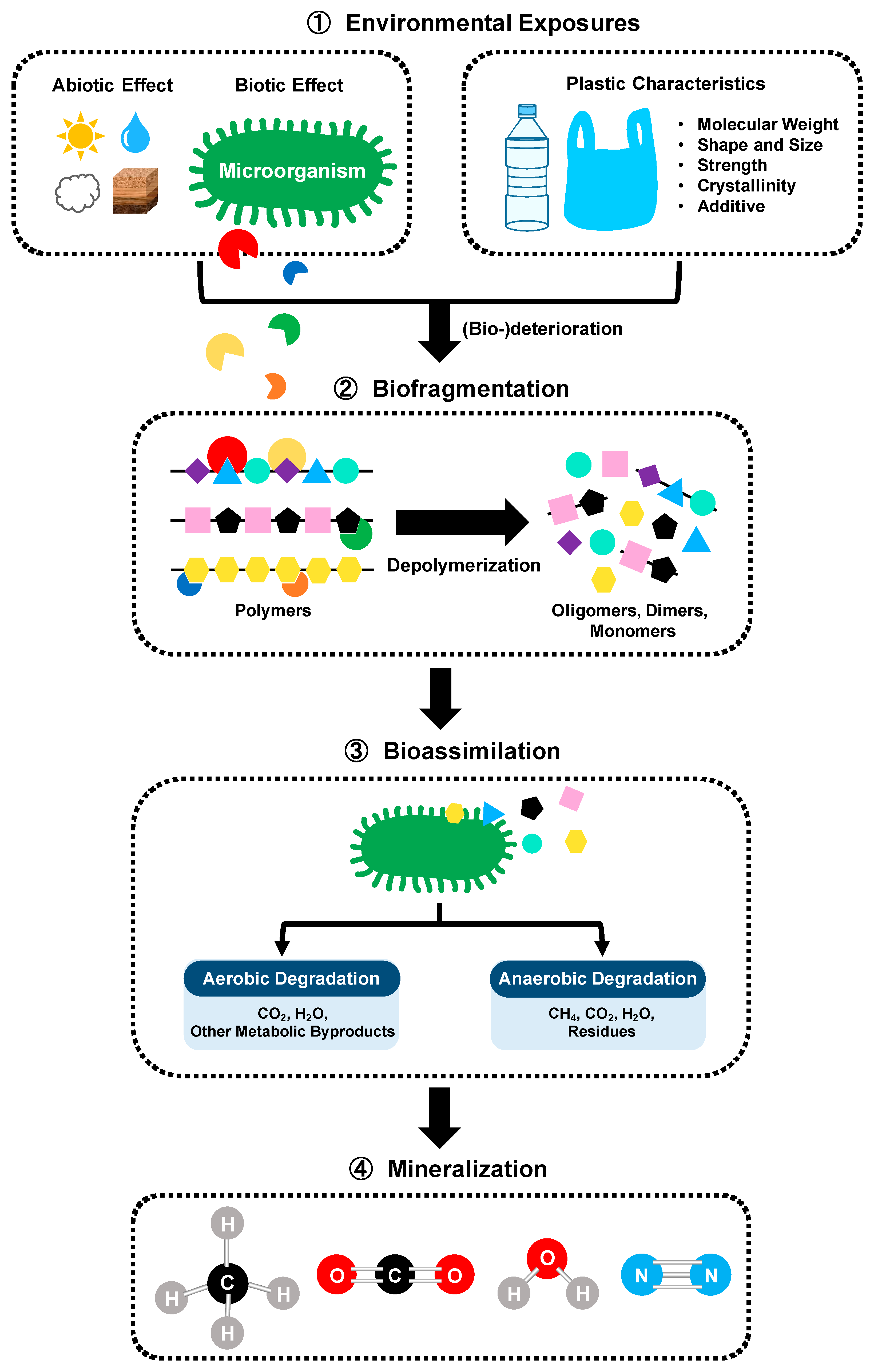
4. PET Recycling via Microbial Degradation
5. PET Recycling via Enzymatic Degradation
5.1. PET-Hydrolyzing Enzymes from Actinobacteria
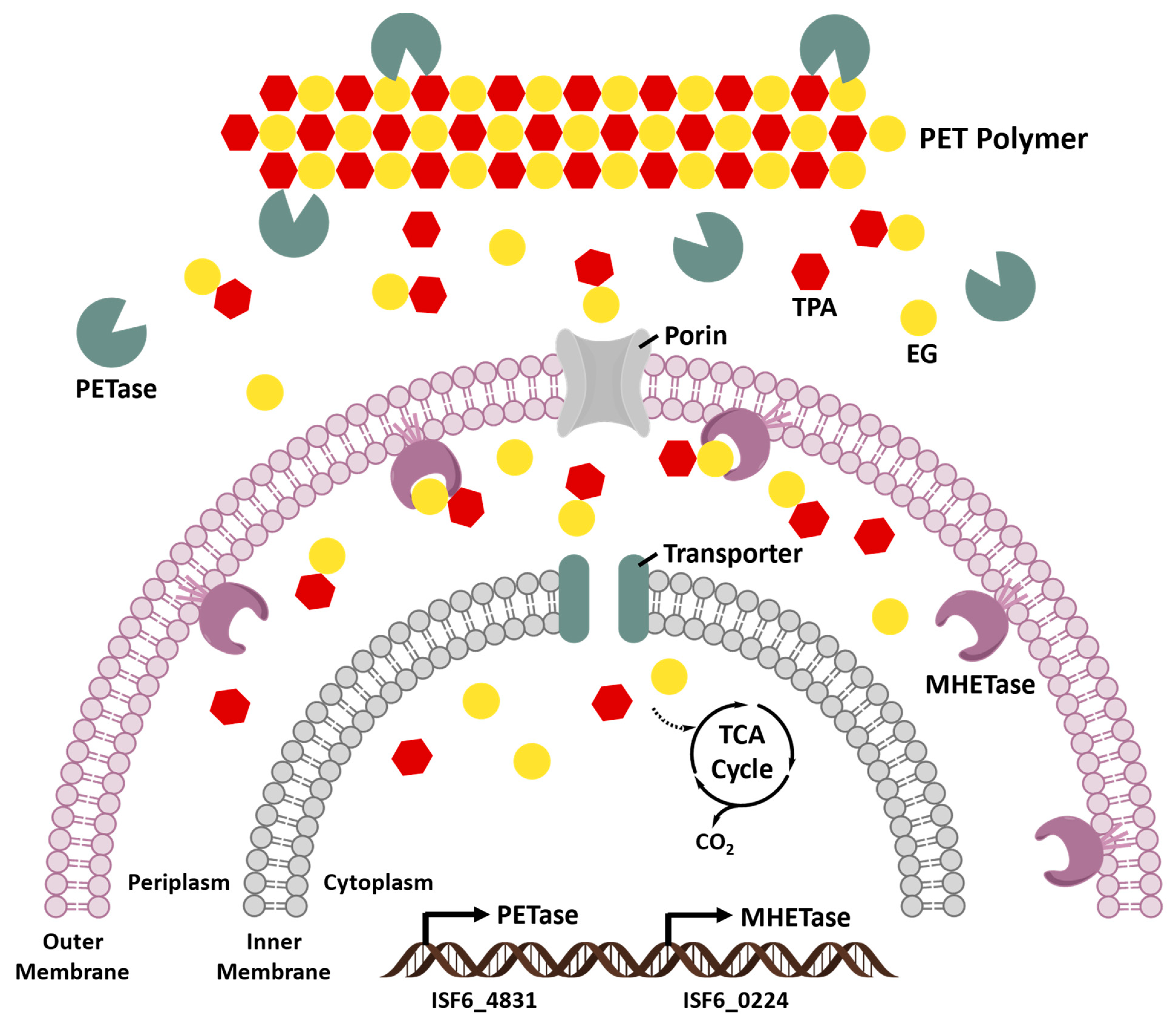
5.2. Ideonella sakaiensis Enzymes
5.2.1. Microbial Consortium No. 46
5.2.2. Ideonella sakaiensis 201-F6 from Microbial Consortium No. 46
5.2.3. Identification of PETase and MHETase in Ideonella sakaiensis 201-F6
5.3. PET-Hydrolyzing Enzymes from Fungi
5.4. Metagenome-Derived PET-Hydrolyzing Enzymes
6. Future Opportunities and Challenges for Biorecycling of PET Plastic
7. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Saminathan, P.; Sripriya, A.; Nalini, K.; Sivakumar, T.; Thangapandian, V. Biodegradation of plastics by Pseudomonas putida isolated from garden soil samples. J. Adv. Bot. Zool. 2014, 1, 34–38. [Google Scholar]
- Ahmed, T.; Shahid, M.; Azeem, F.; Rasul, I.; Shah, A.A.; Noman, M.; Hameed, A.; Manzoor, N.; Manzoor, I.; Muhammad, S. Biodegradation of plastics: Current scenario and future prospects for environmental safety. Environ. Sci. Pollut. Res. 2018, 25, 7287–7298. [Google Scholar] [CrossRef] [PubMed]
- Andrady, A.L. Plastics and Environmental Sustainability; John Wiley & Sons: Hoboken, NJ, USA, 2015. [Google Scholar]
- Nkwachukwu, O.I.; Chima, C.H.; Ikenna, A.O.; Albert, L. Focus on potential environmental issues on plastic world towards a sustainable plastic recycling in developing countries. Int. J. Ind. Chem. 2013, 4, 34. [Google Scholar] [CrossRef] [Green Version]
- Geyer, R.; Jambeck, J.R.; Law, K.L. Production, use, and fate of all plastics ever made. Sci. Adv. 2017, 3, e1700782. [Google Scholar] [CrossRef] [Green Version]
- Hopewell, J.; Dvorak, R.; Kosior, E. Plastics recycling: Challenges and opportunities. Philos. Trans. R. Soc. B Biol. Sci. 2009, 364, 2115–2126. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Zhao, S.; Wang, T.; Zhu, L.; Xu, P.; Wang, X.; Gao, L.; Li, D. Analysis of suspended microplastics in the Changjiang Estuary: Implications for riverine plastic load to the ocean. Water Res. 2019, 161, 560–569. [Google Scholar] [CrossRef] [PubMed]
- Hundertmark, T.; Mayer, M.; McNally, C.; Simons, T.J.; Witte, C. How Plastics Waste Recycling Could Transform the Chemical Industry. Available online: https://search-ebscohost-com.umasslowell.idm.oclc.org/login.aspx?direct=true&db=ply&AN=20190602564&site=eds-live (accessed on 26 December 2021).
- Rochman, C.M.; Browne, M.A.; Halpern, B.S.; Hentschel, B.T.; Hoh, E.; Karapanagioti, H.K.; Rios-Mendoza, L.M.; Takada, H.; Teh, S.; Thompson, R.C. Classify plastic waste as hazardous. Nature 2013, 494, 169–171. [Google Scholar] [CrossRef] [PubMed]
- Rochman, C.M.; Hoh, E.; Kurobe, T.; Teh, S.J. Ingested plastic transfers hazardous chemicals to fish and induces hepatic stress. Sci. Rep. 2013, 3, 3263. [Google Scholar] [CrossRef] [PubMed]
- Wilcox, C.; Van Sebille, E.; Hardesty, B.D. Threat of plastic pollution to seabirds is global, pervasive, and increasing. Proc. Natl. Acad. Sci. USA 2015, 112, 11899–11904. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Sussarellu, R.; Suquet, M.; Thomas, Y.; Lambert, C.; Fabioux, C.; Pernet, M.E.J.; Le Goïc, N.; Quillien, V.; Mingant, C.; Epelboin, Y. Oyster reproduction is affected by exposure to polystyrene microplastics. Proc. Natl. Acad. Sci. USA 2016, 113, 2430–2435. [Google Scholar] [CrossRef] [Green Version]
- Narancic, T.; O’Connor, K.E. Plastic waste as a global challenge: Are biodegradable plastics the answer to the plastic waste problem? Microbiology 2019, 165, 129–137. [Google Scholar] [CrossRef] [PubMed]
- Webb, H.K.; Arnott, J.; Crawford, R.J.; Ivanova, E.P. Plastic degradation and its environmental implications with special reference to poly (ethylene terephthalate). Polymers 2013, 5, 1–18. [Google Scholar] [CrossRef] [Green Version]
- Kint, D.; Muñoz-Guerra, S. A review on the potential biodegradability of poly (ethylene terephthalate). Polym. Int. 1999, 48, 346–352. [Google Scholar] [CrossRef]
- Awaja, F.; Pavel, D. Recycling of PET. Eur. Polym. J. 2005, 41, 1453–1477. [Google Scholar] [CrossRef]
- Karayannidis, G.P.; Psalida, E.A. Chain extension of recycled poly (ethylene terephthalate) with 2,2′-(1,4-phenylene)bis(2-oxazoline). J. Appl. Polym. Sci. 2000, 77, 2206–2211. [Google Scholar] [CrossRef]
- Scheirs, J. Polymer Recycling: Science, Technology and Applications; John Wiley & Sons Ltd.: Chichester, UK, 1998; p. 591. [Google Scholar]
- Caldicott, R.J. The basics of stretch blow molding PET containers. Plast. Eng. 1999, 55, 35–39. [Google Scholar]
- Rosato, D.V. Plastics Processing Data Handbook; Springer Science & Business Media: Berlin/Heidelberg, Germany, 2012. [Google Scholar]
- Hiraga, K.; Taniguchi, I.; Yoshida, S.; Kimura, Y.; Oda, K. Biodegradation of waste PET: A sustainable solution for dealing with plastic pollution. EMBO Rep. 2019, 20, e49365. [Google Scholar] [CrossRef] [PubMed]
- Barnes, D.K.; Galgani, F.; Thompson, R.C.; Barlaz, M. Accumulation and fragmentation of plastic debris in global environments. Philos. Trans. R. Soc. B Biol. Sci. 2009, 364, 1985–1998. [Google Scholar] [CrossRef] [Green Version]
- Gregory, M.R. Environmental implications of plastic debris in marine settings—Entanglement, ingestion, smothering, hangers-on, hitch-hiking and alien invasions. Philos. Trans. R. Soc. B Biol. Sci. 2009, 364, 2013–2025. [Google Scholar] [CrossRef] [PubMed]
- Oehlmann, J.; Schulte-Oehlmann, U.; Kloas, W.; Jagnytsch, O.; Lutz, I.; Kusk, K.O.; Wollenberger, L.; Santos, E.M.; Paull, G.C.; Van Look, K.J. A critical analysis of the biological impacts of plasticizers on wildlife. Philos. Trans. R. Soc. B Biol. Sci. 2009, 364, 2047–2062. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Jambeck, J.R.; Geyer, R.; Wilcox, C.; Siegler, T.R.; Perryman, M.; Andrady, A.; Narayan, R.; Law, K.L. Plastic waste inputs from land into the ocean. Science 2015, 347, 768–771. [Google Scholar] [CrossRef] [PubMed]
- McArthur, E. The New Plastics Economy: Rethinking the Future of Plastics & Catalysing Action; Ellen MacArthur Foundation: Cowes, UK, 2017; p. 68. [Google Scholar]
- Zhang, H.; Wen, Z.-G. The consumption and recycling collection system of PET bottles: A case study of Beijing, China. Waste Manag. 2014, 34, 987–998. [Google Scholar] [CrossRef] [PubMed]
- Prata, J.C.; Silva, A.L.P.; Da Costa, J.P.; Mouneyrac, C.; Walker, T.R.; Duarte, A.C.; Rocha-Santos, T. Solutions and integrated strategies for the control and mitigation of plastic and microplastic pollution. Int. J. Environ. Res. Public Health 2019, 16, 2411. [Google Scholar] [CrossRef] [Green Version]
- Schneider, D.R.; Ragossnig, A. Recycling and incineration, contradiction or coexistence? Waste Manag. Res. 2015, 33, 693–695. [Google Scholar] [PubMed] [Green Version]
- Calcott, P.; Walls, M. Can downstream waste disposal policies encourage upstream “design for environment”? Am. Econ. Rev. 2000, 90, 233–237. [Google Scholar] [CrossRef]
- Teuten, E.L.; Saquing, J.M.; Knappe, D.R.; Barlaz, M.A.; Jonsson, S.; Björn, A.; Rowland, S.J.; Thompson, R.C.; Galloway, T.S.; Yamashita, R. Transport and release of chemicals from plastics to the environment and to wildlife. Philos. Trans. R. Soc. B Biol. Sci. 2009, 364, 2027–2045. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Liu, Z.; Adams, M.; Walker, T.R. Are exports of recyclables from developed to developing countries waste pollution transfer or part of the global circular economy? Resour. Conserv. Recycl. 2018, 136, 22–23. [Google Scholar] [CrossRef]
- Arena, U. Process and technological aspects of municipal solid waste gasification. A review. Waste Manag. 2012, 32, 625–639. [Google Scholar] [CrossRef] [PubMed]
- Lombardi, L.; Carnevale, E.; Corti, A. A review of technologies and performances of thermal treatment systems for energy recovery from waste. Waste Manag. 2015, 37, 26–44. [Google Scholar] [CrossRef]
- Zevenhoven, R.; Karlsson, M.; Hupa, M.; Frankenhaeuser, M. Combustion and gasification properties of plastics particles. J. Air Waste Manag. Assoc. 1997, 47, 861–870. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Gilpin, R.K.; Wagel, D.J.; Solch, J.G. Production, Distribution, and Fate of Polychlorinated Dibenzo-p-dioxins, Dibenzofurans and Related Organohalogens in the Environment. In Dioxins and Health; John Wiley & Sons, Inc.: Hoboken, NJ, USA, 2003; pp. 55–87. [Google Scholar]
- Curlee, T.R.; Das, S. Plastic Wastes: Management, Control, Recycling and Disposal; William Andrew Publishing/Noyes Data Corp.: Park Ridge, NJ, USA, 1991. [Google Scholar]
- Okan, M.; Aydin, H.M.; Barsbay, M. Current approaches to waste polymer utilization and minimization: A review. J. Chem. Technol. Biotechnol. 2019, 94, 8–21. [Google Scholar] [CrossRef] [Green Version]
- Royte, E. Is Burning Plastic Waste a Good Idea? Available online: https://www.nationalgeographic.com/environment/article/should-we-burn-plastic-waste (accessed on 26 December 2021).
- Achilias, D.S. Chemical recycling of polymers. The case of poly (methyl methacrylate). In Proceedings of the International Conference on Energy & Environmental Systems, Chalkida, Greece, 8–10 May 2006. [Google Scholar]
- Saphire, D.; Bluestone, M. Case Reopened: Reassessing Refillable Bottles; Inform: Atlanta, GA, USA, 1994. [Google Scholar]
- Feron, V.; Jetten, J.; De Kruijf, N.; Van Den Berg, F. Polyethylene terephthalate bottles (PRBs): A health and safety assessment. Food Addit. Contam. 1994, 11, 571–594. [Google Scholar] [CrossRef] [PubMed]
- Schmidt, L. PET recycling: The view from NAPCOR. Resour. Recycl. 1998, 17, 37–42. [Google Scholar]
- Albrecht, P.; Brodersen, J.; Horst, D.; Scherf, M. Reuse and Recycling Systems for Selected Beverage Packaging from a Sustainability Perspective; PricewaterhouseCoopers AG WPG: Berlin, Germany, 2011. [Google Scholar]
- Paszun, D.; Spychaj, T. Chemical recycling of poly (ethylene terephthalate). Ind. Eng. Chem. Res. 1997, 36, 1373–1383. [Google Scholar] [CrossRef]
- Burillo, G.; Clough, R.L.; Czvikovszky, T.; Guven, O.; Le Moel, A.; Liu, W.; Singh, A.; Yang, J.; Zaharescu, T. Polymer recycling: Potential application of radiation technology. Radiat. Phys. Chem. 2002, 64, 41–51. [Google Scholar] [CrossRef]
- Majauskienė, R.; Miknius, L. Properties of residual marine fuel produced by thermolysis from polypropylene waste. Mater. Sci. 2015, 21, 249–254. [Google Scholar]
- Dimitris, S.; Achilias, L. Recent advances in the chemical recycling of polymers (PP, PS, LDPE, HDPE, PVC, PC, Nylon, PMMA). Mater. Recycl. Trends Perspect 2014, 3, 64. [Google Scholar]
- Jagtap, M.; Khatavkar, M.; Quazi, T. Methods for waste plastic recycling. Int. J. Recent Technol. Mech. Electr. Eng. 2015, 2, 120–122. [Google Scholar]
- Sinha, V.; Patel, M.R.; Patel, J.V. PET waste management by chemical recycling: A review. J. Polym. Environ. 2010, 18, 8–25. [Google Scholar] [CrossRef]
- Garcia, J.M.; Robertson, M.L. The future of plastics recycling. Science 2017, 358, 870–872. [Google Scholar] [CrossRef] [PubMed]
- Al-Salem, S.M.; Lettieri, P.; Baeyens, J. Recycling and recovery routes of plastic solid waste (PSW): A review. Waste Management 2009, 29, 2625–2643. [Google Scholar] [CrossRef] [PubMed]
- Schulte, P.A.; McKernan, L.T.; Heidel, D.S.; Okun, A.H.; Dotson, G.S.; Lentz, T.J.; Geraci, C.L.; Heckel, P.E.; Branche, C.M. Occupational safety and health, green chemistry, and sustainability: A review of areas of convergence. Environ. Health 2013, 12, 1–9. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Idumah, C.I.; Nwuzor, I.C. Novel trends in plastic waste management. SN Appl. Sci. 2019, 1, 1402. [Google Scholar] [CrossRef] [Green Version]
- Mourshed, M.; Masud, M.H.; Rashid, F.; Joardder, M.U.H. Towards the effective plastic waste management in Bangladesh: A review. Environ. Sci. Pollut. Res. 2017, 24, 27021–27046. [Google Scholar] [CrossRef] [PubMed]
- Glaser, J.A. Biological degradation of polymers in the environment. In Plastics in the Environment; IntechOpen: London, UK, 2019. [Google Scholar]
- Khoo, K.S.H.; Ho, L.Y.; Lim, H.R.; Leong, H.Y.; Chew, K.W. Plastic waste associated with the COVID-19 pandemic: Crisis or opportunity? J. Hazard. Mater. 2021, 417, 126108. [Google Scholar] [CrossRef] [PubMed]
- Chia, W.Y.; Ying Tang, D.Y.; Khoo, K.S.; Kay Lup, A.N.; Chew, K.W. Nature’s fight against plastic pollution: Algae for plastic biodegradation and bioplastics production. Environ. Sci. Ecotechnol. 2020, 4, 100065. [Google Scholar] [CrossRef]
- Lucas, N.; Bienaime, C.; Belloy, C.; Queneudec, M.; Silvestre, F.; Nava-Saucedo, J.-E. Polymer biodegradation: Mechanisms and estimation techniques—A review. Chemosphere 2008, 73, 429–442. [Google Scholar] [CrossRef] [PubMed]
- Dussud, C.; Ghiglione, J.-F. Bacterial degradation of synthetic plastics. In CIESM Workshop Monographs; CIESM Publisher: Monaco City, Monaco, 2014. [Google Scholar]
- Jaiswal, S.; Sharma, B.; Shukla, P. Integrated approaches in microbial degradation of plastics. Environ. Technol. Innov. 2020, 17, 100567. [Google Scholar] [CrossRef]
- Shah, A.A.; Hasan, F.; Hameed, A.; Ahmed, S. Biological degradation of plastics: A comprehensive review. Biotechnol. Adv. 2008, 26, 246–265. [Google Scholar] [CrossRef] [PubMed]
- Barth, M.; Wei, R.; Oeser, T.; Then, J.; Schmidt, J.; Wohlgemuth, F.; Zimmermann, W. Enzymatic hydrolysis of polyethylene terephthalate films in an ultrafiltration membrane reactor. J. Membr. Sci. 2015, 494, 182–187. [Google Scholar] [CrossRef]
- Yoshida, S.; Hiraga, K.; Takehana, T.; Taniguchi, I.; Yamaji, H.; Maeda, Y.; Toyohara, K.; Miyamoto, K.; Kimura, Y.; Oda, K. A bacterium that degrades and assimilates poly (ethylene terephthalate). Science 2016, 351, 1196–1199. [Google Scholar] [CrossRef] [PubMed]
- Nowak, B.; Pająk, J.; Łabużek, S.; Rymarz, G.; Talik, E. Biodegradation of poly (ethylene terephthalate) modified with polyester “Bionolle®” by Penicillium funiculosum. Polimery 2011, 56, 35–44. [Google Scholar] [CrossRef]
- Sharon, C.; Sharon, M. Studies on biodegradation of polyethylene terephthalate: A synthetic polymer. J. Microbiol. Biotechnol. Res. 2012, 2, 248–257. [Google Scholar]
- Kawai, F.; Kawabata, T.; Oda, M. Current state and perspectives related to the polyethylene terephthalate hydrolases available for biorecycling. ACS Sustain. Chem. Eng. 2020, 8, 8894–8908. [Google Scholar] [CrossRef]
- Frazee, R.W.; Livingston, D.; LaPorte, D.; Lipscomb, J. Cloning, sequencing, and expression of the Pseudomonas putida protocatechuate 3, 4-dioxygenase genes. J. Bacteriol. 1993, 175, 6194–6202. [Google Scholar] [CrossRef] [Green Version]
- Kasai, D.; Fujinami, T.; Abe, T.; Mase, K.; Katayama, Y.; Fukuda, M.; Masai, E. Uncovering the protocatechuate 2, 3-cleavage pathway genes. J. Bacteriol. 2009, 191, 6758–6768. [Google Scholar] [CrossRef] [Green Version]
- Maruyama, K.; Shibayama, T.; Ichikawa, A.; Sakou, Y.; Yamada, S.; Sugisaki, H. Cloning and characterization of the genes encoding enzymes for the protocatechuate meta-degradation pathway of Pseudomonas ochraceae NGJ1. Biosci. Biotechnol. Biochem. 2004, 68, 1434–1441. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Johnson, C.W.; Salvachúa, D.; Khanna, P.; Smith, H.; Peterson, D.J.; Beckham, G.T. Enhancing muconic acid production from glucose and lignin-derived aromatic compounds via increased protocatechuate decarboxylase activity. Metab. Eng. Commun. 2016, 3, 111–119. [Google Scholar] [CrossRef] [PubMed]
- Salvador, M.; Abdulmutalib, U.; Gonzalez, J.; Kim, J.; Smith, A.A.; Faulon, J.-L.; Wei, R.; Zimmermann, W.; Jimenez, J.I. Microbial genes for a circular and sustainable bio-PET economy. Genes 2019, 10, 373. [Google Scholar] [CrossRef] [Green Version]
- Trifunović, D.; Schuchmann, K.; Müller, V. Ethylene glycol metabolism in the acetogen Acetobacterium woodii. J. Bacteriol. 2016, 198, 1058–1065. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Mückschel, B.; Simon, O.; Klebensberger, J.; Graf, N.; Rosche, B.; Altenbuchner, J.; Pfannstiel, J.; Huber, A.; Hauer, B. Ethylene glycol metabolism by Pseudomonas putida. Appl. Environ. Microbiol. 2012, 78, 8531–8539. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Franden, M.A.; Jayakody, L.N.; Li, W.-J.; Wagner, N.J.; Cleveland, N.S.; Michener, W.E.; Hauer, B.; Blank, L.M.; Wierckx, N.; Klebensberger, J. Engineering Pseudomonas putida KT2440 for efficient ethylene glycol utilization. Metab. Eng. 2018, 48, 197–207. [Google Scholar] [CrossRef] [PubMed]
- Kumar, R.V.; Kanna, G.R.; Elumalai, S. Biodegradation of polyethylene by green photosynthetic microalgae. J. Bioremediat. Biodegrad. 2017, 8, 2. [Google Scholar]
- Zheng, Y.; Yanful, E.K.; Bassi, A.S. A review of plastic waste biodegradation. Crit. Rev. Biotechnol. 2005, 25, 243–250. [Google Scholar] [CrossRef] [PubMed]
- Krueger, M.C.; Harms, H.; Schlosser, D. Prospects for microbiological solutions to environmental pollution with plastics. Appl. Microbiol. Biotechnol. 2015, 99, 8857–8874. [Google Scholar] [CrossRef] [PubMed]
- Nishida, H.; Tokiwa, Y. Distribution of poly (β-hydroxybutyrate) and poly (ε-caprolactone) aerobic degrading microorganisms in different environments. J. Environ. Polym. Degrad. 1993, 1, 227–233. [Google Scholar] [CrossRef]
- Parikh, M.; Gross, R. The effect of crystalline morphology on enzymatic degradation kinetics. Polym. Mater. Sci. Eng. (PMSE) 1992, 66, 408–410. [Google Scholar]
- Marten, E.; Müller, R.-J.; Deckwer, W.-D. Studies on the enzymatic hydrolysis of polyesters I. Low molecular mass model esters and aliphatic polyesters. Polym. Degrad. Stab. 2003, 80, 485–501. [Google Scholar] [CrossRef]
- Marten, E.; Müller, R.-J.; Deckwer, W.-D. Studies on the enzymatic hydrolysis of polyesters. II. Aliphatic–aromatic copolyesters. Polym. Degrad. Stab. 2005, 88, 371–381. [Google Scholar] [CrossRef]
- Tokiwa, Y.; Calabia, B.P.; Ugwu, C.U.; Aiba, S. Biodegradability of plastics. Int. J. Mol. Sci. 2009, 10, 3722–3742. [Google Scholar] [CrossRef]
- Wei, R.; Zimmermann, W. Microbial enzymes for the recycling of recalcitrant petroleum-based plastics: How far are we? Microb. Biotechnol. 2017, 10, 1308–1322. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Vertommen, M.; Nierstrasz, V.; Van Der Veer, M.; Warmoeskerken, M. Enzymatic surface modification of poly (ethylene terephthalate). J. Biotechnol. 2005, 120, 376–386. [Google Scholar] [CrossRef] [PubMed]
- Müller, R.J.; Schrader, H.; Profe, J.; Dresler, K.; Deckwer, W.D. Enzymatic degradation of poly (ethylene terephthalate): Rapid hydrolyse using a hydrolase from T. fusca. Macromol. Rapid Commun. 2005, 26, 1400–1405. [Google Scholar] [CrossRef]
- Mohanan, N.; Montazer, Z.; Sharma, P.K.; Levin, D.B. Microbial and Enzymatic Degradation of Synthetic Plastics. Front. Microbiol. 2020, 11, 2837. [Google Scholar] [CrossRef] [PubMed]
- Austin, H.P.; Allen, M.D.; Donohoe, B.S.; Rorrer, N.A.; Kearns, F.L.; Silveira, R.L.; Pollard, B.C.; Dominick, G.; Duman, R.; El Omari, K. Characterization and engineering of a plastic-degrading aromatic polyesterase. Proc. Natl. Acad. Sci. USA 2018, 115, E4350–E4357. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Wei, R.; Oeser, T.; Then, J.; Kühn, N.; Barth, M.; Schmidt, J.; Zimmermann, W. Functional characterization and structural modeling of synthetic polyester-degrading hydrolases from Thermomonospora curvata. AMB Express 2014, 4, 44. [Google Scholar] [CrossRef] [Green Version]
- Remington, S.; Franken, S.; Sussman, J.; Frolow, F.; Ollis, D.; Verschueren, K.; Cheah, E.; Cygler, M.; Dijkstra, B.; Harel, M. The alpha/beta hydrolase fold. Protein Eng. 1992, 5, 197–211. [Google Scholar]
- Barth, M.; Oeser, T.; Wei, R.; Then, J.; Schmidt, J.; Zimmermann, W. Effect of hydrolysis products on the enzymatic degradation of polyethylene terephthalate nanoparticles by a polyester hydrolase from Thermobifida fusca. Biochem. Eng. J. 2015, 93, 222–228. [Google Scholar] [CrossRef]
- Sulaiman, S.; Yamato, S.; Kanaya, E.; Kim, J.-J.; Koga, Y.; Takano, K.; Kanaya, S. Isolation of a novel cutinase homolog with polyethylene terephthalate-degrading activity from leaf-branch compost by using a metagenomic approach. Appl. Environ. Microbiol. 2012, 78, 1556–1562. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Kawabata, T.; Oda, M.; Kawai, F. Mutational analysis of cutinase-like enzyme, Cut190, based on the 3D docking structure with model compounds of polyethylene terephthalate. J. Biosci. Bioeng. 2017, 124, 28–35. [Google Scholar] [CrossRef]
- Tournier, V.; Topham, C.; Gilles, A.; David, B.; Folgoas, C.; Moya-Leclair, E.; Kamionka, E.; Desrousseaux, M.-L.; Texier, H.; Gavalda, S. An engineered PET depolymerase to break down and recycle plastic bottles. Nature 2020, 580, 216–219. [Google Scholar] [CrossRef] [PubMed]
- Kim, H.R.; Song, W.S. Lipase treatment of polyester fabrics. Fibers Polym. 2006, 7, 339–343. [Google Scholar] [CrossRef]
- Guebitz, G.M.; Cavaco-Paulo, A. Enzymes go big: Surface hydrolysis and functionalisation of synthetic polymers. Trends Biotechnol. 2008, 26, 32–38. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Ribitsch, D.; Herrero Acero, E.; Greimel, K.; Dellacher, A.; Zitzenbacher, S.; Marold, A.; Rodriguez, R.D.; Steinkellner, G.; Gruber, K.; Schwab, H. A new esterase from Thermobifida halotolerans hydrolyses polyethylene terephthalate (PET) and polylactic acid (PLA). Polymers 2012, 4, 617–629. [Google Scholar] [CrossRef]
- Sales, J.C.S.; de Castro, A.M.; Ribeiro, B.D.; Coelho, M.A.Z. Supplementation of watermelon peels as an enhancer of lipase and esterase production by Yarrowia lipolytica in solid-state fermentation and their potential use as biocatalysts in poly (ethylene terephthalate)(PET) depolymerization reactions. Biocatal. Biotransform. 2020, 38, 457–468. [Google Scholar] [CrossRef]
- Roth, C.; Wei, R.; Oeser, T.; Then, J.; Föllner, C.; Zimmermann, W.; Sträter, N. Structural and functional studies on a thermostable polyethylene terephthalate degrading hydrolase from Thermobifida fusca. Appl. Microbiol. Biotechnol. 2014, 98, 7815–7823. [Google Scholar] [CrossRef] [PubMed]
- Sulaiman, S.; You, D.-J.; Kanaya, E.; Koga, Y.; Kanaya, S. Crystal structure and thermodynamic and kinetic stability of metagenome-derived LC-cutinase. Biochemistry 2014, 53, 1858–1869. [Google Scholar] [CrossRef] [PubMed]
- Then, J.; Wei, R.; Oeser, T.; Barth, M.; Belisário-Ferrari, M.R.; Schmidt, J.; Zimmermann, W. Ca2+ and Mg2+ binding site engineering increases the degradation of polyethylene terephthalate films by polyester hydrolases from Thermobifida fusca. Biotechnol. J. 2015, 10, 592–598. [Google Scholar] [CrossRef]
- Thumarat, U.; Nakamura, R.; Kawabata, T.; Suzuki, H.; Kawai, F. Biochemical and genetic analysis of a cutinase-type polyesterase from a thermophilic Thermobifida alba AHK119. Appl. Microbiol. Biotechnol. 2012, 95, 419–430. [Google Scholar] [CrossRef] [PubMed]
- Herrero Acero, E.; Ribitsch, D.; Steinkellner, G.; Gruber, K.; Greimel, K.; Eiteljoerg, I.; Trotscha, E.; Wei, R.; Zimmermann, W.; Zinn, M. Enzymatic surface hydrolysis of PET: Effect of structural diversity on kinetic properties of cutinases from Thermobifida. Macromolecules 2011, 44, 4632–4640. [Google Scholar] [CrossRef] [Green Version]
- Kawai, F.; Oda, M.; Tamashiro, T.; Waku, T.; Tanaka, N.; Yamamoto, M.; Mizushima, H.; Miyakawa, T.; Tanokura, M. A novel Ca2+-activated, thermostabilized polyesterase capable of hydrolyzing polyethylene terephthalate from Saccharomonospora viridis AHK190. Appl. Microbiol. Biotechnol. 2014, 98, 10053–10064. [Google Scholar] [CrossRef] [PubMed]
- Purdy, R.; Kolattukudy, P. Hydrolysis of plant cuticle by plant pathogens. Purification, amino acid composition, and molecular weight of two isoenzymes of cutinase and a nonspecific esterase from Fusarium solani f. pisi. Biochemistry 1975, 14, 2824–2831. [Google Scholar] [CrossRef] [PubMed]
- Furukawa, M.; Kawakami, N.; Tomizawa, A.; Miyamoto, K. Efficient degradation of poly (ethylene terephthalate) with Thermobifida fusca cutinase exhibiting improved catalytic activity generated using mutagenesis and additive-based approaches. Sci. Rep. 2019, 9, 16038. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Ronkvist, Å.M.; Xie, W.; Lu, W.; Gross, R.A. Cutinase-Catalyzed Hydrolysis of Poly(ethylene terephthalate). Macromolecules 2009, 42, 5128–5138. [Google Scholar] [CrossRef]
- Zimmermann, W.; Billig, S. Enzymes for the biofunctionalization of poly (ethylene terephthalate). In Biofunctionalization of Polymers and Their Applications; Springer: Berlin/Heidelberg, Germany, 2010; pp. 97–120. [Google Scholar]
- Eberl, A.; Heumann, S.; Brückner, T.; Araujo, R.; Cavaco-Paulo, A.; Kaufmann, F.; Kroutil, W.; Guebitz, G.M. Enzymatic surface hydrolysis of poly (ethylene terephthalate) and bis (benzoyloxyethyl) terephthalate by lipase and cutinase in the presence of surface active molecules. J. Biotechnol. 2009, 143, 207–212. [Google Scholar] [CrossRef] [PubMed]
- Ribitsch, D.; Heumann, S.; Trotscha, E.; Herrero Acero, E.; Greimel, K.; Leber, R.; Birner-Gruenberger, R.; Deller, S.; Eiteljoerg, I.; Remler, P. Hydrolysis of polyethyleneterephthalate by p-nitrobenzylesterase from Bacillus subtilis. Biotechnol. Prog. 2011, 27, 951–960. [Google Scholar] [CrossRef] [PubMed]
- Kleeberg, I.; Hetz, C.; Kroppenstedt, R.M.; Müller, R.-J.; Deckwer, W.-D. Biodegradation of aliphatic-aromatic copolyesters by Thermomonospora fusca and other thermophilic compost isolates. Appl. Environ. Microbiol. 1998, 64, 1731–1735. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Kleeberg, I.; Welzel, K.; VandenHeuvel, J.; Müller, R.-J.; Deckwer, W.-D. Characterization of a New Extracellular Hydrolase from Thermobifida fusca Degrading Aliphatic−Aromatic Copolyesters. Biomacromolecules 2005, 6, 262–270. [Google Scholar] [CrossRef] [PubMed]
- Chen, S.; Su, L.; Billig, S.; Zimmermann, W.; Chen, J.; Wu, J. Biochemical characterization of the cutinases from Thermobifida fusca. J. Mol. Catal. B Enzym. 2010, 63, 121–127. [Google Scholar] [CrossRef]
- Oda, M.; Yamagami, Y.; Inaba, S.; Oida, T.; Yamamoto, M.; Kitajima, S.; Kawai, F. Enzymatic hydrolysis of PET: Functional roles of three Ca 2+ ions bound to a cutinase-like enzyme, Cut190*, and its engineering for improved activity. Appl. Microbiol. Biotechnol. 2018, 102, 10067–10077. [Google Scholar] [CrossRef] [PubMed]
- Cruz, H.; Pérez, C.; Wellington, E.; Castro, C.; Servín-González, L. Sequence of the Streptomyces albus G lipase-encoding gene reveals the presence of a prokaryotic lipase family. Gene 1994, 144, 141–142. [Google Scholar] [CrossRef]
- Pérez, C.; Juárez, K.; García-Castells, E.; Soberón, G.; Servín-González, L. Cloning, characterization, and expression in Streptomyces lividans 66 of an extracellular lipase-encoding gene from Streptomyces sp. M11. Gene 1993, 123, 109–114. [Google Scholar] [CrossRef]
- Lykidis, A.; Mavromatis, K.; Ivanova, N.; Anderson, I.; Land, M.; DiBartolo, G.; Martinez, M.; Lapidus, A.; Lucas, S.; Copeland, A.; et al. Genome sequence and analysis of the soil cellulolytic actinomycete Thermobifida fusca YX. J. Bacteriol. 2007, 189, 2477–2486. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Chen, V.B.; Arendall, W.B.; Headd, J.J.; Keedy, D.A.; Immormino, R.M.; Kapral, G.J.; Murray, L.W.; Richardson, J.S.; Richardson, D.C. MolProbity: All-atom structure validation for macromolecular crystallography. Acta Crystallogr. Sect. D Biol. Crystallogr. 2010, 66, 12–21. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Alisch-Mark, M.; Herrmann, A.; Zimmermann, W. Increase of the hydrophilicity of polyethylene terephthalate fibres by hydrolases from Thermomonospora fusca and Fusarium solani f. sp. pisi. Biotechnol. Lett. 2006, 28, 681–685. [Google Scholar] [CrossRef] [PubMed]
- Feuerhack, A.; Alisch-Mark, M.; Kisner, A.; Pezzin, S.; Zimmermann, W.; Andreaus, J. Biocatalytic surface modification of knitted fabrics made of poly (ethylene terephthalate) with hydrolytic enzymes from Thermobifida fusca KW3b. Biocatal. Biotransform. 2008, 26, 357–364. [Google Scholar] [CrossRef]
- Hegde, K.; Veeranki, V.D. Production optimization and characterization of recombinant cutinases from Thermobifida fusca sp. NRRL B-8184. Appl. Biochem. Biotechnol. 2013, 170, 654–675. [Google Scholar] [CrossRef] [PubMed]
- Chertkov, O.; Sikorski, J.; Nolan, M.; Lapidus, A.; Lucas, S.; Del Rio, T.G.; Tice, H.; Cheng, J.-F.; Goodwin, L.; Pitluck, S.; et al. Complete genome sequence of Thermomonospora curvata type strain (B9). Stand. Genom. Sci. 2011, 4, 13–22. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Arima, J.; Uesugi, Y.; Uraji, M.; Yatsushiro, S.; Tsuboi, S.; Iwabuchi, M.; Hatanaka, T. Modulation of Streptomyces leucine aminopeptidase by calcium: Identification and functional analysis of key residues in activation and stabilization by calcium. J. Biol. Chem. 2006, 281, 5885–5894. [Google Scholar] [CrossRef] [Green Version]
- Almeida, E.L.; Carrillo Rincón, A.F.; Jackson, S.A.; Dobson, A.D.W. In silico Screening and Heterologous Expression of a Polyethylene Terephthalate Hydrolase (PETase)-Like Enzyme (SM14est) with Polycaprolactone (PCL)-Degrading Activity, From the Marine Sponge-Derived Strain Streptomyces sp. SM14. Front. Microbiol. 2019, 10, 2187. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Jabloune, R.; Khalil, M.; Ben Moussa, I.E.; Simao-Beaunoir, A.-M.; Lerat, S.; Brzezinski, R.; Beaulieu, C. Enzymatic Degradation of p-Nitrophenyl Esters, Polyethylene Terephthalate, Cutin, and Suberin by Sub1, a Suberinase Encoded by the Plant Pathogen Streptomyces scabies. Microbes Environ. 2020, 35, ME19086. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Alves, N.; Mano, J.F.; Balaguer, E.; Dueñas, J.M.; Ribelles, J.G. Glass transition and structural relaxation in semi-crystalline poly (ethylene terephthalate): A DSC study. Polymer 2002, 43, 4111–4122. [Google Scholar] [CrossRef]
- Kikkawa, Y.; Fujita, M.; Abe, H.; Doi, Y. Effect of water on the surface molecular mobility of poly (lactide) thin films: An atomic force microscopy study. Biomacromolecules 2004, 5, 1187–1193. [Google Scholar] [CrossRef]
- Taniguchi, I.; Yoshida, S.; Hiraga, K.; Miyamoto, K.; Kimura, Y.; Oda, K. Biodegradation of PET: Current status and application aspects. ACS Catal. 2019, 9, 4089–4105. [Google Scholar] [CrossRef]
- Dimarogona, M.; Nikolaivits, E.; Kanelli, M.; Christakopoulos, P.; Sandgren, M.; Topakas, E. Structural and functional studies of a Fusarium oxysporum cutinase with polyethylene terephthalate modification potential. Biochim. Biophys. Acta (BBA) Gen. Subj. 2015, 1850, 2308–2317. [Google Scholar] [CrossRef] [PubMed]
- Gamerith, C.; Vastano, M.; Ghorbanpour, S.M.; Zitzenbacher, S.; Ribitsch, D.; Zumstein, M.T.; Sander, M.; Herrero Acero, E.; Pellis, A.; Guebitz, G.M. Enzymatic degradation of aromatic and aliphatic polyesters by P. pastoris expressed cutinase 1 from Thermobifida cellulosilytica. Front. Microbiol. 2017, 8, 938. [Google Scholar] [CrossRef]
- Tanasupawat, S.; Takehana, T.; Yoshida, S.; Hiraga, K.; Oda, K. Ideonella sakaiensis sp. nov., isolated from a microbial consortium that degrades poly (ethylene terephthalate). Int. J. Syst. Evol. Microbiol. 2016, 66, 2813–2818. [Google Scholar] [CrossRef] [PubMed]
- Uchida, H.; Shigeno-Akutsu, Y.; Nomura, N.; Nakahara, T.; Nakajima-Kambe, T. Cloning and sequence analysis of poly (tetramethylene succinate) depolymerase from Acidovorax delafieldii strain BS-3. J. Biosci. Bioeng. 2002, 93, 245–247. [Google Scholar] [CrossRef]
- Tang, B.; Yu, Y.; Zhang, Y.; Zhao, G.; Ding, X. Complete genome sequence of the glidobactin producing strain [Polyangium] brachysporum DSM 7029. J. Biotechnol. 2015, 210, 83–84. [Google Scholar] [CrossRef]
- Anantharaman, K.; Brown, C.T.; Hug, L.A.; Sharon, I.; Castelle, C.J.; Probst, A.J.; Thomas, B.C.; Singh, A.; Wilkins, M.J.; Karaoz, U. Thousands of microbial genomes shed light on interconnected biogeochemical processes in an aquifer system. Nat. Commun. 2016, 7, 1–11. [Google Scholar] [CrossRef]
- Silva, C.M.; Carneiro, F.; O’Neill, A.; Fonseca, L.P.; Cabral, J.S.; Guebitz, G.; Cavaco-Paulo, A. Cutinase—A new tool for biomodification of synthetic fibers. J. Polym. Sci. Part A Polym. Chem. 2005, 43, 2448–2450. [Google Scholar] [CrossRef] [Green Version]
- Sasoh, M.; Masai, E.; Ishibashi, S.; Hara, H.; Kamimura, N.; Miyauchi, K.; Fukuda, M. Characterization of the terephthalate degradation genes of Comamonas sp. strain E6. Appl. Environ. Microbiol. 2006, 72, 1825–1832. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Hosaka, M.; Kamimura, N.; Toribami, S.; Mori, K.; Kasai, D.; Fukuda, M.; Masai, E. Novel tripartite aromatic acid transporter essential for terephthalate uptake in Comamonas sp. strain E6. Appl. Environ. Microbiol. 2013, 79, 6148–6155. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Pérez-Pantoja, D.; Donoso, R.; Agulló, L.; Córdova, M.; Seeger, M.; Pieper, D.H.; González, B. Genomic analysis of the potential for aromatic compounds biodegradation in Burkholderiales. Environ. Microbiol. 2012, 14, 1091–1117. [Google Scholar] [CrossRef] [PubMed]
- Wilkes, R.A.; Aristilde, L. Degradation and metabolism of synthetic plastics and associated products by Pseudomonas sp.: Capabilities and challenges. J. Appl. Microbiol. 2017, 123, 582–593. [Google Scholar] [CrossRef] [Green Version]
- Li, W.J.; Jayakody, L.N.; Franden, M.A.; Wehrmann, M.; Daun, T.; Hauer, B.; Blank, L.M.; Beckham, G.T.; Klebensberger, J.; Wierckx, N. Laboratory evolution reveals the metabolic and regulatory basis of ethylene glycol metabolism by Pseudomonas putida KT2440. Environ. Microbiol. 2019, 21, 3669–3682. [Google Scholar] [CrossRef]
- Tiwari, V. In vitro engineering of novel bioactivity in the natural enzymes. Front. Chem. 2016, 4, 39. [Google Scholar] [CrossRef] [Green Version]
- Fecker, T.; Galaz-Davison, P.; Engelberger, F.; Narui, Y.; Sotomayor, M.; Parra, L.P.; Ramírez-Sarmiento, C.A. Active site flexibility as a hallmark for efficient PET degradation by I. sakaiensis PETase. Biophys. J. 2018, 114, 1302–1312. [Google Scholar] [CrossRef] [Green Version]
- Liu, B.; He, L.; Wang, L.; Li, T.; Li, C.; Liu, H.; Luo, Y.; Bao, R. Protein crystallography and site-direct mutagenesis analysis of the poly (ethylene terephthalate) hydrolase PETase from Ideonella sakaiensis. ChemBioChem 2018, 19, 1471–1475. [Google Scholar] [CrossRef]
- Joo, S.; Cho, I.J.; Seo, H.; Son, H.F.; Sagong, H.-Y.; Shin, T.J.; Choi, S.Y.; Lee, S.Y.; Kim, K.-J. Structural insight into molecular mechanism of poly (ethylene terephthalate) degradation. Nat. Commun. 2018, 9, 382. [Google Scholar] [CrossRef] [Green Version]
- Chen, S.; Tong, X.; Woodard, R.W.; Du, G.; Wu, J.; Chen, J. Identification and characterization of bacterial cutinase. J. Biol. Chem. 2008, 283, 25854–25862. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Palm, G.J.; Reisky, L.; Böttcher, D.; Müller, H.; Michels, E.A.; Walczak, M.C.; Berndt, L.; Weiss, M.S.; Bornscheuer, U.T.; Weber, G. Structure of the plastic-degrading Ideonella sakaiensis MHETase bound to a substrate. Nat. Commun. 2019, 10, 1717. [Google Scholar] [CrossRef] [PubMed]
- Matak, M.Y.; Moghaddam, M.E. The role of short-range Cys171–Cys178 disulfide bond in maintaining cutinase active site integrity: A molecular dynamics simulation. Biochem. Biophys. Res. Commun. 2009, 390, 201–204. [Google Scholar] [CrossRef] [PubMed]
- Egmond, M.R.; De Vlieg, J. Fusarium solani pisi cutinase. Biochimie 2000, 82, 1015–1021. [Google Scholar] [CrossRef]
- Cunha, M.T.; Costa, M.J.; Calado, C.R.; Fonseca, L.P.; Aires-Barros, M.R.; Cabral, J.M. Integration of production and aqueous two-phase systems extraction of extracellular Fusarium solani pisi cutinase fusion proteins. J. Biotechnol. 2003, 100, 55–64. [Google Scholar] [CrossRef]
- Longhi, S.; Cambillau, C. Structure-activity of cutinase, a small lipolytic enzyme. Biochim. Biophys. Acta 1999, 1441, 185–196. [Google Scholar] [CrossRef]
- Gindro, K.; Pezet, R. Purification and characterization of a 40.8-kDa cutinase in ungerminated conidia of Botrytis cinerea Pers.: Fr. FEMS Microbiol. Lett. 1999, 171, 239–243. [Google Scholar] [CrossRef]
- Chen, Z.; Franco, C.F.; Baptista, R.P.; Cabral, J.M.S.; Coelho, A.V.; Rodrigues, C.J.; Melo, E.P. Purification and identification of cutinases from Colletotrichum kahawae and Colletotrichum gloeosporioides. Appl. Microbiol. Biotechnol. 2007, 73, 1306–1313. [Google Scholar] [CrossRef]
- Wang, G.Y.; Michailides, T.J.; Hammock, B.D.; Lee, Y.M.; Bostock, R.M. Molecular cloning, characterization, and expression of a redox-responsive cutinase from Monilinia fructicola (Wint.) Honey. Fungal Genet. Biol. 2002, 35, 261–276. [Google Scholar] [CrossRef] [Green Version]
- Wang, G.Y.; Michailides, T.J.; Hammock, B.D.; Lee, Y.M.; Bostock, R.M. Affinity purification and characterization of a cutinase from the fungal plant pathogen Monilinia fructicola (Wint.) honey. Arch. Biochem. Biophys. 2000, 382, 31–38. [Google Scholar] [CrossRef] [Green Version]
- Sweigard, J.A.; Carroll, A.M.; Farrall, L.; Chumley, F.G.; Valent, B. Magnaporthe grisea pathogenicity genes obtained through insertional mutagenesis. Mol. Plant Microbe Interact. 1998, 11, 404–412. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Maeda, H.; Yamagata, Y.; Abe, K.; Hasegawa, F.; Machida, M.; Ishioka, R.; Gomi, K.; Nakajima, T. Purification and characterization of a biodegradable plastic-degrading enzyme from Aspergillus oryzae. Appl. Microbiol. Biotechnol. 2005, 67, 778–788. [Google Scholar] [CrossRef] [PubMed]
- Xu, H.; Yan, Q.; Duan, X.; Yang, S.; Jiang, Z. Characterization of an acidic cold-adapted cutinase from Thielavia terrestris and its application in flavor ester synthesis. Food Chem. 2015, 188, 439–445. [Google Scholar] [CrossRef] [PubMed]
- Yang, S.; Xu, H.; Yan, Q.; Liu, Y.; Zhou, P.; Jiang, Z. A low molecular mass cutinase of Thielavia terrestris efficiently hydrolyzes poly(esters). J. Ind. Microbiol. Biotechnol. 2013, 40, 217–226. [Google Scholar] [CrossRef]
- Rogers, L.M.; Flaishman, M.A.; Kolattukudy, P.E. Cutinase gene disruption in Fusarium solani f sp pisi decreases its virulence on pea. Plant Cell 1994, 6, 935–945. [Google Scholar] [PubMed] [Green Version]
- Li, D.; Ashby, A.M.; Johnstone, K. Molecular Evidence that the Extracellular Cutinase Pbc1 Is Required for Pathogenicity of Pyrenopeziza brassicae on Oilseed Rape. Mol. Plant-Microbe Interact. 2003, 16, 545–552. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Liu, T.; Hou, J.; Wang, Y.; Jin, Y.; Borth, W.; Zhao, F.; Liu, Z.; Hu, J.; Zuo, Y. Genome-wide identification, classification and expression analysis in fungal–plant interactions of cutinase gene family and functional analysis of a putative ClCUT7 in Curvularia lunata. Mol. Genet. Genom. 2016, 291, 1105–1115. [Google Scholar] [CrossRef] [PubMed]
- Wang, Y.; Chen, J.; Li, D.-W.; Zheng, L.; Huang, J. CglCUT1 gene required for cutinase activity and pathogenicity of Colletotrichum gloeosporioides causing anthracnose of Camellia oleifera. Eur. J. Plant Pathol. 2017, 147, 103–114. [Google Scholar] [CrossRef]
- Sweigard, J.A.; Chumley, F.G.; Valent, B. Disruption of a Magnaporthe grisea cutinase gene. Mol. Gen. Genet. 1992, 232, 183–190. [Google Scholar] [CrossRef]
- Skamnioti, P.; Gurr, S.J. Magnaporthe grisea Cutinase2 Mediates Appressorium Differentiation and Host Penetration and Is Required for Full Virulence. Plant Cell 2007, 19, 2674–2689. [Google Scholar] [CrossRef] [Green Version]
- Wilson, R.A.; Talbot, N.J. Under pressure: Investigating the biology of plant infection by Magnaporthe oryzae. Nat. Rev. Microbiol. 2009, 7, 185–195. [Google Scholar] [CrossRef] [PubMed]
- Martinez, C.; De Geus, P.; Lauwereys, M.; Matthyssens, G.; Cambillau, C. Fusarium solani cutinase is a lipolytic enzyme with a catalytic serine accessible to solvent. Nature 1992, 356, 615–618. [Google Scholar] [CrossRef] [PubMed]
- Nikolaivits, E.; Kanelli, M.; Dimarogona, M.; Topakas, E. A Middle-Aged Enzyme Still in Its Prime: Recent Advances in the Field of Cutinases. Catalysts 2018, 8, 612. [Google Scholar] [CrossRef] [Green Version]
- Nishida, H.; Tokiwa, Y. Effects of higher-order structure of poly (3-hydroxybutyrate) on its biodegradation. II. Effects of crystal structure on microbial degradation. J. Environ. Polym. Degrad. 1993, 1, 65–80. [Google Scholar] [CrossRef]
- Abe, H.; Doi, Y. Structural effects on enzymatic degradabilities for poly[(R)-3-hydroxybutyric acid] and its copolymers. Int. J. Biol. Macromol. 1999, 25, 185–192. [Google Scholar] [CrossRef]
- Seretoudi, G.; Bikiaris, D.N.; Panayiotou, C. Synthesis, characterization and biodegradability of poly(ethylene succinate)/poly+µ-caprolactone) block copolymers. Polymer 2002, 43, 5405–5415. [Google Scholar] [CrossRef]
- Pudack, C.; Stepanski, M.; Fässler, P. PET Recycling—Contributions of Crystallization to Sustainability. Chem. Ing. Tech. 2020, 92, 452–458. [Google Scholar] [CrossRef] [Green Version]
- Mateo, C.; Palomo, J.M.; Fernandez-Lorente, G.; Guisan, J.M.; Fernandez-Lafuente, R. Improvement of enzyme activity, stability and selectivity via immobilization techniques. Enzym. Microb. Technol. 2007, 40, 1451–1463. [Google Scholar] [CrossRef]
- Kumari, V.; Kumar, S.; Kaur, I.; Bhalla, T.C. Graft copolymerization of acrylamide on chitosan-co-chitin and its application for immobilization of Aspergillus sp. RL2Ct cutinase. Bioorganic Chem. 2017, 70, 34–43. [Google Scholar] [CrossRef] [PubMed]
- Sousa, I.T.; Lourenço, N.M.T.; Afonso, C.A.M.; Taipa, M.A. Protein stabilization with a dipeptide-mimic triazine-scaffolded synthetic affinity ligand. J. Mol. Recognit. 2013, 26, 104–112. [Google Scholar] [CrossRef]
- Wang, Z.; Su, T.; Zhao, J. Immobilization of Fusarium solani Cutinase onto Magnetic Genipin-Crosslinked Chitosan Beads. Catalysts 2021, 11, 1158. [Google Scholar] [CrossRef]
- Zhang, C.; Zeng, G.; Huang, D.; Lai, C.; Huang, C.; Li, N.; Xu, P.; Cheng, M.; Zhou, Y.; Tang, W. Combined removal of di (2-ethylhexyl) phthalate (DEHP) and Pb (II) by using a cutinase loaded nanoporous gold-polyethyleneimine adsorbent. Rsc Adv. 2014, 4, 55511–55518. [Google Scholar] [CrossRef]
- Obregón, W.D.; Cisneros, J.S.; Ceccacci, F.; Quiroga, E. A highly stable biocatalyst obtained from covalent immobilization of a non-commercial cysteine phytoprotease. J. Bioprocess. Biotech. 2015, 5, 1000211. [Google Scholar]
- Gupta, M. Thermostabilization of proteins. Biotechnol. Appl. Biochem. (USA) 1991, 14, 1–11. [Google Scholar]
- Ma, M.M.; Wang, L.Y.; Zhu, H.Y. Enzymatic degradation of polyester-nanoparticles by lipases and adsorption of lipases on the polyester-nanoparticles. In Advanced Materials Research; Trans Tech Publications: Stafa-Zurich, Switzerland, 2012. [Google Scholar]
- Wang, X.; Lu, D.; Jönsson, L.J.; Hong, F. Preparation of a PET-Hydrolyzing Lipase from Aspergillus oryzae by the Addition of Bis(2-hydroxyethyl) Terephthalate to the Culture Medium and Enzymatic Modification of PET Fabrics. Eng. Life Sci. 2008, 8, 268–276. [Google Scholar] [CrossRef]
- Carniel, A.; Valoni, É.; Nicomedes, J.; Gomes, A.d.C.; Castro, A.M.d. Lipase from Candida antarctica (CALB) and cutinase from Humicola insolens act synergistically for PET hydrolysis to terephthalic acid. Process Biochem. 2017, 59, 84–90. [Google Scholar] [CrossRef]
- de Castro, A.M.; Carniel, A.; Nicomedes Junior, J.; da Conceição Gomes, A.; Valoni, É. Screening of commercial enzymes for poly(ethylene terephthalate) (PET) hydrolysis and synergy studies on different substrate sources. J. Ind. Microbiol. Biotechnol. 2017, 44, 835–844. [Google Scholar] [CrossRef] [PubMed]
- Henderson, B.; Curtis, M.; Seymour, R.; Donos, N. Periodontal Medicine and Systems Biology; John Wiley & Sons: Hoboken, NJ, USA, 2009. [Google Scholar]
- Garza, D.R.; Dutilh, B.E. From cultured to uncultured genome sequences: Metagenomics and modeling microbial ecosystems. Cell. Mol. Life Sci. 2015, 72, 4287–4308. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Parages, M.L.; Gutiérrez-Barranquero, J.A.; Reen, F.J.; Dobson, A.D.; O’Gara, F. Integrated (Meta) Genomic and Synthetic Biology Approaches to Develop New Biocatalysts. Mar. Drugs 2016, 14, 62. [Google Scholar] [CrossRef]
- Danso, D.; Schmeisser, C.; Chow, J.; Zimmermann, W.; Wei, R.; Leggewie, C.; Li, X.; Hazen, T.; Streit, W.R.; Parales, R.E. New Insights into the Function and Global Distribution of Polyethylene Terephthalate (PET)-Degrading Bacteria and Enzymes in Marine and Terrestrial Metagenomes. Appl. Environ. Microbiol. 2018, 84, e02773-17. [Google Scholar] [CrossRef] [Green Version]
- Meilleur, C.; Hupé, J.-F.; Juteau, P.; Shareck, F. Isolation and characterization of a new alkali-thermostable lipase cloned from a metagenomic library. J. Ind. Microbiol. Biotechnol. 2009, 36, 853–861. [Google Scholar] [CrossRef] [PubMed]
- Baumann, P.; Baumann, L.; Bang, S.S.; Woolkalis, M.J. Reevaluation of the taxonomy ofVibrio, beneckea, and Photobacterium: Abolition of the genus Beneckea. Curr. Microbiol. 1980, 4, 127–132. [Google Scholar] [CrossRef]
- Dodd, D.; Mackie, R.I.; Cann, I.K.O. Xylan degradation, a metabolic property shared by rumen and human colonic Bacteroidetes. Mol. Microbiol. 2011, 79, 292–304. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Thomas, F.; Hehemann, J.-H.; Rebuffet, E.; Czjzek, M.; Michel, G. Environmental and Gut Bacteroidetes: The Food Connection. Front. Microbiol. 2011, 2, 93. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Foley, M.H.; Cockburn, D.W.; Koropatkin, N.M. The Sus operon: A model system for starch uptake by the human gut Bacteroidetes. Cell. Mol. Life Sci. 2016, 73, 2603–2617. [Google Scholar] [CrossRef] [Green Version]
- Zhou, D.; Chen, J.; Wu, J.; Yang, J.; Wang, H. Biodegradation and catalytic-chemical degradation strategies to mitigate microplastic pollution. Sustain. Mater. Technol. 2021, 28, e00251. [Google Scholar] [CrossRef]
- Furukawa, M.; Kawakami, N.; Oda, K.; Miyamoto, K. Acceleration of enzymatic degradation of poly (ethylene terephthalate) by surface coating with anionic surfactants. ChemSusChem 2018, 11, 4018–4025. [Google Scholar] [CrossRef] [PubMed]

Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Soong, Y.-H.V.; Sobkowicz, M.J.; Xie, D. Recent Advances in Biological Recycling of Polyethylene Terephthalate (PET) Plastic Wastes. Bioengineering 2022, 9, 98. https://doi.org/10.3390/bioengineering9030098
Soong Y-HV, Sobkowicz MJ, Xie D. Recent Advances in Biological Recycling of Polyethylene Terephthalate (PET) Plastic Wastes. Bioengineering. 2022; 9(3):98. https://doi.org/10.3390/bioengineering9030098
Chicago/Turabian StyleSoong, Ya-Hue Valerie, Margaret J. Sobkowicz, and Dongming Xie. 2022. "Recent Advances in Biological Recycling of Polyethylene Terephthalate (PET) Plastic Wastes" Bioengineering 9, no. 3: 98. https://doi.org/10.3390/bioengineering9030098