1. Introduction
Acute myeloid leukemia (AML) is the deadliest of the four types of leukemia, accounting for 11,000 annual deaths in the US with an average five-year survival rate of 28.7% [
1]. AML is characterized by the overproduction and accumulation of immature leukocytes, specifically myeloid precursors, in the bone marrow and peripheral blood. The immature white blood cells prevent the functions of the bone marrow, including the production of red blood cells and platelets, which makes the immune system vulnerable [
2,
3]. Detecting and classifying immature leukocytes is crucial for the diagnosis of AML.
Progressing rapidly, AML can be fatal within months or even weeks if not diagnosed and treated immediately [
4]. Hence, accurate and quick diagnosis is necessary for AML patients. Microscopic examination of peripheral blood smears is the standard procedure for the diagnosis of leukemia, but other procedures are also used [
5]. Manual blood smear examination is labor intensive and time consuming [
6]. Moreover, manual examination is prone to considerable inter- and intra-observer variation of standards, as well as biases such as tiredness and operator experience [
7]. Depending on the experience of the hematologist, manual examination has an error rate of 30% to 40% [
8].
In developing countries such as Nicaragua, diagnosis takes 29 days to be reached due to lack of access to healthcare and physician delay [
9,
10,
11]. The current method of diagnosis is unsatisfactory and a quick, accurate method is required. An automated approach will enable standardized and efficient screening for immature leukocytes, thus overcoming the limitations of the current manual method for diagnosis, especially in developing countries. Since different types of leukocytes vary in cytomorphology, detection and classification of immature leukocytes can be formulated as a machine learning classification task based on morphological features [
12]. Previous studies on the computer-aided detection of leukemia have mainly focused on acute lymphoblastic leukemia (ALL) [
12]. Abdeldaim et al. [
13] used the ALL Image Database (ALL-IDB) [
14] to train a k-nearest neighbors (k-NN) classifier with 95.99% accuracy for classification of ALL subtypes. Classification performance was improved by Shafique and Tehsin [
15], who applied a convolutional neural network (CNN) on images from the ALL-IDB repository and achieved 99.50% accuracy for detection of ALL with 96.74% accuracy for classification. Research aiming to detect and classify AML has obtained lower performance compared to studies on ALL due to the high diversity in cytomorphology of AML cells [
12].
A multitude of segmentation techniques, morphological features, and machine learning classifiers have been employed in the literature. Kazemi et al. [
16] segmented 165 images of four subtypes of AML cells with k-means clustering. Using a support vector machine (SVM), 95% accuracy was obtained for detection of AML cells and 87% accuracy was obtained for classification into four of the eight French-American-British (FAB) subtypes of AML. The study utilized 60 cytomorphological features for classification, yet the most important features were not established. E.S. Wiharto et al. [
17] selected three morphological features and calculated the importance of each feature. Evidently, there is a lack of standardization in the number and type of features used for selection, which needs to be addressed. Classification into two FAB subtypes was obtained with 67.28% accuracy [
17]. Harjoko et al. [
18] used active contour segmentation, extracted six morphological features, and used the momentum backpropagation neural network to classify three subtypes of AML with 93.57% accuracy. Despite the high accuracy, the proposed model was limited by precision and sensitivity values below 85%. W. Wiharto et al. [
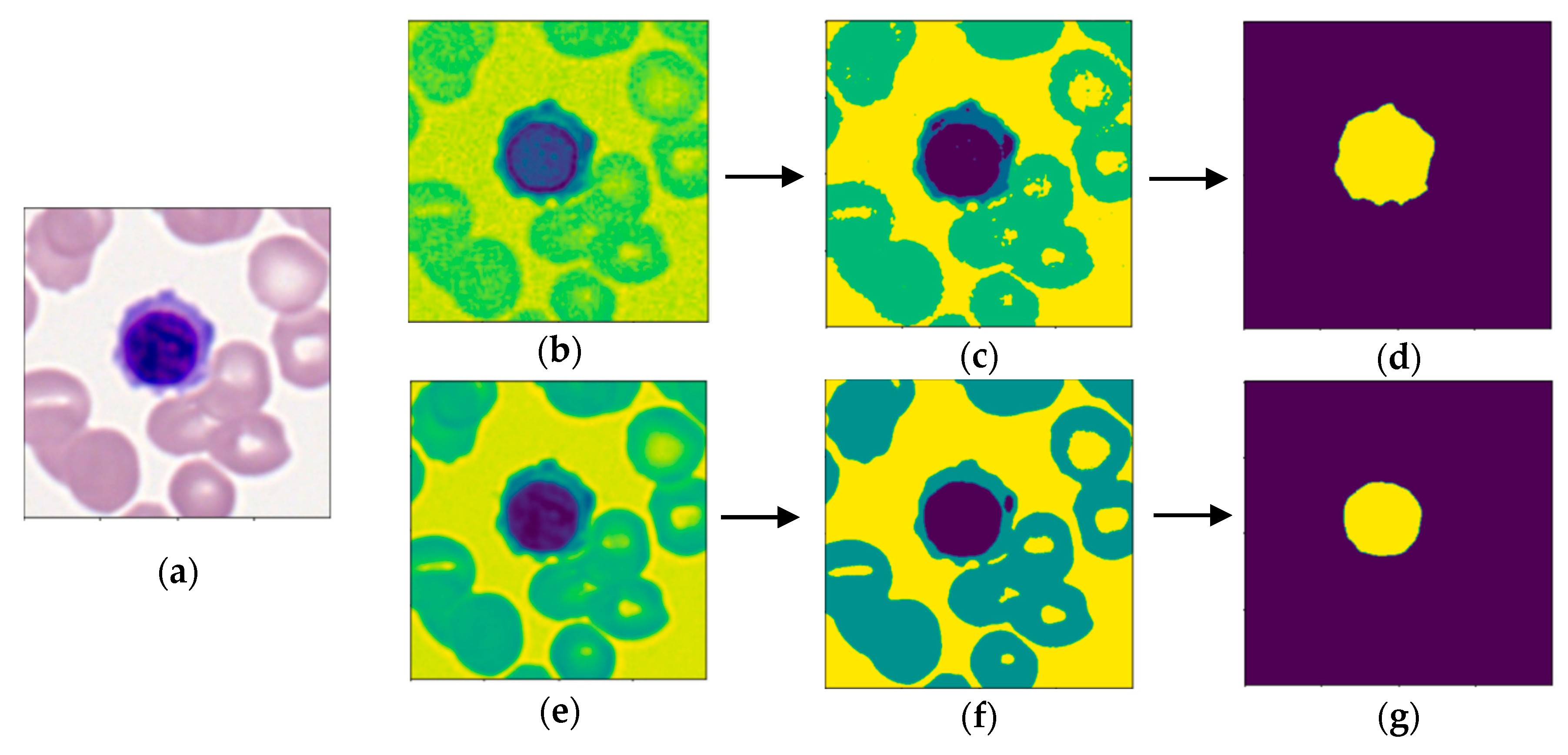
19] classified three immature leukocytes in AML cells from a small dataset of 50 images. After segmentation through Otsu thresholding, which is a common method used in literature, three morphological characteristics were extracted and ranked based on importance for classification. To overcome the imbalance of data, synthetic minority oversampling technique (SMOTE) was employed with a random forest algorithm, which obtained 90% accuracy. The research displayed that imbalanced data, which has limited many previous models, can be overcome through selection and tuning of a random forest classifier. Matek et al. [
12] assembled an image dataset of 18,365 leukocytes [
20,
21] and employed a CNN for classification. For binary classification between immature and mature blood cells, the CNN obtained an area under curve (AUC) of the receiver operating characteristic (ROC) of 0.992, which is the current state of art. Despite high performance in detection of immature leukocytes, the CNN achieved precision of below 65% for the majority of immature leukocyte classes, which was attributed to the imbalance of data across different classes. Overall, previous studies are limited by the use of small data sets, which may lead to overfitting, and imbalance across classes. In addition, given the importance of feature selection in machine learning classifiers, the lack of uniformity in the type of features used for classification of AML cells still needs to be addressed by identifying the most important features.
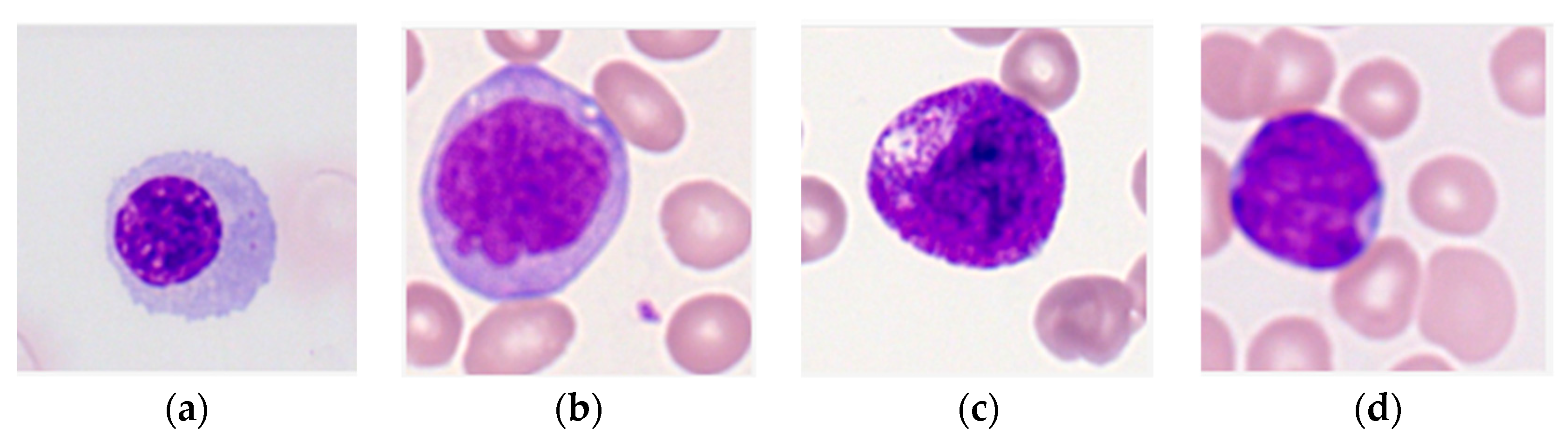
The purpose of this research is to develop a model capable of accurately detecting and classifying immature leukocytes in AML cells from an imbalanced dataset into four types (erythroblasts, monoblasts, promyelocytes, and myeloblasts) with a random forest algorithm. Detection and classification of immature leukocytes will greatly aid the clinical diagnosis of AML. To add to the limited set of color features used for classification of leukocytes [
16], two new features for classification of leukocytes, specifically the average and standard deviation of nucleus color intensity in the B channel of LAB color space, are proposed and demonstrated to be discriminative. Furthermore, the most important features for both detection and classification are calculated and ranked using the Gini importance, which is defined as the loss of Gini impurity caused by each feature in the random forest. To the best of the authors’ knowledge, this is the first study that calculates the Gini importance of a multitude of morphological features for classification of leukocytes in AML.
4. Conclusions
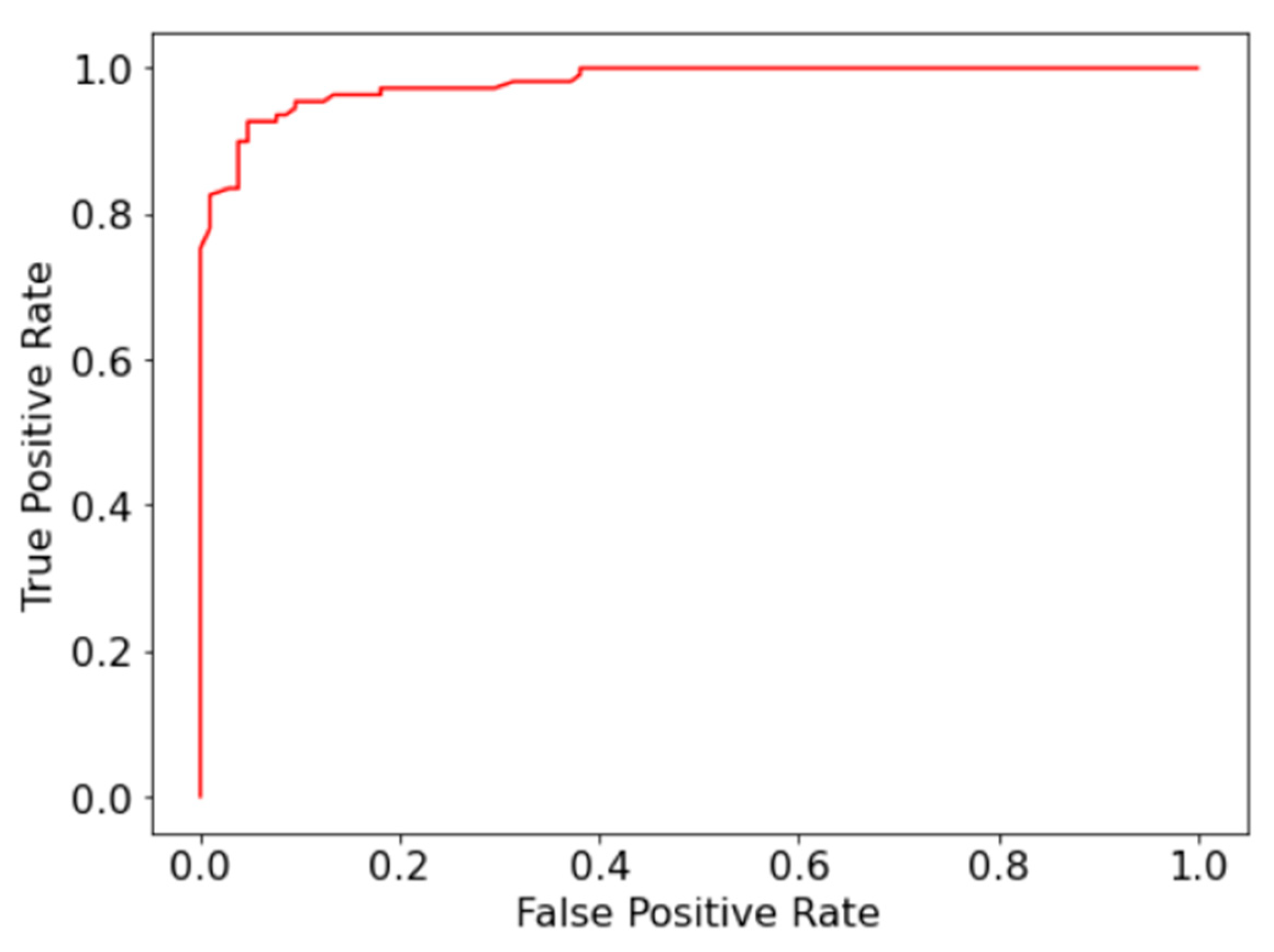
To overcome the limitations of the manual diagnosis methodology for AML, a random forest model for automatic detection and classification of immature leukocytes was presented. The model was capable of detecting immature leukocytes with 93% accuracy and 0.98 AUC-ROC, which is on par with the current state of art [
12]. Furthermore, the model achieved precision of above 65% for each of the four immature leukocyte classes during multiclass classification, despite imbalance in numbers across classes, which is an improvement over previous research. Using Gini importance, N:C ratio was determined to be significant for both detection and classification, while the proposed color features of the nucleus in the B channel of LAB color space were calculated to be important for classification.
Applications of the study are two-fold. While the proposed model cannot diagnose AML alone, it can be used as an effective support tool for doctors to reduce the time and cost required for the diagnosis of AML. The high accuracy of the model in binary classification demonstrates that the model can serve as an efficient screening tool, which can rapidly identify potentially cancerous cells for further examination by a doctor [
36,
37,
38]. The proposed model can expedite the detection of AML by identifying immature leukocytes, especially in developing countries where diagnosis takes numerous weeks, and potentially save lives because early diagnosis is vital for treatment success in AML patients [
9,
39]. In addition, the precise classification of immature leukocytes can aid in treatment and prognosis decisions, which differ based on the type of cancerous cell [
40,
41]. The second application of this study is in future research, where the features calculated to be most important and the proposed features can be used to elevate the classification performance.
An important future direction is to gather a comprehensive dataset and develop a machine learning classifier that can classify all the types of immature leukocytes and work with imbalanced data. Future studies can expand on this work by calculating and ranking the importance of additional morphological features for the classification of leukocytes. Improving the discrimination between similar cell types, such as myeloblasts and promyelocytes, is also an avenue for future work. The difficulty of differentiating myeloblasts and promyelocytes can potentially be overcome by identifying features that are especially discriminative for the two cell types and training a specialized model to discriminate between the two cell types. Research on leukemia detection has obtained very promising results, and further work is required to develop systems that can be completely integrated into the clinical diagnosis method. Contributions of this study are an accurate model for detecting and classifying immature leukocytes, as well as calculation of the most important morphological features, which provide a basis for future research on computer-aided diagnosis of leukemia.