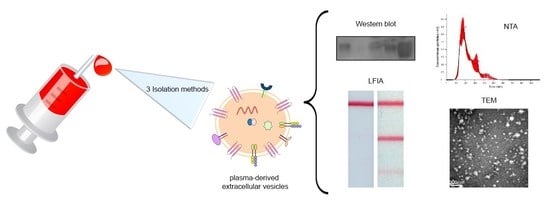
Characterization of Plasma-Derived Extracellular Vesicles Isolated by Different Methods: A Comparison Study
Abstract
1. Introduction
2. Materials and Methods
2.1. Plasma Samples
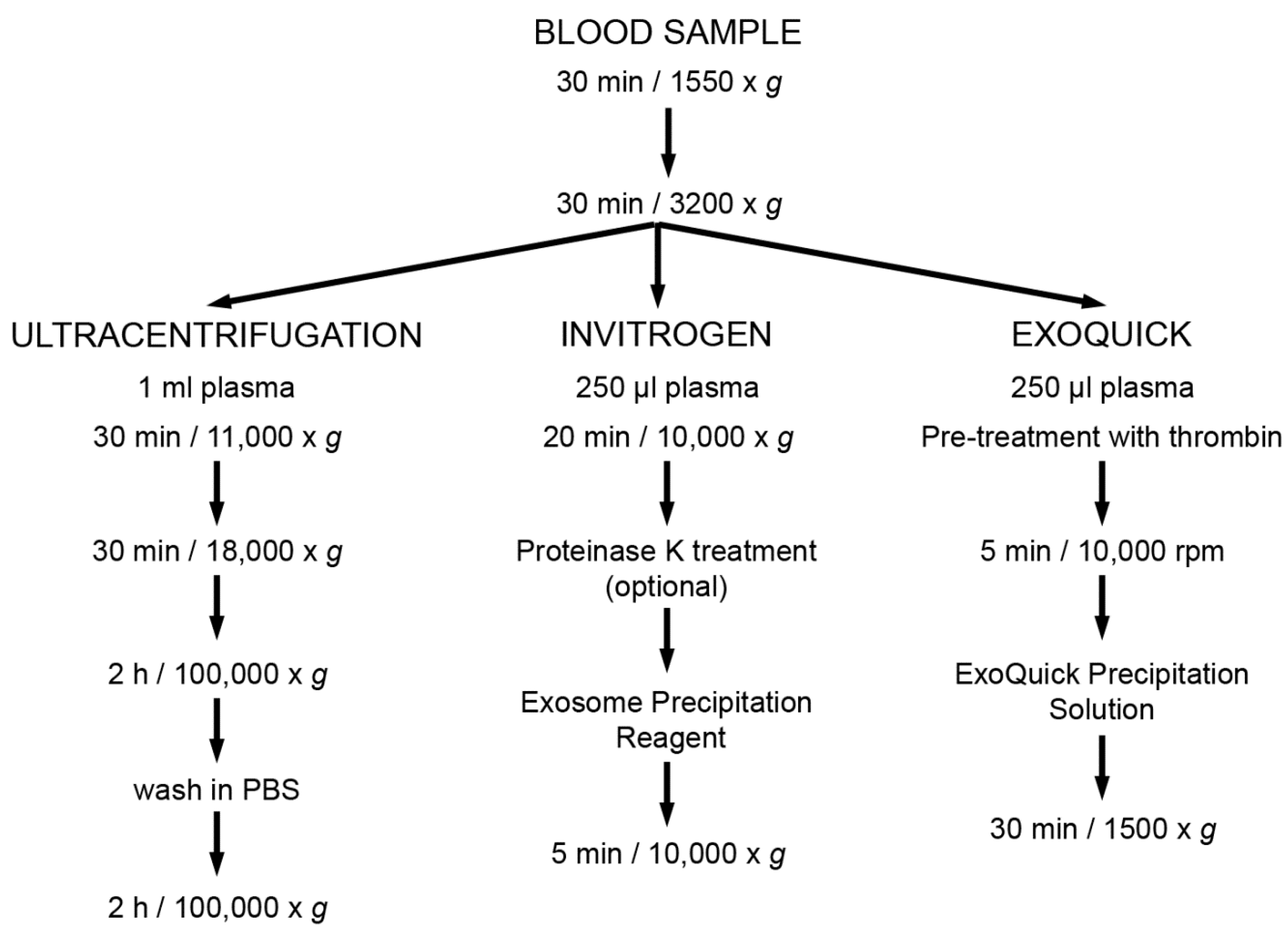
2.2. Enrichment of Extracellular Vesicles
2.2.1. Ultracentrifugation
2.2.2. EV Enrichment Based on Precipitation Reagents
2.3. Assessment of Total Protein Content
2.4. Western Blot
2.5. Dynamic Light Scattering (DLS) and Nanoparticle Tracking Analysis (NTA)
2.6. Transmission Electron Microscopy (TEM)
2.7. Lateral Flow Immunoassay (LFIA)
2.8. Statistical Analysis
3. Results
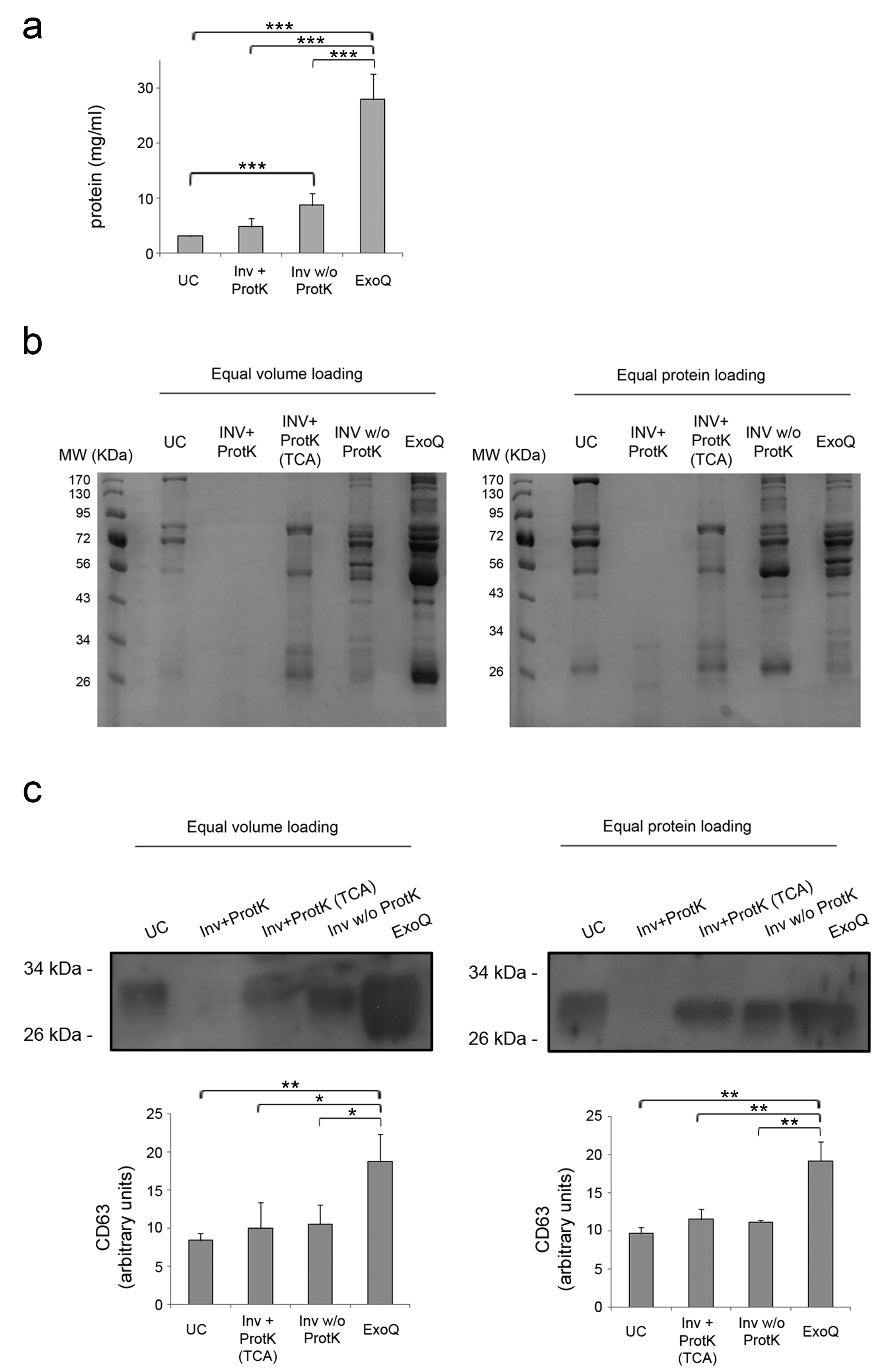
3.1. Yield of Extracellular Vesicles Enrichment from Plasma by Ultracentrifugation and Precipitation Reagents
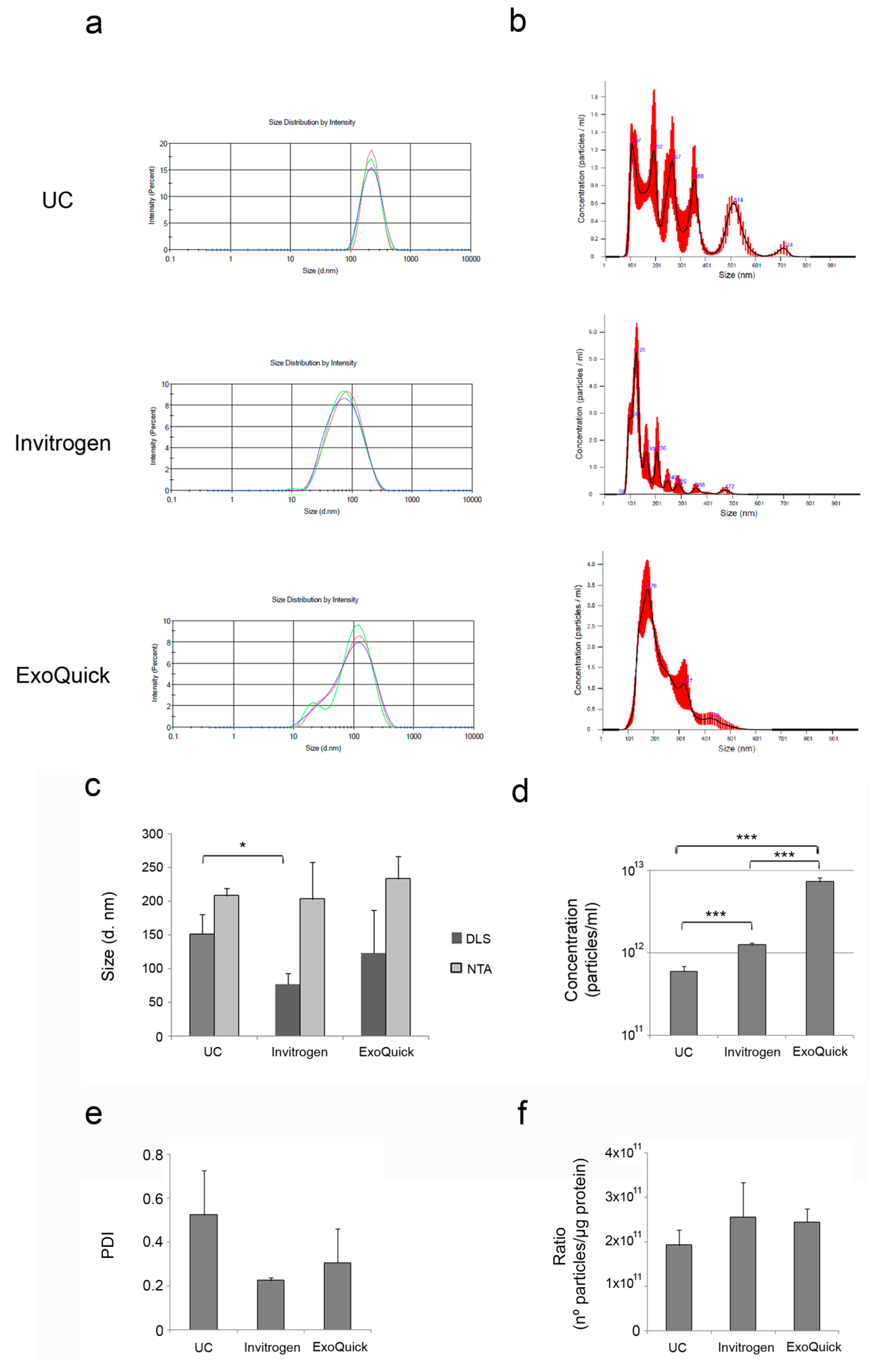
3.2. Characterization of Plasma-Derived Extracellular Vesicles
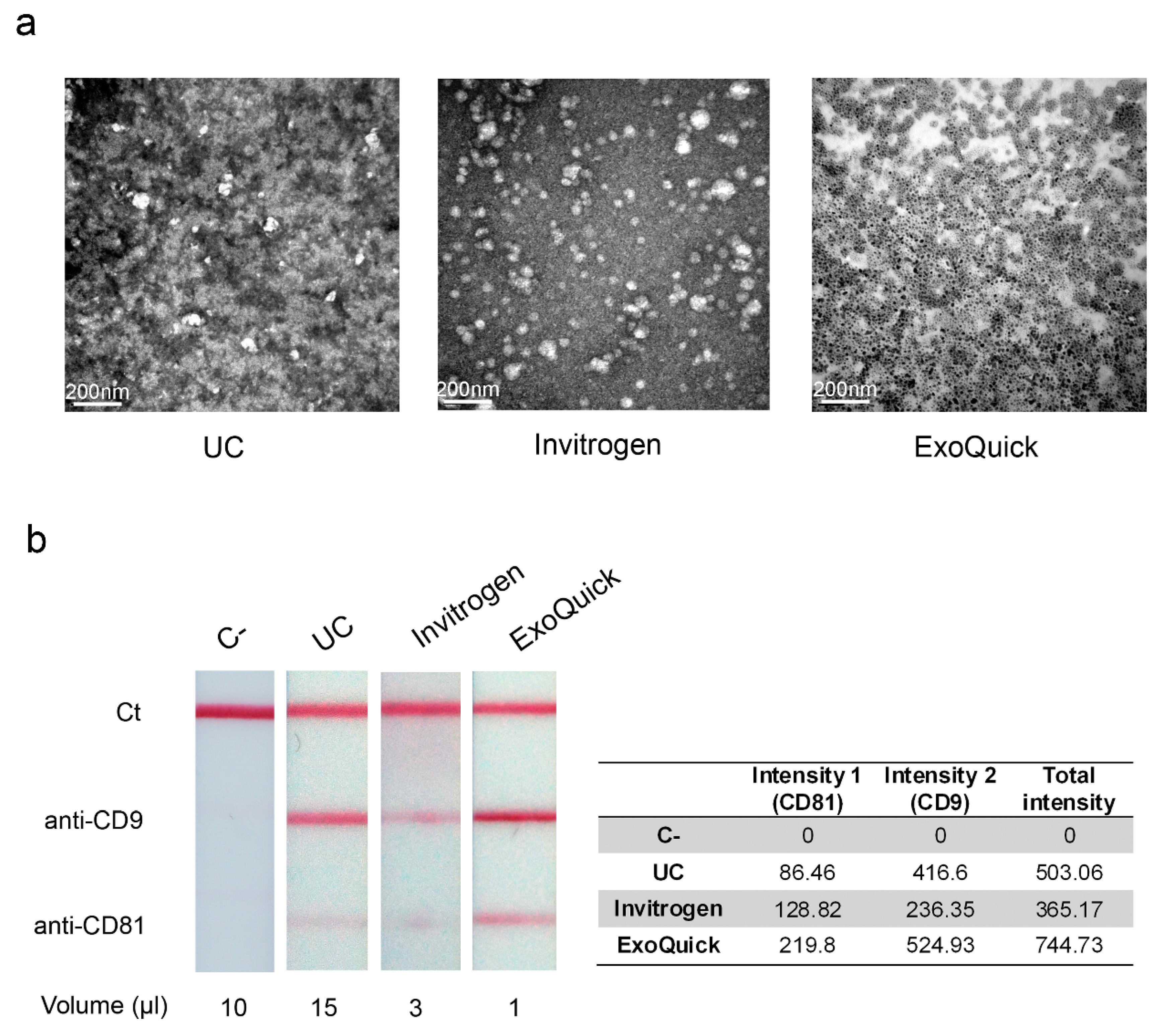
3.3. Electron Microscopy Observation and Detection of Extracellular Vesicles by LFIA
4. Discussion
Author Contributions
Funding
Conflicts of Interest
Abbreviations
References
- Raposo, G.; Stoorvogel, W. Extracellular vesicles: Exosomes, microvesicles, and friends. J. Cell Biol. 1999, 200, 373–383. [Google Scholar] [CrossRef] [PubMed]
- El Andaloussi, S.; Mäger, I.; Breakefield, X.O.; Wood, M.J. Extracellular vesicles: Biology and emerging therapeutic opportunities. Nat. Rev. Drug Discov. 2013, 12, 347–357. [Google Scholar] [CrossRef] [PubMed]
- Cocucci, E.; Meldolesi, J. Ectosomes and exosomes: Shedding the confusion between extracellular vesicles. Trends Cell Biol. 2015, 25, 364–372. [Google Scholar] [CrossRef] [PubMed]
- Xu, R.; Greening, D.W.; Zhu, H.J.; Takahashi, N.; Simpson, R.J. Extracellular vesicle isolation and characterization: Toward clinical application. J. Clin. Investig. 2016, 126, 1152–1162. [Google Scholar] [CrossRef] [PubMed]
- Van Dommelen, S.M.; Vader, P.; Lakhal, S.; Kooijman, S.A.A.; van Solinge, W.W.; Wood, M.J.A.; Schiffelers, R.M. Microvesicles and exosomes: Opportunities for cell-derived membrane vesicles in drug delivery. J. Control Release 2012, 161, 635–644. [Google Scholar] [CrossRef]
- Batrakova, E.V.; Kim, M.S. Using exosomes, naturally-equipped nanocarriers, for drug delivery. J. Control Release 2015, 219, 396–405. [Google Scholar] [CrossRef]
- Baranyai, T.; Herczeg, K.; Onódi, Z.; Voszka, I.; Módos, K.; Marton, M.; Nagy, G.; Mäger, I.; Wood, M.J.; El Andaloussi, S.; et al. Isolation of exosomes from blood plasma: Qualitative and quantitative comparison of ultracentrifugation and size exclusion chromatography methods. PLoS ONE 2015, 10, e0145686. [Google Scholar] [CrossRef]
- Taylor, D.D.; Gercel-Taylor, C. MicroRNA signatures of tumor-derived exosomes as diagnostic biomarkers of ovarian cancer. Gynecol. Oncol. 2008, 110, 13–21. [Google Scholar]
- Shi, M.; Liu, C.; Cook, T.J.; Bullock, K.M.; Zhao, Y.; Ginghina, C.; Li, Y.; Aro, P.; Dator, R.; He, C.; et al. Plasma exosomal α-synuclein is likely CNS-derived and increased in Parkinson’s disease. Acta Neuropathol. 2014, 128, 639–650. [Google Scholar] [CrossRef]
- Helwa, I.; Cai, J.; Drewry, M.D.; Zimmerman, A.; Dinkins, M.B.; Khaled, M.L.; Seremwe, M.; Dismuke, W.M.; Bieberich, E.; Stamer, W.D.; et al. A comparative study of serum exosome isolation using differential ultracentrifugation and three commercial reagents. PLoS ONE 2017, 12, e0170628. [Google Scholar] [CrossRef]
- Li, P.; Kaslan, M.; Lee, S.H.; Yao, J.; Gao, Z. Progress in exosome isolation techniques. Theranostics 2017, 7, 789–804. [Google Scholar] [CrossRef]
- Silva, J.; García, V.; Rodríguez, M.; Compte, M.; Cisneros, E.; Veguillas, P.; Garcia, J.M.; Dominguez, G.; Campos-Martin, Y.; Cuevas, J.; et al. Analysis of exosome release and its prognostic value in human colorectal cancer. Genes Chromosomes Cancer 2012, 51, 409–418. [Google Scholar] [CrossRef] [PubMed]
- Duijvesz, D.; Verslui, C.Y.; van der Fels, C.A.; den Berg, M.S.; Leivo, J.; Peltola, M.T.; Bangma, C.H.; Pettersson, K.S.; Jenster, G. Immuno-based detection of extracellular vesicles in urine as diagnostic marker for prostate cancer. Int. J. Cancer 2015, 137, 2869–2878. [Google Scholar] [CrossRef]
- Skog, J.; Wurdinger, T.; van Rijn, S.; Meijer, D.; Gainche, L.; Sena-Esteves, M.; Carter, B.S.; Krichevsky, A.M.; Breakefield, X.O. Glioblastoma microvesicles transport RNA and proteins that promote tumour growth and provide diagnostic biomarkers. Nat. Cell Biol. 2008, 10, 1470–1476. [Google Scholar] [CrossRef] [PubMed]
- Li, P.; Qin, C. Elevated circulating VE-cadherin+CD144+endothelial microparticles in ischemic cerebrovascular disease. Thromb. Res. 2015, 135, 375–381. [Google Scholar] [CrossRef] [PubMed]
- Canella, A.; Harshman, S.W.; Radomska, H.S.; Freitas, M.A.; Pichiorri, F. The potential diagnostic power of extracellular vesicle analysis for multiple myeloma. Expert Rev. Mol. Diagn. 2016, 16, 277–284. [Google Scholar] [CrossRef] [PubMed]
- Wang, J.W.; Gijsberts, C.M.; Seneviratna, A.; de Hoog, V.C.; Vrijenhoek, J.E.P.; Schoneveld, A.H.; Chan, M.Y.; Lam, C.S.P.; Richards, A.M.; Lee, C.N.; et al. Plasma extracellular vesicle protein content for diagnosis and prognosis of global cardiovascular disease. Neth. Heart J. 2013, 21, 467–471. [Google Scholar] [CrossRef]
- Vaidyanathan, R.; Naghibosadat, M.; Rauf, S.; Korbie, D.; Carrascosa, L.G.; Shiddiky, M.J.; Trau, M. Detecting exosomes specifically: A multiplexed device based on alternating current electrohydrodynamic induced nanoshearing. Anal. Chem. 2014, 86, 11125–11132. [Google Scholar] [CrossRef]
- Jeong, S.; Park, J.; Panatia, D.; Castro, C.M.; Weissleder, R.; Lee, H. Integrated magneto–electrochemical sensor for exosome analysis. ACS Nano 2016, 10, 1802–1809. [Google Scholar] [CrossRef]
- Oliveira-Rodríguez, M.; Serrano-Pertierra, E.; García, A.C.; López-Martín, S.; Yañez-Mo, M.; Cernuda-Morollón, E.; Blanco-López, M.D.C. Point-of-care detection of extracellular vesicles: Sensitivity optimization and multiple-target detection. Biosens. Bioelectron. 2017, 87, 38–45. [Google Scholar] [CrossRef]
- Soo, C.Y.; Song, Y.; Zheng, Y.; Campbell, E.C.; Riches, A.C.; Gunn-Moore, F.; Powis, S.J. Nanoparticle tracking analysis monitors microvesicle and exosome secretion from immune cells. Immunology 2012, 136, 192–197. [Google Scholar] [CrossRef]
- Goldburg, W.I. Dynamic light scattering. Am. J. Phys. 1999, 8, 1152–1160. [Google Scholar] [CrossRef]
- Webber, J.; Clayton, A. How pure are your vesicles? J. Extracell. Vesicles 2013, 2, 19861. [Google Scholar] [CrossRef] [PubMed]
- Lobb, R.J.; Becker, M.; Wen, S.W.; Wong, C.S.F.; Wiegmans, A.P.; Leimgruber, A.; Möller, A. Optimized exosome isolation protocol for cell culture supernatant and human plasma. J. Extracell. Vesicles 2015, 4, 27031. [Google Scholar] [CrossRef] [PubMed]
- Oliveira-Rodríguez, M.; López-Cobo, S.; Reyburn, H.T.; Costa-García, A.; López-Martín, S.; Yáñez-Mó, M.; Cernuda-Morollón, E.; Paschen, A.; Valés-Gómez, M.; Blanco-López, M.C. Development of a rapid lateral flow immunoassay test for detection of exosomes previously enriched from cell culture medium and body fluids. J. Extracell. Vesicles 2016, 5, 31803. [Google Scholar] [CrossRef] [PubMed]
- Momen-Heravi, F.; Balaj, L.; Alian, S.; Mantel, P.; Halleck, A.E.; Trachtenberg, A.J.; Soria, C.E.; Oquin, S.; Bonebreak, C.M.; Saracoglu, E.; et al. Current methods for the isolation of extracellular vesicles. Biol. Chem. 2013, 394, 1253–1262. [Google Scholar] [CrossRef] [PubMed]
- Szatanek, R.; Baran, J.; Siedla, M.; Baj-Krzyworzeka, M. Isolation of extracellular vesicles: Determining the correct approach. Int. J. Mol. Med. 2015, 36, 11–17. [Google Scholar] [CrossRef] [PubMed]
- Sunkara, V.; Woo, H.K.; Cho, Y.K. Emerging techniques in the isolation and characterization of extracellular vesicles and their roles in cancer diagnostics and prognostics. Analyst 2016, 141, 371–381. [Google Scholar] [CrossRef] [PubMed]
- Royo, F.; Zuñiga-Garcia, P.; Sanchez-Mosquera, P.; Egia, A.; Perez, A.; Loizaga, A.; Arceo, R.; Lacasa, I.; Rabade, A.; Arrieta, E.; et al. Different EV enrichment methods suitable for clinical settings yield different subpopulations of urinary extracellular vesicles from human samples. J. Extracell. Vesicles 2016, 5, 29497. [Google Scholar] [CrossRef]
- Linares, R.; Tan, S.; Gounou, C.; Arraud, N.; Brisson, A.R. High-speed centrifugation induces aggregation of extracellular vesicles. J. Extracell. Vesicles 2015, 4, 29509. [Google Scholar] [CrossRef]
- Gercel-Taylor, C.; Atay, S.; Tullis, R.H.; Kesimer, M.; Taylor, D.D. Nanoparticle analysis of circulating cell-derived vesicles in ovarian cancer patients. Anal. Biochem. 2012, 428, 44–53. [Google Scholar] [CrossRef]
- Tauro, B.J.; Greening, D.W.; Mathias, R.A.; Ji, H.; Mathivanan, S.; Scott, A.M.; Simpson, R.J. Comparison of ultracentrifugation, density gradient separation, and immunoaffinity capture methods for isolating human colon cancer cell line LIM1863-derived exosomes. Methods 2012, 56, 293–304. [Google Scholar] [CrossRef] [PubMed]
- Caradec, J.; Kharmate, G.; Hosseini-Beheshti, E.; Adomat, H.; Gleave, M.; Guns, E. Reproducibility and efficiency of serum-derived exosome extraction methods. Clin. Biochem. 2014, 47, 1286–1292. [Google Scholar] [CrossRef] [PubMed]
- Taylor, D.D.; Zacharias, W.; Gercel-Taylor, C. Exosome isolation for proteomic analyses and RNA profiling. Methods Mol. Biol. 2011, 728, 235–246. [Google Scholar]
- Castro-Marrero, J.; Serrano-Pertierra, E.; Oliveira-Rodríguez, M.; Zaragozá, M.C.; Martínez-Martínez, A.; Blanco-López, M.C.; Alegre, J. Circulating extracellular vesicles as potential biomarkers in chronic fatigue syndrome/myalgic encephalomyelitis: An exploratory pilot study. J. Extracell. Vesicles 2018, 7, 1453730. [Google Scholar] [CrossRef] [PubMed]
| UC | Invitrogen | ExoQuick | |
|---|---|---|---|
| Starting volume | 1 mL | 250 µL | 250 µL |
| Time requirements | ~5–6 h | ~1.5 h | ~2 h |
| Protein [mg/mL] | 3.05 ± 0.19 | 4.76 ± 2.09 | 27.98 ± 4.66 |
| Suitable for SDS-PAGE/Western blot | Yes | No * | Yes |
| Suitable for dynamic light scattering (DLS) (mean diameter, nm) | Yes 152.09 ± 29.38 | Yes 76.64±16.17 | Yes 123.55 ± 63.04 |
| Suitable for nanoparticle tracking analysis (NTA) (mean diameter, nm) [particles/mL] | Yes 208.5 ± 10.60 ~1011 | Yes 203.67 ± 55.41 ~1012 | Yes 233.97 ± 33.73 ~1012 |
| Suitable for TEM | Yes | Yes | No |
| Suitable for lateral flow immunoassay (LFIA) | Yes | Yes | Yes |
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Serrano-Pertierra, E.; Oliveira-Rodríguez, M.; Rivas, M.; Oliva, P.; Villafani, J.; Navarro, A.; Blanco-López, M.C.; Cernuda-Morollón, E. Characterization of Plasma-Derived Extracellular Vesicles Isolated by Different Methods: A Comparison Study. Bioengineering 2019, 6, 8. https://doi.org/10.3390/bioengineering6010008
Serrano-Pertierra E, Oliveira-Rodríguez M, Rivas M, Oliva P, Villafani J, Navarro A, Blanco-López MC, Cernuda-Morollón E. Characterization of Plasma-Derived Extracellular Vesicles Isolated by Different Methods: A Comparison Study. Bioengineering. 2019; 6(1):8. https://doi.org/10.3390/bioengineering6010008
Chicago/Turabian StyleSerrano-Pertierra, Esther, Myriam Oliveira-Rodríguez, Montserrat Rivas, Pedro Oliva, Javier Villafani, Ana Navarro, M. Carmen Blanco-López, and Eva Cernuda-Morollón. 2019. "Characterization of Plasma-Derived Extracellular Vesicles Isolated by Different Methods: A Comparison Study" Bioengineering 6, no. 1: 8. https://doi.org/10.3390/bioengineering6010008
APA StyleSerrano-Pertierra, E., Oliveira-Rodríguez, M., Rivas, M., Oliva, P., Villafani, J., Navarro, A., Blanco-López, M. C., & Cernuda-Morollón, E. (2019). Characterization of Plasma-Derived Extracellular Vesicles Isolated by Different Methods: A Comparison Study. Bioengineering, 6(1), 8. https://doi.org/10.3390/bioengineering6010008