MARU-MTL: A Mamba-Enhanced Multi-Task Learning Framework for Continuous Blood Pressure Estimation Using Radar Pulse Waves
Abstract
1. Introduction
2. Materials and Methods
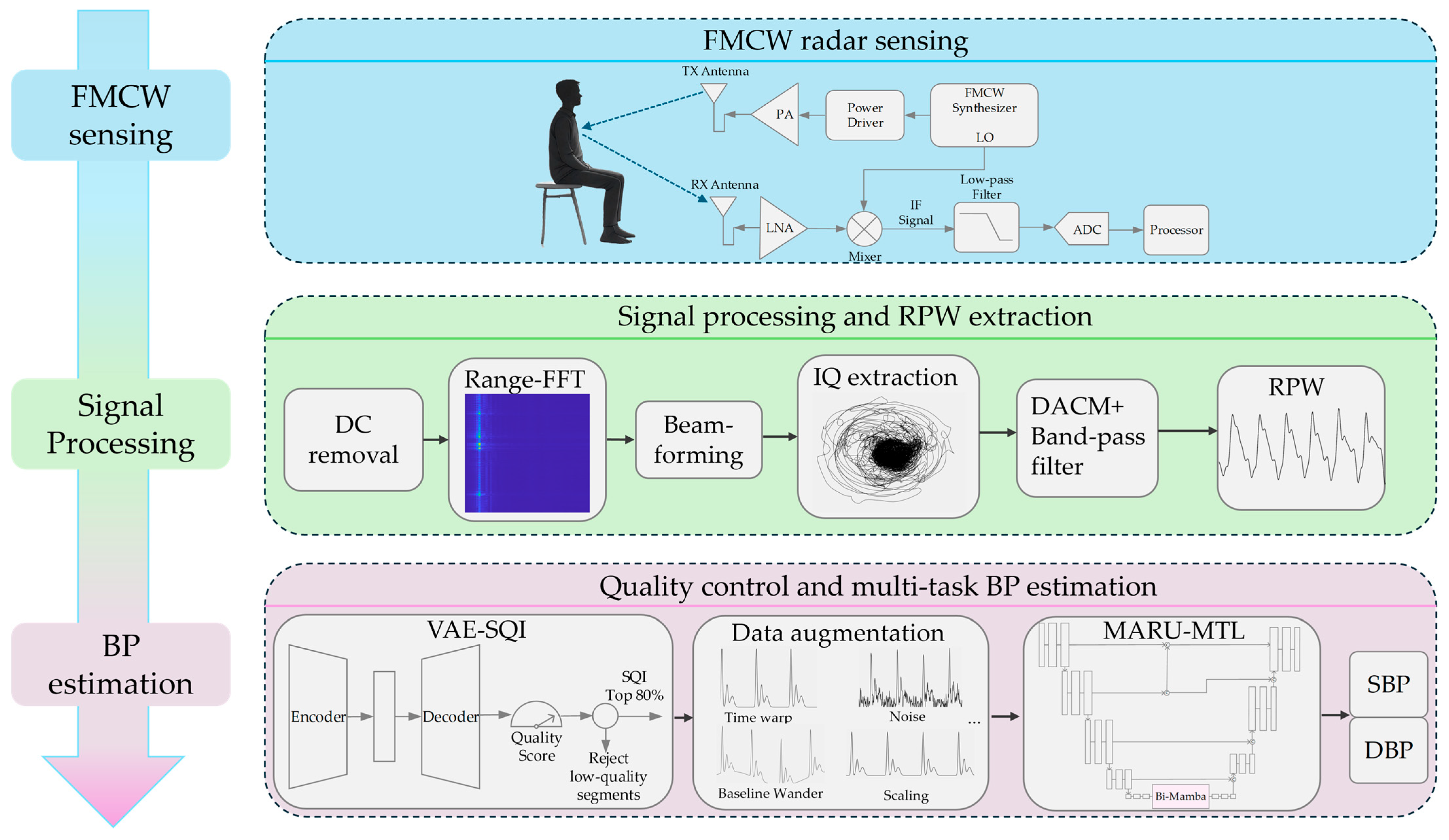
2.1. FMCW Radar Principle
2.2. Signal Processing
2.3. Unsupervised RPW Quality Assessment and Screening Based on VAE-SQI
2.4. Data Augmentation
2.5. Blood Pressure Estimation Model
2.5.1. Encoder
2.5.2. Multi-Scale Attention Bottleneck
2.5.3. Bidirectional Mamba Block
2.5.4. Decoder and Attention-Guided Skip Connections
2.5.5. Multi-Task Prediction Heads
2.6. Loss Function
3. Experiments and Results
3.1. Dataset Description
3.2. Experimental Setup
3.3. Main Results
4. Discussion
4.1. Comparison with State-of-the-Art Methods
4.2. Ablation Study
4.2.1. Effect of Bi-Mamba Module
4.2.2. Effect of Multi-Task Learning
4.2.3. Effect of VAE-SQI Screening
4.2.4. Effect of Multi-Scale Attention Bottleneck
4.3. Cross-Dataset Evaluation
4.4. Limitations and Future Work
5. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Roth, G.A.; Mensah, G.A.; Johnson, C.O.; Addolorato, G.; Ammirati, E.; Baddour, L.M.; Barengo, N.C.; Beaton, A.Z.; Benjamin, E.J.; Benziger, C.P. Global burden of cardiovascular diseases and risk factors, 1990–2019: Update from the GBD 2019 study. J. Am. Coll. Cardiol. 2020, 76, 2982–3021. [Google Scholar] [CrossRef]
- Whelton, P.K.; Carey, R.M.; Aronow, W.S.; Casey, D.E.; Collins, K.J.; Dennison Himmelfarb, C.; DePalma, S.M.; Gidding, S.; Jamerson, K.A.; Jones, D.W. 2017 ACC/AHA/AAPA/ABC/ACPM/AGS/APhA/ASH/ASPC/NMA/PCNA guideline for the prevention, detection, evaluation, and management of high blood pressure in adults: A report of the American College of Cardiology/American Heart Association Task Force on Clinical Practice Guidelines. J. Am. Coll. Cardiol. 2018, 71, e127–e248. [Google Scholar] [PubMed]
- Zhou, B.; Carrillo-Larco, R.M.; Danaei, G.; Riley, L.M.; Paciorek, C.J.; Stevens, G.A.; Gregg, E.W.; Bennett, J.E.; Solomon, B.; Singleton, R.K. Worldwide trends in hypertension prevalence and progress in treatment and control from 1990 to 2019: A pooled analysis of 1201 population-representative studies with 104 million participants. Lancet 2021, 398, 957–980. [Google Scholar] [CrossRef] [PubMed]
- Weldegiorgis, M.; Woodward, M. The impact of hypertension on chronic kidney disease and end-stage renal disease is greater in men than women: A systematic review and meta-analysis. BMC Nephrol. 2020, 21, 506. [Google Scholar]
- Saugel, B.; Kouz, K.; Meidert, A.S.; Schulte-Uentrop, L.; Romagnoli, S. How to measure blood pressure using an arterial catheter: A systematic 5-step approach. Crit. Care 2020, 24, 172. [Google Scholar] [CrossRef] [PubMed]
- Schults, J.A.; Young, E.R.; Marsh, N.; Larsen, E.; Corley, A.; Ware, R.S.; Murgo, M.; Alexandrou, E.; McGrail, M.; Gowardman, J. Risk factors for arterial catheter failure and complications during critical care hospitalisation: A secondary analysis of a multisite, randomised trial. J. Intensive Care 2024, 12, 12. [Google Scholar] [CrossRef] [PubMed]
- Mukkamala, R.; Hahn, J.-O. Toward ubiquitous blood pressure monitoring via pulse transit time: Predictions on maximum calibration period and acceptable error limits. IEEE Trans. Biomed. Eng. 2017, 65, 1410–1420. [Google Scholar] [CrossRef] [PubMed]
- Nyvad, J.; Christensen, K.L.; Buus, N.H.; Reinhard, M. The cuffless SOMNOtouch NIBP device shows poor agreement with a validated oscillometric device during 24-h ambulatory blood pressure monitoring. J. Clin. Hypertens. 2021, 23, 61–70. [Google Scholar] [CrossRef] [PubMed]
- Pour Ebrahim, M.; Heydari, F.; Wu, T.; Walker, K.; Joe, K.; Redoute, J.-M.; Yuce, M.R. Blood pressure estimation using on-body continuous wave radar and photoplethysmogram in various posture and exercise conditions. Sci. Rep. 2019, 9, 16346. [Google Scholar] [CrossRef]
- Li, C.; Lubecke, V.M.; Boric-Lubecke, O.; Lin, J. A review on recent advances in Doppler radar sensors for noncontact healthcare monitoring. IEEE Trans. Microw. Theory Tech. 2013, 61, 2046–2060. [Google Scholar] [CrossRef]
- Hu, W.; Zhao, Z.; Wang, Y.; Zhang, H.; Lin, F. Noncontact accurate measurement of cardiopulmonary activity using a compact quadrature Doppler radar sensor. IEEE Trans. Biomed. Eng. 2013, 61, 725–735. [Google Scholar] [CrossRef] [PubMed]
- Buxi, D.; Redouté, J.-M.; Yuce, M.R. Cuffless blood pressure estimation from the carotid pulse arrival time using continuous wave radar. In Proceedings of the 2015 37th Annual International Conference of the IEEE Engineering in Medicine and Biology Society (EMBC), Milan, Italy, 25–29 August 2015; IEEE: Piscataway, NJ, USA, 2015; pp. 5704–5707. [Google Scholar]
- Lauteslager, T.; Tommer, M.; Lande, T.S.; Constandinou, T.G. Coherent UWB radar-on-chip for in-body measurement of cardiovascular dynamics. IEEE Trans. Biomed. Circuits Syst. 2019, 13, 814–824. [Google Scholar] [CrossRef]
- Geng, F.; Bai, Z.; Zhang, H.; Yao, Y.; Liu, C.; Wang, P.; Chen, X.; Du, L.; Li, X.; Han, B. Contactless and continuous blood pressure measurement according to caPTT obtained from millimeter wave radar. Measurement 2023, 218, 113151. [Google Scholar] [CrossRef]
- Zhao, H.; Gu, X.; Hong, H.; Li, Y.; Zhu, X.; Li, C. Non-contact beat-to-beat blood pressure measurement using continuous wave Doppler radar. In Proceedings of the 2018 IEEE/MTT-S International Microwave Symposium-IMS, Philadelphia, PA, USA, 10–15 June 2018; IEEE: Piscataway, NJ, USA, 2018; pp. 1413–1415. [Google Scholar]
- Tseng, T.-J.; Tseng, C.-H. Noncontact wrist pulse waveform detection using 24-GHz continuous-wave radar sensor for blood pressure estimation. In Proceedings of the 2020 IEEE/MTT-S International Microwave Symposium (IMS), Los Angeles, CA, USA, 4–6 August 2020; IEEE: Piscataway, NJ, USA, 2020; pp. 647–650. [Google Scholar]
- Jiang, X.; Zhang, J.; Mu, W.; Wang, K.; Li, L.; Zhang, L. TRCCBP: Transformer network for radar-based contactless continuous blood pressure monitoring. Sensors 2023, 23, 9680. [Google Scholar] [CrossRef] [PubMed]
- Qiu, Y.; Ma, X.; Li, X.; Fan, S.; Deng, Z.; Huang, X. Non-contact blood pressure estimation from radar signals by a stacked deformable convolution network. IEEE J. Biomed. Health Inform. 2024, 28, 4553–4564. [Google Scholar] [CrossRef] [PubMed]
- Zhao, H.; Wang, Z.; Ding, C.; Xue, B.; Xu, D.; Wang, Z.; Hong, H.; Zhu, X. Non-Contact Blood Pressure Estimation from the Carotid Pulse Wave Signal Using a Modified ResNet-18 Network with LFMCW Radar. IEEE Sens. J. 2024, 24, 39383–39393. [Google Scholar] [CrossRef]
- Wang, P.; Yang, M.; Zhang, X.; Wang, J.; Wang, C.; Jia, H. Non-Contact Blood Pressure Monitoring Using Radar Signals: A Dual-Stage Deep Learning Network. Bioengineering 2025, 12, 252. [Google Scholar] [CrossRef] [PubMed]
- Rezende, D.J.; Mohamed, S.; Wierstra, D. Stochastic backpropagation and approximate inference in deep generative models. In Proceedings of the International Conference on Machine Learning, Beijing, China, 21–26 June 2014; PMLR: Cambridge, MA, USA, 2014; pp. 1278–1286. [Google Scholar]
- Ronneberger, O.; Fischer, P.; Brox, T. U-net: Convolutional networks for biomedical image segmentation. In Proceedings of the International Conference on Medical Image Computing and Computer-Assisted Intervention, Munich, Germany, 5–9 October 2015; Springer: Berlin/Heidelberg, Germany, 2015; pp. 234–241. [Google Scholar]
- He, K.; Zhang, X.; Ren, S.; Sun, J. Deep residual learning for image recognition. In Proceedings of the IEEE Conference on Computer Vision and Pattern Recognition, Las Vegas, NV, USA, 26 June–1 July 2016; pp. 770–778. [Google Scholar]
- Gu, A.; Dao, T. Mamba: Linear-time sequence modeling with selective state spaces. In Proceedings of the First Conference on Language Modeling, 2024, Philadelphia, PA, USA, 7–9 October 2024. [Google Scholar]
- Schellenberger, S.; Shi, K.; Steigleder, T.; Malessa, A.; Michler, F.; Hameyer, L.; Neumann, N.; Lurz, F.; Weigel, R.; Ostgathe, C. A dataset of clinically recorded radar vital signs with synchronised reference sensor signals. Sci. Data 2020, 7, 291. [Google Scholar] [CrossRef] [PubMed]
- Stergiou, G.S.; Alpert, B.; Mieke, S.; Asmar, R.; Atkins, N.; Eckert, S.; Frick, G.; Friedman, B.; Graßl, T.; Ichikawa, T. A universal standard for the validation of blood pressure measuring devices: Association for the Advancement of Medical Instrumentation/European Society of Hypertension/International Organization for Standardization (AAMI/ESH/ISO) Collaboration Statement. Hypertension 2018, 71, 368–374. [Google Scholar] [CrossRef] [PubMed]
- O’Brien, E.; Petrie, J.; Littler, W.; De Swiet, M.; Padfield, P.L.; Altman, D.; Bland, M.; Coats, A.; Atkins, N. The British Hypertension Society protocol for the evaluation of blood pressure measuring devices. J Hypertens. 1993, 11, S43–S62. [Google Scholar]
- Wang, P.; Wang, C.; Yang, M.; Jia, H. Advanced Radar-Based Non-Contact Blood Pressure Monitoring: A Machine Learning Approach. In Proceedings of the 2024 IEEE 12th International Conference on Information, Communication and Networks (ICICN), Guilin, China, 21–24 August 2024; IEEE: Piscataway, NJ, USA, 2024; pp. 607–612. [Google Scholar]
- Liu, L.; Cui, H.; Chen, H.; Xiao, W. Contactless and continuous blood pressure measurement with wrist pulse signals obtained from millimeter wave frequency-modulated continuous wave radar. Meas. Sci. Technol. 2025, 36, 055701. [Google Scholar] [CrossRef]
- Liu, L.; Zhang, J.; Feng, Z.; Zhang, S.; Zhang, Z.; Xiao, W. Contactless and Continuous Blood Pressure Detection with FMCW Millimeter-Wave Radar. IEEE Sens. J. 2025, 25, 15489–15501. [Google Scholar] [CrossRef]
- Shi, Z.; Gu, T.; Zhang, Y.; Zhang, X. mmBP+: Contact-free Blood Pressure Measurement using Millimeter-wave Radar. IEEE Trans. Mob. Comput. 2025, 25, 518–532. [Google Scholar] [CrossRef]
- Wang, Y.; Wang, S.; Fang, C.; Zhou, M.; Yang, X.; Zhang, Q.; Pang, Y. Radar based Contactless Blood Pressure Estimation System using Signal Decomposition and Deep Neural Network. IEEE Trans. Instrum. Meas. 2025, 74, 8504812. [Google Scholar] [CrossRef]













| Component | Definition | Weight |
|---|---|---|
| Reconstruction consistency | Measures conformity to VAE’s dominant mode based on reconstruction error | 0.35 |
| Latent space consistency | Computes Euclidean distance from latent mean to high-quality center | 0.15 |
| Morphological features | Quantifies RPW plausibility based on peak count, post-reconstruction peak preservation, and waveform smoothness | 0.3 |
| Physiological priors | Directly adopts the pre-filtering score | 0.2 |
| Metric | Value |
|---|---|
| ROC-AUC | 0.717 |
| AP | 0.89 |
| F1-Score | 0.864 |
| Pearson r | −0.316 (p < 0.01) |
| Spearman ρ | −0.291 (p < 0.01) |
| Parameter | Value |
|---|---|
| Tx | 1 |
| Rx | 4 |
| Start frequency | 77 GHz |
| Bandwidth | 3 GHz |
| Frequency slope | 50 MHz/μs |
| Idle time | 15 μs |
| Ramp end time | 60 μs |
| Sample points | 64 |
| Sample rate | 2 MHz |
| Accuracy | SBP [95% CI] | DBP [95% CI] |
|---|---|---|
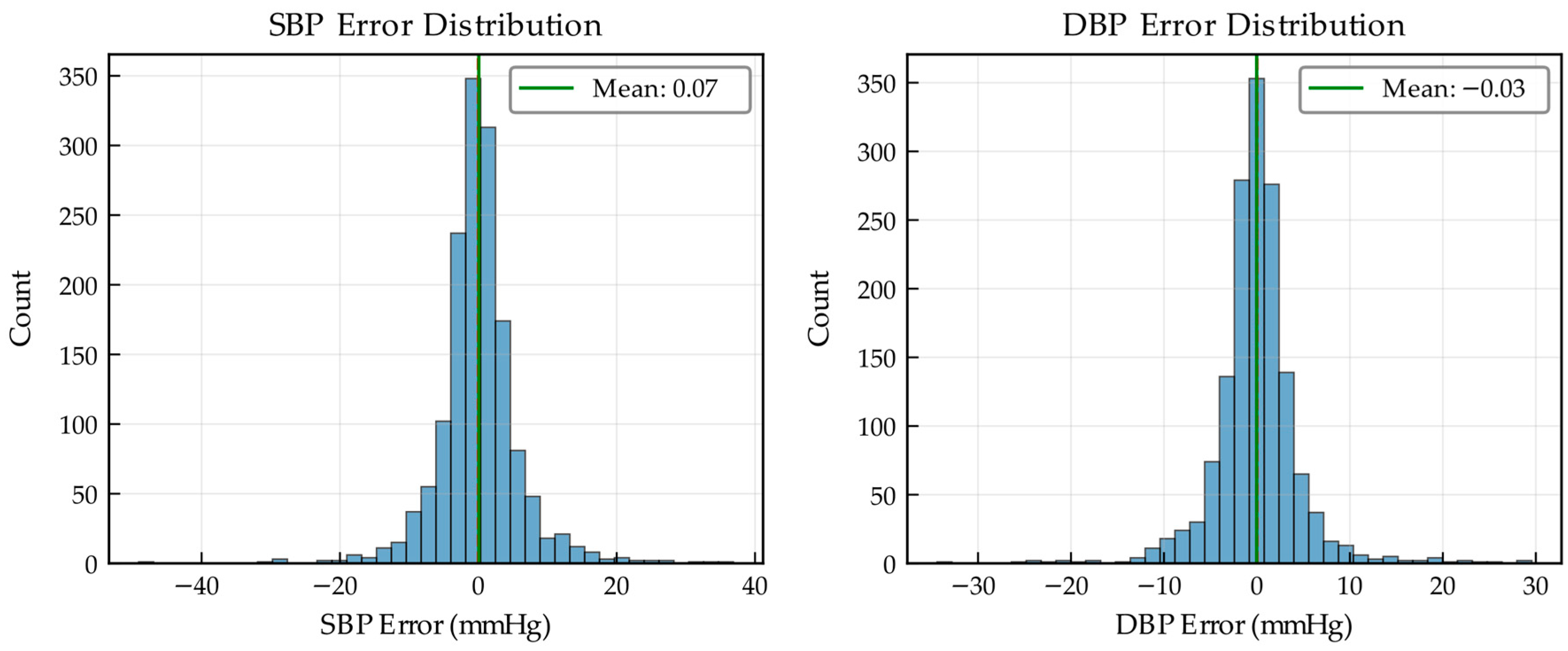
| ME (mmHg) | 0.07 [−0.22, 0.36] | –0.03 [−0.27, 0.20] |
| SD (mmHg) | 5.90 [5.43, 6.40] | 4.61 [4.23, 5.02] |
| MAE (mmHg) | 3.87 [3.64, 4.09] | 2.93 [2.75, 3.11] |
| RMSE (mmHg) | 5.90 [5.43, 6.41] | 4.61 [4.23, 5.02] |
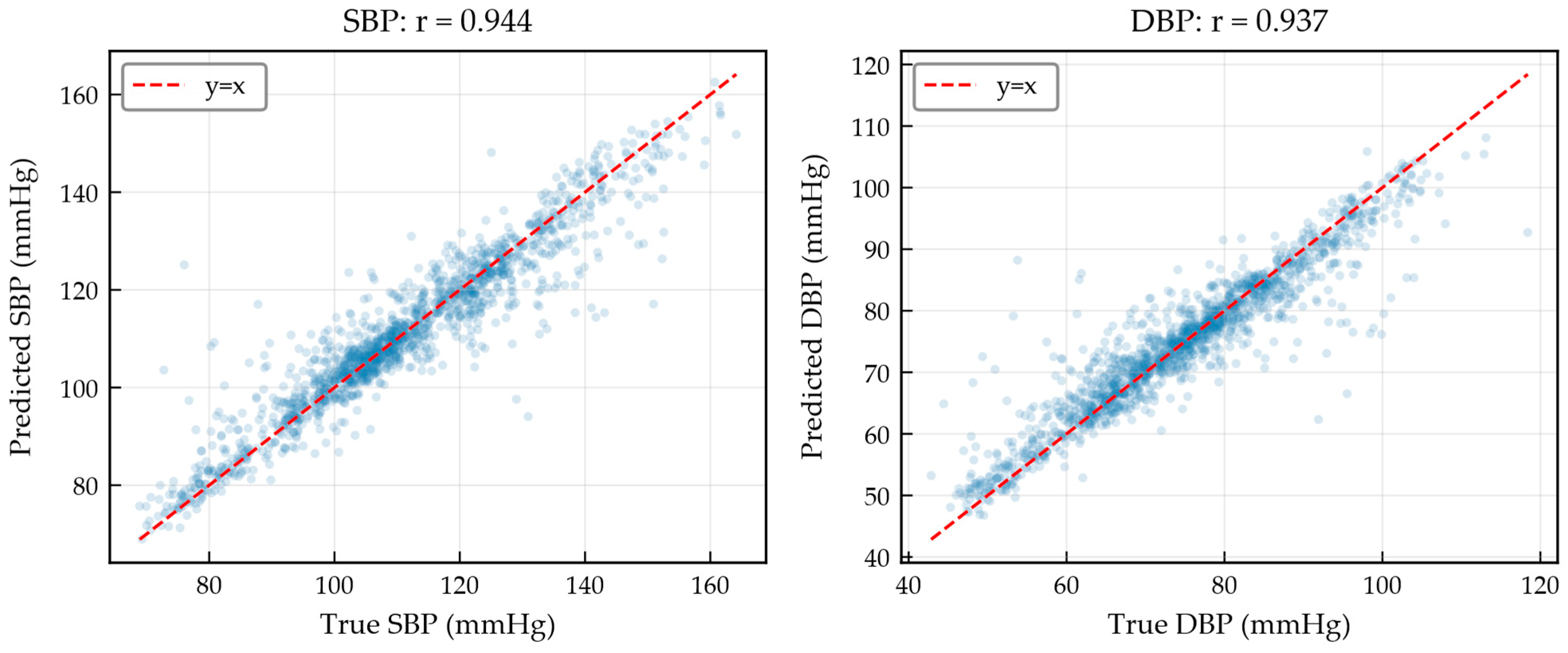
| r | 0.944 [0.934, 0.953] | 0.937 [0.925, 0.947] |
| ≤5 mmHg (%) | 75.7% | 83.9% |
| ≤10 mmHg (%) | 92.1% | 95.8% |
| ≤15 mmHg (%) | 97.0% | 98.3% |
| Dataset | SBP (ME/SD/MAE/RMSE) | SBP r | DBP (ME/SD/MAE/RMSE) | DBP r | SBP/DBP BHS | SBP/DBP AAMI |
|---|---|---|---|---|---|---|
| Public (N = 30) | 0.09/6.12/4.05/6.12 | 0.928 | −0.05/4.82/3.08/4.82 | 0.924 | A/A | Yes |
| In-house (N = 25) | 0.25/6.55/4.42/6.55 | 0.901 | −0.21/5.25/3.45/5.25 | 0.895 | B/A | Yes |
| Pooled (N = 55) | 0.07/5.90/3.87/5.90 | 0.944 | −0.03/4.61/2.93/4.61 | 0.937 | A/A | Yes |
| Method | Measurement Site | Subject | SBP (ME/SD/MAE/RMSE) | SBP r | DBP (ME/SD/MAE/RMSE) | DBP r |
|---|---|---|---|---|---|---|
| CNN + LSTM [28] | Chest | 30 | 2.04/12.65/9.30/NR | 0.80 | 0.48/8.16/5.91/NR | 0.85 |
| TRCCBP [17] | Chest | 31 | −0.94/6.28/4.95/NR | NR | −0.62/6.46/4.53/NR | NR |
| CCBP [29] | Wrist | 15 | −1.30/6.17/4.42/NR | 0.919 | −3.10/4.93/4.33/NR | 0.892 |
| mmRBP [30] | Wrist | 15 | 0.87/6.12/NR/6.00 | 0.88 | 0.59/3.78/NR/4.00 | 0.86 |
| RSD-Net [18] | Chest | 30 | −0.32/6.14/4.61/6.14 | 0.84 | −0.20/5.50/4.42/5.50 | 0.80 |
| Two-stage model [20] | Chest | 30 | −1.09/5.15/5.00/6.24 | 0.933 | −0.26/4.35/3.96/4.98 | 0.934 |
| mmBP+ [31] | Wrist | 33 | 0.65/3.92/NR/NR | NR | 1.31/3.99/NR/NR | NR |
| DSFNN-BP [32] | Neck | 40 | 0.49/5.19/4.16/NR | NR | −0.32/5.11/3.87/NR | NR |
| Ours | Chest | 55 | 0.07/5.90/3.87/5.90 | 0.944 | −0.03/4.61/2.93/4.61 | 0.937 |
| Configuration | SBP MAE | SBP SD | SBP r | SBP BHS | DBP MAE | DBP SD | DBP r | DBP BHS | AAMI |
|---|---|---|---|---|---|---|---|---|---|
| without MTL | 5.05 | 7.75 | 0.902 | B | 3.83 | 5.91 | 0.949 | A | Yes |
| λ = 0.5 | 10.35 | 13.52 | 0.657 | D | 8.33 | 10.73 | 0.574 | D | No |
| λ = 0.2 | 7.02 | 9.66 | 0.842 | C | 5.74 | 7.99 | 0.795 | B | No |
| λ = 0.1 | 7.10 | 10.32 | 0.817 | C | 6.09 | 8.57 | 0.756 | B | No |
| λ = 0.05 | 6.11 | 8.72 | 0.874 | B | 4.79 | 6.94 | 0.848 | A | No |
| λ = 0.01 (Proposed) | 3.87 | 5.90 | 0.944 | A | 2.93 | 4.61 | 0.937 | A | Yes |
| Configuration | SBP MAE | SBP SD | SBP r | SBP BHS | DBP MAE | DBP SD | DBP r | DBP BHS | AAMI |
|---|---|---|---|---|---|---|---|---|---|
| Without SQI filtering | 5.33 | 7.61 | 0.856 | B | 4.57 | 6.92 | 0.851 | B | No |
| With SQI filtering (90%) | 4.31 | 6.69 | 0.891 | B | 3.64 | 5.39 | 0.883 | A | Yes |
| With SQI filtering (80%) | 3.87 | 5.90 | 0.944 | A | 2.93 | 4.61 | 0.937 | A | Yes |
| With SQI filtering (70%) | 3.68 | 5.69 | 0.948 | A | 2.81 | 4.57 | 0.939 | A | Yes |
| Method | ROC-AUC | AP | F1 | Pearson r | Spearman ρ | SBP MAE | SBP MAE | SBP BHS | DBP MAE | DBP SD | DBP BHS | AAMI |
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| SNR threshold | 0.394 | 0.765 | 0.775 | 0.120 | 0.161 | 5.06 | 7.14 | B | 4.12 | 5.89 | A | Yes |
| Pre-screening score | 0.692 | 0.874 | 0.856 | −0.292 | −0.279 | 4.31 | 6.39 | A | 3.40 | 5.19 | A | Yes |
| VAE-SQI (proposed) | 0.717 | 0.890 | 0.864 | −0.316 | −0.291 | 3.87 | 5.90 | A | 2.93 | 4.61 | A | Yes |
| Configuration | SBP MAE | SBP SD | SBP r | SBP BHS | DBP MAE | DBP SD | DBP r | DBP BHS | AAMI |
|---|---|---|---|---|---|---|---|---|---|
| Single-scale without CTA | 4.67 | 6.99 | 0.911 | A | 3.64 | 5.55 | 0.892 | A | Yes |
| Multi-scale without CTA | 4.26 | 6.47 | 0.920 | A | 3.18 | 4.90 | 0.917 | A | Yes |
| Multi-scale + CTA | 3.87 | 5.90 | 0.944 | A | 2.93 | 4.61 | 0.937 | A | Yes |
| Training Set/Test Set | SBP (ME/SD/MAE/RMSE) | SBP r | DBP (ME/SD/MAE/RMSE) | DBP r | SBP/DBP BHS | SBP/DBP AAMI |
|---|---|---|---|---|---|---|
| Public/In-house | 0.85/7.12/5.25/7.17 | 0.865 | 0.42/5.65/4.18/5.67 | 0.852 | B/B | Yes |
| In-house/Public | 5.42/8.35/7.15/9.95 | 0.784 | 3.15/8.12/6.05/8.71 | 0.765 | C/B | No |
| Pooled (proposed) | 0.07/5.90/3.87/5.90 | 0.944 | −0.03/4.61/2.93/4.61 | 0.937 | A/A | Yes |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license.
Share and Cite
Xie, J.; Huang, J.; Xu, C.; Wan, H.; Zuo, X.; Dong, G. MARU-MTL: A Mamba-Enhanced Multi-Task Learning Framework for Continuous Blood Pressure Estimation Using Radar Pulse Waves. Bioengineering 2026, 13, 320. https://doi.org/10.3390/bioengineering13030320
Xie J, Huang J, Xu C, Wan H, Zuo X, Dong G. MARU-MTL: A Mamba-Enhanced Multi-Task Learning Framework for Continuous Blood Pressure Estimation Using Radar Pulse Waves. Bioengineering. 2026; 13(3):320. https://doi.org/10.3390/bioengineering13030320
Chicago/Turabian StyleXie, Jinke, Juhua Huang, Chongnan Xu, Hongtao Wan, Xuetao Zuo, and Guanfang Dong. 2026. "MARU-MTL: A Mamba-Enhanced Multi-Task Learning Framework for Continuous Blood Pressure Estimation Using Radar Pulse Waves" Bioengineering 13, no. 3: 320. https://doi.org/10.3390/bioengineering13030320
APA StyleXie, J., Huang, J., Xu, C., Wan, H., Zuo, X., & Dong, G. (2026). MARU-MTL: A Mamba-Enhanced Multi-Task Learning Framework for Continuous Blood Pressure Estimation Using Radar Pulse Waves. Bioengineering, 13(3), 320. https://doi.org/10.3390/bioengineering13030320







