ICIsc: A Deep Learning Framework for Predicting Immune Checkpoint Inhibitor Response by Integrating scRNA-Seq and Protein Language Models
Abstract
1. Introduction
2. Materials and Methods
2.1. Data Collection and Preprocessing
2.2. Patient Feature Encoder
2.3. ICI Drug Feature Encoder
2.4. Bilinear Attention Module
2.5. Construction of Single-Sample Networks
2.6. Graph Attention Network v2 Module
2.7. Predictors
2.8. Model Training
3. Results
3.1. ICIsc Framework Overview
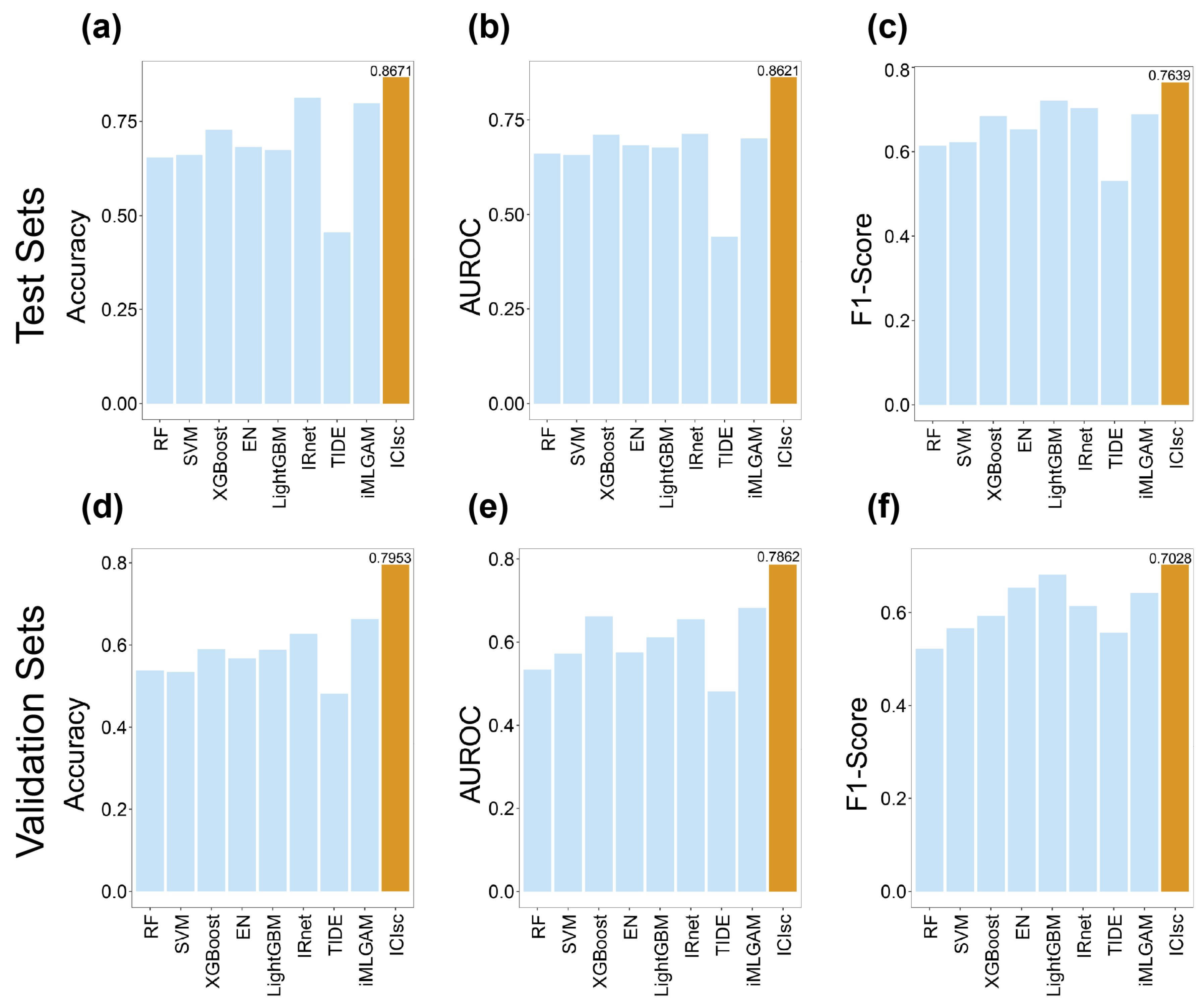
3.2. Evaluation of ICIs Model Performance on the Bulk Cohort
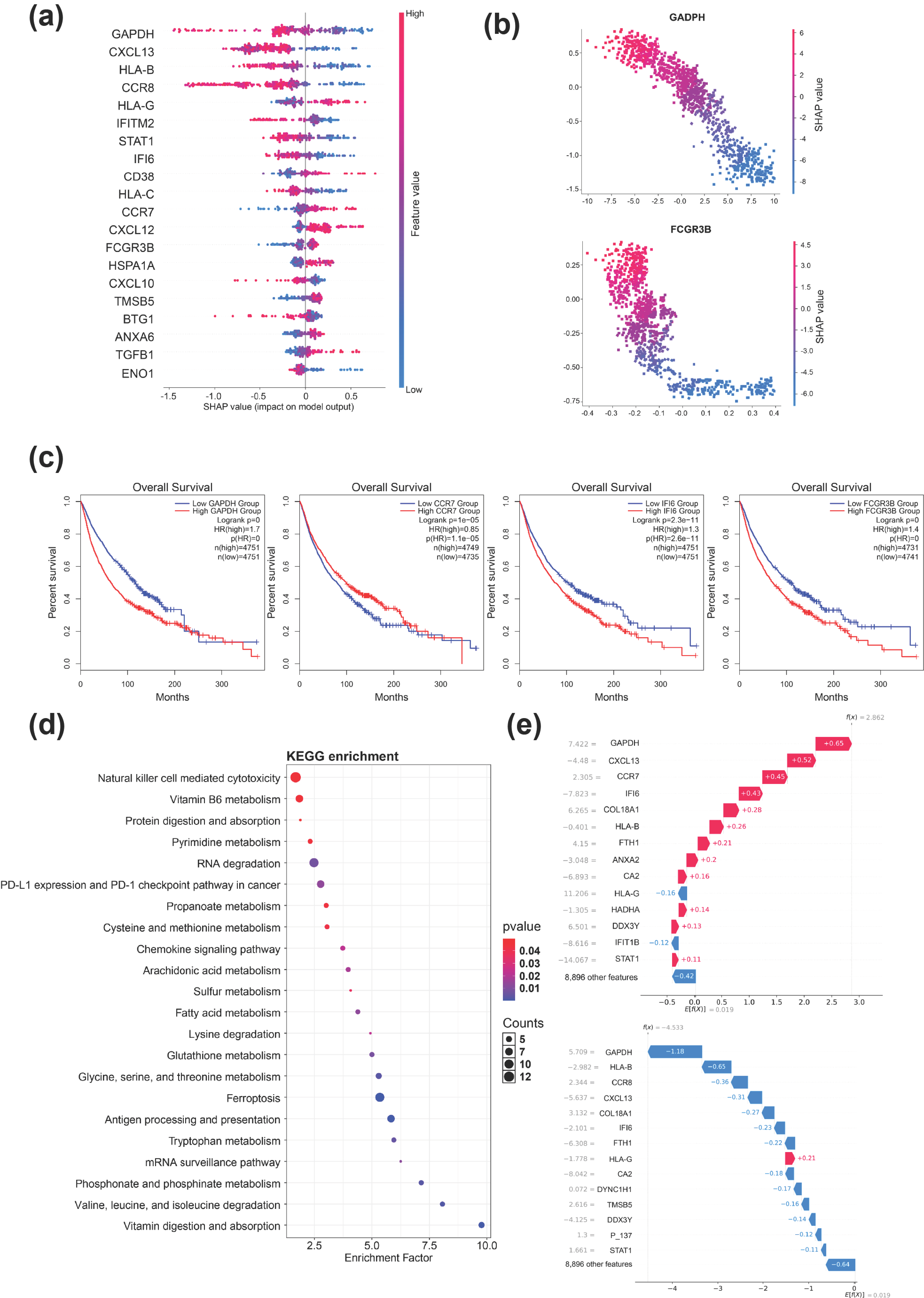
3.3. SHAP Values Reveal Key Feature Genes and Pathways
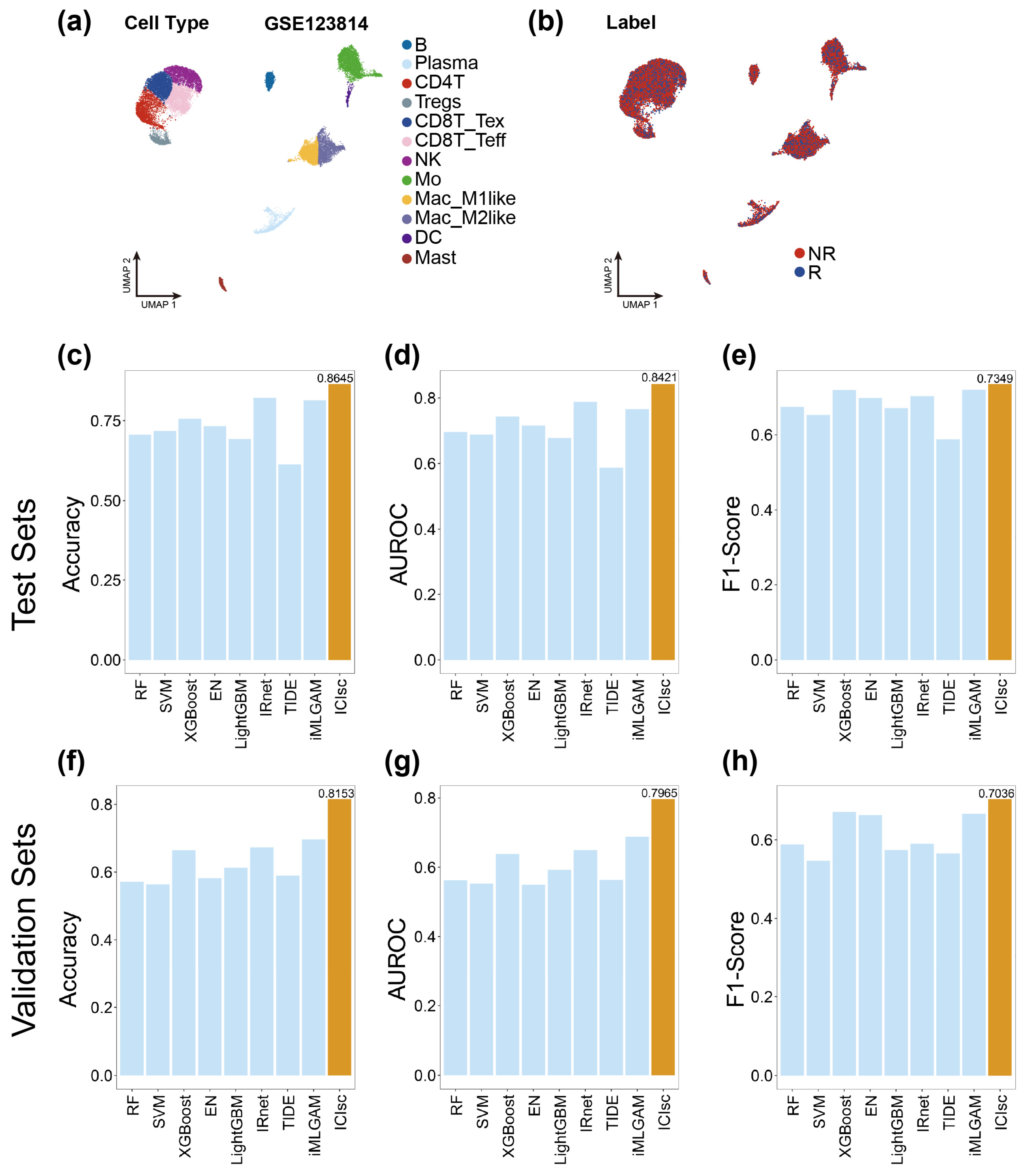
3.4. Evaluation of ICIs Model Performance on the Single-Cell Cohort
3.5. Analysis of Interactions Among Different Cell Types in Single-Cell Data
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
Abbreviations
| ICI | Immune checkpoint inhibitor |
| scRNA-seq | Single-cell RNA sequencing |
| ESM2 | Evolutionary Scale Modeling 2 |
| SSN | Single-sample network |
| PD-1 | Programmed cell death 1 |
| CTLA-4 | Cytotoxic T-lymphocyte-associated antigen 4 |
| FDA | Food and Drug Administration |
| TME | Tumor microenvironment |
| SVM | Support vector machine |
| GATv2 | Graph attention network v2 |
| RF | Random forest |
| EN | Elastic Net Regression |
| LightGBM | Light Gradient Boosting Machine |
| XGBoost | Extreme gradient boosting |
| GSVA | Gene Set Variation Analysis |
| RECIST | Response Evaluation Criteria in Solid Tumors |
| KS | Kolmogorov–Smirnov |
| MLP | Multi-layer perceptron |
| ReLU | Rectified linear unit |
| AUROC | The area under the receiver operating characteristic curve |
| SHAP | SHapley Additive exPlanations |
| OS | Overall survival |
| PFE | Patient feature encoder |
References
- Pardoll, D.M. The Blockade of Immune Checkpoints in Cancer Immunotherapy. Nat. Rev. Cancer 2012, 12, 252–264. [Google Scholar] [CrossRef]
- Sharma, P.; Hu-Lieskovan, S.; Wargo, J.A.; Ribas, A. Primary, Adaptive, and Acquired Resistance to Cancer Immunotherapy. Cell 2017, 168, 707–723. [Google Scholar] [CrossRef]
- Nowicki, T.S.; Hu-Lieskovan, S.; Ribas, A. Mechanisms of Resistance to PD-1 and PD-L1 Blockade. Cancer J. 2018, 24, 47–53. [Google Scholar] [CrossRef]
- Ribas, A.; Wolchok, J.D. Cancer Immunotherapy Using Checkpoint Blockade. Science 2018, 359, 1350–1355. [Google Scholar] [CrossRef] [PubMed]
- Sharma, P.; Goswami, S.; Raychaudhuri, D.; Siddiqui, B.A.; Singh, P.; Nagarajan, A.; Liu, J.; Subudhi, S.K.; Poon, C.; Gant, K.L.; et al. Immune Checkpoint Therapy—Current Perspectives and Future Directions. Cell 2023, 186, 1652–1669. [Google Scholar] [CrossRef]
- Nixon, N.A.; Blais, N.; Ernst, S.; Kollmannsberger, C.; Bebb, G.; Butler, M.; Smylie, M.; Verma, S. Current Landscape of Immunotherapy in the Treatment of Solid Tumours, with Future Opportunities and Challenges. Curr. Oncol. 2018, 25, 373–384. [Google Scholar] [CrossRef] [PubMed]
- Gridelli, C.; Rossi, A.; Carbone, D.P.; Guarize, J.; Karachaliou, N.; Mok, T.; Petrella, F.; Spaggiari, L.; Rosell, R. Non-Small-Cell Lung Cancer. Nat. Rev. Dis. Primers 2015, 1, 15009. [Google Scholar] [CrossRef]
- Xu, Y.; Wan, B.; Chen, X.; Zhan, P.; Zhao, Y.; Zhang, T.; Liu, H.; Afzal, M.Z.; Dermime, S.; Hochwald, S.N.; et al. The Association of PD-L1 Expression with the Efficacy of Anti-PD-1/PD-L1 Immunotherapy and Survival of Non-Small Cell Lung Cancer Patients: A Meta-Analysis of Randomized Controlled Trials. Transl. Lung Cancer Res. 2019, 8, 413–428. [Google Scholar] [CrossRef] [PubMed]
- Strickler, J.H.; Hanks, B.A.; Khasraw, M. Tumor Mutational Burden as a Predictor of Immunotherapy Response: Is More Always Better? Clin. Cancer Res. 2021, 27, 1236–1241. [Google Scholar] [CrossRef]
- Marabelle, A.; Fakih, M.; Lopez, J.; Shah, M.; Shapira-Frommer, R.; Nakagawa, K.; Chung, H.C.; Kindler, H.L.; Lopez-Martin, J.A.; Miller, W.H.; et al. Association of Tumour Mutational Burden with Outcomes in Patients with Advanced Solid Tumours Treated with Pembrolizumab: Prospective Biomarker Analysis of the Multicohort, Open-Label, Phase 2 KEYNOTE-158 Study. Lancet Oncol. 2020, 21, 1353–1365. [Google Scholar] [CrossRef]
- Wu, T.; Dai, Y. Tumor Microenvironment and Therapeutic Response. Cancer Lett. 2017, 387, 61–68. [Google Scholar] [CrossRef] [PubMed]
- Toor, S.M.; Sasidharan Nair, V.; Decock, J.; Elkord, E. Immune Checkpoints in the Tumor Microenvironment. Semin. Cancer Biol. 2020, 65, 1–12. [Google Scholar] [CrossRef]
- Auslander, N.; Zhang, G.; Lee, J.S.; Frederick, D.T.; Miao, B.; Moll, T.; Tian, T.; Wei, Z.; Madan, S.; Sullivan, R.J.; et al. Robust Prediction of Response to Immune Checkpoint Blockade Therapy in Metastatic Melanoma. Nat. Med. 2018, 24, 1545–1549. [Google Scholar] [CrossRef]
- Jiang, P.; Gu, S.; Pan, D.; Fu, J.; Sahu, A.; Hu, X.; Li, Z.; Traugh, N.; Bu, X.; Li, B.; et al. Signatures of T Cell Dysfunction and Exclusion Predict Cancer Immunotherapy Response. Nat. Med. 2018, 24, 1550–1558. [Google Scholar] [CrossRef]
- Kong, J.; Ha, D.; Lee, J.; Kim, I.; Park, M.; Im, S.-H.; Shin, K.; Kim, S. Network-Based Machine Learning Approach to Predict Immunotherapy Response in Cancer Patients. Nat. Commun. 2022, 13, 3703. [Google Scholar] [CrossRef] [PubMed]
- Zhao, L.; Qi, X.; Chen, Y.; Qiao, Y.; Bu, D.; Wu, Y.; Luo, Y.; Wang, S.; Zhang, R.; Zhao, Y. Biological Knowledge Graph-Guided Investigation of Immune Therapy Response in Cancer with Graph Neural Network. Brief. Bioinform. 2023, 24, bbad023. [Google Scholar]
- Kang, Y.; Vijay, S.; Gujral, T.S. Deep Neural Network Modeling Identifies Biomarkers of Response to Immune-Checkpoint Therapy. iScience 2022, 25, 104228. [Google Scholar] [CrossRef]
- Ma, Q.; Xu, D. Deep Learning Shapes Single-Cell Data Analysis. Nat. Rev. Mol. Cell Biol. 2022, 23, 303–304. [Google Scholar] [CrossRef]
- Erfanian, N.; Heydari, A.A.; Feriz, A.M.; Iañez, P.; Derakhshani, A.; Ghasemigol, M.; Farahpour, M.; Razavi, S.M.; Nasseri, S.; Safarpour, H.; et al. Deep Learning Applications in Single-Cell Genomics and Transcriptomics Data Analysis. Biomed. Pharmacother. 2023, 165, 115077. [Google Scholar] [CrossRef]
- Dong, Y.; Chen, Z.; Yang, F.; Wei, J.; Huang, J.; Long, X. Prediction of Immunotherapy Responsiveness in Melanoma through Single-Cell Sequencing-Based Characterization of the Tumor Immune Microenvironment. Transl. Oncol. 2024, 43, 101910. [Google Scholar] [CrossRef]
- Hänzelmann, S.; Castelo, R.; Guinney, J. GSVA: Gene Set Variation Analysis for Microarray and RNA-Seq Data. BMC Bioinform. 2013, 14, 7. [Google Scholar] [CrossRef]
- Huang, Y.; Chang, X.; Zhang, Y.; Chen, L.; Liu, X. Disease Characterization Using a Partial Correlation-Based Sample-Specific Network. Brief. Bioinform. 2021, 22, bbaa062. [Google Scholar]
- Aran, D.; Looney, A.P.; Liu, L.; Wu, E.; Fong, V.; Hsu, A.; Chak, S.; Naikawadi, R.P.; Wolters, P.J.; Abate, A.R.; et al. Reference-Based Analysis of Lung Single-Cell Sequencing Reveals a Transitional Profibrotic Macrophage. Nat. Immunol. 2019, 20, 163–172. [Google Scholar] [CrossRef]
- Lin, Z.; Akin, H.; Rao, R.; Hie, B.; Zhu, Z.; Lu, W.; Smetanin, N.; Verkuil, R.; Kabeli, O.; Shmueli, Y.; et al. Evolutionary-Scale Prediction of Atomic Level Protein Structure with a Language Model. Science 2023, 379, 1123–1130. [Google Scholar] [CrossRef]
- Jiang, Y.; Immadi, M.S.; Wang, D.; Zeng, S.; On Chan, Y.; Zhou, J.; Xu, D.; Joshi, T. IRnet: Immunotherapy Response Prediction Using Pathway Knowledge-Informed Graph Neural Network. J. Adv. Res. 2025, 72, 319–331. [Google Scholar] [CrossRef]
- Ye, B.; Fan, J.; Xue, L.; Zhuang, Y.; Luo, P.; Jiang, A.; Xie, J.; Li, Q.; Liang, X.; Tan, J.; et al. iMLGAM: Integrated Machine Learning and Genetic Algorithm-driven Multiomics Analysis for Pan-cancer Immunotherapy Response Prediction. iMeta 2025, 4, e70011. [Google Scholar] [CrossRef] [PubMed]
- Lundberg, S.M.; Lee, S.-I. A Unified Approach to Interpreting Model Predictions. In NIPS’17: Proceedings of the 31st International Conference on Neural Information Processing Systems; Curran Associates Inc.: Long Beach, CA, USA, 2017; pp. 4768–4777. [Google Scholar]
- Wu, P.; Zhao, L.; Chen, Y.; Xin, Z.; Lin, M.; Hao, Z.; Chen, X.; Chen, D.; Wu, D.; Chai, Y. CD38 Identifies Pre-Activated CD8+ T Cells Which Can Be Reinvigorated by Anti-PD-1 Blockade in Human Lung Cancer. Cancer Immunol. Immunother. 2021, 70, 3603–3616. [Google Scholar] [CrossRef]
- Lv, L.-L.; Zhai, J.-W.; Wu, J.-J.; Fan, G.-Q.; Zhang, Y.-X.; Shen, Y.; Qu, Q.-X.; Chen, C. High CD38 Expression Defines a Mitochondrial Function-Adapted CD8+ T Cell Subset with Implications for Lung Cancer Immunotherapy. Cancer Immunol. Immunother. 2025, 74, 49. [Google Scholar] [CrossRef] [PubMed]
- Ng, H.H.M.; Lee, R.Y.; Goh, S.; Tay, I.S.Y.; Lim, X.; Lee, B.; Chew, V.; Li, H.; Tan, B.; Lim, S.; et al. Immunohistochemical Scoring of CD38 in the Tumor Microenvironment Predicts Responsiveness to Anti-PD-1/PD-L1 Immunotherapy in Hepatocellular Carcinoma. J. Immunother. Cancer 2020, 8, e000987. [Google Scholar] [CrossRef] [PubMed]
- Chen, L.; Diao, L.; Yang, Y.; Yi, X.; Rodriguez, B.L.; Li, Y.; Villalobos, P.A.; Cascone, T.; Liu, X.; Tan, L.; et al. CD38-Mediated Immunosuppression as a Mechanism of Tumor Cell Escape from PD-1/PD-L1 Blockade. Cancer Discov. 2018, 8, 1156–1175. [Google Scholar] [CrossRef]
- Oshi, M.; Tokumaru, Y.; Asaoka, M.; Yan, L.; Satyananda, V.; Matsuyama, R.; Matsuhashi, N.; Futamura, M.; Ishikawa, T.; Yoshida, K.; et al. M1 Macrophage and M1/M2 Ratio Defined by Transcriptomic Signatures Resemble Only Part of Their Conventional Clinical Characteristics in Breast Cancer. Sci. Rep. 2020, 10, 16554. [Google Scholar] [CrossRef]
- Liu, F.; Liu, J.; Zhang, J.; Shi, J.; Gui, L.; Xu, G. Expression of STAT1 Is Positively Correlated with PD-L1 in Human Ovarian Cancer. Cancer Biol. Ther. 2020, 21, 963–971. [Google Scholar] [CrossRef]
- Horowitch, B.; Lee, D.Y.; Ding, M.; Martinez-Morilla, S.; Aung, T.N.; Ouerghi, F.; Wang, X.; Wei, W.; Damsky, W.; Sznol, M.; et al. Subsets of IFN Signaling Predict Response to Immune Checkpoint Blockade in Patients with Melanoma. Clin. Cancer Res. 2023, 29, 2908–2918. [Google Scholar] [CrossRef]
- Meng, F.; Zhang, S.; Xie, J.; Zhou, Y.; Wu, Q.; Lu, B.; Zhou, S.; Zhao, X.; Li, Y. Leveraging CD16 Fusion Receptors to Remodel the Immune Response for Enhancing Anti-Tumor Immunotherapy in iPSC-Derived NK Cells. J. Hematol. Oncol. 2023, 16, 62. [Google Scholar] [CrossRef] [PubMed]
- Aguilar, O.A.; Gonzalez-Hinojosa, M.D.R.; Arakawa-Hoyt, J.S.; Millan, A.J.; Gotthardt, D.; Nabekura, T.; Lanier, L.L. The CD16 and CD32b Fc-Gamma Receptors Regulate Antibody-Mediated Responses in Mouse Natural Killer Cells. J. Leukoc. Biol. 2023, 113, 27–40. [Google Scholar] [CrossRef] [PubMed]
- Park, J.-E.; Kim, S.-E.; Keam, B.; Park, H.-R.; Kim, S.; Kim, M.; Kim, T.M.; Doh, J.; Kim, D.-W.; Heo, D.S. Anti-Tumor Effects of NK Cells and Anti-PD-L1 Antibody with Antibody-Dependent Cellular Cytotoxicity in PD-L1-Positive Cancer Cell Lines. J. Immunother. Cancer 2020, 8, e000873. [Google Scholar] [CrossRef]
- Peng, C.; Ye, H.; Yi, Z. GAPDH: Unveiling Its Impact as a Key Hypoxia-Related Player in Head and Neck Squamous Cell Carcinoma Tumor Progression, Prognosis, and Therapeutic Potential. Am. J. Cancer Res. 2023, 13, 5846–5867. [Google Scholar] [PubMed]
- Wang, J.; Yu, X.; Cao, X.; Tan, L.; Jia, B.; Chen, R.; Li, J. GAPDH: A Common Housekeeping Gene with an Oncogenic Role in Pan-Cancer. Comput. Struct. Biotechnol. J. 2023, 21, 4056–4069. [Google Scholar] [CrossRef]
- Tang, Z.; Kang, B.; Li, C.; Chen, T.; Zhang, Z. GEPIA2: An Enhanced Web Server for Large-Scale Expression Profiling and Interactive Analysis. Nucleic Acids Res. 2019, 47, W556–W560. [Google Scholar] [CrossRef]
- Liu, B.; Zhang, Y.; Wang, D.; Hu, X.; Zhang, Z. Single-Cell Meta-Analyses Reveal Responses of Tumor-Reactive CXCL13+ T Cells to Immune-Checkpoint Blockade. Nat. Cancer 2022, 3, 1123–1136. [Google Scholar] [CrossRef]
- Chow, A.; Perica, K.; Klebanoff, C.A.; Wolchok, J.D. Clinical Implications of T Cell Exhaustion for Cancer Immunotherapy. Nat. Rev. Clin. Oncol. 2022, 19, 775–790. [Google Scholar] [CrossRef]
- Liu, Z.; Yang, Z.; Wu, J.; Zhang, W.; Sun, Y.; Zhang, C.; Bai, G.; Yang, L.; Fan, H.; Chen, Y.; et al. A Single-Cell Atlas Reveals Immune Heterogeneity in Anti-PD-1-Treated Non-Small Cell Lung Cancer. Cell 2025, 188, 3081–3096.e19. [Google Scholar] [CrossRef]
- Yamaguchi, Y.; Gibson, J.; Ou, K.; Lopez, L.S.; Ng, R.H.; Leggett, N.; Jonsson, V.D.; Zarif, J.C.; Lee, P.P.; Wang, X.; et al. PD-L1 Blockade Restores CAR T Cell Activity through IFN-γ-Regulation of CD163+ M2 Macrophages. J. Immunother. Cancer 2022, 10, e004400. [Google Scholar] [CrossRef]
- Wang, D.-R.; Wu, X.-L.; Sun, Y.-L. Therapeutic Targets and Biomarkers of Tumor Immunotherapy: Response versus Non-Response. Signal Transduct. Target. Ther. 2022, 7, 331. [Google Scholar] [CrossRef] [PubMed]
- Knopf, P.; Stowbur, D.; Hoffmann, S.H.L.; Hermann, N.; Maurer, A.; Bucher, V.; Poxleitner, M.; Tako, B.; Sonanini, D.; Krishnamachary, B.; et al. Acidosis-Mediated Increase in IFN-γ-Induced PD-L1 Expression on Cancer Cells as an Immune Escape Mechanism in Solid Tumors. Mol. Cancer 2023, 22, 207. [Google Scholar] [CrossRef]
- Pustylnikov, S.; Costabile, F.; Beghi, S.; Facciabene, A. Targeting Mitochondria in Cancer: Current Concepts and Immunotherapy Approaches. Transl. Res. 2018, 202, 35–51. [Google Scholar] [CrossRef]
- Huang, L.; Xu, H.; Peng, G. TLR-Mediated Metabolic Reprogramming in the Tumor Microenvironment: Potential Novel Strategies for Cancer Immunotherapy. Cell. Mol. Immunol. 2018, 15, 428–437. [Google Scholar] [CrossRef] [PubMed]
- Zhao, H.; Raines, L.N.; Huang, S.C.-C. Carbohydrate and Amino Acid Metabolism as Hallmarks for Innate Immune Cell Activation and Function. Cells 2020, 9, 562. [Google Scholar] [CrossRef] [PubMed]
- Gong, D.; Chen, M.; Wang, Y.; Shi, J.; Hou, Y. Role of Ferroptosis on Tumor Progression and Immunotherapy. Cell Death Discov. 2022, 8, 427. [Google Scholar] [CrossRef]
- Wang, W.; Green, M.; Choi, J.E.; Gijón, M.; Kennedy, P.D.; Johnson, J.K.; Liao, P.; Lang, X.; Kryczek, I.; Sell, A.; et al. CD8+ T Cells Regulate Tumour Ferroptosis during Cancer Immunotherapy. Nature 2019, 569, 270–274. [Google Scholar] [CrossRef]
- Dai, Z.; Liu, J.; Zeng, L.; Shi, K.; Peng, X.; Jin, Z.; Zheng, R.; Zeng, C. Targeting Ferroptosis in Cancer Therapy: Mechanisms, Strategies, and Clinical Applications. Cell Investig. 2025, 1, 100049. [Google Scholar] [CrossRef]
- Gao, M.; Yu, W.; Xi, Z.; Zhang, Z.; Fan, X.; Wang, X. Recent Update on the Discovery of Indoleamine-2,3-Dioxygenase 1 Inhibitors Targeting Cancer Immunotherapy. Eur. J. Med. Chem. 2025, 298, 118017. [Google Scholar] [CrossRef] [PubMed]
- Jin, S.; Plikus, M.V.; Nie, Q. CellChat for Systematic Analysis of Cell–Cell Communication from Single-Cell Transcriptomics. Nat. Protoc. 2025, 20, 180–219. [Google Scholar] [CrossRef] [PubMed]
- Liang, Y.; Zhang, R.; Zhang, L.; Xie, P. DrugChat: Towards Enabling ChatGPT-like Capabilities on Drug Molecule Graphs. arXiv 2023, arXiv:2309.03907. [Google Scholar]
- Liu, T.; Chu, T.; Luo, X.; Zhao, H. Building a Unified Model for Drug Synergy Analysis Powered by Large Language Models. Nat. Commun. 2025, 16, 4537. [Google Scholar] [CrossRef] [PubMed]
- Sang, S.; Sun, Z.; Zheng, W.; Wang, W.; Islam, M.T.; Chen, Y.; Yuan, Q.; Cheng, C.; Xi, S.; Han, Z.; et al. TME-Guided Deep Learning Predicts Chemotherapy and Immunotherapy Response in Gastric Cancer with Attention-Enhanced Residual Swin Transformer. Cell Rep. Med. 2025, 6, 102242. [Google Scholar] [CrossRef]
- Li, X.; Pan, B.; He, Y.; Wang, Z.; Tang, Y.; Zhang, Y.; Wang, L.; Han, J. PathHDNN: A Pathway Hierarchical-Informed Deep Neural Network Framework for Predicting Immunotherapy Response and Mechanism Interpretation. Genome Med. 2025, 17, 152. [Google Scholar] [CrossRef]
- Min, J.H.; Chen, P.-J.; Qureshi, T.A.; Javed, S.; Xie, Y.; Azab, L.; Wang, L.; Kim, H.; Li, D.; Yang, J.D. Prediction of Immunotherapy Response in Hepatocellular Carcinoma Patients Using Pretreatment CT Images. Diagnostics 2025, 15, 2090. [Google Scholar] [CrossRef]


Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license.
Share and Cite
Jin, Z.; Zhang, D.; Chen, L. ICIsc: A Deep Learning Framework for Predicting Immune Checkpoint Inhibitor Response by Integrating scRNA-Seq and Protein Language Models. Bioengineering 2026, 13, 187. https://doi.org/10.3390/bioengineering13020187
Jin Z, Zhang D, Chen L. ICIsc: A Deep Learning Framework for Predicting Immune Checkpoint Inhibitor Response by Integrating scRNA-Seq and Protein Language Models. Bioengineering. 2026; 13(2):187. https://doi.org/10.3390/bioengineering13020187
Chicago/Turabian StyleJin, Zhenyu, Di Zhang, and Luonan Chen. 2026. "ICIsc: A Deep Learning Framework for Predicting Immune Checkpoint Inhibitor Response by Integrating scRNA-Seq and Protein Language Models" Bioengineering 13, no. 2: 187. https://doi.org/10.3390/bioengineering13020187
APA StyleJin, Z., Zhang, D., & Chen, L. (2026). ICIsc: A Deep Learning Framework for Predicting Immune Checkpoint Inhibitor Response by Integrating scRNA-Seq and Protein Language Models. Bioengineering, 13(2), 187. https://doi.org/10.3390/bioengineering13020187




