AIMS: An Automatic Semantic Machine Learning Microservice Framework to Support Biomedical and Bioengineering Research
Abstract
:1. Introduction
2. Related Work, Limitations, and Technology Background
2.1. Related Work and Current Limitations
2.2. Multiple-Task AI System Research
- The information acquired from different tasks may present value that can be used as the basis to build new ML models for new tasks without requiring high-cost processing to recapture the same feature characteristics.
- Updating knowledge through validation is a relatively consistent process that will be less prone to bias from noisy data.
- The knowledge definition is too narrow and only uses the generated neural network as the knowledge limits the capability of recording all valuable outcomes through the learning experience.
- There is no unified knowledge representation structure for knowledge inference (machine thinking).
2.3. Knowledge Representation and Reasoning
2.4. Services and Machine Learning Ontologies
- Directly extracting the services description file (e.g., WSDL) and Quality of Services (QoS) into a mathematical model with a logical framework for composing services such as a linear logic approach [30] and genetic algorithms [31,32]. The major limitation is that there are no formal specifications for modeling and reasoning. Therefore, the processes are mostly hard-coded to match the logic framework.
2.5. Generative AI
2.6. The Gaps
- Self-supervised knowledge generation during the machine learning process and solution creation: In the past, knowledge generation system mainly referred to expert systems that acquire knowledge from human expertise or systems that transform existing knowledge from one presentation to the other. Enabling the understanding of common knowledge in the biomedical domain is crucial. Reference [29] presents an automatic process of disease causality knowledge generation from HTML-text documents. However, it still does not fully address the problem of how to automatically learn valuable knowledge from the whole task–solution–evaluation machine learning life cycle. Considering human-level intelligence, we always learn either directly from problem-solving or indirectly through other human expertise (e.g., reading a book or watching a video), or a combination of both (e.g., reflecting on the opinions of others).
- Provisioning a knowledge-guided auto-ML solution: In contrast to the first gap, there are no significant research works on using knowledge to assist in providing an AI solution. Again, compared with human-level intelligence, we always try to apply acquired knowledge or knowledge-based reasoning to solve a problem. We can consider that the transformer process [37] is a step forward in this direction. We can treat well-trained AI models as a type of knowledge to apply to different tasks in a similar problem domain. However, there is still no defined framework that can specify what knowledge is required and how to use the knowledge to find a solution to new tasks [16].
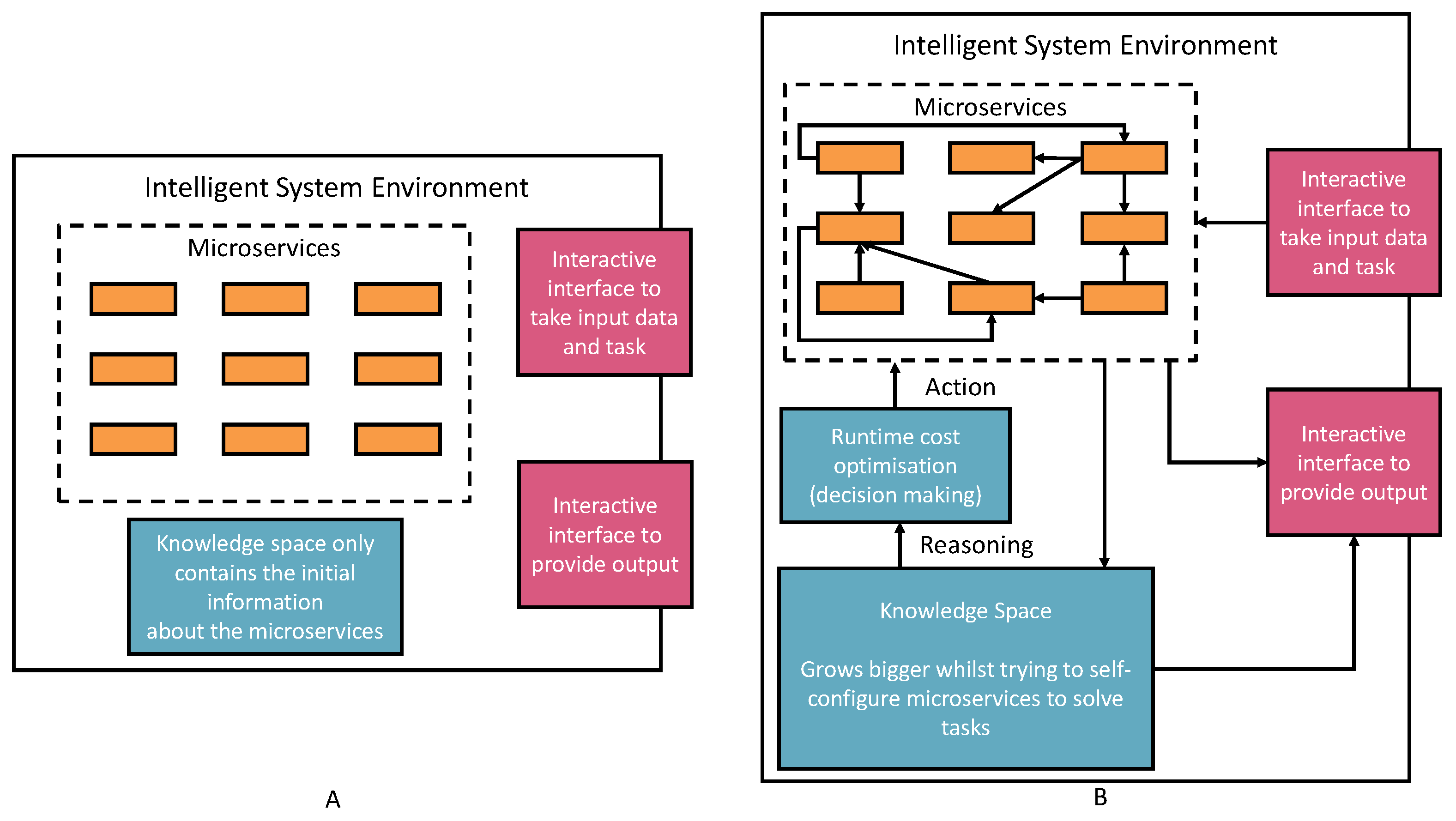
3. The Framework Architecture
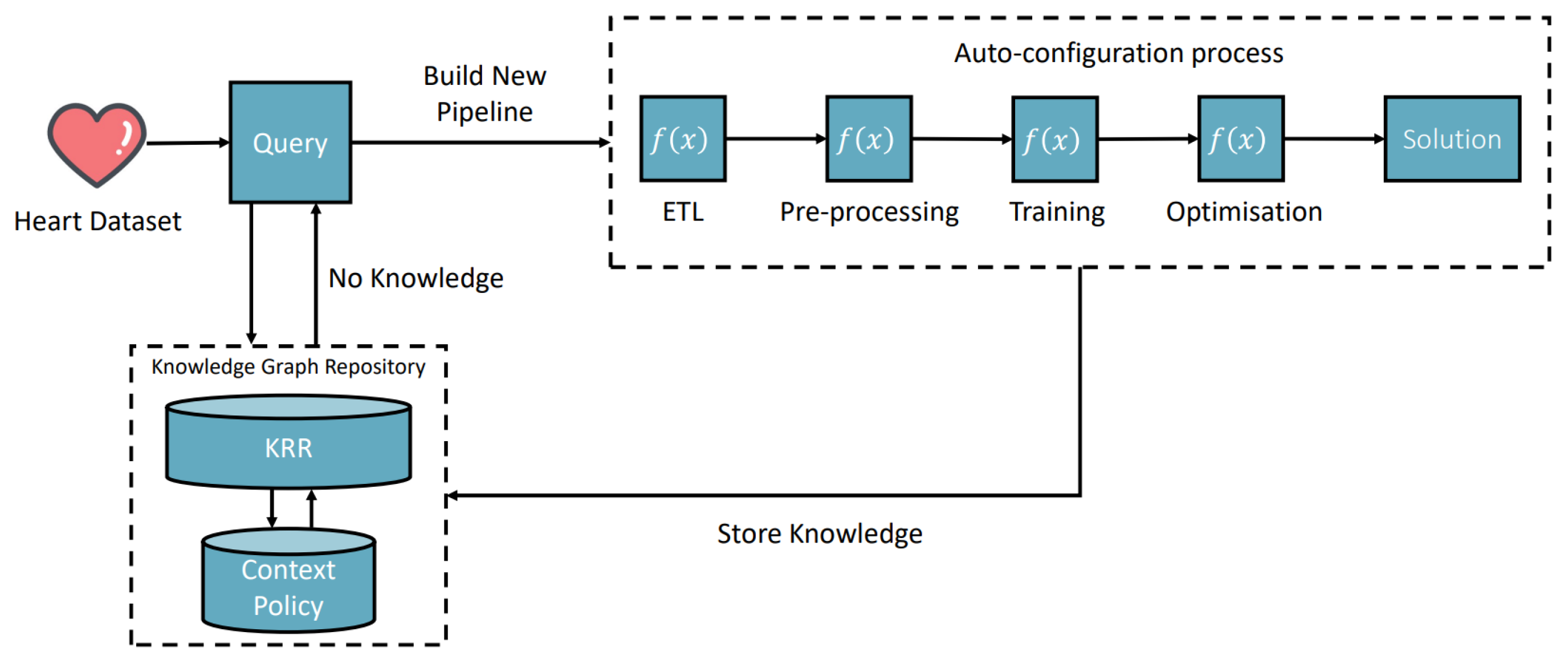
- Registering new AIMSs (Automatic Semantic Machine Learning Microservices) from outside the environment. The registration process is through the interactive interface according to the defined microservice ontology (see Figure 2). Therefore, human involvement in machine learning microservice engineering is a core part of this vision, which defines humans as educators to teach basic skills and capabilities to deal with different tasks. Then, the environment will reuse these skills and capabilities to acquire knowledge. The knowledge will provide powerful reasoning sources to independently deal with complex tasks, decision making, and creating new pipelines. Specific to the biomedical research, the registration ontology can refer to biomedical engineering ontologies, including Disease Ontology, Foundational Model of Anatomy (FMA), Human Phenotype Ontology (HPO), and many others [38].
- Taking tasks with a variety of inputs, such as CSV data files, images, text, and audio data: The environment autoconfigures on the default AIMSs and provides solutions to the tasks. The success or failure outcome will be recorded as knowledge. Microservice autoconfiguration refers to the automated setup and configuration of individual microservices in a pipeline to serve a machine learning task in our context. The microservice human engineering process will start if there are no suitable AIMSs to deal with the task.
- The environment can compose multiple AIMSs to complete a task if one single microservice cannot achieve it.
- The environment can start learning, representing, and storing knowledge in the knowledge space as knowledge graph data. The knowledge is derived from processing input data, the autoconfiguration process, and task outcomes. The knowledge size will increase and thus provide better optimizations, autoconfiguration, and feedback to the system user.
4. Self-Supervised Knowledge Learning for Solution Generation
5. Experimental Implementation
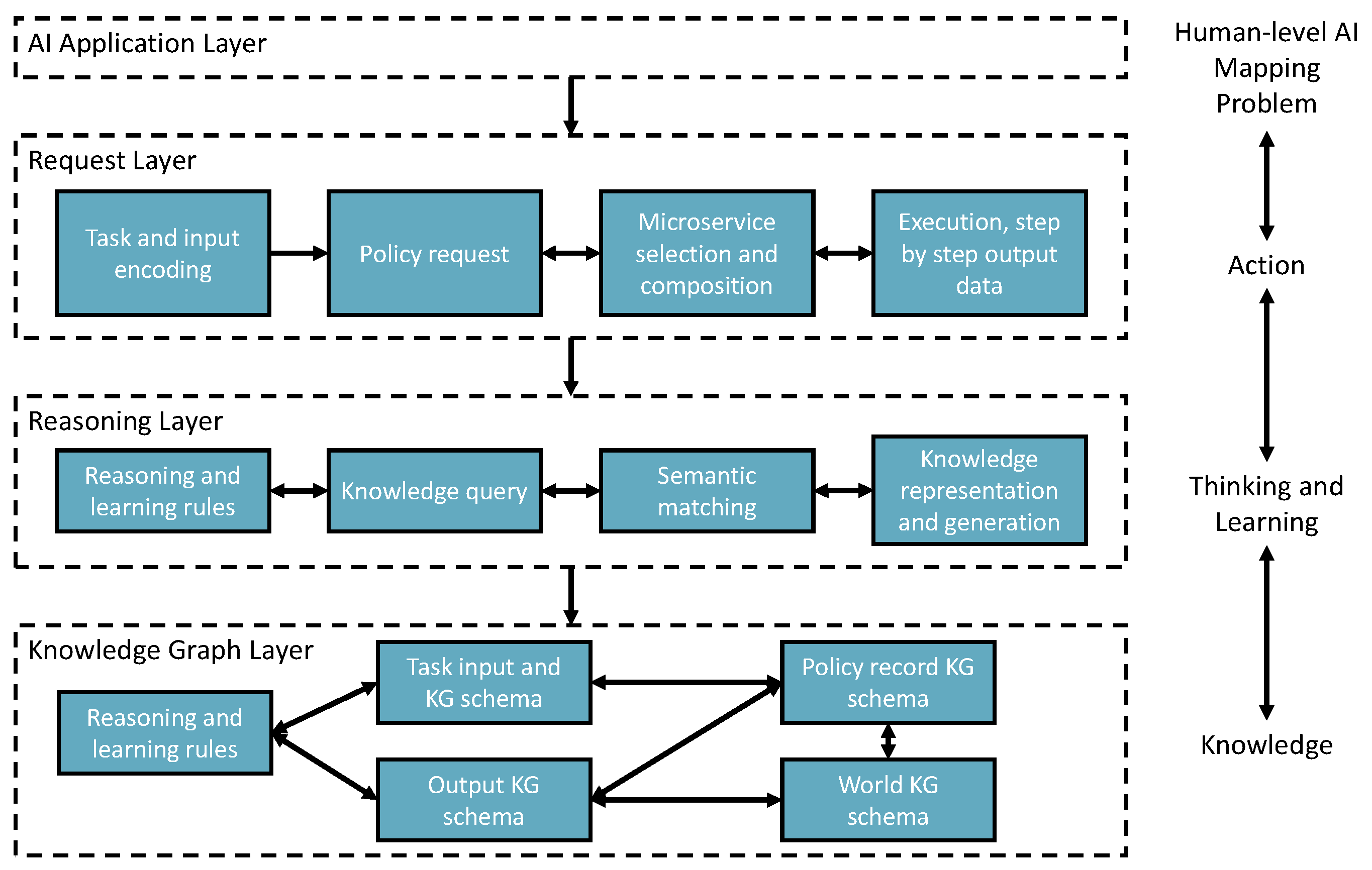
- The request layer takes tasks and inputs from AI applications to trigger the solution searching and self-learning processes. Task context is semantically encoded to enable starting the policy knowledge to explore the environment for learning, creating, or finding solutions.
- The reasoning layer takes the request layer’s semantic reasoning tasks for semantic matching, reasoning, and performing the reinforcement learning mechanism. Finally, the policy will be recorded in the knowledge graph layer. In addition, the newly added AIMSs are registered to the environment with semantic annotations through knowledge registration and generation components.
- The knowledge graph layer remembers the knowledge data in the knowledge graph triple store based on different types of knowledge schemata.
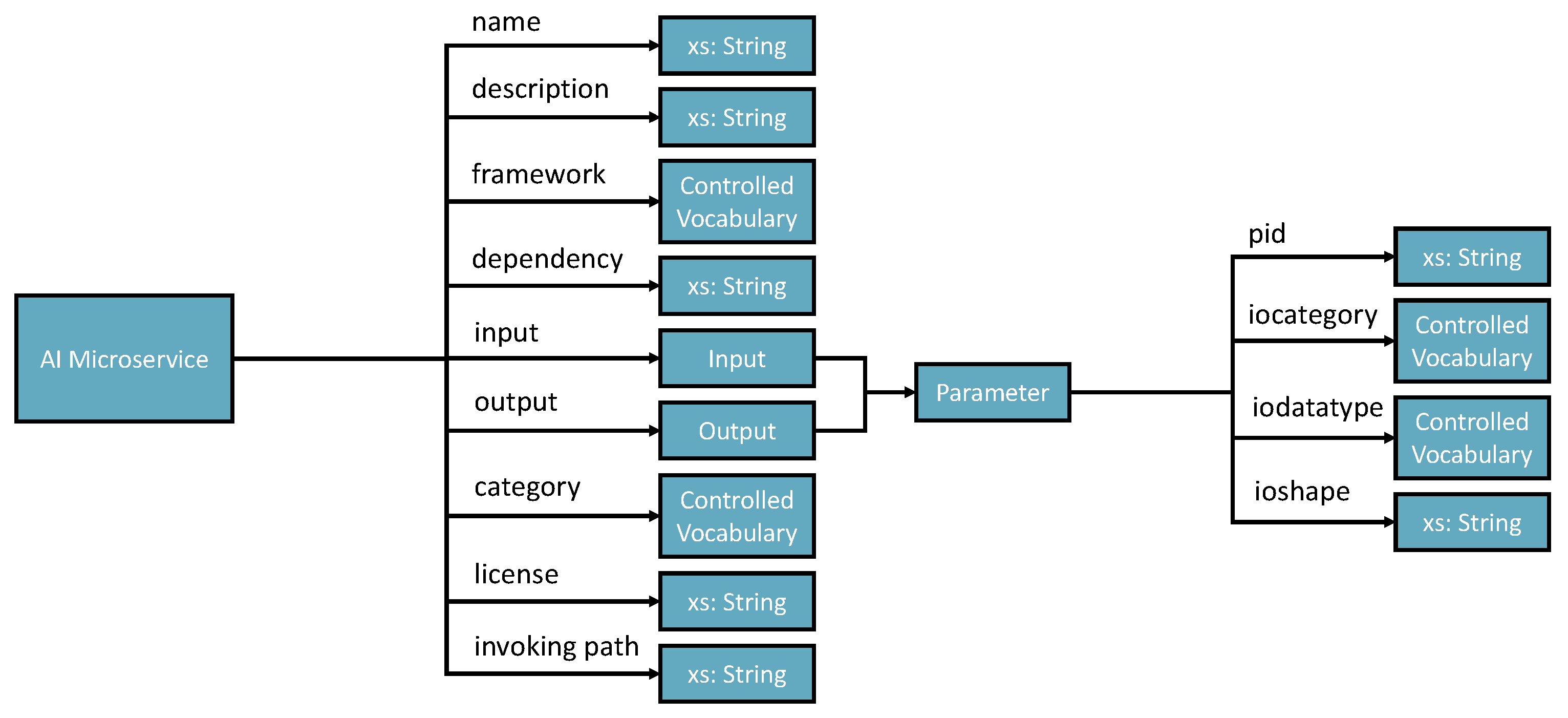
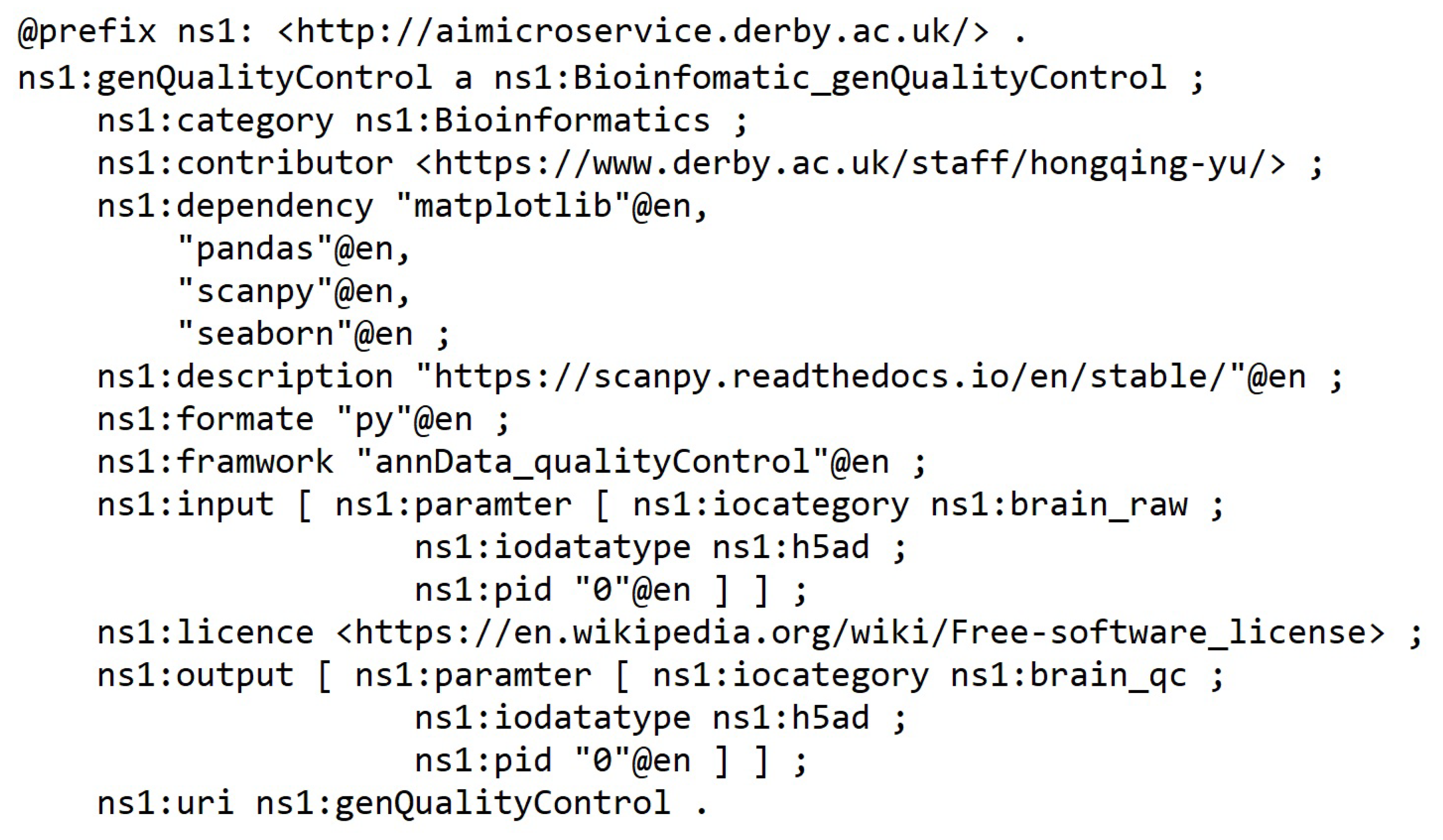
5.1. Knowledge Ontology Implementation
- Name— Must be no duplication in the system, and the registration process will check the name’s legibility.
- Description—A short presentation of the AIMS for human understanding.
- Framework—Indicates the programming framework used to develop the AIMS. Normally, it should be just one framework as AIMS is designed to be decoupled and ideally single responsibility.
- Dependency—Describes the required programming libraries that need to be preconfigured to enable the AIMS to work. The schema includes id, library install port URI and version.
- Input and output—Specifies the parameters that should be in the input and output messages.
- Category—Tells what AI-related domain the ms works on, such as supervised classification, unsupervised clustering, image classification base model, and more.
- License—Identifies the use conditions and copyright of the AIMS.
- Invoke path—Contains the portal for accessing the AIMS. The path can be a local path or URI of a restful API.
- The type of input data—A controlled variable that majorly includes normal dataset (e.g., tableau data stored in a CSV file, image, or text).
- Task domain—Free text to record the specific application domain.
- Desire output type—Records the output required to complete the task successfully.
- The parameters—The dataset or data presents the initial characters of the input data. For example, the number of columns and column names of the tableau data will be remembered as part of the context knowledge of the task.
- Policy context—Links to a task-context.
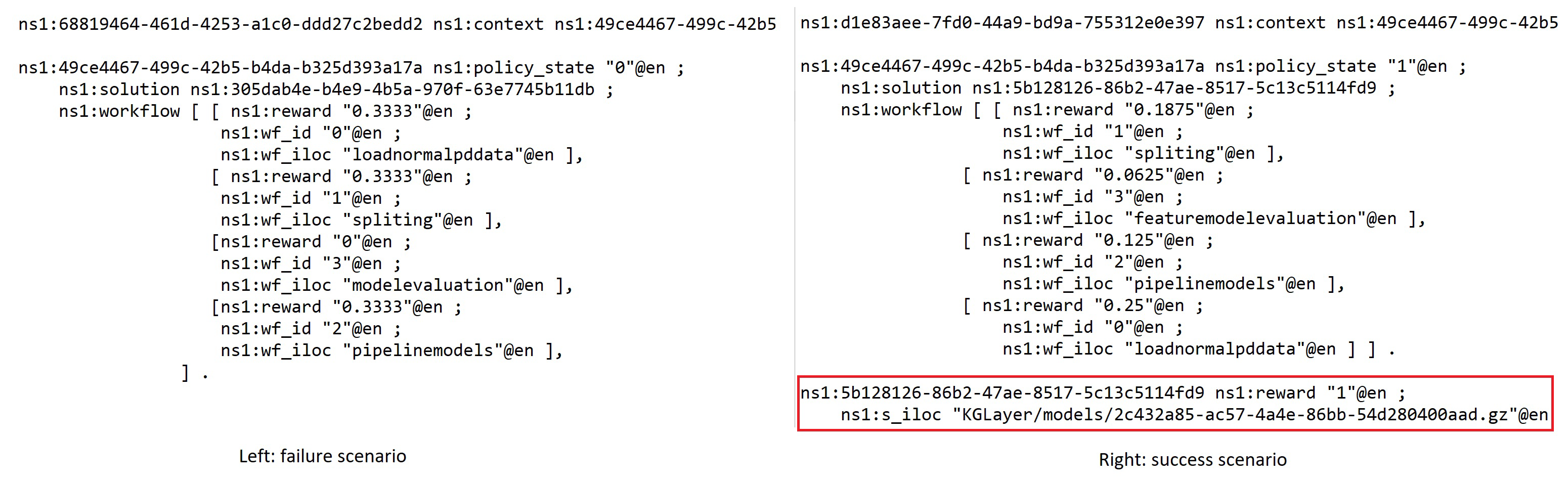
- Policy state—1 is success and 0 is failure, the binary state only presents whether the whole workflow is work or not but inside the workflow context that shows the continuous measure of individual component’s potential contribution toward to success in other possible solutions (see Figure 5).
- Solution iloc—The location where the solution can be loaded and executed.
- Workflow—Presents a pipeline solution that composes multiple AIMSs.
- Solution reward—The reward value stored for the policy that can be the recommended guidance for supporting the creation of a new task solution.
- Feature optimization outcomes—The features selected in the optimization are valuable, and these features will be reused to create a classification model if the new dataset features are the same.
- Answers for a certain text topic—A generated text answer for a question. The answer quality will be reported as a reward value feedback from humans back to the policy knowledge.
- Image RBG vectors—Map to a classification label. The reward process is the same as the answers.
5.2. Environment Initialization
- 1.
- Data processing AIMSs that include CSV files to a Panda service, data training and split services, data quality control services, data normalization services, image process services, and data quality control services.
- 2.
- 3.
- RFECV optimization services.
6. Scenario Evaluation and Lessons Learned
6.1. Heart Disease Classification Scenario
- Input: 335 clinical CSV heart disease files labeled 0 (no disease) and 1 (confirmed disease).
- Domain: Medical.
- Desire output: An optimized classification pipeline model.
- Age: The person’s age in years.
- Sex: The person’s sex (1 = male, 0 = female).
- cp: Chest pain type—value 0: asymptomation, value 1: atypical angina, value 2: nonanginal pain, value 3: typical angina.
- trestbps: The person’s resting blood pressure (mm Hg on admission to the hospital).
- chol: The person’s cholesterol measurement in mg/dL.
- fbs: The person’s fasting blood sugar (>120 mg/dL, 1 = true; 0 = false).
- restecg: Resting electrocardiographic results.
- thalach: The person’s maximum heart rate achieved.
- exang: Exercise-induced angina (1 = yes; 0 = no).
- oldpeak: ST depression induced by exercise relative to rest (ST relates to positions on the ECG plot.
- Slope: The slope of the peak exercise ST segment—0: downsloping; 1: flat; 2: upsloping.
- ca: The number of major vessels (0–3).
- thal: A blood disorder called thalassemia.
- target Heart disease (1 = no, 0 = yes).
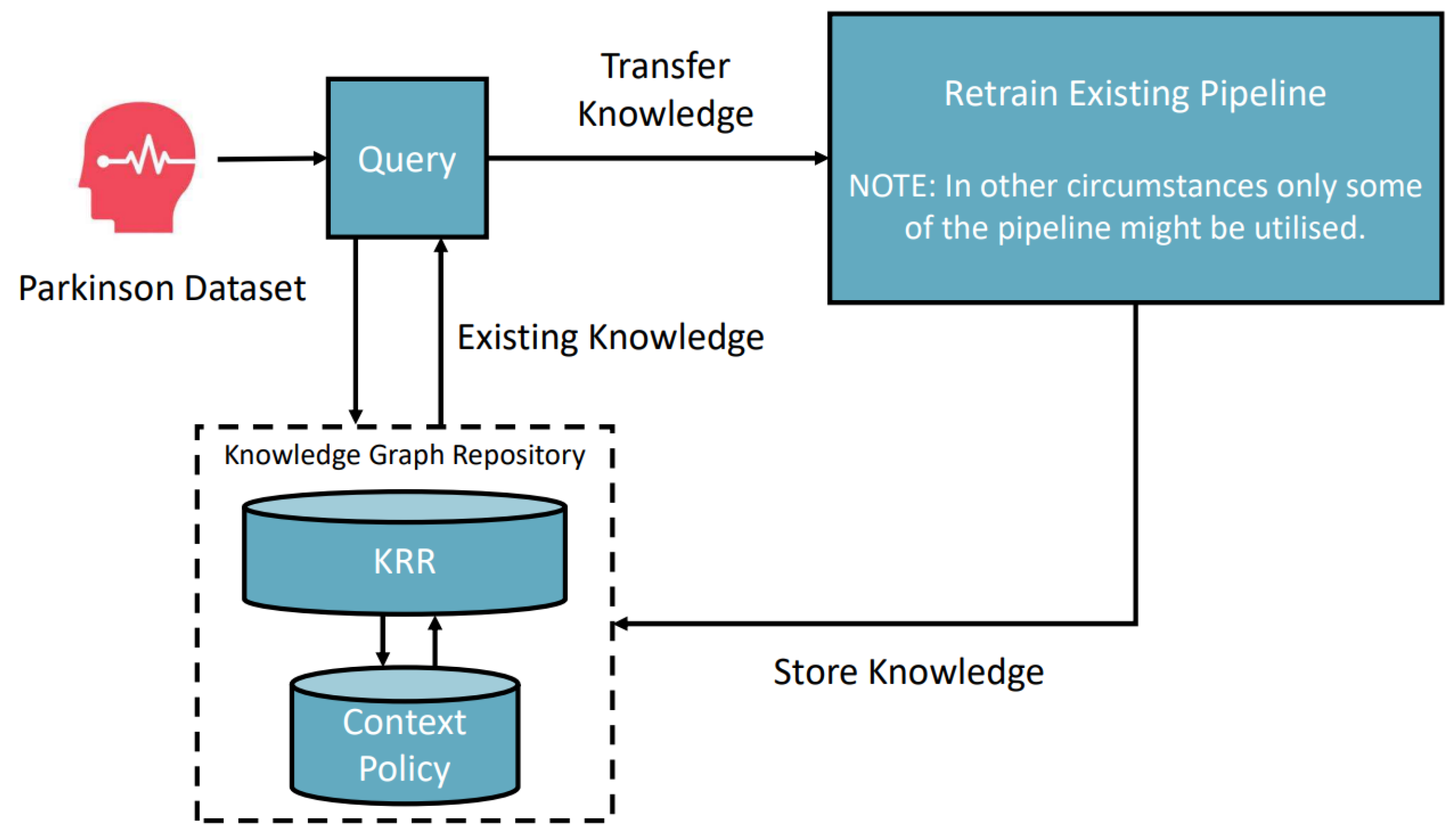
6.2. Parkinson Disease Classification Scenario
- Input: CSV Parkinson’s disease clinical example data with labels 0 (no disease) and 1 (confirmed disease).
- Domain: Medical.
- Desired output: An optimized classification pipeline model.
6.3. A Complex Scenario: Mouse Brain Single-Cell RNASeq Downstream Analysis
- Data quality control: This aims to find and remove the poor-quality cell observation data which were not detected in the previous processing of the raw data. The low-quality cell data may potentially introduce analysis noise and obscure the biological signals of interest in the downstream analysis.
- Data normalization: Dimensionality reduction and scaling of the data. Biologically, dimensional reduction is valuable and appropriate since cells respond to their environment by turning on regulatory programs that result in the expression of modules of genes. As a result, gene expression displays structured coexpression, and dimensionality reduction by the algorithm such as principle component analysis can group these covarying genes into principle components, ordered by how much variation they explained.
- Data feature embedding: Further dimensionality reduction using advanced algorithms, such as t-SNE and UMAP. They are powerful tools for visualizing and understanding big and high-dimensional datasets.
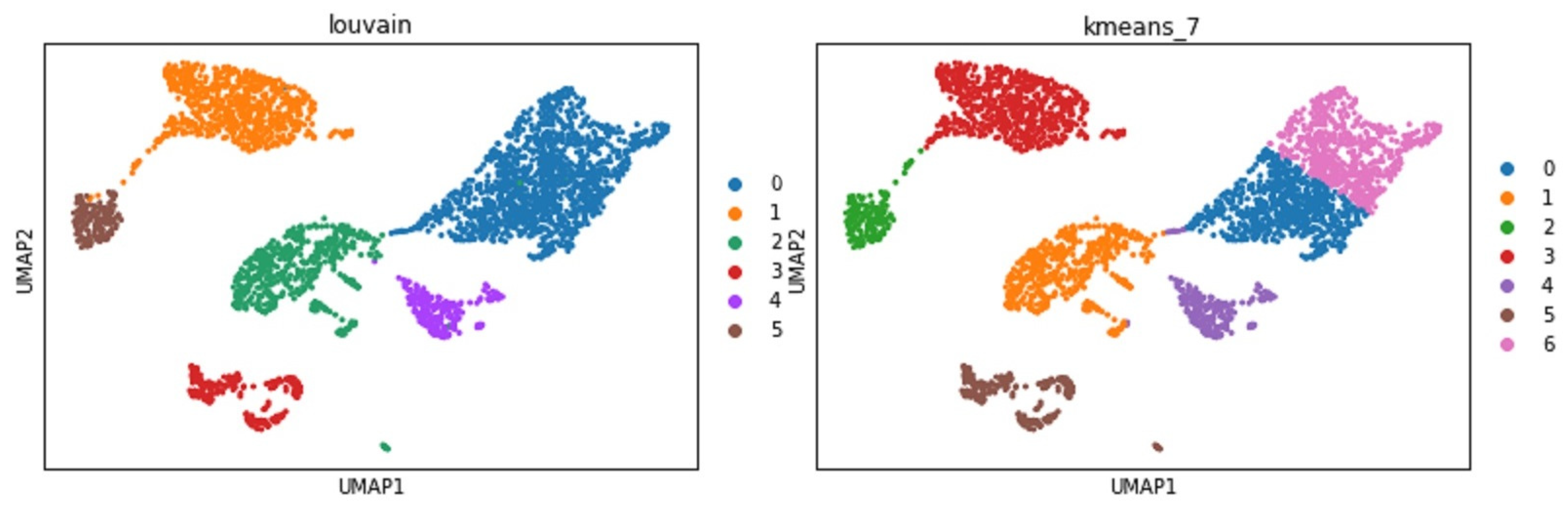
- Clustering analysis: Groups cells into different clusters based on the embedded features.

7. Discussion
- The advantage of using a triple KG structure to encode KRR elements lies in its unification, standardization, and adaptability across diverse applications. Nonetheless, as the KG expands, its referencing efficiency diminishes, particularly with intricate graph queries. This inefficiency is exacerbated when different knowledge types are stored separately, making union queries on the graph resource-intensive. A proposed solution is to embed the knowledge graph into a more efficient vector space [44]. To achieve this, we plan on investigating state-of-the-art embedding techniques, such as graph neural networks, that can maintain the relationships between entities while offering efficient querying.
- The current system architecture does not support multimodal inputs pertaining to a singular task (multimodal machine learning). While humans can seamlessly integrate visual, auditory, and other sensory data to accomplish tasks, machines struggle to synthesize multiple data types [45,46,47]. Moving forward, we aim to explore fusion techniques, both at the feature and decision levels, to facilitate more comprehensive input processing.
- During the initial stages of our manuscript’s preparation, Google released research papers detailing the mutation of neural networks (NNs) to handle diverse image classification tasks [16,17]. These papers have illuminated the potential of not just mutating data or services but also the possibility of adding or removing NN hidden layers as a form of knowledge storage for future considerations. Our intent is to delve deeper into the dynamics of such mutations and explore frameworks that allow for flexible and dynamic architectural changes in neural networks.
8. Conclusions
Author Contributions
Funding
Data Availability Statement
Conflicts of Interest
References
- Obermeyer, Z.; Emanuel, E.J. Predicting the future—Big data, machine learning, and clinical medicine. N. Engl. J. Med. 2016, 375, 1216–1219. [Google Scholar] [CrossRef] [PubMed]
- Miotto, R.; Wang, F.; Wang, S.; Jiang, X.; Dudley, J.T. Deep learning for healthcare: Review, opportunities and challenges. Briefings Bioinform. 2017, 19, 1236–1246. [Google Scholar] [CrossRef] [PubMed]
- Waring, J.; Lindvall, C.; Umeton, R. Automated machine learning: Review of the state-of-the-art and opportunities for healthcare. Artif. Intell. Med. 2020, 104, 101822. [Google Scholar] [CrossRef]
- Holzinger, A.; Langs, G.; Denk, H.; Zatloukal, K.; Müller, H. Causability and explainability of artificial intelligence in medicine. Wiley Interdiscip. Rev. Data Min. Knowl. Discov. 2019, 9, e1312. [Google Scholar] [CrossRef] [PubMed]
- Zheng, W.; Lin, H.; Liu, X.; Xu, B. A document level neural model integrated domain knowledge for chemical-induced disease relations. BMC Bioinform. 2018, 19, 328. [Google Scholar] [CrossRef] [PubMed]
- Abadi, M.; Agarwal, A.; Barham, P.; Brevdo, E.; Chen, Z.; Citro, C.; Corrado, G.S.; Davis, A.; Dean, J.; Devin, M.; et al. TensorFlow: Large-Scale Machine Learning on Heterogeneous Systems. 2015. Available online: tensorflow.org (accessed on 7 May 2023).
- Pedregosa, F.; Varoquaux, G.; Gramfort, A.; Michel, V.; Thirion, B.; Grisel, O.; Blondel, M.; Prettenhofer, P.; Weiss, R.; Dubourg, V.; et al. Scikit-learn: Machine Learning in Python. J. Mach. Learn. Res. 2011, 12, 2825–2830. [Google Scholar]
- Google LLC. Google Cloud AutoML. 2021. Available online: https://cloud.google.com/automl/docs (accessed on 24 September 2023).
- Le, T.T.; Fu, W.; Moore, J.H. Scaling tree-based automated machine learning to biomedical big data with a feature set selector. Bioinformatics 2020, 36, 250–256. [Google Scholar] [CrossRef]
- H2O.ai. H2O AutoML. 2023. Available online: https://docs.h2o.ai/h2o/latest-stable/h2o-docs/automl.html (accessed on 24 September 2023).
- LeDell, E.; Poirier, S. H2O AutoML: Scalable Automatic Machine Learning. In Proceedings of the 7th ICML Workshop on Automated Machine Learning (AutoML), Vienna, Austria, 12–8 July 2020. [Google Scholar]
- Ramsundar, B.; Eastman, P.; Walters, P.; Pande, V. Deep Learning for the Life Sciences: Applying Deep Learning to Genomics, Microscopy, Drug Discovery, and More; O’Reilly Media: Sebastopol, CA, USA, 2019. [Google Scholar]
- He, J.; Baxter, S.L.; Xu, J.; Xu, J.; Zhou, X.; Zhang, K. The practical implementation of artificial intelligence technologies in medicine. Nat. Med. 2019, 25, 30–36. [Google Scholar] [CrossRef]
- Mustafa, A.; Rahimi Azghadi, M. Automated Machine Learning for Healthcare and Clinical Notes Analysis. Computers 2021, 10, 24. [Google Scholar] [CrossRef]
- Ntoutsi, E.; Fafalios, P.; Gadiraju, U.; Iosifidis, V.; Nejdl, W.; Vidal, M.E.; Ruggieri, S.; Turini, F.; Papadopoulos, S.; Krasanakis, E.; et al. Bias in data-driven artificial intelligence systems—An introductory survey. WIREs Data Min. Knowl. Discov. 2020, 10, e1356. [Google Scholar] [CrossRef]
- Gesmundo, A.; Dean, J. An Evolutionary Approach to Dynamic Introduction of Tasks in Large-scale Multitask Learning Systems. arXiv 2022, arXiv:2205.12755. [Google Scholar]
- Gesmundo, A.; Dean, J. muNet: Evolving Pretrained Deep Neural Networks into Scalable Auto-tuning Multitask Systems. arXiv 2022, arXiv:2205.10937. [Google Scholar]
- LeCun, Y. A Path Towards Autonomous Machine Intelligence. Open Review. 27 June 2022. Available online: https://openreview.net/pdf?id=BZ5a1r-kVsf (accessed on 24 September 2023).
- Yao, Q.; Wang, M.; Escalante, H.J.; Guyon, I.; Hu, Y.; Li, Y.; Tu, W.; Yang, Q.; Yu, Y. Taking Human out of Learning Applications: A Survey on Automated Machine Learning. arXiv 2018, arXiv:1810.13306. [Google Scholar]
- Jin, H.; Song, Q.; Hu, X. Auto-Keras: An Efficient Neural Architecture Search System. arXiv 2018, arXiv:1806.10282. [Google Scholar]
- Sharma, L.; Garg, P.K. Knowledge representation in artificial intelligence: An overview. In Artificial Intelligence; CRC: Boca Raton, FL, USA, 2021; pp. 19–28. [Google Scholar]
- Cozman, F.G.; Munhoz, H.N. Some thoughts on knowledge-enhanced machine learning. Int. J. Approx. Reason. 2021, 136, 308–324. [Google Scholar] [CrossRef]
- Hu, Z.; Yang, Z.; Salakhutdinov, R.; Xing, E. Deep neural networks with massive learned knowledge. In Proceedings of the 2016 Conference on Empirical Methods in Natural Language Processing, Austin, TX, USA, 1–5 November 2016; pp. 1670–1679. [Google Scholar]
- Chen, X.; Jia, S.; Xiang, Y. A review: Knowledge reasoning over knowledge graph. Expert Syst. Appl. 2020, 141, 112948. [Google Scholar] [CrossRef]
- Berners-Lee, T.; Hendler, J.; Lassila, O. The semantic web. Sci. Am. 2001, 284, 34–43. [Google Scholar] [CrossRef]
- Baader, F.; Horrocks, I.; Lutz, C.; Sattler, U. Introduction to Description Logic; Cambridge University Press: Cambridge, UK, 2017. [Google Scholar]
- Zhang, Q. Dynamic Uncertain Causality Graph for Knowledge Representation and Probabilistic Reasoning: Directed Cyclic Graph and Joint Probability Distribution. IEEE Trans. Neural Netw. Learn. Syst. 2015, 26, 1503–1517. [Google Scholar] [CrossRef]
- Botha, L.; Meyer, T.; Peñaloza, R. The Probabilistic Description Logic. Theory Pract. Log. Program. 2021, 21, 404–427. [Google Scholar] [CrossRef]
- Yu, H.Q.; Reiff-Marganiec, S. Learning Disease Causality Knowledge From the Web of Health Data. Int. J. Semant. Web Inf. Syst. (IJSWIS) 2022, 18, 1–19. [Google Scholar] [CrossRef]
- Zhao, X.; Liu, E.; Yu, H.Q.; Clapworthy, G.J. A Linear Logic Approach to the Composition of RESTful Web Services. Int. J. Web Eng. Technol. 2015, 10, 245–271. [Google Scholar] [CrossRef]
- Allameh Amiri, M.; Serajzadeh, H. QoS aware web service composition based on genetic algorithm. In Proceedings of the 2010 5th International Symposium on Telecommunications, Kauai, HI, USA, 5–8 January 2010; pp. 502–507. [Google Scholar] [CrossRef]
- Qiang, B.; Liu, Z.; Wang, Y.; Xie, W.; Xina, S.; Zhao, Z. Service composition based on improved genetic algorithm and analytical hierarchy process. Int. J. Robot. Autom. 2018. [Google Scholar] [CrossRef]
- Yu, H.Q.; Zhao, X.; Reiff-Marganiec, S.; Domingue, J. Linked Context: A Linked Data Approach to Personalised Service Provisioning. In Proceedings of the 2012 IEEE 19th International Conference on Web Services, Honolulu, HI, USA, 24–29 June 2012; pp. 376–383. [Google Scholar] [CrossRef]
- Dong, H.; Hussain, F.; Chang, E. Semantic Web Service matchmakers: State of the art and challenges. Concurr. Comput. Pract. Exp. 2013, 25, 961–988. [Google Scholar] [CrossRef]
- Publio, G.C.; Esteves, D.; Lawrynowicz, A.; Panov, P.; Soldatova, L.N.; Soru, T.; Vanschoren, J.; Zafar, H. ML-Schema: Exposing the Semantics of Machine Learning with Schemas and Ontologies. arXiv 2018, arXiv:1807.05351. [Google Scholar]
- Braga, J.; Dias, J.; Regateiro, F. A machine learning ontology. Frenxiv Pap. 2020, preprint. [Google Scholar] [CrossRef]
- Tan, C.; Sun, F.; Kong, T.; Zhang, W.; Yang, C.; Liu, C. A Survey on Deep Transfer Learning. In Proceedings of the Artificial Neural Networks and Machine Learning–ICANN 2018: 27th International Conference on Artificial Neural Networks, Rhodes, Greece, 4–7 October 2018; Proceedings, Part III 27. Kůrková, V., Manolopoulos, Y., Hammer, B., Iliadis, L., Maglogiannis, I., Eds.; Springer International Publishing: Cham, Switzerland, 2018; pp. 270–279. [Google Scholar]
- Filice, R.W.; Kahn, C.E.J. Biomedical Ontologies to Guide AI Development in Radiology. J. Digit. Imaging 2021, 34, 1331–1341. [Google Scholar] [CrossRef]
- Black, S.; Leo, G.; Wang, P.; Leahy, C.; Biderman, S. GPT-Neo: Large Scale Autoregressive Language Modeling with Mesh-Tensorflow (1.0). Zenodo 2021. [Google Scholar] [CrossRef]
- Gao, L.; Biderman, S.; Black, S.; Golding, L.; Hoppe, T.; Foster, C.; Phang, J.; He, H.; Thite, A.; Nabeshima, N.; et al. The Pile: An 800GB Dataset of Diverse Text for Language Modeling. arXiv 2020, arXiv:2101.00027. [Google Scholar]
- Bhojanapalli, S.; Chakrabarti, A.; Glasner, D.; Li, D.; Unterthiner, T.; Veit, A. Understanding Robustness of Transformers for Image Classification. In Proceedings of the IEEE/CVF International Conference on Computer Vision (ICCV), Montreal, BC, Canada, 11–17 October 2021; pp. 10231–10241. [Google Scholar]
- Luecken, M.D.; Theis, F.J. Current best practices in single-cell RNA-seq analysis: A tutorial. J. Mol. Syst. Biol. 2019, 15, e8746. [Google Scholar] [CrossRef]
- Cannoodt, R. Anndata: ‘Anndata’ for R. 2022. Available online: https://anndata.readthedocs.io/en/latest/ (accessed on 24 September 2023).
- Le, T.; Huynh, N.; Le, B. Link Prediction on Knowledge Graph by Rotation Embedding on the Hyperplane in the Complex Vector Space. In Proceedings of the Artificial Neural Networks and Machine Learning–ICANN 2021: 30th International Conference on Artificial Neural Networks, Bratislava, Slovakia, 14–17 September 2021; Proceedings, Part III 30. Farkaš, I., Masulli, P., Otte, S., Wermter, S., Eds.; Springer International Publishing: Cham, Switzerland, 2021; pp. 164–175. [Google Scholar]
- Ramachandram, D.; Taylor, G.W. Deep multimodal learning: A survey on recent advances and trends. IEEE Signal Process. Mag. 2017, 34, 96–108. [Google Scholar] [CrossRef]
- Baltrušaitis, T.; Ahuja, C.; Morency, L.P. Multimodal machine learning: A survey and taxonomy. IEEE Trans. Pattern Anal. Mach. Intell. 2018, 41, 423–443. [Google Scholar] [CrossRef] [PubMed]
- Li, J.; Hong, D.; Gao, L.; Yao, J.; Zheng, K.; Zhang, B.; Chanussot, J. Deep learning in multimodal remote sensing data fusion: A comprehensive review. arXiv 2022, arXiv:2205.01380. [Google Scholar] [CrossRef]

Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2023 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Yu, H.Q.; O’Neill, S.; Kermanizadeh, A. AIMS: An Automatic Semantic Machine Learning Microservice Framework to Support Biomedical and Bioengineering Research. Bioengineering 2023, 10, 1134. https://doi.org/10.3390/bioengineering10101134
Yu HQ, O’Neill S, Kermanizadeh A. AIMS: An Automatic Semantic Machine Learning Microservice Framework to Support Biomedical and Bioengineering Research. Bioengineering. 2023; 10(10):1134. https://doi.org/10.3390/bioengineering10101134
Chicago/Turabian StyleYu, Hong Qing, Sam O’Neill, and Ali Kermanizadeh. 2023. "AIMS: An Automatic Semantic Machine Learning Microservice Framework to Support Biomedical and Bioengineering Research" Bioengineering 10, no. 10: 1134. https://doi.org/10.3390/bioengineering10101134
APA StyleYu, H. Q., O’Neill, S., & Kermanizadeh, A. (2023). AIMS: An Automatic Semantic Machine Learning Microservice Framework to Support Biomedical and Bioengineering Research. Bioengineering, 10(10), 1134. https://doi.org/10.3390/bioengineering10101134







