Hematotoxic Effect of Respiratory Exposure to PHMG-p and Its Integrated Genetic Analysis
Abstract
1. Background
2. Methods and Materials
2.1. Animals
2.2. Experimental Design
2.3. Blood Samples
2.4. Reagents
2.5. Histologic and Immunohistochemical Examination
2.6. RNA Isolation
2.7. Library Preparation and Sequencing
2.8. Data Analysis
2.9. Statistical Analyses
3. Results
3.1. Altered Blood Composition Induced by PHMG-p Respiratory Exposure
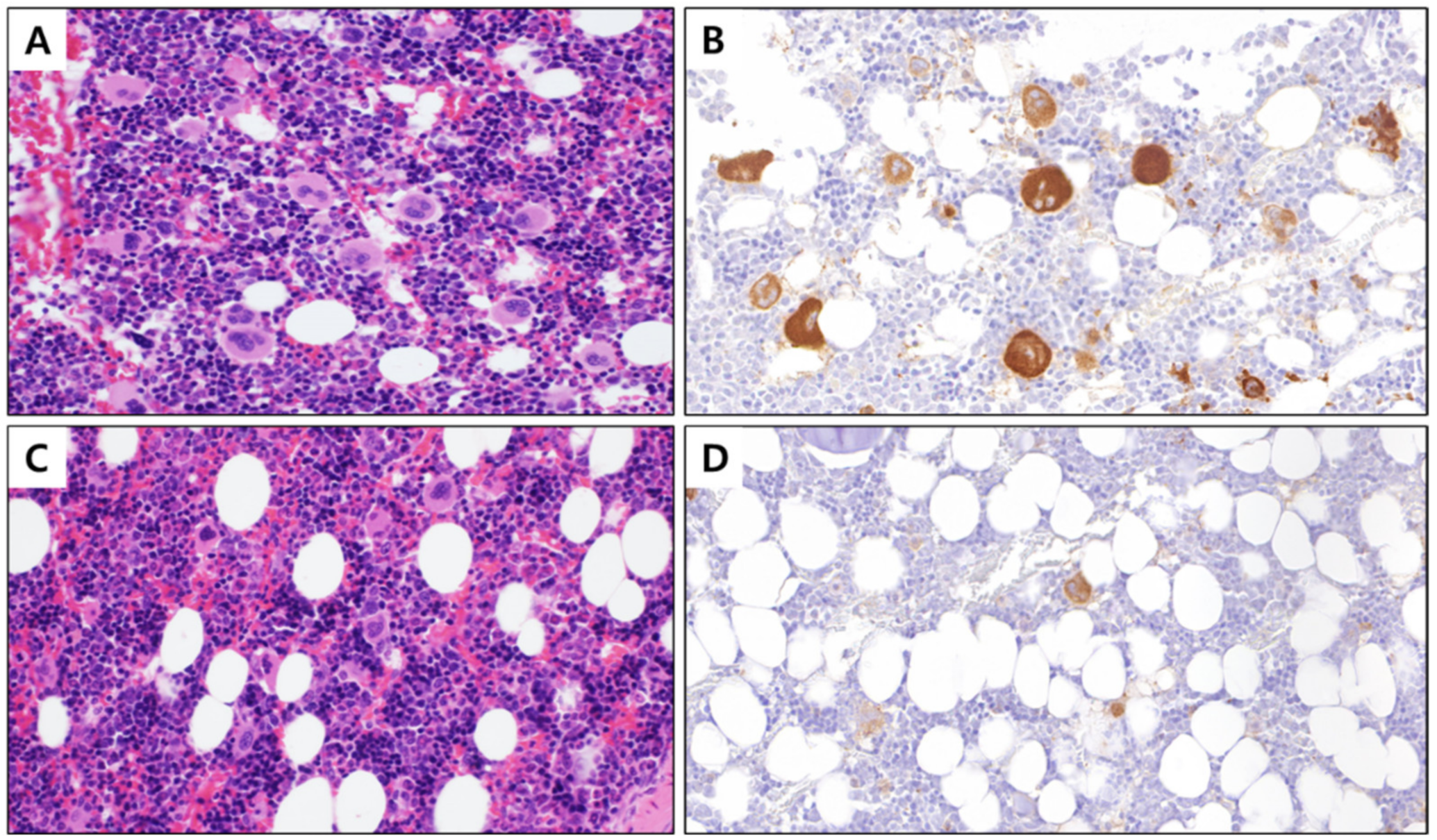
3.2. Number of Megakaryocytes in PHMG-p-Exposed Group Selected Based on Decreased PLT Count
3.3. Identification of Altered Genes Expression in BM with Reduced Megakaryocytes
3.4. Predicted Top Canonical Pathways Significantly Related to Candidate Genes
3.5. Predicted Top Diseases and Biological Functions Associated with Candidate Genes
3.6. Top Five Annotations Related to Hematological Diseases and Biofunctions
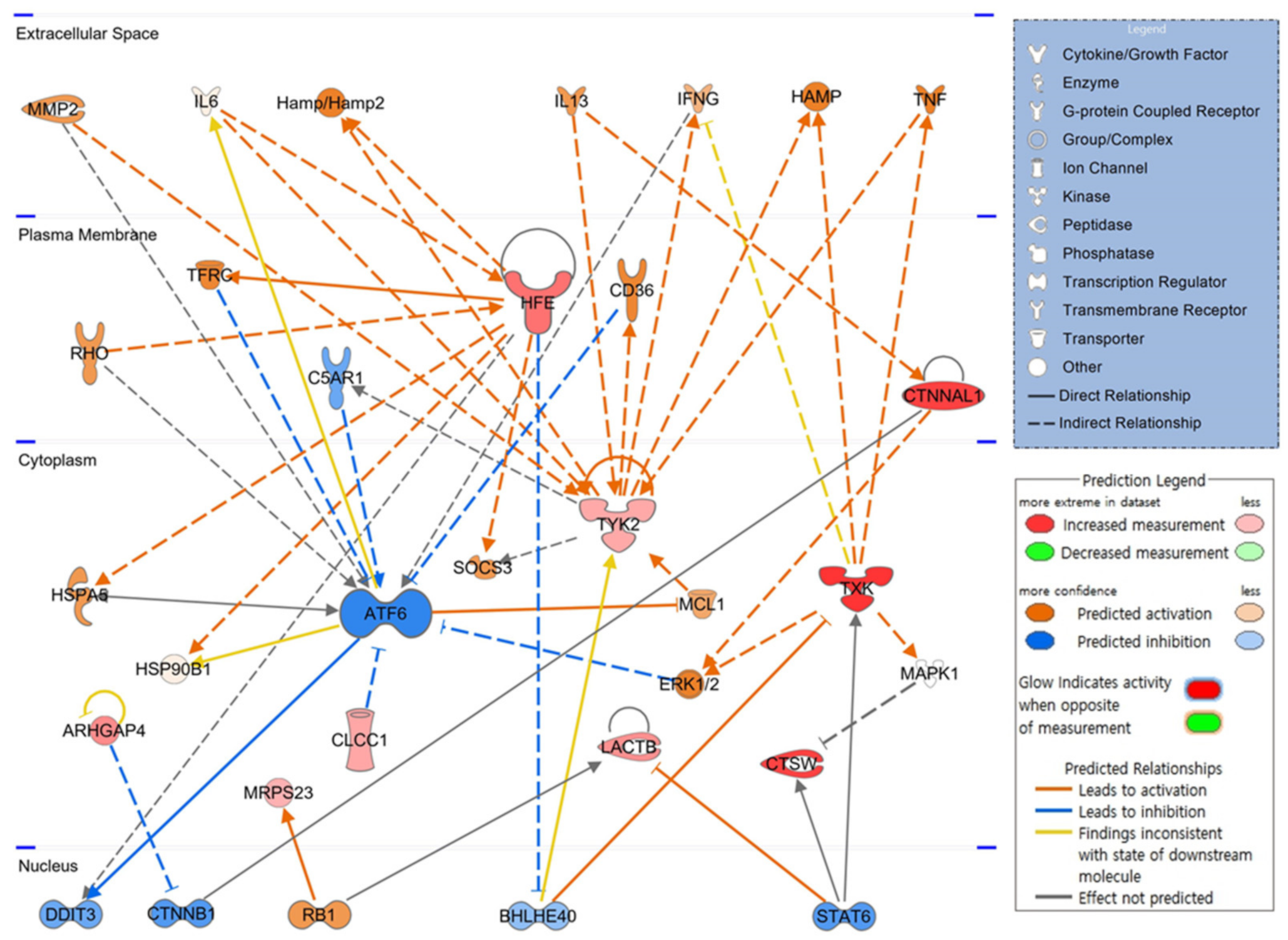
3.7. Schematic of Hematopoietic Signaling Pathways and Predicted Key Molecules via IPA
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
Abbreviations
References
- Paek, D.; Koh, Y.; Park, D.-U.; Cheong, H.-K.; Do, K.-H.; Lim, C.-M.; Hong, S.-J.; Kim, Y.-H.; Leem, J.-H.; Chung, K.H. Nationwide study of humidifier disinfectant lung injury in South Korea, 1994–2011. Incidence and dose-response relationships. Ann. Am. Thorac. Soc. 2015, 12, 1813–1821. [Google Scholar] [CrossRef] [PubMed]
- Yoon, J.; Cho, H.-J.; Lee, E.; Choi, Y.J.; Kim, Y.-H.; Lee, J.L.; Lee, Y.J.; Hong, S.-J. Rate of humidifier and humidifier disinfectant usage in Korean children: A nationwide epidemiologic study. Environ. Res. 2017, 155, 60–63. [Google Scholar] [CrossRef] [PubMed]
- Yoon, J.; Kang, M.; Jung, J.; Ju, M.J.; Jeong, S.H.; Yang, W.; Choi, Y.H. Humidifier disinfectant consumption and humidifier disinfectant-associated lung injury in South Korea: A nationwide population-based study. Int. J. Environ. Res. Public Health 2021, 18, 6136. [Google Scholar] [CrossRef] [PubMed]
- Kim, H.R.; Lee, K.; Park, C.W.; Song, J.A.; Shin, D.Y.; Park, Y.J.; Chung, K.H. Polyhexamethylene guanidine phosphate aerosol particles induce pulmonary inflammatory and fibrotic responses. Arch. Toxicol. 2015, 90, 617–632. [Google Scholar] [CrossRef]
- Choi, Y.; Paek, D. Humidifier disinfectants, unfinished stories. Environ. Health Toxicol. 2016, 31, e2016004. [Google Scholar] [CrossRef]
- Kim, J.Y.; Kim, H.H.; Cho, K.H. Acute cardiovascular toxicity of sterilizers, PHMG, and PGH: Severe inflammation in human cells and heart failure in zebrafish. Cardiovasc. Toxicol. 2013, 13, 148–160. [Google Scholar] [CrossRef]
- Snyder, R. Leukemia and benzene. Int. J. Environ. Res. Public Health 2012, 9, 2875–2893. [Google Scholar] [CrossRef]
- Wang, L.; He, X.; Bi, Y.; Ma, Q. Stem cell and benzene-induced malignancy and hematotoxicity. Chem. Res. Toxicol. 2012, 25, 1303–1315. [Google Scholar] [CrossRef]
- Kido, T.; Tamagawa, E.; Bai, N.; Suda, K.; Yang, H.-H.C.; Li, Y.; Chiang, G.; Yatera, K.; Mukae, H.; Sin, D.D. Particulate matter induces translocation of IL-6 from the lung to the systemic circulation. Am. J. Respir. Cell Mol. Biol. 2011, 44, 197–204. [Google Scholar] [CrossRef]
- Tan, W.C.; Qiu, D.; Liam, B.L.; Ng, T.P.; Lee, S.H.; van Eeden, S.F.; D’Yachkova, Y.; Hogg, J.C. The human bone marrow response to acute air pollution caused by forest fires. Am. J. Respir. Crit. Care Med. 2000, 161, 1213–1217. [Google Scholar] [CrossRef]
- Goto, Y.; Ishii, H.; Hogg, J.C.; Shih, C.-H.; Yatera, K.; Vincent, R.; van Eeden, S.F. Particulate matter air pollution stimulates monocyte release from the bone marrow. Am. J. Respir. Crit. Care Med. 2004, 170, 891–897. [Google Scholar] [CrossRef] [PubMed]
- Breivik, K.; Alcock, R.; Li, Y.-F.; Bailey, R.E.; Fiedler, H.; Pacyna, J.M. Primary sources of selected POPs: Regional and global scale emission inventories. Environ. Pollut. 2003, 128, 3–16. [Google Scholar] [CrossRef] [PubMed]
- Gasiewicz, T.A.; Singh, K.P.; Casado, F.L. The aryl hydrocarbon receptor has an important role in the regulation of hematopoiesis: Implications for benzene-induced hematopoietic toxicity. Chem. Biol. Interact. 2010, 184, 246–251. [Google Scholar] [CrossRef] [PubMed]
- Kim, C.; Jeong, S.H.; Kim, J.; Lee, K.Y.; Cha, J.; Lee, C.H.; Park, E.-K.; Lee, J.-H. Evaluation of polyhexamethylene guanidine-induced lung injuries by chest CT, pathologic examination, and RNA sequencing in a rat model. Sci. Rep. 2021, 11, 6318. [Google Scholar] [CrossRef] [PubMed]
- Simon, A. FastQC, version 0.11.9. Babraham Bioinformatics Last Modified 8 January 2019. Babraham Institute: Cambridge, UK, 2010.
- Hannon Lab. FASTX-Toolkit, version 0.0.13. FASTQ A Short-Reads Pre-Processing Tools, Last Modified 2 February 2010. Hannon Lab: Cambridge, UK, 2014. Available online: http://hannonlab.cshl.edu/fastx_toolkit/ (accessed on 9 December 2021).
- Bushnell, B. BBMap: BBMap Short Read Aligner, and Other Bioinformatic Tools, Last Modified 11 August 2021; SourceFORGE: San Diego, CA, USA, 2014; Available online: https://sourceforge.net/projects/bbmap/ (accessed on 9 December 2021).
- Trapnell, C.; Pachter, L.; Salzberg, S.L. TopHat: Discovering splice junctions with RNA-Seq. Bioinformatics 2009, 25, 1105–1111. [Google Scholar] [CrossRef] [PubMed]
- Roberts, A.; Trapnell, C.; Donaghey, J.; Rinn, J.L.; Pachter, L. Improving RNA-Seq expression estimates by correcting for fragment bias. Genome Biol. 2011, 12, R22. [Google Scholar] [CrossRef] [PubMed]
- R Core Team. R: A Language and Environment for Statistical Computing; R Foundation for Statistical Computing: Vienna, Austria, 2018. [Google Scholar]
- He, Q.; Su, G.; Liu, K.; Zhang, F.; Jiang, Y.; Gao, J.; Liu, L.; Jiang, Z.; Jin, M.; Xie, H. Sex-specific reference intervals of hematologic and biochemical analytes in Sprague-Dawley rats using the nonparametric rank percentile method. PLoS ONE 2017, 12, e0189837. [Google Scholar] [CrossRef]
- Alemáan, C.L.; Más, R.M.; Rodeiro, I.; Noa, M.; Hernández, C.; Menéndez, R.; Gámez, R. Reference database of the main physiological parameters in Sprague-Dawley rats from 6 to 32 months. Lab. Anim. 1998, 32, 457–466. [Google Scholar] [CrossRef]
- Han, Z.-Z.; Xu, H.-D.; Kim, K.-H.; Ahn, T.-H.; Bae, J.-S.; Lee, J.-Y.; Gil, K.-H.; Lee, J.-Y.; Woo, S.-J.; Yoo, H.-J. Reference data of the main physiological parameters in control Sprague-Dawley rats from pre-clinical toxicity studies. Lab. Anim. Res. 2010, 26, 153–164. [Google Scholar] [CrossRef]
- Lu, Y.C.; Sanada, C.; Xavier-Ferrucio, J.; Wang, L.; Zhang, P.X.; Grimes, H.L.; Venkatasubramanian, M.; Chetal, K.; Aronow, B.; Salomonis, N. The molecular signature of megakaryocyte-erythroid progenitors reveals a role for the cell cycle in fate specification. Cell Rep. 2018, 25, 2083–2093.e4. [Google Scholar] [CrossRef]
- Rieger, M.A.; Schroeder, T. Hematopoiesis. Cold Spring Harb. Perspect. Biol. 2012, 4, a008250. [Google Scholar] [CrossRef] [PubMed]
- Thiagarajan, P.; Parker, C.J.; Prchal, J.T. How do red blood cells die? Front. Physiol. 2021, 12, 655393. [Google Scholar] [CrossRef] [PubMed]
- Ballard, H.S. The hematological complications of alcoholism. Alcohol Health Res. World 1997, 21, 42–52. [Google Scholar] [CrossRef] [PubMed]
- McHale, C.M.; Zhang, L.; Smith, M.T. Current understanding of the mechanism of benzene-induced leukemia in humans: Implications for risk assessment. Carcinogenesis 2011, 33, 240–252. [Google Scholar] [CrossRef]
- Lopez, J.J.; Palazzo, A.; Chaabane, C.; Albarran, L.; Polidano, E.; Lebozec, K.; Dally, S.; Nurden, P.; Enouf, J.; Debili, N. Crucial role for endoplasmic reticulum stress during megakaryocyte maturation. Arterioscler. Thromb. Vasc. Biol. 2013, 33, 2750–2758. [Google Scholar] [CrossRef]
- Ye, J.; Rawson, R.B.; Komuro, R.; Chen, X.; Davé, U.P.; Prywes, R.; Brown, M.S.; Goldstein, J.L. ER stress induces cleavage of membrane-bound ATF6 by the same proteases that process SREBPs. Mol. Cell 2000, 6, 1355–1364. [Google Scholar] [CrossRef]
- Yoshida, H.; Matsui, T.; Yamamoto, A.; Okada, T.; Mori, K. XBP1 mRNA is induced by ATF6 and spliced by IRE1 in response to ER stress to produce a highly active transcription factor. Cell 2001, 107, 881–891. [Google Scholar] [CrossRef]
- Yan, D.; Wang, H.-W.; Bowman, R.L.; Joyce, J.A. STAT3 and STAT6 signaling pathways synergize to promote cathepsin secretion from macrophages via IRE1alpha activation. Cell Rep. 2016, 16, 2914–2927. [Google Scholar] [CrossRef]
- Macaulay, I.C.; Thon, J.N.; Tijssen, M.R.; Steele, B.M.; Macdonald, B.T.; Meade, G.; Burns, P.; Rendon, A.; Salunkhe, V.; Murphy, R.P. Canonical Wnt signaling in megakaryocytes regulates proplatelet formation. Blood 2013, 121, 188–196. [Google Scholar] [CrossRef]
- Pina, C.; Teles, J.; Fugazza, C.; May, G.; Wang, D.; Guo, Y.; Soneji, S.; Brown, J.; Edén, P.; Ohlsson, M. Single-cell network analysis identifies DDIT3 as a nodal lineage regulator in hematopoiesis. Cell Rep. 2015, 11, 1503–1510. [Google Scholar] [CrossRef]
- Wang, Y.-L.; Qian, J.; Lin, J.; Yao, D.-M.; Qian, Z.; Zhu, Z.-H.; Li, J.-Y. Methylation status of DDIT3 gene in chronic myeloid leukemia. J. Exp. Clin. Cancer Res. 2010, 29, 54. [Google Scholar] [CrossRef] [PubMed]
- Akashi, K.; Traver, D.; Miyamoto, T.; Weissman, I.L. A clonogenic common myeloid progenitor that gives rise to all myeloid lineages. Nature 2000, 404, 193–197. [Google Scholar] [CrossRef] [PubMed]
- Sun, X.-J.; Man, N.; Tan, Y.; Nimer, S.D.; Wang, L. The role of histone acetyltransferases in normal and malignant hematopoiesis. Front. Oncol. 2015, 5, 108. [Google Scholar] [CrossRef] [PubMed]
- Zimmer, S.N.; Zhou, Q.; Zhou, T.; Cheng, Z.; Abboud-Werner, S.L.; Horn, D.; Lecocke, M.; White, R.; Krivtsov, A.V.; Armstrong, S.A. Crebbp haploinsufficiency in mice alters the bone marrow microenvironment, leading to loss of stem cells and excessive myelopoiesis. Blood 2011, 118, 69–79. [Google Scholar] [CrossRef]
- Geddis, A.E. The regulation of proplatelet production. Haematologica 2009, 94, 756–759. [Google Scholar] [CrossRef][Green Version]
- Pleines, I.; Dütting, S.; Cherpokova, D.; Eckly, A.; Meyer, I.; Morowski, M.; Krohne, G.; Schulze, H.; Gachet, C.; Debili, N. Defective tubulin organization and proplatelet formation in murine megakaryocytes lacking Rac1 and Cdc42. Blood 2013, 122, 3178–3187. [Google Scholar] [CrossRef]
- Cabrera, M.; Ungermann, C. Guanine nucleotide exchange factors (GEFs) have a critical but not exclusive role in organelle localization of Rab GTPases. J. Biol. Chem. 2013, 288, 28704–28712. [Google Scholar] [CrossRef]
- Kobielak, A.; Fuchs, E. Alpha-catenin: At the junction of intercellular adhesion and actin dynamics. Nat. Rev. Mol. Cell Biol. 2004, 5, 614–625. [Google Scholar] [CrossRef]
- Acar, M.; Kocherlakota, K.S.; Murphy, M.M.; Peyer, J.G.; Oguro, H.; Inra, C.N.; Christabel, J.; Zhao, Z.; Luby-Phelps, K.; Morrison, S.J. Deep imaging of bone marrow shows non-dividing stem cells are mainly perisinusoidal. Nature 2015, 526, 126–130. [Google Scholar] [CrossRef]

| Blood Test Items | Units | Normal Saline | PHMG-p | z-Scores | p-Value | Reference Interval | Ref. | ||
|---|---|---|---|---|---|---|---|---|---|
| Mean | SD | Mean | SD | ||||||
| WBC | 103/µL | 6.50 | 1.46 | 6.70 | 1.91 | −0.581 | 0.561 | 6.11 ± 3.11 | [21] |
| Band Neutrophil | % | 32.60 | 18.56 | 41.24 | 21.15 | −1.141 | 0.254 | 32.48 ± 13.41 | [22] |
| Lymphocyte | % | 59.57 | 17.50 | 52.05 | 20.98 | −0.892 | 0.373 | 63.24 ± 13.06 | [22] |
| Monocyte | % | 3.50 | 0.73 | 2.40 | 0.88 | −2.970 | 0.003 * | 3.49 ± 1.46 | [23] |
| Eosinophil | % | 2.57 | 1.75 | 2.45 | 1.81 | −0.208 | 0.835 | 1.42 ± 1.94 | [22] |
| Basophil | % | 1.23 | 0.89 | 2.03 | 1.56 | −1.746 | 0.081 | 1.16 ± 1.32 | [23] |
| RBC | 106/µL | 8.94 | 0.44 | 9.83 | 1.43 | −1.556 | 0.120 | 8.79 ± 0.39 | [23] |
| Hemoglobin (Hb) | g/dL | 15.86 | 0.69 | 17.61 | 2.61 | −1.973 | 0.049 * | 15.69 ± 0.65 | [23] |
| Hematocrit (Hct) | % | 57.50 | 5.02 | 64.82 | 9.56 | −2.261 | 0.024 * | 46.19 ± 3.53 | [23] |
| Platelet | 103/µL | 928.67 | 142.71 | 703.07 | 179.99 | −3.132 | 0.001 * | 971.45 ± 126.77 | [23] |
| Blood Test Items | Units | Normal Saline | PHMG-p | z-Scores | p-Value | Reference Interval | Ref. | ||
|---|---|---|---|---|---|---|---|---|---|
| Mean | SD | Mean | SD | ||||||
| Total protein | g/dL | 6.25 | 0.51 | 5.79 | 0.31 | −2.503 | 0.012 * | 6.50 ± 0.52 | [23] |
| Albumin | g/dL | 3.87 | 0.38 | 3.74 | 0.16 | −0.773 | 0.439 | 3.15 ± 0.21 | [23] |
| BUN | mg/dL | 22.07 | 3.73 | 26.40 | 7.60 | −1.585 | 0.113 | 16.97 ± 2.78 | [23] |
| Creatinine | mg/dL | 0.59 | 0.14 | 0.67 | 0.13 | −1.495 | 0.135 | 0.55 ± 0.07 | [23] |
| CRP | mg/dL | <0.06 | - | <0.06 | - | - | - | - | - |
| Information | Units | Normal Saline | PHMG-p | z-Scores | p-Value | ||
|---|---|---|---|---|---|---|---|
| Mean | SD | Mean | SD | ||||
| Platelet count * | 103/µL | 1068.6 | 69.16 | 503.4 | 112.4 | −2.611 | 0.009 * |
| Megakaryocyte count * | 10 HPF | 204.4 | 33.49 | 133.8 | 40.33 | −2.611 | 0.009 * |
| Spleen weight | g | 0.962 | 0.344 | 0.710 | 0.152 | −1.149 | 0.251 |
| Liver weight | g | 17.98 | 3.199 | 15.36 | 4.717 | −0.940 | 0.347 |
| Body weight | g | 507.0 | 10.42 | 496.4 | 63.40 | −0.419 | 0.675 |
| Spleen/Body weight | - | 0.0019 | 0.0007 | 0.0014 | 0.0002 | −1.149 | 0.251 |
| Liver/Body weight | - | 0.0354 | 0.0058 | 0.0305 | 0.0059 | −1.149 | 0.251 |
| Symbol | Entrez Gene Name | Transcript ID | Fold Change | p-Value |
|---|---|---|---|---|
| Txk | TXK tyrosine kinase, transcript variant X1 | NM_001024255 | 13.712 | 0.049 |
| Ctnnal1 | catenin alpha-like 1, transcript variant X1 | NM_001106649 | 12.744 | 0.045 |
| Ctsw | cathepsin W | NM_001024242 | 12.621 | 0.010 |
| Hfe | hemochromatosis, transcript variant 1 | NM_053301 | 9.440 | 0.044 |
| Arhgap4 | Rho GTPase-activating protein 4, transcript variant X1 | NM_144740 | 7.775 | 0.038 |
| Lactb | lactamase, beta, transcript variant X1 | NM_001106833 | 7.416 | 0.021 |
| Clcc1 | chloride channel CLIC-like 1, transcript variant X1 | NM_133414 | 5.885 | 0.036 |
| Tyk2 | tyrosine kinase 2, transcript variant X2 | NM_001257347 | 5.764 | 0.018 |
| Mrps23 | mitochondrial ribosomal protein S23, transcript variant X1 | NM_001108289 | 5.290 | 0.046 |
| Slc11a1 | solute carrier family 11 member 1, transcript variant X1 | NM_001031658 | 4.966 | 0.027 |
| Ap5z1 | adaptor-related protein complex 5, zeta 1 subunit | NM_001037220 | 4.807 | 0.040 |
| Gstz1 | glutathione S-transferase zeta 1 | NM_001109445 | 4.739 | 0.031 |
| Rheb | Ras homolog enriched in brain | NM_013216 | 4.671 | 0.026 |
| Lypd3 | Ly6/Plaur domain-containing 3 | NM_021759 | 4.667 | 0.046 |
| Relt | RELT tumor necrosis factor receptor | NM_001108495 | 4.529 | 0.016 |
| Lsm4 | LSM4 homolog, U6 small nuclear RNA and mRNA degradation associated | NM_001106073 | 4.512 | 0.025 |
| Ttpal | alpha tocopherol transfer protein like, transcript variant X1 | NM_001106537 | 4.420 | 0.035 |
| Ly6e | lymphocyte antigen 6 complex, locus E, transcript variant X4 | NM_001017467 | 4.412 | 0.037 |
| Taf1 | TATA-box binding protein associated factor 1 | NM_001191723 | 4.391 | 0.010 |
| Nrros | negative regulator of reactive oxygen species, transcript variant X1 | NM_001024995 | 4.388 | 0.008 |
| Pank3 | pantothenate kinase 3 | NM_001108272 | 4.309 | 0.040 |
| RGD1565784 | RGD1565784 | NM_001109028 | 4.267 | 0.013 |
| Fchsd2 | FCH and double SH3 domains 2, transcript variant X4 | NM_001107539 | 4.158 | 0.017 |
| Ogfr | opioid growth factor receptor | NM_053340 | 4.141 | 0.047 |
| Eif3k | eukaryotic translation initiation factor 3, subunit K | NM_001106242 | 3.910 | 0.012 |
| Mphosph6 | M phase phosphoprotein 6 | NM_001129881 | 3.820 | 0.046 |
| Pnkp | polynucleotide kinase 3′-phosphatase, transcript variant X1 | NM_001004259 | 3.714 | 0.016 |
| Lgals5 | lectin, galactose binding, soluble 5 | NM_012976 | 3.688 | 0.035 |
| Snx4 | sorting nexin 4 | NM_001127550 | 3.682 | 0.040 |
| Fmo1 | flavin-containing monooxygenase 1, transcript variant X2 | NM_012792 | 3.622 | 0.035 |
| Sik3 | SIK family kinase 3 | NM_001271216 | 3.603 | 0.010 |
| Phkg2 | phosphorylase kinase catalytic subunit gamma 2, transcript variant X2 | NM_080584 | 3.537 | 0.047 |
| Pip5k1b | phosphatidylinositol-4-phosphate 5-kinase type 1 beta | NM_001012743 | 3.505 | 0.041 |
| Pld3 | phospholipase D family, member 3, transcript variant X3 | NM_001012167 | 3.429 | 0.005 |
| Limk2 | LIM domain kinase 2, transcript variant X1 | NM_024135 | 3.397 | 0.017 |
| Cirbp | cold-inducible RNA-binding protein, transcript variant X4 | NM_031147 | 3.216 | 0.026 |
| Pip5k1c | phosphatidylinositol-4-phosphate 5-kinase type 1 gamma, transcript variant X1 | NM_001033970 | 3.186 | 0.004 |
| LOC501110 | similar to Glutathione S-transferase A1 (GTH1) (HA subunit 1) (GST-epsilon) (GSTA1-1) (GST class-alpha) | NM_001024361 | 3.137 | 0.023 |
| Plcl2 | phospholipase C-like 2 | NM_001106880 | 3.121 | 0.041 |
| Arhgap11a | Rho GTPase-activating protein 11A | NM_001168524 | 2.892 | 0.036 |
| Polr2g | polymerase (RNA) II subunit G, transcript variant X1 | NM_053948 | 2.867 | 0.042 |
| Ptpn12 | protein tyrosine phosphatase, non-receptor type 12 | NM_057115 | 2.860 | 0.041 |
| Emc2 | ER membrane protein complex subunit 2 | NM_001113785 | 2.845 | 0.014 |
| Lrrc14 | leucine-rich repeat containing 14, transcript variant X1 | NM_001024354 | 2.834 | 0.001 |
| LOC102549726 | uncharacterized LOC102549726 | NR_110713 | 2.803 | 0.017 |
| Psmd9 | proteasome 26S subunit, non-ATPase 9 | NM_130430 | 2.797 | 0.010 |
| Lsm2 | LSM2 homolog, U6 small nuclear RNA and mRNA degradation associated, transcript variant 2 | NM_001165922 | 2.742 | 0.016 |
| Rcsd1 | RCSD domain containing 1, transcript variant X1 | NM_001108349 | 2.740 | 0.004 |
| Cnot1 | CCR4-NOT transcription complex, subunit 1 | NM_001134840 | 2.720 | 0.014 |
| Shisa5 | shisa family member 5, transcript variant X1 | NM_001006989 | 2.701 | 0.017 |
| Eif6 | eukaryotic translation initiation factor 6, transcript variant X2 | NM_001037352 | 2.546 | 0.023 |
| Junb | JunB proto-oncogene, AP-1 transcription factor subunit | NM_021836 | 2.518 | 0.045 |
| Ube2i | ubiquitin-conjugating enzyme E2I, transcript variant X2 | NM_013050 | 2.498 | 0.014 |
| Kat6a | lysine acetyltransferase 6A | NM_001100570 | 2.430 | 0.028 |
| Adck4 | aarF domain-containing kinase 4, transcript variant X1 | NM_001012065 | 2.429 | 0.006 |
| C1qc | complement C1q C chain, transcript variant X1 | NM_001008524 | 2.414 | 0.030 |
| Zdhhc18 | zinc finger, DHHC-type containing 18 | NM_001039339 | 2.352 | 0.048 |
| Snx5 | sorting nexin 5 | NM_001106518 | 2.316 | 0.004 |
| Psmc3 | proteasome 26S subunit, ATPase 3 | NM_031595 | 2.252 | 0.002 |
| Mmp9 | matrix metallopeptidase 9 | NM_031055 | 2.205 | 0.020 |
| Abhd4 | abhydrolase domain-containing 4, transcript variant X2 | NM_001108866 | 2.162 | 0.008 |
| Etfb | electron transfer flavoprotein beta subunit | NM_001004220 | 2.153 | 0.028 |
| Cd74 | CD74 molecule, transcript variant X1 | NM_013069 | 2.132 | 0.005 |
| Tor1aip2 | torsin 1A interacting protein 2, transcript variant X3 | NM_001165896 | 2.086 | 0.039 |
| Ehd1 | EH-domain containing 1 | NM_001011939 | 2.082 | 0.048 |
| Yipf3 | Yip1 domain family, member 3, transcript variant X1 | NM_001007801 | 2.046 | 0.046 |
| Mxi1 | MAX interactor 1, dimerization protein, transcript variant X1 | NM_013160 | 2.028 | 0.037 |
| Hpgds | hematopoietic prostaglandin D synthase, transcript variant X2 | NM_031644 | −20.212 | 0.036 |
| Dr1 | down-regulator of transcription 1, TBP-binding (negative cofactor 2) | NM_001011914 | −19.743 | 0.014 |
| Cabin1 | calcineurin binding protein 1 | NM_053575 | −12.615 | 0.010 |
| Ccnl1 | cyclin L1, transcript variant X1 | NM_053662 | −11.615 | 0.009 |
| Dtnb | dystrobrevin, beta, transcript variant X3 | NM_001012191 | −10.521 | 0.044 |
| Tti1 | TELO2 interacting protein 1, transcript variant X1 | NM_001134619 | −9.680 | 0.025 |
| Zfp64 | zinc finger protein 64 | NM_001012093 | −9.322 | 0.015 |
| Atp5a1 | ATP synthase, H+ transporting, mitochondrial F1 complex, alpha subunit 1, cardiac muscle | NM_023093 | −9.259 | 0.047 |
| Irf7 | interferon regulatory factor 7, transcript variant X2 | NM_001033691 | −9.214 | 0.047 |
| Ergic2 | ERGIC and Golgi 2 | NM_001024984 | −8.686 | 0.014 |
| Med13 | mediator complex subunit 13 | NM_001107035 | −8.430 | 0.004 |
| Lap3 | leucine aminopeptidase 3 | NM_001011910 | −8.196 | 0.049 |
| Arhgef3 | Rho guanine nucleotide exchange factor 3, transcript variant X7 | NM_001106061 | −7.714 | 0.021 |
| Pacsin2 | protein kinase C and casein kinase substrate in neurons 2, transcript variant X12 | NM_130740 | −7.330 | 0.034 |
| Rpp21 | ribonuclease P/MRP 21 subunit | NM_001002831 | −6.889 | 0.015 |
| Tceb3 | transcription elongation factor B subunit 3 | NM_017103 | −6.772 | 0.028 |
| Ipo7 | importin 7 | NM_001107545 | −6.583 | 0.041 |
| Fam203a | family with sequence similarity 203, member A | NM_001007707 | −6.267 | 0.032 |
| Ascc3 | activating signal cointegrator 1 complex subunit 3, transcript variant X3 | NM_001277057 | −6.076 | 0.001 |
| Ipo13 | importin 13 | NM_053778 | −5.873 | 0.003 |
| Hsd17b1 | hydroxysteroid (17-beta) dehydrogenase 1 | NM_012851 | −5.758 | 0.024 |
| Ppib | peptidylprolyl isomerase B | NM_022536 | −5.672 | 0.031 |
| RGD735065 | similar to GI:13385412-like protein splice form I | NM_199379 | −5.650 | 0.023 |
| Snx27 | sorting nexin family member 27, transcript variant 1 | NM_001110151 | −5.484 | 0.020 |
| Vat1 | vesicle amine transport 1 | NM_001033683 | −5.279 | 0.031 |
| Atp6v1e1 | ATPase H+ transporting V1 subunit E1 | NM_198745 | −5.101 | 0.039 |
| Rcor1 | REST corepressor 1, transcript variant X1 | NM_001305463 | −5.030 | 0.009 |
| Slc35c1 | solute carrier family 35 member C1, transcript variant X1 | NM_001107748 | −4.994 | 0.031 |
| Fbxo33 | F-box protein 33, transcript variant X1 | NM_001108023 | −4.989 | 0.040 |
| Tsc22d1 | TSC22 domain family, member 1, transcript variant X1 | NM_001109912 | −4.892 | 0.023 |
| Polb | polymerase (DNA) beta | NM_017141 | −4.427 | 0.011 |
| Tmem109 | transmembrane protein 109, transcript variant X2 | NM_001007736 | −4.008 | 0.016 |
| Klf16 | Kruppel-like factor 16 | NM_001127604 | −3.909 | 0.042 |
| Rpia | ribose 5-phosphate isomerase A | NM_001108632 | −3.843 | 0.020 |
| Tbcc | tubulin folding cofactor C | NM_001108200 | −3.836 | 0.023 |
| Morf4l2 | mortality factor 4 like 2 | NM_001007714 | −3.746 | 0.033 |
| Plagl2 | PLAG1 like zinc finger 2, transcript variant X1 | NM_001106528 | −3.733 | 0.020 |
| Pde4b | phosphodiesterase 4B | NM_017031 | −3.718 | 0.019 |
| Atg10 | autophagy related 10 | NM_001109505 | −3.473 | 0.029 |
| Nos1ap | nitric oxide synthase 1 adaptor protein | NM_138922 | −3.455 | 0.005 |
| Ncor1 | nuclear receptor co-repressor 1 | NM_001271103 | −3.368 | 0.007 |
| Slmo2 | slowmo homolog 2 | NM_001009543 | −3.319 | 0.020 |
| Ino80 | INO80 complex subunit | NM_001261404 | −3.199 | 0.002 |
| Phyh | phytanoyl-CoA 2-hydroxylase | NM_053674 | −3.183 | 0.046 |
| Edc4 | enhancer of mRNA decapping 4 | NM_001033068 | −3.141 | 0.034 |
| Ccar1 | cell division cycle and apoptosis regulator 1 | NM_001108535 | −3.052 | 0.008 |
| Larp7 | La ribonucleoprotein domain family, member 7 | NM_001044290 | −3.040 | 0.033 |
| Cks1b | CDC28 protein kinase regulatory subunit 1B | NM_001135749 | −3.020 | 0.026 |
| Gps2 | G protein pathway suppressor 2, transcript variant X1 | NM_001017477 | −3.014 | 0.008 |
| Baz2a | bromodomain adjacent to zinc finger domain, 2A | NM_001107158 | −3.006 | 0.027 |
| Mrpl14 | mitochondrial ribosomal protein L14, transcript variant 2 | NM_001106890 | −3.001 | 0.008 |
| Kif2a | kinesin family member 2A | NM_053376 | −2.953 | 0.011 |
| Snx8 | sorting nexin 8, transcript variant X1 | NM_001105912 | −2.936 | 0.030 |
| Eif3f | eukaryotic translation initiation factor 3, subunit F | NM_001277302 | −2.900 | 0.010 |
| Ep400 | E1A binding protein p400 | NM_001107149 | −2.898 | 0.008 |
| Baz1a | bromodomain adjacent to zinc finger domain, 1A | NM_001170568 | −2.887 | 0.002 |
| Zrsr1 | zinc finger (CCCH type), RNA-binding motif and serine/arginine-rich 1 | NM_001017504 | −2.883 | 0.046 |
| Thg1l | tRNA-histidine guanylyltransferase 1-like, transcript variant X1 | NM_001013966 | −2.817 | 0.038 |
| LOC294154 | similar to chromosome 6 open reading frame 106 isoform a, transcript variant X1 | NM_001039607 | −2.644 | 0.010 |
| Cdca8 | cell division cycle associated 8, transcript variant X1 | NM_001025050 | −2.599 | 0.012 |
| St3gal2 | ST3 beta-galactoside alpha-2,3-sialyltransferase 2, transcript variant X3 | NM_031695 | −2.539 | 0.010 |
| Psmd7 | proteasome 26S subunit, non-ATPase 7 | NM_001107426 | −2.518 | 0.011 |
| Ebf1 | early B-cell factor 1, transcript variant X5 | NM_053820 | −2.513 | 0.007 |
| Ublcp1 | ubiquitin-like domain-containing CTD phosphatase 1, transcript variant X1 | NM_001014117 | −2.437 | 0.029 |
| Zfp703 | zinc finger protein 703 | NM_001109425 | −2.320 | 0.037 |
| Luc7l3 | LUC7-like 3 pre-mRNA splicing factor | NM_001108291 | −2.217 | 0.015 |
| Kmt2e | lysine methyltransferase 2E | NM_001100851 | −2.201 | 0.009 |
| Dhx15 | DEAH-box helicase 15, transcript variant X1 | NM_001191597 | −2.189 | 0.029 |
| Crebbp | CREB binding protein, transcript variant X2 | NM_133381 | −2.177 | 0.020 |
| Metap2 | methionyl aminopeptidase 2 | NM_022539 | −2.142 | 0.016 |
| Srrm1 | serine and arginine repetitive matrix 1, transcript variant X8 | NM_001107986 | −2.103 | 0.043 |
| Dmtf1 | cyclin D binding myb-like transcription factor 1, transcript variant X1 | NM_053693 | −2.090 | 0.029 |
| RatNP-3b | defensin RatNP-3 precursor | NM_001079898 | −2.074 | 0.032 |
| Brd4 | bromodomain containing 4 | NM_001100903 | −2.052 | 0.013 |
| Tax1bp1 | Tax1 binding protein 1, transcript variant X1 | NM_001004199 | −2.014 | 0.047 |
| Category | p-Value Range | No. of Molecules |
|---|---|---|
| Diseases and disorders | ||
| Cancer | 1.66 × 10−2–4.98 × 10−8 | 127 |
| Organismal injury and abnormalities | 1.67 × 10−2–4.98 × 10−8 | 129 |
| Gastrointestinal disease | 1.67 × 10−2–6.60 × 10−6 | 113 |
| Hematological disease | 1.65 × 10−2–1.30 × 10−4 | 50 |
| Endocrine system disorders | 1.38 × 10−2–2.44 × 10−4 | 101 |
| Physiological system development and function | ||
| Organismal survival | 1.46 × 10−2–9.36 × 10−7 | 52 |
| Connective tissue development and function | 1.67 × 10−2–3.82 × 10−5 | 30 |
| Skeletal and muscular system development and function | 1.24 × 10−2–3.82 × 10−5 | 13 |
| Tissue development | 1.67 × 10−2–3.82 × 10−5 | 36 |
| Hematological system development and function | 1.67 × 10−2–4.49 × 10−5 | 39 |
| Molecular and cellular functions | ||
| Cell cycle | 1.61 × 10−2–4.31 × 10−7 | 43 |
| Gene expression | 1.55 × 10−2–3.56 × 10−6 | 45 |
| Post-translational modification | 5.53 × 10−3–7.34 × 10−5 | 4 |
| DNA replication, recombination, and repair | 1.52 × 10−2–9.30 × 10−5 | 15 |
| Cellular development | 1.64 × 10−2–1.28 × 10−5 | 51 |
| Disease or Bio Function Annotation | p-Value | Molecules | # Mol. |
|---|---|---|---|
| Hematological disease | |||
| Abnormal morphology of bone marrow | 1.30 × 10−4 | CREBBP, EBF1, EP400, KAT6A, TYK2 | 5 |
| Bone marrow neoplasm | 7.41 × 10−4 | ATP5F1A, ATP6V1E1, CCNL1, CD74, CKS1B, CREBBP, CTSW, DHX15, DTNB, EDC4, EP400, IPO7, JUNB, KAT6A, KMT2E, LY6E, LYPD3, MMP9, NCOR1, PDE4B, PLAGL2, PLCL2, POLB, PSMD9, TAF1, TYK2, UBE2I, ZFP64 | 28 |
| Central nervous system leukemia | 1.09 × 10−3 | MMP9, POLB | 2 |
| T acute lymphoblastic leukemia | 1.35 × 10−3 | CREBBP, EBF1, NCOR1, POLB, TAX1BP1, TYK2 | 6 |
| Incidence of lymphoma | 1.74 × 10−3 | CD74, DMTF1, INO80, JUNB, MXI1, TYK2 | 6 |
| Hematological system development and function | |||
| Morphology of lymphoid tissue | 4.49 × 10−5 | BRD4, CD74, CKS1B, CREBBP, GSTZ1, HPGDS, IRF7, KAT6A, KMT2E, MXI1, NCOR1, SLC35C1, SNX27, ST3GAL2, TYK2 | 15 |
| Leukopoiesis | 1.28 × 10−4 | C1QC, CD74, CREBBP, DMTF1, EBF1, EIF6, GPS2, IRF7, JUNB, KMT2E, NCOR1, NRROS, PIP5K1C, PLCL2, POLB, PTPN12, RHEB, TXK, TYK2 | 19 |
| Development of hematopoietic cells | 4.23 × 10−4 | CD74, CREBBP, EBF1, EIF6, KMT2E, MMP9, NCOR1, RCOR1, TYK2 | 9 |
| Conjugation of T lymphocytes | 6.41 × 10−4 | PIP5K1C, TXK | 2 |
| Quantity of erythroid cells | 1.36 × 10−3 | CREBBP, GPS2 | 2 |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Sung, H.J.; Jeong, S.H.; Kang, J.Y.; Kim, C.; Nam, Y.J.; Kim, J.Y.; Choi, J.Y.; Lee, H.J.; Lee, Y.S.; Kim, E.Y.; et al. Hematotoxic Effect of Respiratory Exposure to PHMG-p and Its Integrated Genetic Analysis. Toxics 2022, 10, 694. https://doi.org/10.3390/toxics10110694
Sung HJ, Jeong SH, Kang JY, Kim C, Nam YJ, Kim JY, Choi JY, Lee HJ, Lee YS, Kim EY, et al. Hematotoxic Effect of Respiratory Exposure to PHMG-p and Its Integrated Genetic Analysis. Toxics. 2022; 10(11):694. https://doi.org/10.3390/toxics10110694
Chicago/Turabian StyleSung, Hwa Jung, Sang Hoon Jeong, Ja Young Kang, Cherry Kim, Yoon Jeong Nam, Jae Young Kim, Jin Young Choi, Hye Jin Lee, Yu Seon Lee, Eun Yeob Kim, and et al. 2022. "Hematotoxic Effect of Respiratory Exposure to PHMG-p and Its Integrated Genetic Analysis" Toxics 10, no. 11: 694. https://doi.org/10.3390/toxics10110694
APA StyleSung, H. J., Jeong, S. H., Kang, J. Y., Kim, C., Nam, Y. J., Kim, J. Y., Choi, J. Y., Lee, H. J., Lee, Y. S., Kim, E. Y., Baek, Y. W., Lee, H., & Lee, J. H. (2022). Hematotoxic Effect of Respiratory Exposure to PHMG-p and Its Integrated Genetic Analysis. Toxics, 10(11), 694. https://doi.org/10.3390/toxics10110694






