Determining the Authenticity of Shark Meat Products by DNA Sequencing
Abstract
:1. Introduction
2. Materials and Methods
2.1. Samples Collection
2.2. Fish Labeling Analysis
2.3. DNA Extraction and Purification
2.4. Oligonucleotide Primers and Reference Genes
2.5. COI and NADH2 PCR Assay
2.6. Detection of Amplified Products
2.7. PCR Cleanup
2.8. Cycle Sequencing Reaction
2.9. Analysis of Sequences
2.10. Assessment Conservation Status
3. Results
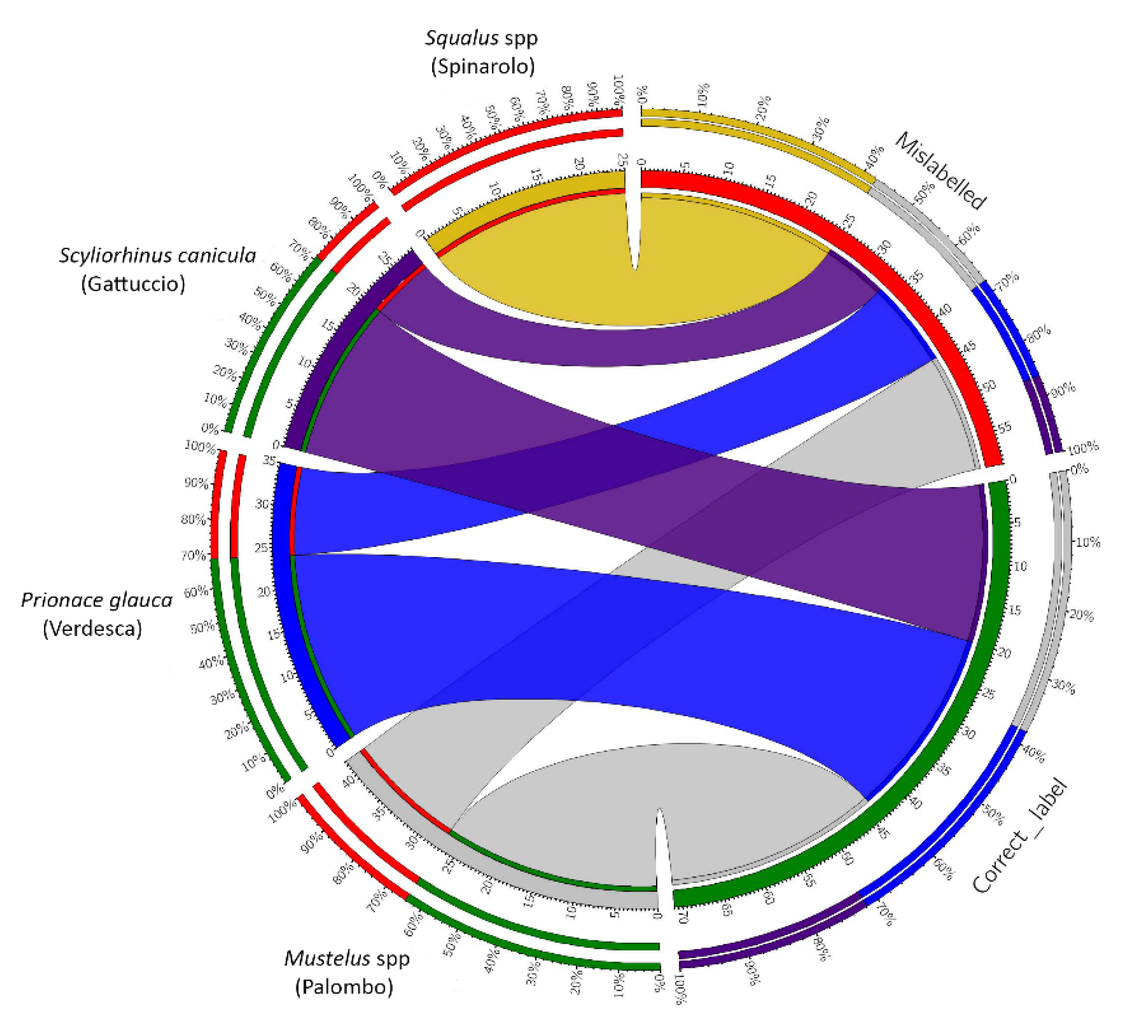
3.1. Analysis of the Fish Labels
3.2. Data Analysis
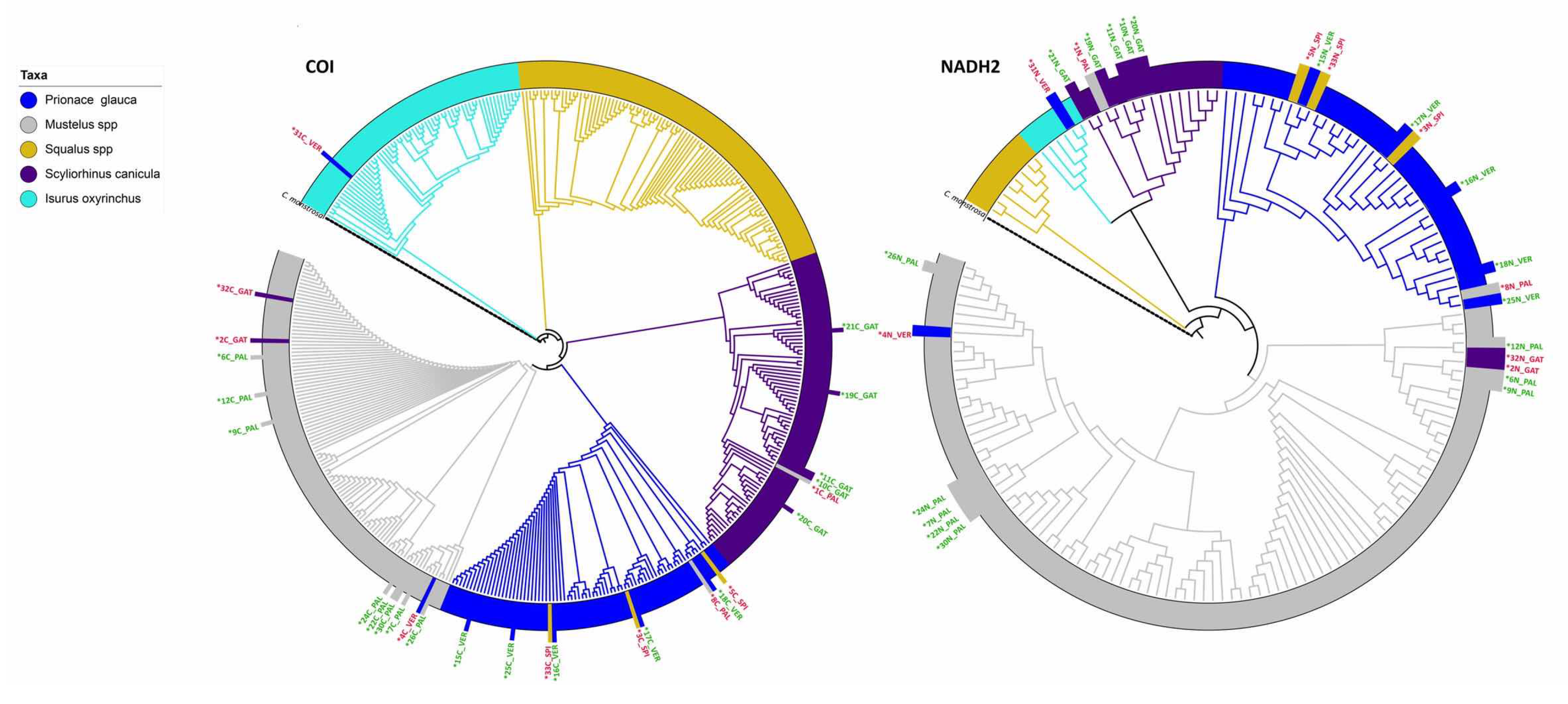
3.3. DNA-Based Species Identification
3.4. Conservation Status of Identified Species
4. Discussion
Supplementary Materials
Author Contributions
Funding
Conflicts of Interest
References
- Hellberg, R.S.; Isaacs, R.B.; Hernandez, E.L. Identification of shark species in commercial products using DNA barcoding. Fish. Res. 2019, 210, 81–88. [Google Scholar] [CrossRef]
- Bradai, M.N.; Saidi, B.; Enajjar, S. Elasmobranchs of the Mediterranean and Black Sea: Status, Ecology and Biology Bibliographic Analysis; FAO: Rome, Italy, 2012; p. 103. [Google Scholar]
- Clarke, S. Shark Product Trade in Hong Kong and Mainland China and Implementation of the CITES Shark Listings; Traffic East Asia: Hong Kong, China, 2004; pp. 1–63. [Google Scholar]
- Dent, F.; Clarke, S. State of the Global Market for Shark Products; Technical Paper No. 590; FAO: Rome, Italy, 2015. [Google Scholar]
- Hobbs, C.A.D.; Potts, R.W.A.; Walsh, M.B.; Usher, J.; Griffiths, A.M. Using DNA Barcoding to investigate patterns of species utilisation in UK shark products reveals threatened species on sale. Sci. Rep. 2019, 9, 1028. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Giovos, I.; Arculeo, M.; Doumpas, N.; Katsada, D.; Maximiadi, M.; Mitsou, E.; Paravas, V.; Aga-Spyridopoulou, R.N.; Stoilas, V.O.; Tiralongo, F. Assessing multiple sources of data to detect illegal fishing, trade and mislabeling of elasmobranchs in Greek markets. Mar. Policy 2020, 112, 103730. [Google Scholar] [CrossRef]
- Pazartzi, T.; Siaperopoulou, S.; Gubili, C.; Maradidou, S.; Loukovitis, D.; Chatzispyro, A.; Griffiths, A.M.; Minos, G.; Imsiridou, A. High levels of mislabeling in shark meat—Investigating patterns of species utilization with DNA barcoding in Greek retailers. Food Control. 2019, 98, 179–186. [Google Scholar] [CrossRef]
- Marko, P.B.; Lee, S.C.; Rice, A.M.; Gramling, J.M.; Fitzhenry, T.M.; McAlister, J.S.; Harper, G.R.; Moran, A.L. Mislabelling of a depleted reef fish. Nature 2004, 430, 309–310. [Google Scholar] [CrossRef]
- Di Pinto, A.; Marchetti, P.; Mottola, A.; Bozzo, G.; Bonerba, E.; Ceci, E.; Bottaro, M.; Tantillo, G. Species identification in fish fillet products using DNA barcoding. Fish. Res. 2015, 170, 9–13. [Google Scholar] [CrossRef]
- Filonzi, L.; Chiesa, S.; Vaghi, M.; Marzano, F.N. Molecular barcoding reveals mislabelling of commercial fish products in Italy. Food Res. Int. 2010, 43, 1383–1388. [Google Scholar] [CrossRef]
- Miller, D.; Jessel, A.; Mariani, S. Seafood mislabeling: Comparisons of two western European case studies assist in defining influencing factors, mechanisms and motives. Fish. Fish. 2012, 13, 345–358. [Google Scholar] [CrossRef]
- IUCN. The IUCN Red List of Threatened Species. Version 2020.1. Available online: www.iucnredlist.org (accessed on 3 June 2020).
- Carr, L.A.; Stier, A.C.; Fietz, K.; Montero, I.; Gallagher, A.J.; Bruno, J.F. Illegal shark fishing in the Galápagos Marine Reserve. Mar. Policy 2013, 39, 317–321. [Google Scholar] [CrossRef]
- European Commission. The EU Food Fraud Network and the System for Administrative Assistance—Food Fraud; Annual Report 2018; Publications office of the EU: Luxembourg, 2018; pp. 1–16. [Google Scholar]
- WWF. Sharks in Crisis: A Call to Action for the Mediterranean; Report 2019; WWF: Gland, Switzerland, 2019; pp. 1–44. [Google Scholar]
- Wilkens, H.; Strecker, U. Convergent evolution of the cavefish Astyanax (Characidae, Teleostei): Genetic evidence from reduced eye-size and pigmentation. Biol. J. Linn. Soc. 2003, 80, 545–554. [Google Scholar] [CrossRef] [Green Version]
- Li, X.; Shen, X.; Chen, X.; Xiang, D.; Murphy, R.W.; Shen, Y. Detection of potential problematic Cytb gene sequences of fishes in GenBank. Front. Genet. 2018, 9, 30. [Google Scholar] [CrossRef] [PubMed]
- Hebert, P.D.N.; Ratnasingham, S.; de Waard, J.R. Barcoding animal life: Cytochrome c oxidase subunit 1 divergences among closely related species. Proc. R. Soc. B Biol. Sci. 2003, 270, S96–S99. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Ratnasingham, S.; Hebert, P.D. bold: The barcode of life data system (http://www.barcodinglife.org). Mol. Ecol. Notes 2007, 7, 355–364. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- FishBase. Available online: www.fishbase.org (accessed on 20 March 2020).
- Ward, R.D. FISH-BOL, a case study for DNA barcodes. Methods Mol. Biol. 2012, 858, 423–439. [Google Scholar] [CrossRef]
- Becker, S.; Hanner, R.; Steinke, D. Five years of FISH-BOL: Brief status report. Mitochondr. DNA 2011, 22, 3–9. [Google Scholar] [CrossRef] [Green Version]
- Steinke, D.; Hanner, R. FISH-BOL. The FISH-BOL collaborators’ protocol. Mitochondr. DNA 2011, 22, 10–14. [Google Scholar] [CrossRef] [Green Version]
- Nneji, L.M.; Adeola, A.C.; Mustapha, M.K.; Oladipo, S.O.; Djagoun, C.A.; Nneji, I.C.; Adedeji, B.E.; Olatunde, O.; Ayoola, A.O.; Okeyoyin, A.O.; et al. DNA barcoding silver butter catfish (Schilbe intermedius) reveals patterns of mitochondrial genetic diversity across African river systems. Sci. Rep. 2020, 10, 1–9. [Google Scholar] [CrossRef]
- Fadli, N.; Nor, S.A.M.; Othman, A.S.; Sofyan, H.; Muchlisin, Z.A. DNA barcoding of commercially important reef fishes in Weh Island, Aceh, Indonesia. PeerJ 2020, 8, e9641. [Google Scholar] [CrossRef]
- Arroyave, J.; Martinez, C.M.; Stiassny, M.L. DNA barcoding uncovers extensive cryptic diversity in the African long-fin tetra Bryconalestes longipinnis (Alestidae: Characiformes). J. Fish. Biol. 2019, 95, 379–392. [Google Scholar] [CrossRef]
- Escobar, R.; Luna-Acosta, A.; Caballero, S. DNA barcoding, fisheries and communities: What do we have? Science and local knowledge to improve resource management in partnership with communities in the Colombian Caribbean. Mar. Policy 2020, 99, 407–413. [Google Scholar] [CrossRef]
- Galal-Khallaf, A.; Ardura, A.; Mohammed-Geba, K.; Borrell, Y.J.; Garcia-Vazquez, E. DNA barcoding reveals a high level of mislabeling in Egyptian fish fillets. Food Control 2014, 46, 441–445. [Google Scholar] [CrossRef]
- Hu, Y.; Huang, S.Y.; Hanner, R.; Levin, J.; Lu, X. Study of fish products in Metro Vancouver using DNA barcoding methods reveals fraudulent labeling. Food Control 2018, 94, 38–47. [Google Scholar] [CrossRef]
- Barendse, J.; Roel, A.; Longo, C.; Andriessen, L.; Webster, L.M.; Ogden, R.; Neat, F. DNA barcoding validates species labelling of certified seafood. Curr. Biol. 2019, 29, R198–R199. [Google Scholar] [CrossRef] [Green Version]
- FDA. Available online: https://www.fda.gov/food/dna-based-seafood-identification/single-laboratory-validated-method-dna-barcoding-species-identification-fish (accessed on 20 March 2020).
- Wong, E.H.K.; Shivji, M.S.; Hanner, R.H. Identifying sharks with DNA barcodes: Assessing the utility of a nucleotide diagnostic approach. Mol. Ecol. Res. 2009, 9, 243–256. [Google Scholar] [CrossRef] [PubMed]
- Bucklin, A.; Steinke, D.; Blanco-Bercial, L. DNA barcoding of marine metazoa. Annu. Rev. Mar. Sci. 2011, 3, 471–508. [Google Scholar] [CrossRef] [PubMed]
- Zanzi, A.; Martinsohn, J.T. FishTrace: A genetic catalogue of European fishes. Database 2017, 2017, bax075. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Jeon, H.B.; Anderson, D.; Won, H.; Lim, H.; Suk, H.Y. Taxonomic characterization of Tanakia species (Acheilognathidae) using DNA barcoding analyses. Mitochondr. DNA 2018, 29, 964–973. [Google Scholar] [CrossRef]
- Horreo, J.L.; Fitze, P.S.; Jiménez-Valverde, A.; Noriega, J.A.; Pelaez, M.L. Amplification of 16S rDNA reveals important fish mislabeling in Madrid restaurants. Food Control 2019, 96, 146–150. [Google Scholar] [CrossRef]
- Valentini, A.; Taberlet, P.; Miaud, C.; Civade, R.; Herder, J.; Thomsen, P.F.; Bellemain, E.; Besnard, A.; Coissac, E.; Boyer, F.; et al. Next-generation monitoring of aquatic biodiversity using environmental DNA metabarcoding. Mol. Ecol. 2016, 25, 929–942. [Google Scholar] [CrossRef] [Green Version]
- Collins, R.A.; Bakker, J.; Wangensteen, O.S.; Soto, A.Z.; Corrigan, L.; Sims, D.W.; Genner, M.J.; Mariani, S. Non-specific amplification compromises environmental DNA metabarcoding with COI. Methods Ecol. Evol. 2019, 10, 1985–2001. [Google Scholar] [CrossRef]
- Moore, A.B.M.; White, W.T.; Ward, R.D.; Naylor, G.J.P.; Peirce, R. Rediscovery and redescription of the smoothtooth blacktip shark, Carcharhinus leiodon (Carcharhinidae), from Kuwait, with notes on its possible conservation status. Mar. Freshw. Res. 2011, 62, 528–539. [Google Scholar] [CrossRef]
- Naylor, G.J.P.; Caira, J.N.; Ensen, K.J.; Rosana, K.A.M.; White, W.T.; Last, P.R. A DNA sequence–based approach to the identification of shark and ray species and its implications for global elasmobranch diversity and parasitology. Bull. Am. Mus. Nat. Hist. 2012, 367, 1–262. [Google Scholar] [CrossRef]
- Henderson, A.C.; Reeve, A.J.; Jabado, R.W.; Naylor, G.J.P. Taxonomic assessment of sharks, rays and guitarfishes (Chondrichthyes: Elasmobranchii) from south-eastern Arabia, using the NADH dehydrogenase subunit 2 (NADH2) gene. Zool. J. Linn. Soc. 2016, 176, 399–442. [Google Scholar] [CrossRef] [Green Version]
- Last, P.R.; Seret, B.; Naylor, G.J. A new species of guitarfish, Rhinobatos borneensis sp. nov. with a redefinition of the family-level classification in the order Rhinopristiformes (Chondrichthyes: Batoidea). Zootaxa 2016, 4117, 451–475. [Google Scholar] [CrossRef] [PubMed]
- Vella, A.; Vella, N.; Schembri, S. A molecular approach towards taxonomic identification of elasmobranch species from Maltese fisheries landings. Mar. Genom. 2017, 36, 17–23. [Google Scholar] [CrossRef]
- Feitosa, L.M.; Martins, A.P.B.; Giarrizzo, T.; Macedo, W.; Monteiro, I.L.; Gemaque, R.; Nunes, J.L.S.; Gomes, F.; Schneider, H.; Sampaio, I.; et al. DNA-based identification reveals illegal trade of threatened shark species in a global elasmobranch conservation hotspot. Sci. Rep. 2018, 8, 3347. [Google Scholar] [CrossRef]
- Cutarelli, A.; Amoroso, M.G.; De Roma, A.; Girardi, S.; Galiero, G.; Guarino, A.; Corrado, F. Italian market fish species identification and commercial frauds revealing by DNA sequencing. Food Control. 2014, 37, 46–50. [Google Scholar] [CrossRef]
- Guardone, L.; Tinacci, L.; Costanzo, F.; Azzarelli, D.; D’Amico, P.; Tasselli, G.; Magni, A.; Guidi, A.; Nucera, D.; Armani, A. DNA barcoding as a tool for detecting mislabeling on incoming fishery products from third countries: An official survey conducted at the Border Inspection Post of Livorno-Pisa (Italy). Food Control 2017, 80, 204–215. [Google Scholar] [CrossRef]
- Sarmiento, K.P.; Pereda, J.M.R.; Ventolero, M.F.H.; Santos, M.D. Not fish in fish balls: Fraud in some processed seafood products detected by using DNA barcoding. Phil. Sci. Lett. 2018, 11, 7. [Google Scholar]
- Zeng, L.; Wen, J.; Fan, S.; Chen, Z.; Xu, Y.; Sun, Y.; Chen, D.; Zhao, J. Species identification of fish maw (Porcupinefish) products sold on the market using DNA sequencing of 16S rRNA and COI genes. Food Control. 2018, 86, 159–162. [Google Scholar] [CrossRef]
- Acutis, P.L.; Cambiotti, V.; Riina, M.V.; Meistro, S.; Maurella, C.; Massaro, M.; Stacchini, P.; Gili, S.; Malandra, R.; Pezzolato, M.; et al. Detection of fish species substitution frauds in Italy: A targeted National Monitoring Plan. Food Control 2019, 101, 151–155. [Google Scholar] [CrossRef]
- Deconinck, D.; Volckaert, F.A.; Hostens, K.; Panicz, R.; Eljasik, P.; Faria, M.; Monteirod, C.S.; Robbens, J.; Derycke, S. A high-quality genetic reference database for European commercial fishes reveals substitution fraud of processed Atlantic cod (Gadus morhua) and common sole (Solea solea) at different steps in the Belgian supply chain. Food Chem. Toxicol. 2020, 111417. [Google Scholar] [CrossRef] [PubMed]
- Kolmann, M.A.; Elbassiouny, A.A.; Liverpool, E.A.; Lovejoy, N.R. DNA barcoding reveals the diversity of sharks in Guyana coastal markets. Neotrop. Ichthyol. 2017, 15, e170097. [Google Scholar] [CrossRef] [Green Version]
- Ministerial Decree n°19105 of 22 September 2017. Italian Name for Fish Species of Commercial Interest -Annex 1; Italian Ministry of Agricultural, Food and Forestry Policies (MiPAAF): Rome, Italy, 2017.
- Regulation (EU) No 1379/2013 of the European Parliament and of the Council of 11 December 2013on the common organization of the markets in fishery and aquaculture products, amending Council Regulations (EC) No 1184/2006 and (EC) No 1224/2009 and repealing Council Regulation (EC) No 104/2000. Off. J. EU 2013, L354, 1–21.
- Handy, S.M.; Deeds, J.R.; Ivanova, N.V.; Hebert, P.D.N.; Hanner, R.H.; Ormos, A.; Weigt, L.A.; Moore, M.M.; Yancy, H.F. A single-laboratory validated method for the generation of DNA barcodes for the identification of fish for regulatory compliance. J. AOAC Int. 2011, 94, 1–10. [Google Scholar] [CrossRef] [Green Version]
- Ward, R.D.; Zemlak, T.S.; Innes, B.H.; Last, P.R.; Hebert, P.D.N. DNA barcoding Australia’s fish species. Philos. Trans. R. Soc. Lond. B. 2005, 360, 1847–1857. [Google Scholar] [CrossRef] [PubMed]
- Schloss, P.D.; Westcott, S.L.; Ryabin, T.; Hall, J.R.; Hartmann, M.; Hollister, E.B.; Lesniewski, R.A.; Oakley, B.B.; Parks, D.H.; Robinson, C.J.; et al. Introducing mothur: Open-source, platform-independent, community-supported software for describing and comparing microbial communities. Appl. Environ. Microbiol. 2009, 75, 7537–7541. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- GenBank Nucleotide Database. Available online: https://www.ncbi.nlm.nih.gov (accessed on 23 July 2020).
- Krzywinski, M.; Schein, J.; Birol, I.; Connors, J.; Gascoyne, R.; Horsman, D.; Marra, M.A. Circos: An information aesthetic for comparative genomics. Genome Res. 2009, 19, 1639–1645. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Katoh, K.; Standley, D.M. MAFFT multiple sequence alignment software version 7: Improvements in performance and usability. Mol. Biol. Evol. 2013, 30, 772–780. [Google Scholar] [CrossRef] [Green Version]
- Gouy, M.; Guindon, S.; Gascuel, O. SeaView version 4: A multiplatform graphical user interface for sequence alignment and phylogenetic tree building. Mol. Biol. Evol. 2010, 27, 221–224. [Google Scholar] [CrossRef] [Green Version]
- Minh, B.Q.; Schmidt, H.A.; Chernomor, O.; Schrempf, D.; Woodhams, M.D.; von Haeseler, A.; Lanfear, R. IQ-TREE 2: New models and efficient methods for phylogenetic inference in the genomic era. Mol. Biol. Evol. 2020, 37, 1530–1534. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Kalyaanamoorthy, S.; Minh, B.Q.; Wong, T.; von Haeseler, A.; Jermiin, L. ModelFinder: Fast model selection for accurate Phylogenetic Estimates. Nat. Methods 2017, 14, 587–589. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- iTOL. Available online: www.itol.embl.de/ (accessed on 23 July 2020).
- Letunic, I.; Bork, P. Interactive Tree of Life (iTOL) v4: Recent updates and new developments. Nucleic Acids Res. 2019, 47, W256–W259. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Pavan-Kumar, A.; Gireesh-Babu, P.; Babu, P.S.; Jaiswar, A.K.; Krishna, V.H.; Prasasd, K.P.; Chaudhari, A.; Raje, S.G.; Chakraborty, S.K.; Krishna, G.; et al. Molecular phylogeny of elasmobranchs inferred from mitochondrial and nuclear markers. Mol. Biol. Rep. 2014, 41, 447–457. [Google Scholar] [CrossRef] [PubMed]
- CITES APPENDIX. Available online: https://cites.org/eng/app/index.php (accessed on 24 June 2020).
- Barcelona Convention for the Protection of the Mediterranean. Available online: https://planbleu.org/sites/default/files/upload/files/Barcelona_convention_and_protocols_2005_eng.pdf (accessed on 24 June 2020).
- Bern Convention (Convention on the Conservation of European Wildlife and Natural Habitats). Available online: https://www.coe.int/en/web/bern-convention/appendices (accessed on 24 June 2020).
- Marino, I.A.M.; Riginella, E.; Cariani, A.; Tinti, F.; Farrell, E.D.; Mazzoldi, C.; Zane, L. New molecular tools for the identification of 2 endangered smooth-hound sharks, Mustelus mustelus and Mustelus Punctulatus. J. Hered. 2015, 106, 123–130. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Oceana Protecting the World’s Oceans: Annu. Report. 2018–2019. pp. 1–58. Available online: https://oceana.org/sites/default/files/oceana_annual_report_2018-2019_website.pdf (accessed on 24 June 2020).
- Barbuto, M.; Galimberti, A.; Ferri, E.; Labra, M.; Malandra, R.; Galli, P.; Casiraghi, M. DNA barcoding reveals fraudulent substitutions in shark seafood products: The Italian case of “palombo” (Mustelus spp.). Food Res. Int. 2010, 43, 376–381. [Google Scholar] [CrossRef]
- Armani, A.; Guardone, L.; Castellana, R.L.; Gianfaldoni, D.; Guidi, A.; Castigliego, L. DNA barcoding reveals commercial and health issues in ethnic seafood sold on the Italian market. Food Control. 2015, 55, 206–214. [Google Scholar] [CrossRef] [Green Version]
- Holmes, B.H.; Steinke, D.; Ward, R.D. Identification of shark and ray fins using DNA barcoding. Fish. Res. 2009, 95, 280–288. [Google Scholar] [CrossRef]
- Muttaqin, E.; Abdullah, A.; Nurilmala, M.; Ichsan, M.; Simeone, B.M.; Yulianto, I.; Booth, H. DNA-barcoding as molecular marker for seafood forensics: Species identification of locally consumed shark fish products in the world’s largest shark fishery. Earth Environ. Sci. 2019, 278, 012049. [Google Scholar] [CrossRef]
- Council Regulation (EC) No 1967/2006 of 21 December 2006 concerning management measures for the sustainable exploitation of fishery resources in the Mediterranean Sea, amending Regulation (EEC) No 2847/93 and repealing Regulation (EC) No 1626/94. Off. J. EU 2006, L36/6, 1–25.
- Regulation (EU) 2019/1241 of the European Parliament and of the Council of 20 June 2019 on the conservation of fisheries resources and the protection of marine ecosystems through technical measures, amending Council Regulations (EC) No 1967/2006, (EC) No 1224/2009 and Regulations (EU) No 1380/2013, (EU) 2016/1139, (EU) 2018/973, (EU) 2019/472 and (EU) 2019/1022 of the European Parliament and of the Council, and repealing Council Regulations (EC) No 894/97, (EC) No 850/98, (EC) No 2549/2000, (EC) No 254/2002, (EC) No 812/2004 and (EC) No 2187/2005. Off. J. EU 2019, L198/105, 1–97.
- Council Regulation (EU) 2019/124 of 30 January 2019 fixing for 2019 the fishing opportunities for certain fish stocks and groups of fish stocks, applicable in Union waters and, for Union fishing vessels, in certain non-Union waters. Off. J. EU, 2019; L29/1, 1–166.
- FAO. The State of World Fisheries and Aquaculture—Meeting Sustainable Development Goals; FAO: Rome, Italy, 2018; Volume 3, pp. 1–227. [Google Scholar]
- Maz-Courrau, A.; López-Vera, C.; Galván-Magaña, F.; Escobar-Sánchez, O.; Rosíles-Martínez, R.; Sanjuán-Muñoz, A. Bioaccumulation and biomagnifications of total mercury in four exploited shark species in the Baja California Penisula, Maexico. Bull. Environ. Contam. Toxicol. 2012, 88, 129–134. [Google Scholar] [CrossRef] [PubMed]
- RASFF. The Rapid Alert System for Food and Feed. Annual Report 2018; Publications Office of the European Union: Luxembourg, 2018; pp. 1–53. [Google Scholar]
- Mull, C.G.; Lyons, K.; Blasius, M.E.; Winkler, C.; O’Sullivan, J.B.; Lowe, C.G. Evidence of maternal offloading of organic contaminants in white sharks (Carcharodon carcharias). PLoS ONE 2013, 8, e62886. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Castro-González, M.I.; Méndez-Armenta, M. Heavy metals: Implications associated to fish consumption. Environ. Toxicol. Pharm. 2008, 26, 263–271. [Google Scholar] [CrossRef]
- Karami, A.; Golieskardi, A.; Ho, Y.B. Microplastics in eviscerated flesh and excised organs of dried fish. Sci. Rep. 2017, 7, 5473. [Google Scholar] [CrossRef]
- Williams, M.; Hernandez-Jover, M.; Shamsi, S. Fish substitutions which may increase human health risks from zoonotic seafood borne parasites: A review. Food Control. 2020, 118, 107429. [Google Scholar] [CrossRef]
- Sadovy de Mitcheson, Y.; Andersson, A.A.; Hofford, A.; Law, C.S.W.; Hau, L.C.Y.; Pauly, D. Out of control means off the menu: The case for ceasing consumption of luxury products from highly vulnerable species when international trade cannot be adequately controlled; shark fin as a case study. Mar. Policy 2018, 98, 115–120. [Google Scholar] [CrossRef]
- Regulation (EC) No. 178/2002 Of the European Parliament and of the Council of 28 January 2002 laying down the general principles and requirements of food law, establishing the European Food Safety Authority and laying down procedures in matters of food saf. Off. J. Eur. Commun. 2002, L31, 1–24.
- Regulation (EC) No 1224/2009 of 20 November 2009 establishing a Community control system for ensuring compliance with the rules of the common fisheries policy, amending Regulations (EC) No 847/96, (EC) No 2371/2002, (EC) No 811/2004, (EC) No 768/2005, (EC) No 2115/2005, (EC) No 2166/2005, (EC) No 388/2006, (EC) No 509/2007, (EC) No 676/2007, (EC) No 1098/2007, (EC) No 1300/2008, (EC) No 1342/2008 and repealing Regulations (EEC) No 2847/93, (EC) No 1627/94 and (EC) No 1966/2006. Off. J. EU 2009, L343, 1–50.
- Regulation (EU) No 625/2017 of the European Parliament and of the Council of 15 March 2017 on official controls and other official activities performed to ensure the application of food and feed law, rules on animal health and welfare, plant health and plant protection products, amending Regulations (EC) No 999/2001, (EC) No 396/2005, (EC) No 1069/2009, (EC) No 1107/2009, (EU) No 1151/2012, (EU) No 652/2014, (EU) 2016/429 and (EU) 2016/2031 of the European Parliament and of the Council, Council Regulations (EC) No 1/2005 and (EC) No 1099/2009 and Council Directives 98/58/EC, 1999/74/EC, 2007/43/EC, 2008/119/EC and 2008/120/EC, and repealing Regulations (EC) No 854/2004 and (EC) No 882/2004 of the European Parliament and of the Council, Council Directives 89/608/EEC, 89/662/EEC, 90/425/EEC, 91/496/EEC, 96/23/EC, 96/93/EC and 97/78/EC and Council Decision 92/438/EEC (Official Controls Regulation)Text with EEA relevance. Off. J. EU 2017, L95, 1–142.
- European Parliament. Report on the Food Crisis, Fraud in the Food Chain and the Control Thereof. 2013. Available online: http://www.europarl.europa.eu/sides/getDoc.do?pubRef=//EP//TEXT+REPORT+A7-2013-0434+0+DOC+XML+V0//EN.www.iucnredlist.org (accessed on 23 August 2020).
- Gao, Z.; Liu, Y.; Wang, X.; Wei, X.; Han, J. DNA mini-barcoding: A derived barcoding method for herbal molecular identification. Front. Plant. Sci. 2019, 10, 987. [Google Scholar] [CrossRef] [PubMed]
- Shokralla, S.; Hellberg, R.S.; Handy, S.M.; King, I.; Hajibabaei, M. A DNA mini-barcoding system for authentication of processed fish products. Sci. Rep. 2015, 5, 15894. [Google Scholar] [CrossRef] [PubMed]
- Staffen, C.F.; Staffen, M.D.; Becker, M.L.; Löfgren, S.E.; Costa Netto Muniz, Y.; Hajenius Aché de Freitas, R.; Marrero, A.R. DNA barcoding reveals the mislabeling of fish in a popular tourist destination in Brazil. PeerJ 2017, 29, e4006. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Cerutti-Pereyra, F.; Meekan, M.G.; Wei, N.-W.V.; O’Shea, O.; Bradshaw, C.J.A.; Austin, C.M. Identification of rays through DNA barcoding: An application for ecologists. PLoS ONE 2012, 7, e36479. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Liu, S.-Y.V.; Chan, C.-L.C.; Lin, O.; Hu, C.-S.; Chen, C.A. DNA barcoding of shark meats identify species composition and CITES-listed species from the markets in Taiwan. PLoS ONE 2013, 8, e79373. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Sato, Y.; Miya, M.; Fukunaga, T.; Sado, T.; Iwasaki, W. MitoFish and MiFish pipeline: A Mitochondrial genome database of fish with an analysis pipeline for environmental DNA metabarcoding. Mol. Biol. Evol. 2018, 35, 1553–1555. [Google Scholar] [CrossRef]
| Target Gene | Primer Sequences (5′–3′) | Amplicon Length (bp) | References |
|---|---|---|---|
| COI | F- TCAACCAACCACAAAGACATTGGCAC | ~655 | [55] |
| R- TAGACTTCTGGGTGGCCAAAGAATCA | |||
| NADH2 | F- AAGGAGCAGTTTGATAGAGT | ~1050 | [40] |
| R- AACGCTTAGCTGTTAATTAA |
| Target Gene | Heat Activation | Denaturation | Annealing | Extension | Cycles | Final Extension |
|---|---|---|---|---|---|---|
| COI | 95 °C/15 min | 94 °C/30 s | 54 °C/50 s | 72 °C/60 s | 35 | 72 °C/10 min |
| NADH2 | 95 °C /15 min | 94 °C/30 s | 48 °C/30 s | 72 °C/90 s | 35 | 72 °C/10 min |
| Haplotype | COI | NADH2 |
|---|---|---|
| H1 | 1C_Pal (9 specimens) | 1N_Pal (9 specimens) |
| H2 | 6C_Pal (4 specimens), 9C_Pal (4 specimens) | 6N_Pal (4 specimens), 9N_Pal (4 specimens) |
| H3 | 7C_Pal (3 specimens) | 7N_Pal (3 specimens) |
| H4 | 8C_Pal (6 specimens), 18C_Verd (5 specimens) | 8N_Pal (6 specimens), 18N_Verd (5 specimens) |
| H5 | 12C_Pal (4 specimens), 32C_Gat (4 specimens) | 12N_Pal (4 specimens), 32N_Gat (4 specimens) |
| H6 | 22C_Pal (3 specimens) | 22N_Pal (3 specimens) |
| H7 | 24C_Pal (3 specimens) | 24N_Pal (3 specimens) |
| H8 | 26C_Pal (3 specimens) | 26N_Pal (3 specimens) |
| H9 | 30C_Pal (3 specimens) | 30N_Pal (3 specimens) |
| H10 | 4C_Verd (6 specimens) | 4N_Verd (6 specimens) |
| H11 | 15C_Verd (5 specimens), 16C_Verd (5 specimens), 25C_Verd (4 specimens), 33C_Spin (7 specimens) | 15N_Verd (5 specimens),33N_Spin (7 specimens) |
| H12 | - | 16N_Verd (5 specimens), 25N_Verd (4 specimens) |
| H13 | 17C_Verd (5 specimens), 3C_Spin (7 specimens) | 17N_Verd (5 specimens) |
| H14 | - | 3N_Spin (7 specimens) |
| H15 | 31C_Verd (5 specimens) | 31N_Verd (5 specimens) |
| H16 | 2C_Gat (4specimens) | 2N_Gat (4 specimens) |
| H17 | 10C_Gat (4 specimens) | 10N_Gat (4 specimens) |
| H18 | 11C_Gat (4 specimens) | 11N_Gat (4 specimens) |
| H19 | 19C_Gat (4 specimens) | 19N_Gat (4 specimens) |
| H20 | 20C_Gat (4specimens) | 20N_Gat (4 specimens) |
| H21 | 21C_Gat (4 specimens) | 21N_Gat (4 specimens) |
| H22 | 5C_Spin (11 specimens) | 5N_Spin (11 specimens) |
| Species | Number of Samples | Conservation Status | |||
|---|---|---|---|---|---|
| IUCN * | CITES | Barcelona Convention ** | Bern Convention | ||
| Mustelus asterias | 21 | LC | -- | Annex III | -- |
| Mustelus punctulatus | 20 | DD | -- | Annex III | -- |
| Prionace glauca | 55 | CR | -- | Annex III | Appendix III |
| Isurus oxyrinchus | 5 | CR | -- | Annex II | Appendix III |
| Scyliorhinus canicula | 29 | LC | -- | -- | -- |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Marchetti, P.; Mottola, A.; Piredda, R.; Ciccarese, G.; Di Pinto, A. Determining the Authenticity of Shark Meat Products by DNA Sequencing. Foods 2020, 9, 1194. https://doi.org/10.3390/foods9091194
Marchetti P, Mottola A, Piredda R, Ciccarese G, Di Pinto A. Determining the Authenticity of Shark Meat Products by DNA Sequencing. Foods. 2020; 9(9):1194. https://doi.org/10.3390/foods9091194
Chicago/Turabian StyleMarchetti, Patrizia, Anna Mottola, Roberta Piredda, Giuseppina Ciccarese, and Angela Di Pinto. 2020. "Determining the Authenticity of Shark Meat Products by DNA Sequencing" Foods 9, no. 9: 1194. https://doi.org/10.3390/foods9091194
APA StyleMarchetti, P., Mottola, A., Piredda, R., Ciccarese, G., & Di Pinto, A. (2020). Determining the Authenticity of Shark Meat Products by DNA Sequencing. Foods, 9(9), 1194. https://doi.org/10.3390/foods9091194








