A Real-Time PCR Screening Assay for Rapid Detection of Listeria Monocytogenes Outbreak Strains
Abstract
1. Introduction
2. Materials and Methods
2.1. Bacterial Isolates DNA Extraction
2.2. Serogroup
2.3. PFGE
2.4. Next Generation Sequencing
2.5. Real-Time PCR
3. Results
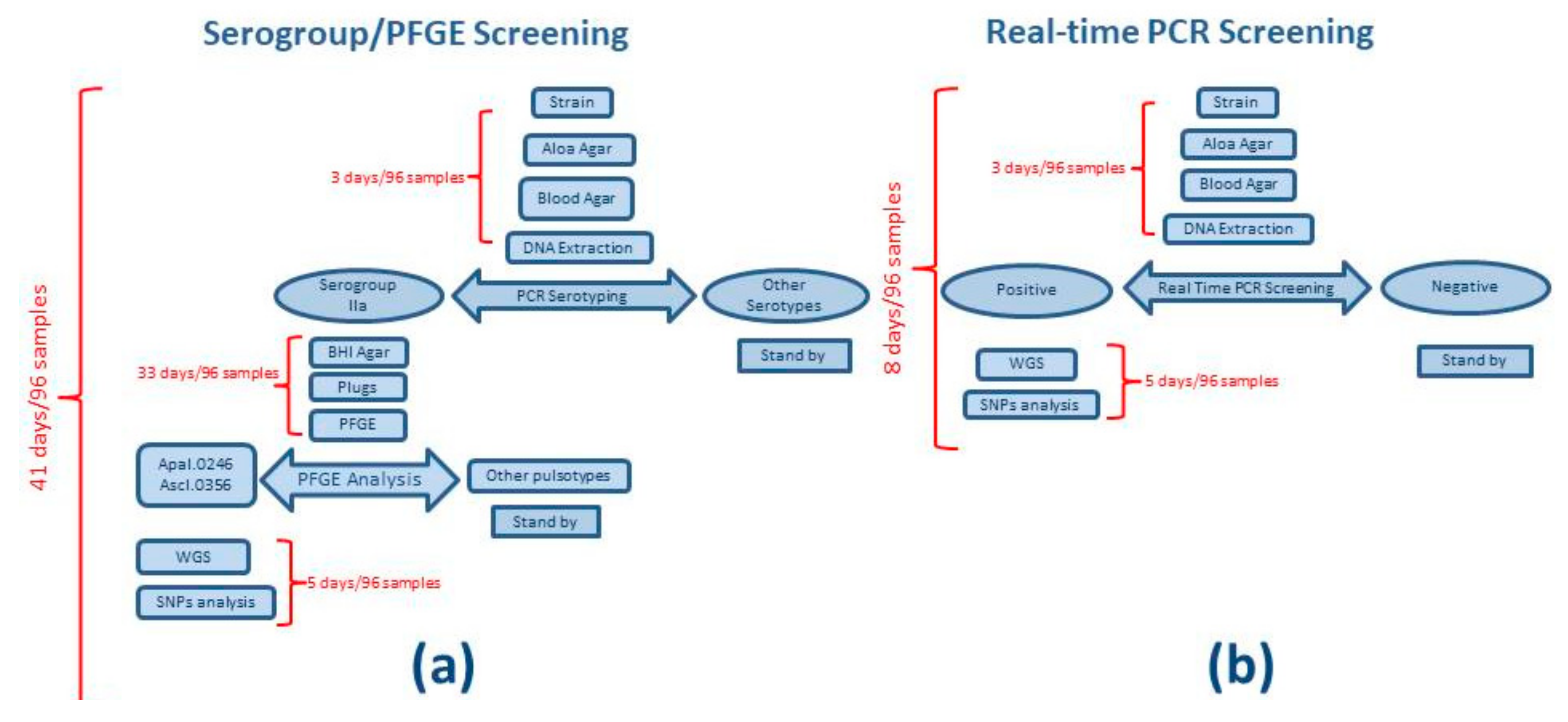
3.1. Workflow
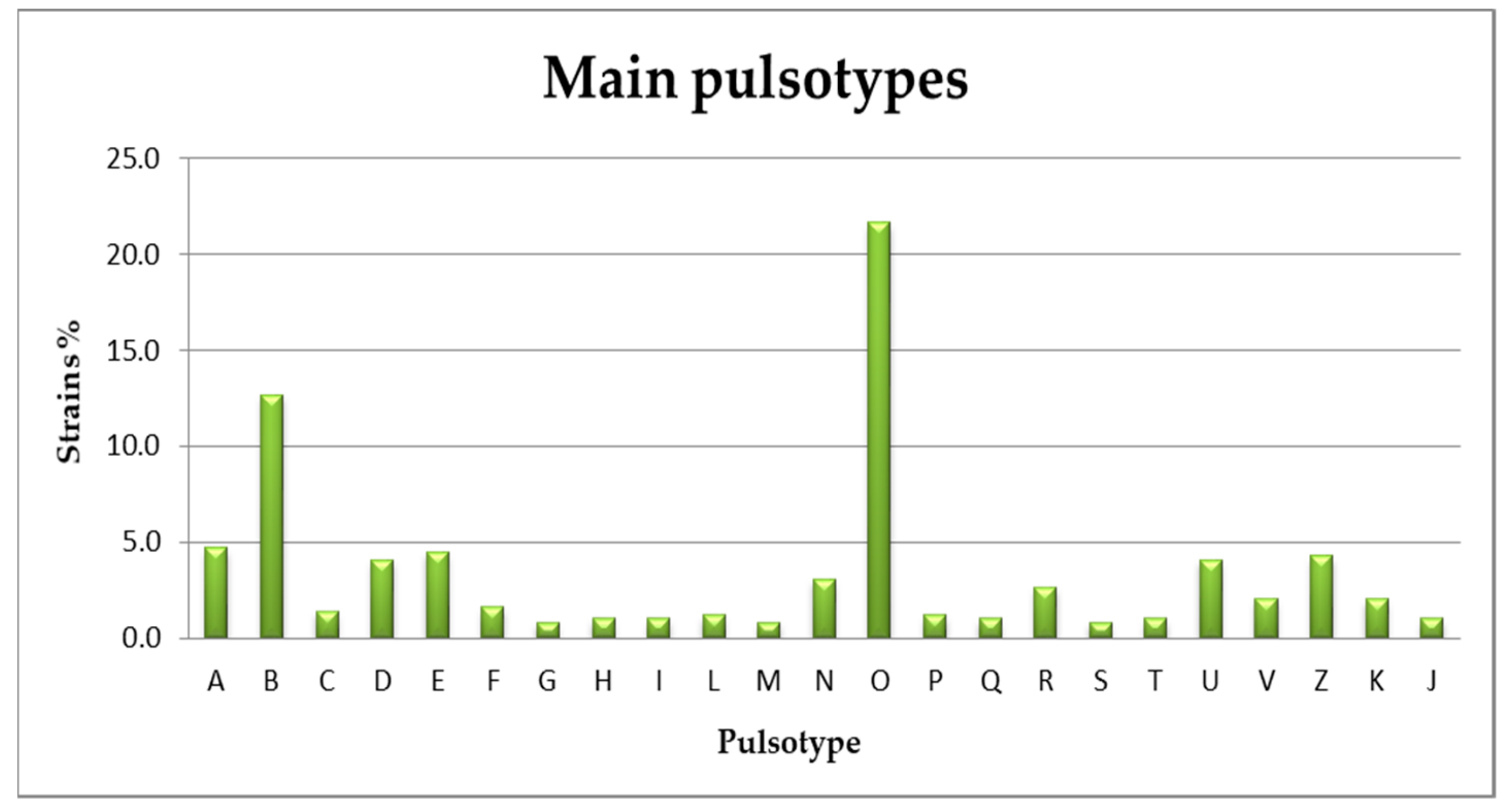
3.2. Serogrouping and PFGE
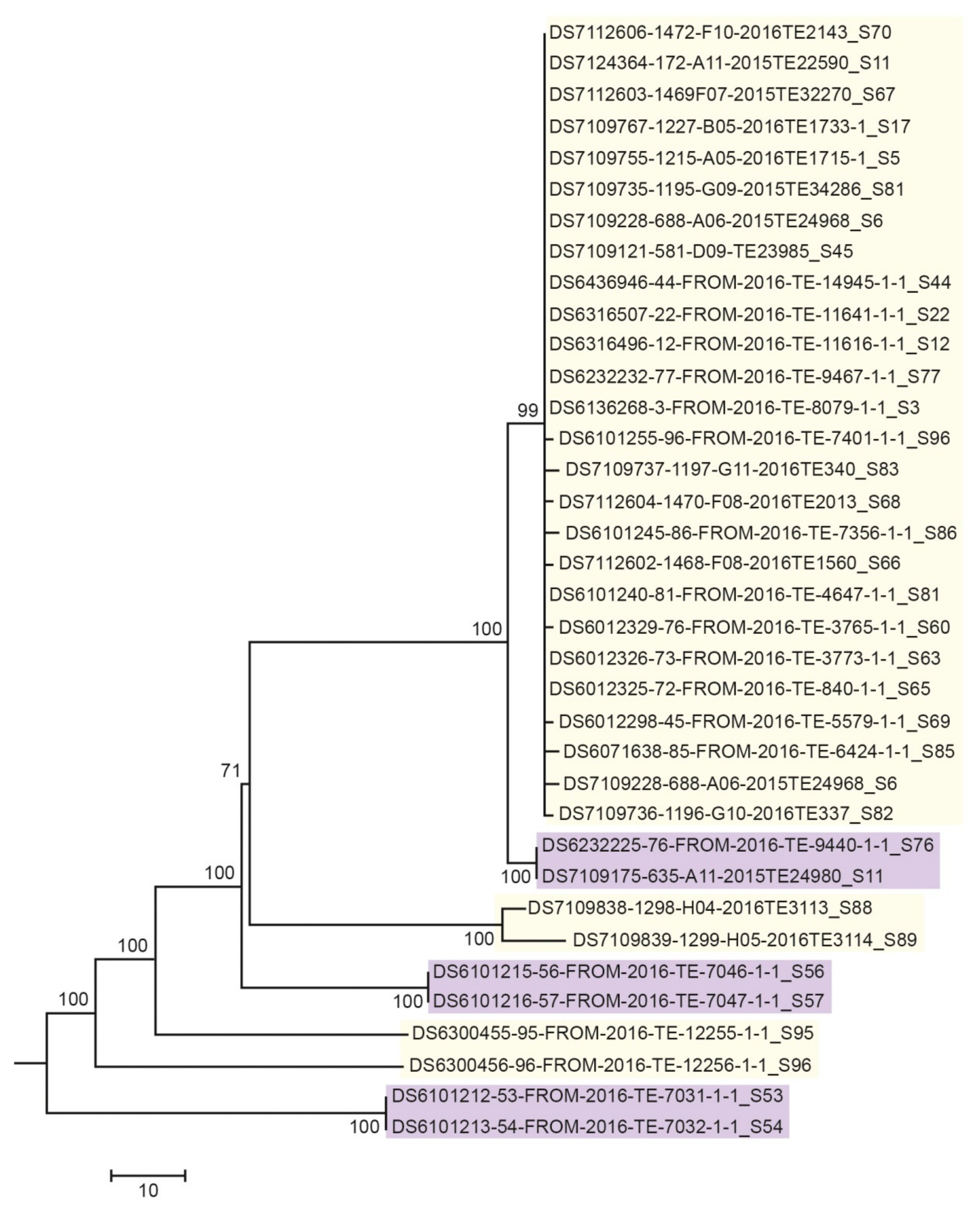
3.3. Next Generation Sequencing
3.4. Real-Time PCR
4. Discussion
5. Conclusions
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Radoshevich, L.; Cossart, P. Listeria monocytogenes: Towards a complete picture of its physiology and pathogenesis. Nat. Rev. Microbiol. 2018, 16, 32–46. [Google Scholar] [CrossRef]
- Chlebicz, A.; Śliżewska, K. Campylobacteriosis, Salmonellosis, Yersiniosis, and Listeriosis as Zoonotic Foodborne Diseases: A Review. Int. J. Environ. Res. Public Health 2018, 15, 863. [Google Scholar] [CrossRef]
- European Food Safety Authority (EFSA) and European Centre for Disease Prevention and Control. (ECDC). The European Union summary report on trends and sources of zoonoses, zoonotic agents and food-borne outbreaks in 2018. EFSA J. 2019, 17, 5926. Available online: https://efsa.onlinelibrary.wiley.com/doi/epdf/10.2903/j.efsa.2019.5926 (accessed on 27 December 2019).
- Duranti, A.; Sabbatucci, M.; Blasi, G.; Acciari, V.A.; Ancora, M.; Bella, A.; Busani, L.; Centorame, P.; Cammà, C.; Conti, F.; et al. A severe outbreak of listeriosis in central Italy with a rare pulsotype associated with processed pork products. J. Med. Microbiol. 2018, 67, 1351–1360. [Google Scholar] [CrossRef] [PubMed]
- Pietzka, A.; Allerberger, F.; Murer, A.; Lennkh, A.; Stöger, A.; Cabal Rosel, A.; Huhulescu, S.; Maritschnik, S.; Springer, B.; Lepuschitz, S.; et al. Whole Genome Sequencing Based Surveillance of L. monocytogenes for Early Detection and Investigations of Listeriosis Outbreaks. Front. Public Health 2019, 7, 139. [Google Scholar] [CrossRef] [PubMed]
- Datta, A.R.; Burall, L.S. Serotype to genotype: The changing landscape of listeriosis outbreak investigations. Food Microb. 2018, 75, 18–27. [Google Scholar] [CrossRef]
- Struelens, M.J.; Brisse, S. From molecular to genomic epidemiology: Transforming surveillance and control of infectious diseases. Eurosurveillance 2013, 18, 20386. [Google Scholar] [CrossRef]
- Quainoo, S.; Coolen, J.P.M.; van Hijum, S.A.F.T.; Huynen, M.A.; Melchers, W.J.G.; van Schaik, W.; Wertheim, H.F.L. Whole-Genome Sequencing of Bacterial Pathogens: The Future of Nosocomial Outbreak Analysis. Clin. Microbiol. Rev. 2017, 30, 1015–1063. [Google Scholar] [CrossRef]
- Doumith, M.; Buchrieser, C.; Glaser, P.; Jacquet, C.; Martin, P. Differentiation of the Major Listeria monocytogenes Serovars by Multiplex PCR. J. Clin. Microbiol. 2004, 42, 3819–3822. [Google Scholar] [CrossRef]
- Kérouanton, A.; Marault, M.; Petit, L.; Grout, J.; Dao, T.T.; Brisabois, A. Evaluation of a multiplex PCR assay as an alternative method for Listeria monocytogenes serotyping. J. Microbiol. Methods 2010, 80, 134–137. [Google Scholar] [CrossRef]
- PulseNet-International. Standard Operating Procedure for Pulsenet PFGE of Listeria monocytogenes. 2013. Available online: http://www.cdc.gov/pulsenet/PDF/listeria-pfge-protocol-508c.pdf (accessed on 21 November 2019).
- Acciari, V.A.; Torresi, M.; Iannetti, L.; Scattolini, S.; Pomilio, F.; Decastelli, L.; Colmegna, S.; Muliari, R.; Bossù, T.; Proroga, Y.; et al. Listeria monocytogenes in smoked salmon and other smoked fish at retail in Italy: Frequency of contamination and strain characterization in products from different manufacturers. J. Food Prot. 2017, 80, 271–278. [Google Scholar] [CrossRef] [PubMed]
- Bolger, A.M.; Lohse, M.; Usadel, B. Trimmomatic: A flexible trimmer for Illumina Sequence Data. Bioinformatics 2014, 30, 2114–2120. [Google Scholar] [CrossRef] [PubMed]
- Bankevich, A.; Nurk, S.; Antipov, D.; Gurevich, A.A.; Dvorkin, M.; Kulikov, A.S.; Lesin, V.M.; Nikolenko, S.I.; Pham, S.; Prjibelski, A.D.; et al. SPAdes: A new genome assembly algorithm and its applications to single-cell sequencing. J. Comput. Biol. 2012, 19, 455–477. [Google Scholar] [CrossRef] [PubMed]
- Seemann, T. Prokka: Rapid prokaryotic genome annotation. Bioinformatics 2014, 30, 2068–2069. [Google Scholar] [CrossRef]
- Page, A.J.; Cummins, C.A.; Hunt, M.; Wong, V.K.; Reuter, S.; Holden, M.T.G.; Fookes, M.; Falush, D.L.; Keane, J.A.; Parkhill, J. Roary: Rapid large-scale prokaryote pan genome analysis. Bioinformatics 2015, 31, 3691–3693. [Google Scholar] [CrossRef]
- Gardner, S.N.; Slezak, T.; Hall, B.G. kSNP3.0: SNP detection and phylogenetic analysis of genomes without genome alignment or reference genomes. Bioinformatics 2015, 31, 2877–2878. [Google Scholar] [CrossRef]
- Morganti, M.; Scaltriti, E.; Cozzolino, P.; Bolzoni, L.; Casadei, G.; Pierantoni, M.; Foni, E.; Pongolini, S. Processing-Dependent and Clonal Contamination Patterns of Listeria monocytogenes in the Cured Ham Food Chain Revealed by Genetic Analysis. Appl. Environ. Microbiol. 2015, 82, 822–831. [Google Scholar] [CrossRef]
- Tamura, K.; Stecher, G.; Peterson, D.; Filipski, A.; Kumar, S. MEGA6: Molecular Evolutionary Genetics Analysis version 6.0. Mol. Biol. Evol. 2013, 2725–2729. [Google Scholar] [CrossRef]
- Larkin, M.A.; Blackshields, G.; Brown, N.P.; Chenna, R.; McGettigan, P.A.; McWilliam, H.; Valentin, F.; Wallace, I.M.; Wilm, A.; Lopez, R.; et al. Clustal W and Clustal X version 2.0. Bioinformatics 2007, 23, 2947–2948. [Google Scholar] [CrossRef]
- Vaerman, J.L.; Saussoy, P.; Ingargiola, I. Evaluation of Real Time PCR data. J. Biol. Regul. Homeost. Agents 2004, 18, 212–214. [Google Scholar]
- Lee, S.; Chen, Y.; Gorski, L.; Ward, T.J.; Osborne, J.; Kathariou, S. Listeria monocytogenes Source Distribution Analysis Indicates Regional Heterogeneity and Ecological Niche Preference among Serotype 4b Clones. mBio 2018, 9. [Google Scholar] [CrossRef] [PubMed]
- Orsi, R.H.; den Bakker, H.C.; Wiedmann, M. Listeria monocytogenes lineages: Genomics, evolution, ecology, and phenotypic characteristics. Int. J. Med. Microbiol. 2011, 301, 79–96. [Google Scholar] [CrossRef] [PubMed]
- Pasquali, F.; Palma, F.; Guillier, L.; Lucchi, A.; De Cesare, A.; Manfreda, G. Listeria monocytogenes Sequence Types 121 and 14 Repeatedly Isolated within One Year of Sampling in a Rabbit Meat Processing Plant: Persistence and Ecophysiology. Front. Microbiol. 2018, 9, 596. [Google Scholar] [CrossRef]
- Michelon, D.; Félix, B.; Vingadassalon, N.; Mariet, J.-F.; Larsson, J.T.; Møller-Nielsen, E.; Roussel, S. PFGE Standard Operating Procedures for Listeria monocytogenes: Harmonizing the Typing of Food and Clinical Strains in Europe. Foodborne Pathog. Dis. 2015, 12, 244–252. [Google Scholar] [CrossRef] [PubMed]
| Oligonucleotide | Sequence 5′-3′ | Size |
|---|---|---|
| Rec-fwd | AAATAATGCGGAGTTAAAAGGTGAA | 74 bp |
| Rec-rev | TGGACTGCATTTGGTATGTGAGT | |
| Rec-probe | FAM-TACGGATTGCCGTCCCCGAAAGT-BHQ1 | |
| Trans-fwd | CTCATTACGTTGATTGGCATACG | 79 bp |
| Trans-rev | GGTTCGTGGTCTCCTTTTACAATAA | |
| Trans-probe | JOE-AACGAAGAAAAGGGAAAAACTCCCACCC-BHQ1 |
| Assay | Slope | R2 | Efficiency |
|---|---|---|---|
| Rec | −3.34 | 0.998 | 99% |
| Trans | −3.47 | 0.999 | 94% |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Torresi, M.; Ruolo, A.; Acciari, V.A.; Ancora, M.; Blasi, G.; Cammà, C.; Centorame, P.; Centorotola, G.; Curini, V.; Guidi, F.; et al. A Real-Time PCR Screening Assay for Rapid Detection of Listeria Monocytogenes Outbreak Strains. Foods 2020, 9, 67. https://doi.org/10.3390/foods9010067
Torresi M, Ruolo A, Acciari VA, Ancora M, Blasi G, Cammà C, Centorame P, Centorotola G, Curini V, Guidi F, et al. A Real-Time PCR Screening Assay for Rapid Detection of Listeria Monocytogenes Outbreak Strains. Foods. 2020; 9(1):67. https://doi.org/10.3390/foods9010067
Chicago/Turabian StyleTorresi, Marina, Anna Ruolo, Vicdalia Aniela Acciari, Massimo Ancora, Giuliana Blasi, Cesare Cammà, Patrizia Centorame, Gabriella Centorotola, Valentina Curini, Fabrizia Guidi, and et al. 2020. "A Real-Time PCR Screening Assay for Rapid Detection of Listeria Monocytogenes Outbreak Strains" Foods 9, no. 1: 67. https://doi.org/10.3390/foods9010067
APA StyleTorresi, M., Ruolo, A., Acciari, V. A., Ancora, M., Blasi, G., Cammà, C., Centorame, P., Centorotola, G., Curini, V., Guidi, F., Marcacci, M., Orsini, M., Pomilio, F., & Di Domenico, M. (2020). A Real-Time PCR Screening Assay for Rapid Detection of Listeria Monocytogenes Outbreak Strains. Foods, 9(1), 67. https://doi.org/10.3390/foods9010067





