Exopolysaccharide (EPS)-Producing Streptococcus thermophilus: Functional and Probiotic Potential
Abstract
1. Introduction
2. Materials and Methods
2.1. Materials
2.2. Bacterial Isolation and Identification
2.3. Technological Characterization of S. thermophilus Strains
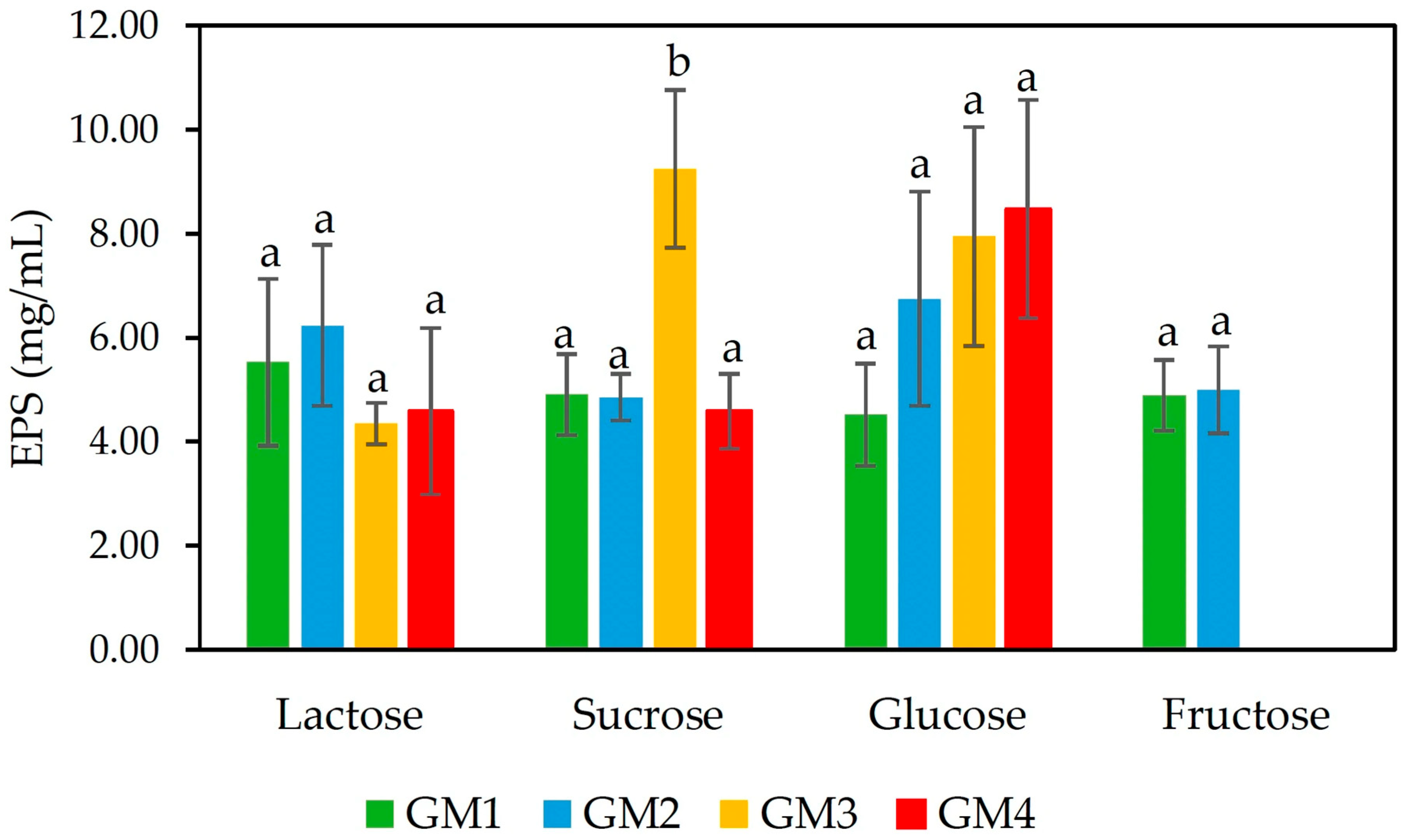
2.3.1. Production of EPS with Different Sugar Substrates
2.3.2. Milk Acidification
2.3.3. Carbohydrate Fermentation
2.3.4. Enzymatic Activity
2.3.5. Proteolytic Activity
2.3.6. Lipolytic Activity
2.4. Safety Evaluation of S. thermophilus Strains
2.4.1. Hemolytic Activity
2.4.2. DNAse Activity
2.4.3. Gelatinase Activity
2.4.4. Antibiotic Susceptibility
2.4.5. Virulence Genes
2.5. Potential Probiotic Properties of S. thermophilus GM4
2.5.1. Gastrointestinal Resistance
2.5.2. Mucin Adhesion
2.5.3. Auto-Aggregation
2.5.4. Co-Aggregation
2.6. Functional Properties of S. thermophilus GM4
2.6.1. β-Galactosidase Activity
2.6.2. Cholesterol-Lowering Ability
2.6.3. DPPH Scavenging Activity
2.6.4. Hydroxyl Scavenging Activity
2.6.5. Production of EPS in Skim Milk
2.7. Purification and Characterization of EPS from S. thermophilus GM4
2.7.1. Purification of EPS
2.7.2. EPS Purity
2.7.3. Carbohydrate Composition
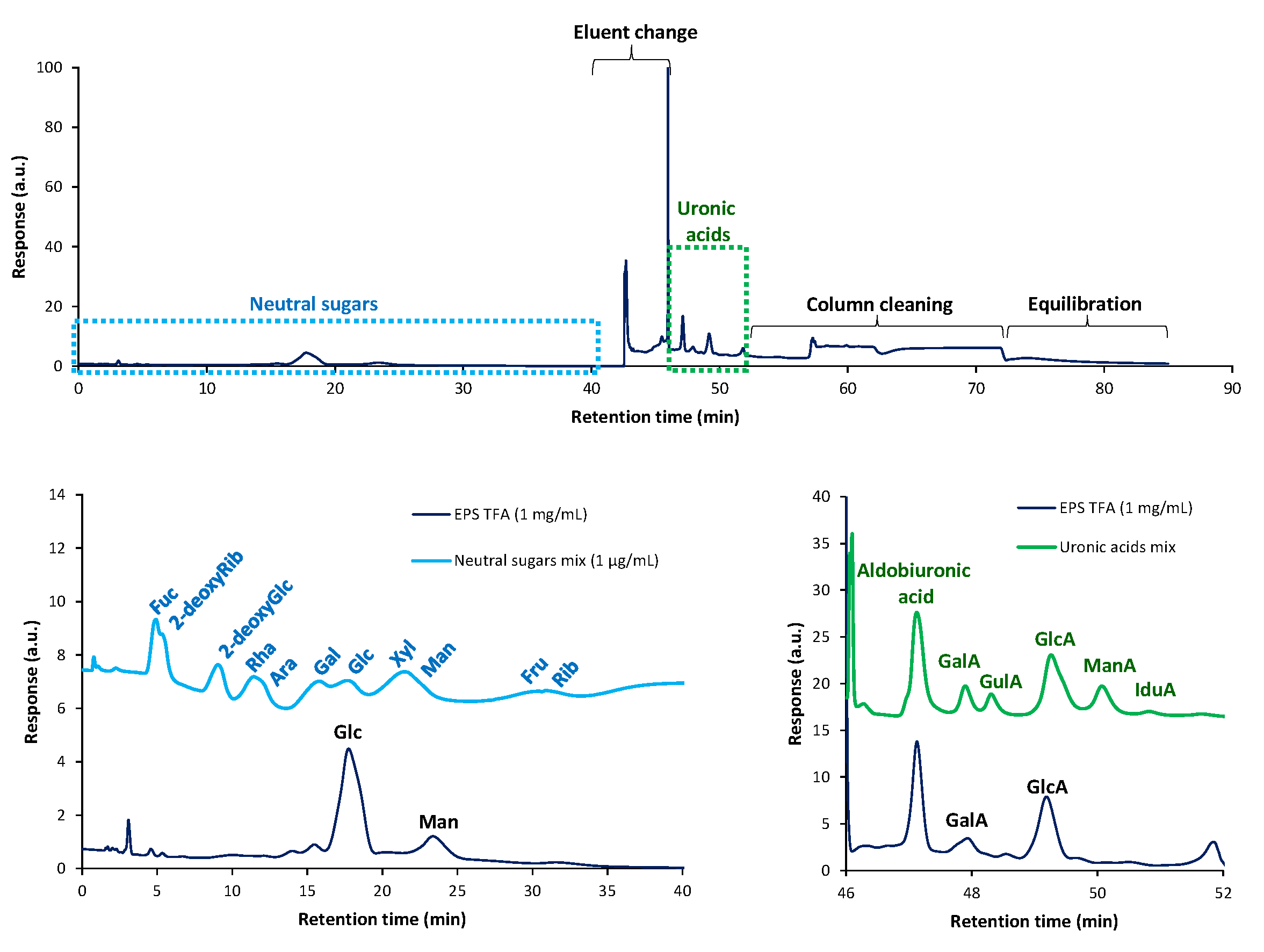
Neutral Sugar Analysis by GC-FID
Uronic Acid Analysis
Carbohydrate Analysis by HPAEC-PAD
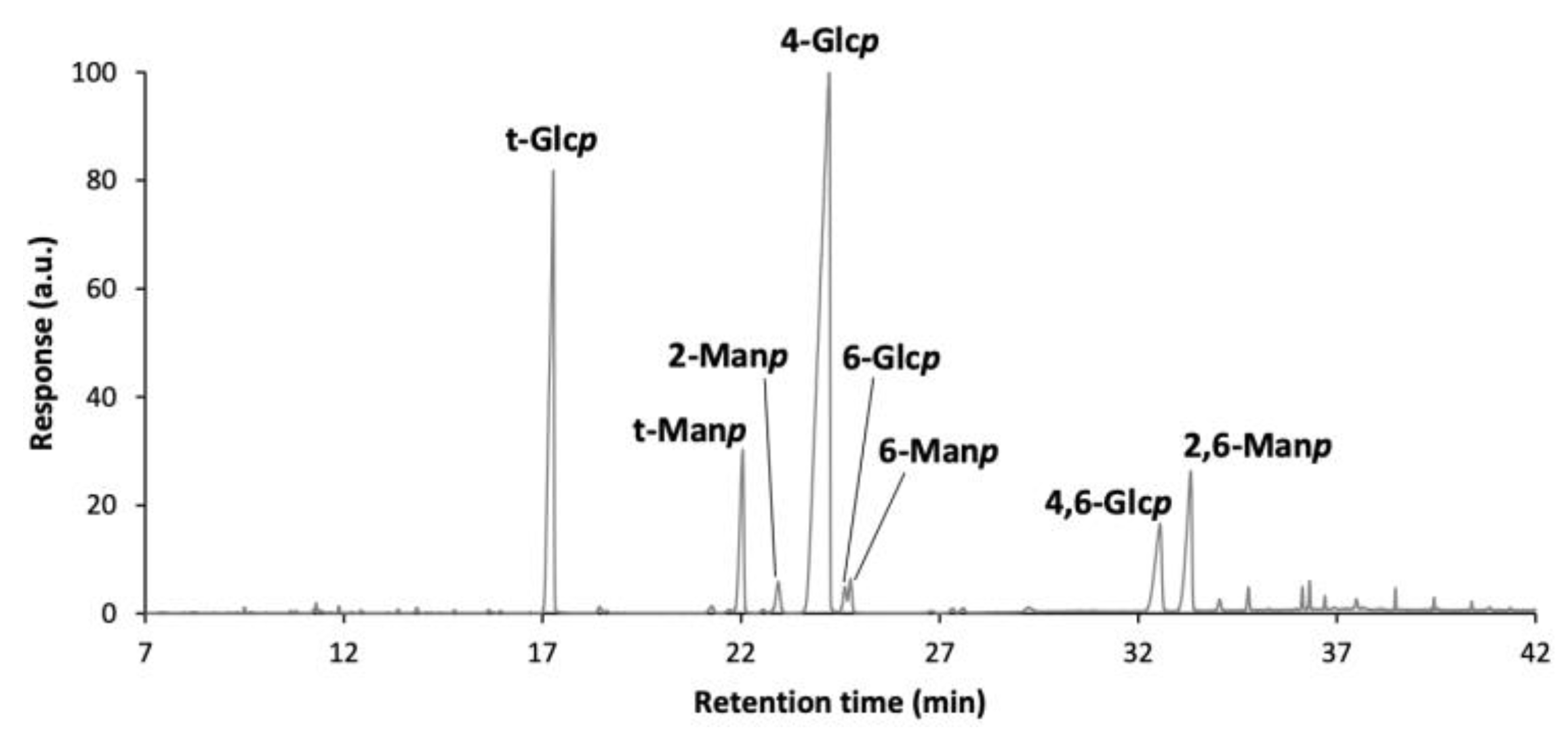
2.7.4. Methylation Analysis
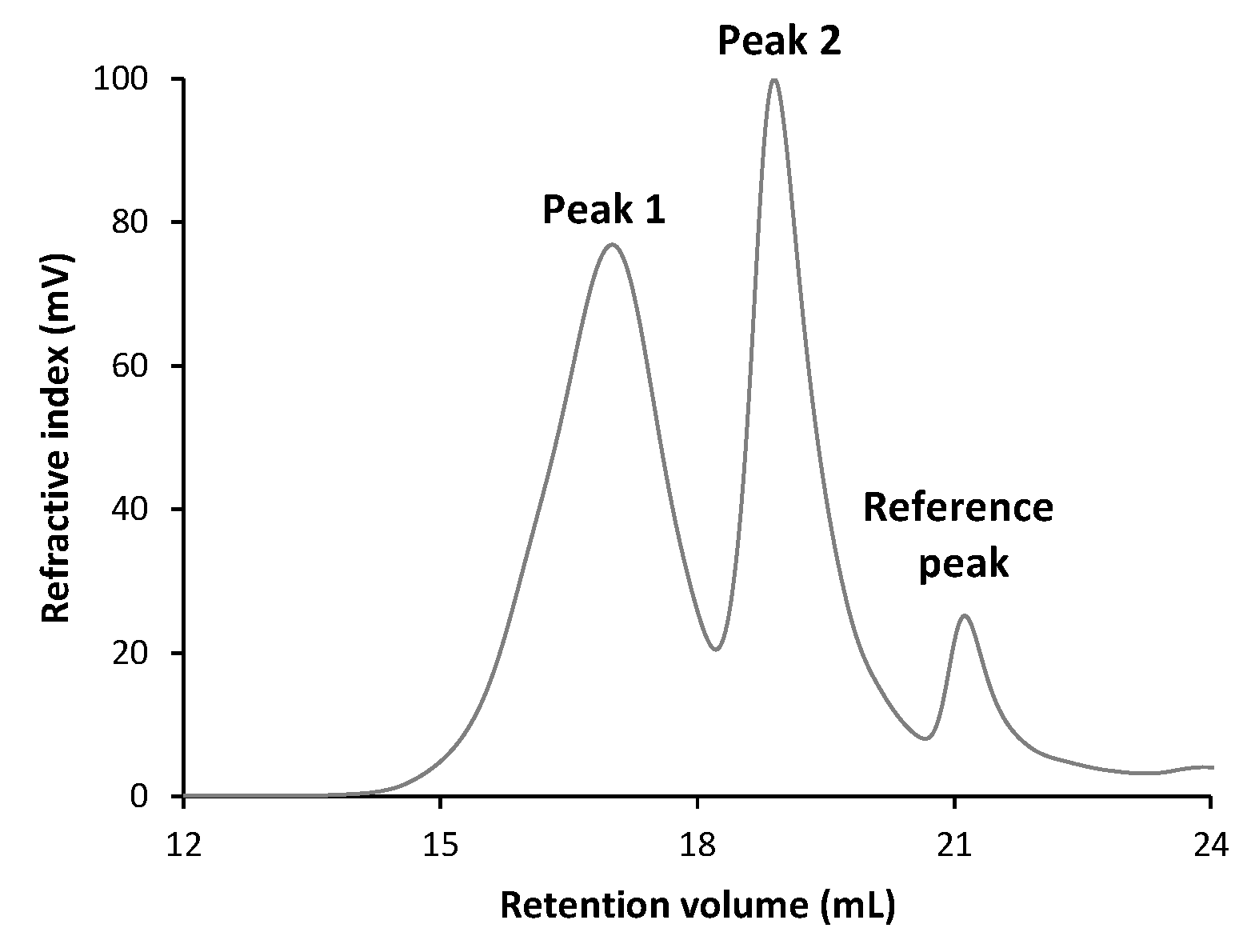
2.7.5. Molecular Weight Distribution and Homogeneity
2.7.6. Dextranase Resistance
2.7.7. DPPH Scavenging Activity
2.7.8. Hydroxyl Scavenging Activity
2.8. Statistical Analysis
3. Results and Discussion
3.1. Technological Characterization of S. thermophilus Strains
3.1.1. Production of EPS with Different Sugar Sources
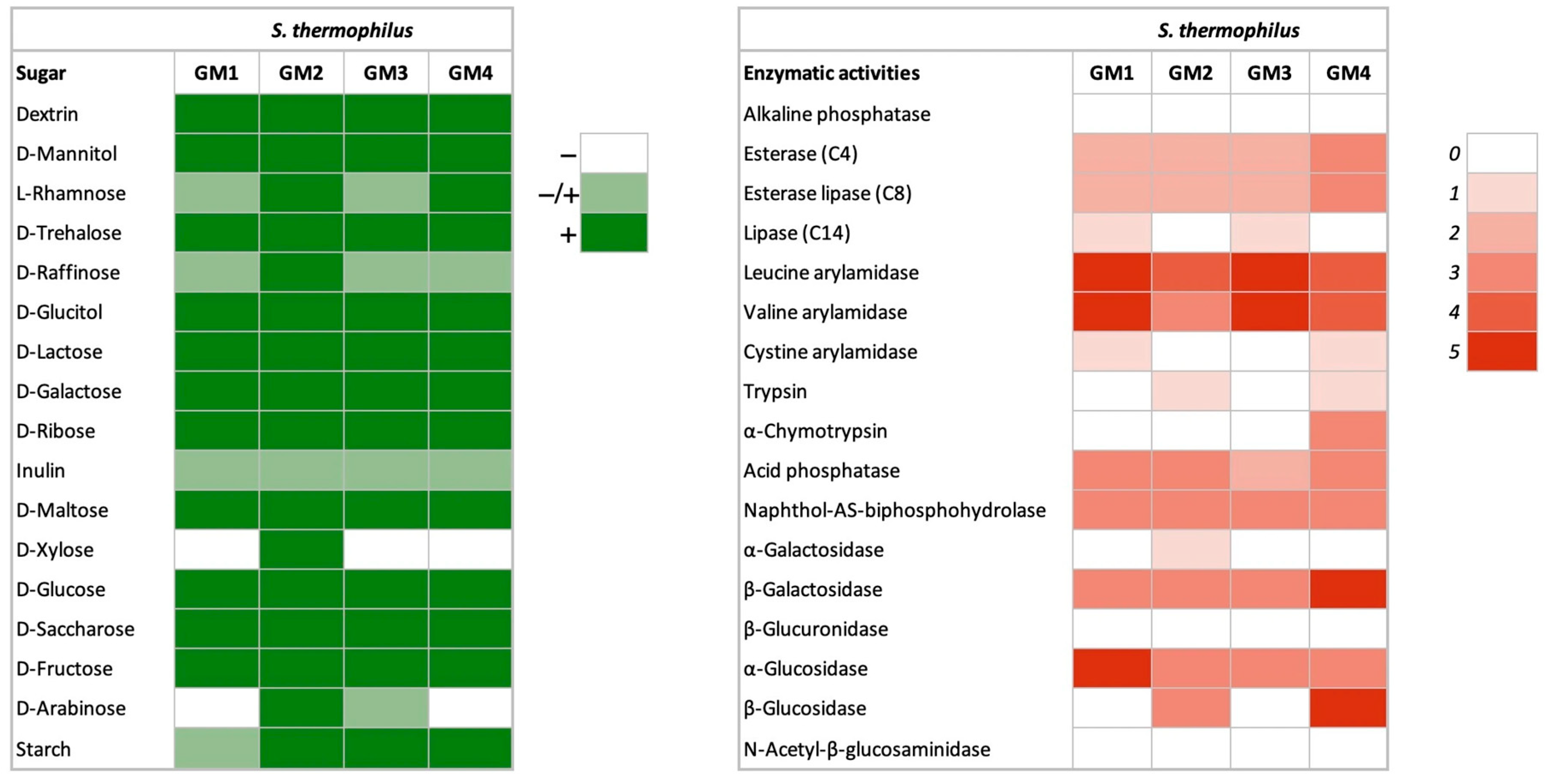
3.1.2. Carbohydrate Fermentation
3.1.3. Enzymatic Activities
3.1.4. Acidification, and Proteolytic and Lipolytic Activities
3.2. Safety Evaluation of S. thermophilus Strains
3.2.1. Hemolysis, DNAse, and Gelatinase Activity
3.2.2. Antibiotic Susceptibility
3.2.3. Virulence Genes
3.3. Probiotic Evaluation of S. thermophilus GM4
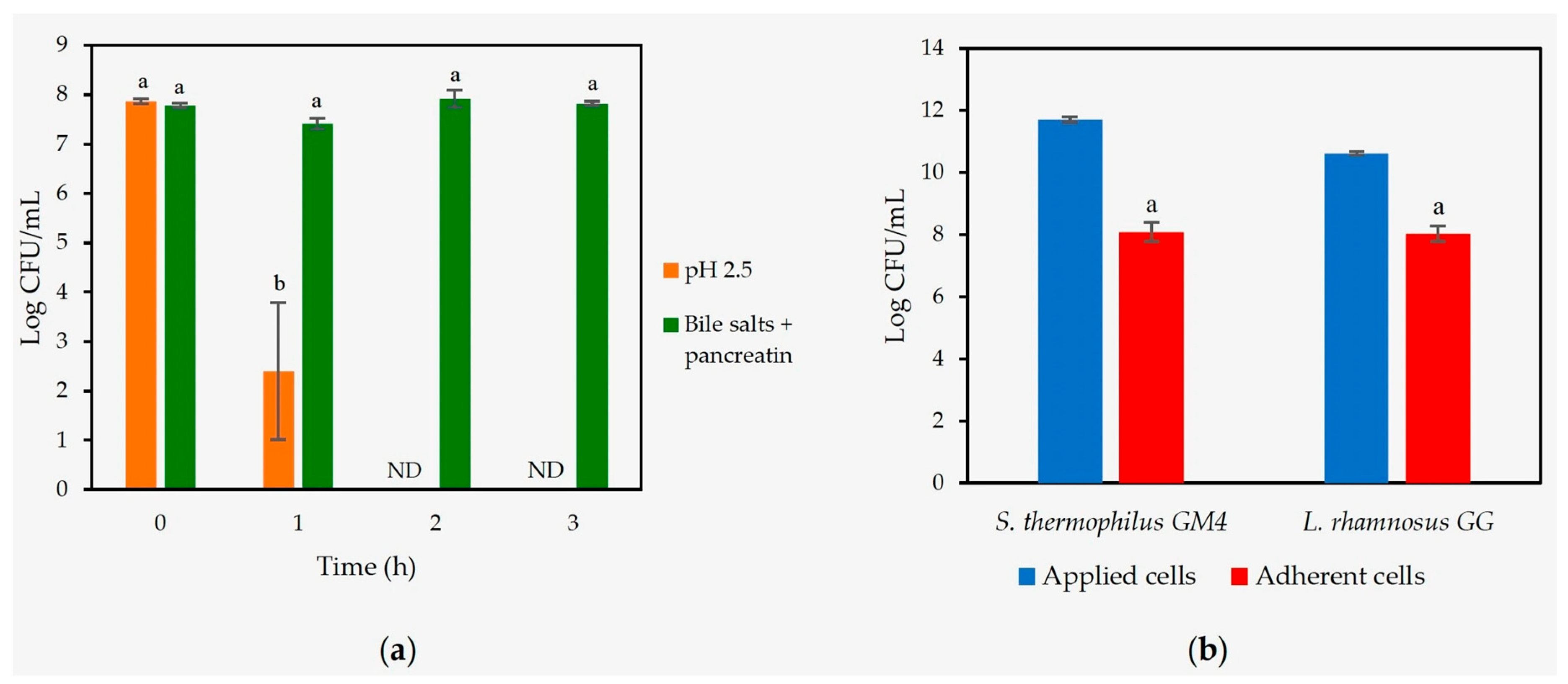
3.3.1. Gastrointestinal Resistance and Adhesion to Mucin
3.3.2. Auto-Aggregation and Co-Aggregation
3.4. Functional Properties of S. thermophilus GM4
3.4.1. β-Galactosidase Activity
3.4.2. Cholesterol-Lowering Ability
3.4.3. Antioxidant Activity
3.4.4. EPS Production During Milk Fermentation
3.5. Characterization of EPS from S. thermophilus GM4
3.5.1. Purity and Carbohydrate Composition of EPS
3.5.2. Molecular Weight Distribution
3.5.3. Dextranase Resistance
3.5.4. Antioxidant Activity
4. Conclusions
Supplementary Materials
Author Contributions
Funding
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Hu, T.; Cui, Y.; Zhang, Y.; Qu, X.; Zhao, C. Genome Analysis and Physiological Characterization of Four Streptococcus thermophilus Strains Isolated from Chinese Traditional Fermented Milk. Front. Microbiol. 2020, 11, 184. [Google Scholar] [CrossRef]
- Grizon, A.; Theil, S.; Callon, C.; Gerber, P.; Helinck, S.; Dugat-Bony, E.; Bonnarme, P.; Chassard, C. Genetic and technological diversity of Streptococcus thermophilus isolated from the Saint-Nectaire PDO cheese-producing area. Front. Microbiol. 2023, 14, 1245510. [Google Scholar] [CrossRef]
- Jurášková, D.; Ribeiro, S.C.; Silva, C.C. Exopolysaccharides produced by lactic acid bacteria: From biosynthesis to healthpromoting properties. Foods 2022, 11, 156. [Google Scholar] [CrossRef]
- Cui, Y.; Xu, T.; Qu, X.; Hu, T.; Jiang, X.; Zhao, C. New Insights into Various Production Characteristics of Streptococcus thermophilus Strains. Int. J. Mol. Sci. 2016, 17, 1701. [Google Scholar] [CrossRef]
- Kapse, N.; Pisu, V.; Dhakephalkar, T.; Margale, P.; Shetty, D.; Wagh, S.; Dagar, S.; Dhakephalkar, P.K. Unveiling the Probiotic Potential of Streptococcus thermophilus MCC0200: Insights from In Vitro Studies Corroborated with Genome Analysis. Microorganisms 2024, 12, 347. [Google Scholar] [CrossRef]
- Huang, Y.-Y.; Lu, Y.-H.; Liu, X.-T.; Wu, W.-T.; Li, W.-Q.; Lai, S.-Q.; Aadil, R.M.; Riaz Rajoka, M.S.; Wang, L.-H.; Zeng, X.-A. Metabolic Properties, Functional Characteristics, and Practical Application of Streptococcus thermophilus. Food Rev. Int. 2024, 40, 792–813. [Google Scholar] [CrossRef]
- Mizuno, H.; Tomotsune, K.; Islam, M.A.; Funabashi, R.; Albarracin, L.; Ikeda-Ohtsubo, W.; Aso, H.; Takahashi, H.; Kimura, K.; Villena, J.; et al. Exopolysaccharides From Streptococcus thermophilus ST538 Modulate the Antiviral Innate Immune Response in Porcine Intestinal Epitheliocytes. Front. Microbiol. 2020, 11, 894. [Google Scholar] [CrossRef]
- Nachtigall, C.; Surber, G.; Wefers, D.; Vogel, C.; Rohm, H.; Jaros, D. Capsular Exopolysaccharides from Two Streptococcus thermophilus Strains Differ in Their Moisture Sorption Behavior. Foods 2023, 12, 596. [Google Scholar] [CrossRef]
- De Vuyst, L.; Zamfir, M.; Mozzi, F.; Adriany, T.; Marshall, V.; Degeest, B.; Vaningelgem, F. Exopolysaccharide-producing Streptococcus thermophilus strains as functional starter cultures in the production of fermented milks. Int. Dairy J. 2003, 13, 707–717. [Google Scholar] [CrossRef]
- Zhang, T.; Zhang, C.; Li, S.; Zhang, Y.; Yang, Z. Growth and exopolysaccharide production by Streptococcus thermophilus ST1 in skim milk. Braz. J. Microbiol. 2011, 42, 1470–1478. [Google Scholar] [CrossRef]
- Wa, Y.; Chanyi, R.M.; Nguyen, H.T.H.; Gu, R.; Day, L.; Altermann, E. Extracellular Polysaccharide Extraction from Streptococcus thermophilus in Fermented Milk. Microbiol. Spectr. 2022, 10, e0228021. [Google Scholar] [CrossRef]
- Vaningelgem, F.; Zamfir, M.; Mozzi, F.; Adriany, T.; Vancanneyt, M.; Swings, J.; De Vuyst, L. Biodiversity of exopolysaccharides produced by Streptococcus thermophilus strains is reflected in their production and their molecular and functional characteristics. Appl. Environ. Microbiol. 2004, 70, 900–912. [Google Scholar] [CrossRef]
- Jurášková, D.; Ribeiro, S.C.; Silva, C.C.G. Identification of ropy phenotype exopolysaccharides in lactic acid bacteria. In 15th Food Chemistry Meeting of Portuguese Chemical Society—Abstract Book; 2021; p. 252. [Google Scholar]
- Pires, V.; Ribeiro, S.C.; Silva, S.P.M.; Juraskova, D.; Silva, C.C.G. Probiotic, technological, and health-related characteristics of lactobacilli isolated from breast milk. J. Appl. Microbiol. 2023, 134, lxad122. [Google Scholar] [CrossRef]
- Cirrincione, S.; Breuer, Y.; Mangiapane, E.; Mazzoli, R.; Pessione, E. ‘Ropy’ phenotype, exopolysaccharides and metabolism: Study on food isolated potential probiotics LAB. Microbiol. Res. 2018, 214, 137–145. [Google Scholar] [CrossRef]
- DuBois, M.; Gilles, K.A.; Hamilton, J.K.; Rebers, P.A.; Smith, F. Colorimetric Method for Determination of Sugars and Related Substances. Anal. Chem. 1956, 28, 350–356. [Google Scholar] [CrossRef]
- Hickey, D.K.; Kilcawley, K.N.; Beresford, T.P.; Wilkinson, M.G. Lipolysis in Cheddar Cheese Made from Raw, Thermized, and Pasteurized Milks. J. Dairy Sci. 2007, 90, 47–56. [Google Scholar] [CrossRef]
- Franciosi, E.; Settanni, L.; Cavazza, A.; Poznanski, E. Biodiversity and technological potential of wild lactic acid bacteria from raw cows’ milk. Int. Dairy J. 2009, 19, 3–11. [Google Scholar] [CrossRef]
- Hantsis-Zacharov, E.; Halpern, M. Culturable psychrotrophic bacterial communities in raw milk and their proteolytic and lipolytic traits. Appl. Environ. Microbiol. 2007, 73, 7162–7168. [Google Scholar] [CrossRef]
- Asteri, I.-A.; Robertson, N.; Kagkli, D.-M.; Andrewes, P.; Nychas, G.; Coolbear, T.; Holland, R.; Crow, V.; Tsakalidou, E. Technological and flavour potential of cultures isolated from traditional Greek cheeses–A pool of novel species and starters. Int. Dairy J. 2009, 19, 595–604. [Google Scholar] [CrossRef]
- Gupta, H.; Malik, R.K. Incidence of virulence in bacteriocin-producing enterococcal isolates. Le Lait 2007, 87, 587–601. [Google Scholar] [CrossRef]
- Terzić-Vidojević, A.; Veljović, K.; Tolinački, M.; Nikolić, M.; Ostojić, M.; Topisirović, L. Characterization of lactic acid bacteria isolated from artisanal Zlatar cheeses produced at two different geographical location. Genetika 2009, 41, 117–136. [Google Scholar] [CrossRef]
- Clinical and Laboratory Standards Institute. Performance Standards for Antimicrobial Susceptibility Testing, 26th ed.; CLSI supplement M100S; Clinical and Laboratory Standards Institute: Malvern, PA, USA, 2016. [Google Scholar]
- Ribeiro, S.C.; Coelho, M.C.; Todorov, S.D.; Franco, B.; Dapkevicius, M.L.E.; Silva, C.C.G. Technological properties of bacteriocin-producing lactic acid bacteria isolated from Pico cheese an artisanal cow’s milk cheese. J. Appl. Microbiol. 2014, 116, 573–585. [Google Scholar] [CrossRef] [PubMed]
- Jurášková, D.; Ribeiro, S.C.; Silva, C.C.G. Prebiotic and Health-Promoting Benefits of Dextran-Type Exopolysaccharide Produced by Leuconostoc mesenteroides SJC113. Foods 2025, 14, 2635. [Google Scholar] [CrossRef] [PubMed]
- Todorov, S.D.; Dicks, L.M. Evaluation of lactic acid bacteria from kefir, molasses and olive brine as possible probiotics based on physiological properties. Ann. Microbiol. 2008, 58, 661–670. [Google Scholar] [CrossRef]
- Chen, T.; Wang, L.; Li, Q.; Long, Y.; Lin, Y.; Yin, J.; Zeng, Y.; Huang, L.; Yao, T.; Abbasi, M.N. Functional probiotics of lactic acid bacteria from Hu sheep milk. BMC Microbiol. 2020, 20, 228. [Google Scholar] [CrossRef]
- Li, W.; Zhao, X.; Zou, S.; Ma, Y.; Zhang, K.; Zhang, M. Scanning assay of β-galactosidase activity. Appl. Biochem. Microbiol. 2012, 48, 603–607. [Google Scholar] [CrossRef]
- Domingos-Lopes, M.; Stanton, C.; Ross, R.; Silva, C. Histamine and cholesterol lowering abilities of lactic acid bacteria isolated from artisanal Pico cheese. J. Appl. Microbiol. 2020, 129, 1428–1440. [Google Scholar] [CrossRef]
- Aarti, C.; Khusro, A. Functional and technological properties of exopolysaccharide producing autochthonous Lactobacillus plantarum strain AAS3 from dry fish based fermented food. LWT 2019, 114, 108387. [Google Scholar] [CrossRef]
- He, Z.; Luo, H.; Cao, C.; Cui, Z. Photometric determination of hydroxyl free radical in Fenton system by brilliant green. Am. J. Chin. Clin. Med. 2004, 6, 236–243. [Google Scholar]
- Domingos-Lopes, M.F.P.; Lamosa, P.; Stanton, C.; Ross, R.P.; Silva, C.C.G. Isolation and characterization of an exopolysaccharide-producing Leuconostoc citreum strain from artisanal cheese. Lett. Appl. Microbiol. 2018, 67, 570–578. [Google Scholar] [CrossRef]
- Ferreira, S.S.; Pereira, R.B.; Bordalo, D.; Barbosa, H.; Ferreira, N.; Correia, A.; Ferreira, P.; Pinto, B.; Rodrigues, A.M.C.; Navalho, J.; et al. Salt pan brine water sulphated polysaccharides retrieved at pilot scale: Ability to stimulate in vitro human macrophages and salmon head kidney cells. Int. J. Biol. Macromol. 2025, 317, 144506. [Google Scholar] [CrossRef]
- Bradford, M.M. A rapid and sensitive method for the quantitation of microgram quantities of protein utilizing the principle of protein-dye binding. Anal. Biochem. 1976, 72, 248–254. [Google Scholar] [CrossRef]
- Circuncisão, A.R.; Ferreira, S.S.; Silva, A.M.S.; Coimbra, M.A.; Cardoso, S.M. Fucus vesiculosus-Rich Extracts as Potential Functional Food Ingredients: A Holistic Extraction Approach. Foods 2024, 13, 540. [Google Scholar] [CrossRef]
- Ferreira, S.S.; Correia, A.; Silva, A.M.S.; Wessel, D.F.; Cardoso, S.M.; Vilanova, M.; Coimbra, M.A. Structure-function relationships of pectic polysaccharides from broccoli by-products with in vitro B lymphocyte stimulatory activity. Carbohydr. Polym. 2023, 303, 120432. [Google Scholar] [CrossRef]
- Passos, C.P.; Cepeda, M.R.; Ferreira, S.S.; Nunes, F.M.; Evtuguin, D.V.; Madureira, P.; Vilanova, M.; Coimbra, M.A. Influence of molecular weight on in vitro immunostimulatory properties of instant coffee. Food Chem. 2014, 161, 60–66. [Google Scholar] [CrossRef]
- Hu, S.-M.; Zhou, J.-M.; Zhou, Q.-Q.; Li, P.; Xie, Y.-Y.; Zhou, T.; Gu, Q. Purification, characterization and biological activities of exopolysaccharides from Lactobacillus rhamnosus ZFM231 isolated from milk. LWT 2021, 147, 111561. [Google Scholar] [CrossRef]
- Zhang, J.; Liu, L.; Chen, F. Production and characterization of exopolysaccharides from Chlorella zofingiensis and Chlorella vulgaris with anti-colorectal cancer activity. Int. J. Biol. Macromol. 2019, 134, 976–983. [Google Scholar] [CrossRef] [PubMed]
- Padmanabhan, A.; Tong, Y.; Wu, Q.; Zhang, J.; Shah, N.P. Transcriptomic Insights Into the Growth Phase- and Sugar-Associated Changes in the Exopolysaccharide Production of a High EPS-Producing Streptococcus thermophilus ASCC 1275. Front. Microbiol. 2018, 9, 1919. [Google Scholar] [CrossRef]
- Fuso, A.; Bancalari, E.; Castellone, V.; Caligiani, A.; Gatti, M.; Bottari, B. Feeding Lactic Acid Bacteria with Different Sugars: Effect on Exopolysaccharides (EPS) Production and Their Molecular Characteristics. Foods 2023, 12, 215. [Google Scholar] [CrossRef] [PubMed]
- Sørensen, H.M.; Rochfort, K.D.; Maye, S.; MacLeod, G.; Brabazon, D.; Loscher, C.; Freeland, B. Exopolysaccharides of Lactic Acid Bacteria: Production, Purification and Health Benefits towards Functional Food. Nutrients 2022, 14, 2938. [Google Scholar] [CrossRef]
- Degeest, B.; Vuyst, L.D. Correlation of Activities of the Enzymes alpha-Phosphoglucomutase, UDP-Galactose 4-Epimerase, and UDP-Glucose Pyrophosphorylase with Exopolysaccharide Biosynthesis by Streptococcus thermophilus LY03. Appl. Environ. Microbiol. 2000, 66, 3519–3527. [Google Scholar] [CrossRef]
- Li, Y.; Wang, Y.; Li, B.; Hou, B.; Hung, W.; He, J.; Jiang, Y.; Zhang, Y.; Man, C. Streptococcus thermophilus JM905—Strain Carbon Source Utilization and Its Fermented Milk Metabolic Profile at Different Fermentation Stages. Foods 2023, 12, 3690. [Google Scholar] [CrossRef]
- Nie, C.; Xiong, Z.; Zhang, H.; Hou, C.; Xie, F.; Song, X.; Xia, Y.; Ai, L. Effect of carbon sources on characterization of exopolysaccharides from Streptococcus thermophilus S-3. Food Biosci. 2024, 61, 104676. [Google Scholar] [CrossRef]
- Erkus, O.; Okuklu, B.; Yenidunya, A.F.; Harsa, S. High genetic and phenotypic variability of Streptococcus thermophilus strains isolated from artisanal Yuruk yoghurts. LWT 2014, 58, 348–354. [Google Scholar] [CrossRef]
- Vin, F.d.; Rådström, P.; Herman, L.; Vuyst, L.D. Molecular and Biochemical Analysis of the Galactose Phenotype of Dairy Streptococcus thermophilus Strains Reveals Four Different Fermentation Profiles. Appl. Environ. Microbiol. 2005, 71, 3659–3667. [Google Scholar] [CrossRef] [PubMed]
- van den Bogaard, P.T.; Hols, P.; Kuipers, O.P.; Kleerebezem, M.; de Vos, W.M. Sugar utilisation and conservation of the gal-lac gene cluster in Streptococcus thermophilus. Syst. Appl. Microbiol. 2004, 27, 10–17. [Google Scholar] [CrossRef]
- Savijoki, K.; Ingmer, H.; Varmanen, P. Proteolytic systems of lactic acid bacteria. Appl. Microbiol. Biotechnol. 2006, 71, 394–406. [Google Scholar] [CrossRef] [PubMed]
- Rul, F.; Monnet, V. Presence of additional peptidases in Streptococcus thermophilus CNRZ 302 compared to Lactococcus lactis. J. Appl. Microbiol. 1997, 82, 695–704. [Google Scholar] [CrossRef]
- Akuzawa, R.; Fox, P.F. Acid phosphatase in cheese. Anim. Sci. J. 2004, 385–391. [Google Scholar] [CrossRef]
- Oak, S.J.; Jha, R. The effects of probiotics in lactose intolerance: A systematic review. Crit. Rev. Food Sci. Nutr. 2019, 59, 1675–1683. [Google Scholar] [CrossRef]
- Śliżewska, K.; Markowiak-Kopeć, P.; Śliżewska, W. The Role of Probiotics in Cancer Prevention. Cancers 2020, 13, 20. [Google Scholar] [CrossRef] [PubMed]
- Delgado, S.; O’Sullivan, E.; Fitzgerald, G.; Mayo, B. Subtractive screening for probiotic properties of lactobacillus species from the human gastrointestinal tract in the search for new probiotics. J. Food Sci. 2007, 72, M310–M315. [Google Scholar] [CrossRef]
- Bujňáková, D.; Kmeť, V. Functional properties of Lactobacillus strains isolated from dairy products. Folia Microbiol. 2012, 57, 263–267. [Google Scholar] [CrossRef]
- Galia, W.; Perrin, C.; Genay, M.; Dary, A. Variability and molecular typing of Streptococcus thermophilus strains displaying different proteolytic and acidifying properties. Int. Dairy J. 2009, 19, 89–95. [Google Scholar] [CrossRef]
- Alexandraki, V.; Kazou, M.; Blom, J.; Pot, B.; Papadimitriou, K.; Tsakalidou, E. Comparative Genomics of Streptococcus thermophilus Support Important Traits Concerning the Evolution, Biology and Technological Properties of the Species. Front. Microbiol. 2019, 10, 2916. [Google Scholar] [CrossRef] [PubMed]
- Hazards, E.P.o.B.; Allende, A.; Alvarez-Ordóñez, A.; Bortolaia, V.; Bover-Cid, S.; De Cesare, A.; Dohmen, W.; Guillier, L.; Jacxsens, L.; Nauta, M. Update of the list of qualified presumption of safety (QPS) recommended microbiological agents intentionally added to food or feed as notified to EFSA 21: Suitability of taxonomic units notified to EFSA until September 2024. EFSA J. 2025, 23, e9169. [Google Scholar] [CrossRef] [PubMed]
- EFSA Panel on Additives and Products or Substances used in Animal Feed (FEEDAP). Guidance on the assessment of bacterial susceptibility to antimicrobials of human and veterinary importance. EFSA J. 2012, 10, 2740. [Google Scholar] [CrossRef]
- Zarzecka, U.; Zadernowska, A.; Chajęcka-Wierzchowska, W. Starter cultures as a reservoir of antibiotic resistant microorganisms. LWT 2020, 127, 109424. [Google Scholar] [CrossRef]
- Flórez, A.B.; Mayo, B. Antibiotic Resistance-Susceptibility Profiles of Streptococcus thermophilus Isolated from Raw Milk and Genome Analysis of the Genetic Basis of Acquired Resistances. Front. Microbiol. 2017, 8, 2608. [Google Scholar] [CrossRef]
- Anderson, A.C.; Jonas, D.; Huber, I.; Karygianni, L.; Wölber, J.; Hellwig, E.; Arweiler, N.; Vach, K.; Wittmer, A.; Al-Ahmad, A. Enterococcus faecalis from Food, Clinical Specimens, and Oral Sites: Prevalence of Virulence Factors in Association with Biofilm Formation. Front. Microbiol. 2015, 6, 1534. [Google Scholar] [CrossRef]
- Fisher, K.; Phillips, C. The ecology, epidemiology and virulence of Enterococcus. Microbiology 2009, 155, 1749–1757. [Google Scholar] [CrossRef]
- Wang, C.; El-Telbany, M.; Lin, Y.; Zhao, J.; Maung, A.T.; Abdelaziz, M.N.S.; Nakayama, M.; Masuda, Y.; Honjoh, K.-i.; Miyamoto, T. Identification of Enterococcus spp. from food sources by matrix-assisted laser desorption ionization-time of flight mass spectrometry and characterization of virulence factors, antibiotic resistance, and biofilm formation. Int. J. Food Microbiol. 2024, 420, 110768. [Google Scholar] [CrossRef]
- Chajęcka-Wierzchowska, W.; Zadernowska, A.; Łaniewska-Trokenheim, Ł. Virulence factors of Enterococcus spp. presented in food. LWT 2017, 75, 670–676. [Google Scholar] [CrossRef]
- Kumari, V.B.C.; Huligere, S.S.; Ramu, R.; Naik Bajpe, S.; Sreenivasa, M.Y.; Silina, E.; Stupin, V.; Achar, R.R. Evaluation of Probiotic and Antidiabetic Attributes of Lactobacillus Strains Isolated From Fermented Beetroot. Front. Microbiol. 2022, 13, 911243. [Google Scholar] [CrossRef]
- Collado, M.C.; Meriluoto, J.; Salminen, S. Adhesion and aggregation properties of probiotic and pathogen strains. Eur. Food Res. Technol. 2008, 226, 1065–1073. [Google Scholar] [CrossRef]
- Mater, D.D.; Drouault-Holowacz, S.; Oozeer, R.; Langella, P.; Anba, J.; Corthier, G. Beta-galactosidase production by Streptococcus thermophilus is higher in the small intestine than in the caecum of human-microbiota-associated mice after lactose supplementation. Br. J. Nutr. 2006, 96, 177–181. [Google Scholar] [CrossRef]
- Savaiano, D.A.; AbouElAnouar, A.; Smith, D.E.; Levitt, M.D. Lactose malabsorption from yogurt, pasteurized yogurt, sweet acidophilus milk, and cultured milk in lactase-deficient individuals. Am. J. Clin. Nutr. 1984, 40, 1219–1223. [Google Scholar] [CrossRef]
- Yang, Y.Q.; Xie, Y.H.; Zhang, H.X.; Liu, H. Studies on the Resistance of Cholesterol-Lowering of Lactic Acid Bacteria to Simulated Gastrointestinal Circumstance In Vitro. Adv. Mater. Res. 2013, 807–809, 2014–2018. [Google Scholar] [CrossRef]
- Ziarno, M. Viability and cholesterol uptake by Streptococcus thermophilus cultures in artificial GIT fluids. Acta Sci. Pol. Technol. Aliment. 2010, 9, 83–94. [Google Scholar]
- Mao, B.; Guo, W.; Chen, M.; Tang, X.; Zhang, Q.; Zhao, J.; Zhang, H.; Cui, S. Effects of Streptococcus thermophilus Fermentation on the Flavors and Antioxidant Properties of Barley Juice. Fermentation 2023, 9, 623. [Google Scholar] [CrossRef]
- Hoffmann, A.; Kleniewska, P.; Pawliczak, R. Antioxidative activity of probiotics. Arch. Med. Sci. AMS 2021, 17, 792–804. [Google Scholar] [CrossRef]
- Hyun, J.H.; Woo, I.K.; Kim, K.T.; Park, Y.S.; Kang, D.K.; Lee, N.K.; Paik, H.D. Heat-Treated Paraprobiotic Latilactobacillus sakei KU15041 and Latilactobacillus curvatus KU15003 Show an Antioxidant and Immunostimulatory Effect. J. Microbiol. Biotechnol. 2024, 34, 358–366. [Google Scholar] [CrossRef] [PubMed]
- DÜz, M.; DoĞan, Y.N.; DoĞan, İ. Antioxidant activitiy of Lactobacillus plantarum, Lactobacillus sake and Lactobacillus curvatus strains isolated from fermented Turkish Sucuk. An. Acad. Bras. Cienc. 2020, 92, e20200105. [Google Scholar] [CrossRef]
- Kanamarlapudi, S.; Muddada, S. Characterization of Exopolysaccharide Produced by Streptococcus thermophilus CC30. Biomed Res. Int. 2017, 2017, 4201809. [Google Scholar] [CrossRef]
- Xu, Z.; Guo, Q.; Zhang, H.; Xiong, Z.; Zhang, X.; Ai, L. Structural characterisation of EPS of Streptococcus thermophilus S-3 and its application in milk fermentation. Int. J. Biol. Macromol. 2021, 178, 263–269. [Google Scholar] [CrossRef] [PubMed]
- de Alexandre Sebastião, F.; Pilarski, F.; Lemos, M.V. Composition of Extracellular Polymeric Substances (EPS) produced by Flavobacterium columnare isolated from tropical fish in Brazil. Braz. J. Microbiol. 2013, 44, 861–864. [Google Scholar] [CrossRef]
- Antunes, S.; Freitas, F.; Alves, V.D.; Grandfils, C.; Reis, M.A. Conversion of cheese whey into a fucose- and glucuronic acid-rich extracellular polysaccharide by Enterobacter A47. J. Biotech. 2015, 210, 1–7. [Google Scholar] [CrossRef]
- Parra-Riofrío, G.; García-Márquez, J.; Casas-Arrojo, V.; Uribe-Tapia, E.; Abdala-Díaz, R.T. Antioxidant and Cytotoxic Effects on Tumor Cells of Exopolysaccharides From Tetraselmis suecica (Kylin) Butcher Grown Under Autotrophic and Heterotrophic Conditions. Mar. Drugs 2020, 18, 534. [Google Scholar] [CrossRef]
- Ahmad, W.; Nasir, A.; Sattar, F.; Ashfaq, I.; Chen, M.H.; Hayat, A.; Rehman, M.U.; Zhao, S.; Khaliq, S.; Ghauri, M.A.; et al. Production of bimodal molecular weight levan by a Lactobacillus reuteri isolate from fish gut. Folia Microbiol. 2022, 67, 21–31. [Google Scholar] [CrossRef]
- Angelov, A.; Georgieva, A.; Petkova, M.; Bartkiene, E.; Rocha, J.M.; Ognyanov, M.; Gotcheva, V. On the Molecular Selection of Exopolysaccharide-Producing Lactic Acid Bacteria from Indigenous Fermented Plant-Based Foods and Further Fine Chemical Characterization. Foods 2023, 12, 3346. [Google Scholar] [CrossRef] [PubMed]
- Chen, C.-C.; Nargotra, P.; Kuo, C.-H.; Liu, Y.-C. High-Molecular-Weight Exopolysaccharides Production from Tuber borchii Cultivated by Submerged Fermentation. Int. J. Mol. Sci. 2023, 24, 4875. [Google Scholar] [CrossRef] [PubMed]
- Ruas-Madiedo, P.; Hugenholtz, J.; Zoon, P. An overview of the functionality of exopolysaccharides produced by lactic acid bacteria. Int. Dairy J. 2002, 12, 163–171. [Google Scholar] [CrossRef]
- Jurášková, D.; Ribeiro, S.C.; Bastos, R.; Coelho, E.; Coimbra, M.A.; Silva, C.C.G. Exopolysaccharide (EPS) Produced by Leuconostoc mesenteroides SJC113: Characterization of Functional and Technological Properties and Application in Fat-Free Cheese. Macromol 2024, 4, 680–696. [Google Scholar] [CrossRef]
- Benattouche, Z.; Bouhadi, D.; Raho, G.B. Antioxidant and antibacterial activities of exopolysaccharides produced by lactic acid bacteria isolated from yogurt. Int. J. Food Stud. 2018, 7, 30–37. [Google Scholar] [CrossRef]
- Adelekan, A.O.; Olurin, T.O.; Ezeani, A.O. Antioxidant activities of exopolysaccharides produced by lactic acid bacteria isolated from commercial yoghurt samples. Adv. Microbiol. 2020, 10, 359–374. [Google Scholar] [CrossRef]
- Vankerckhoven, V.; Van Autgaerden, T.; Vael, C.; Vael, C.; Lammens, C.; Chapelle, S.; Rossi, R.; Jabes, D.; Goossens, H. Development of a multiplex PCR for the detection of asa1, gelE, cylA, esp, and hyl genes in enterococci and survey for virulence determinants among European hospital isolates of Enterococcus faecium. J. Clin. Microbiol. 2004, 42, 4473–4479. [Google Scholar] [CrossRef]
- Martín-Platero, A.M.; Valdivia, E.; Maqueda, M.; Martínez-Bueno, M. Characterization and safety evaluation of enterococci isolated from Spanish goats’ milk cheeses. Int. J. Food Microbiol. 2009, 132, 24–32. [Google Scholar] [CrossRef]
- Rivas, P.; Alonso, J.; Moya, J.; de Gorgolas, M.; Martinell, J.; Fernandez Guerrero, M.L. The impact of hospital-acquired infections on the microbial etiology and prognosis of late-onset prosthetic valve endocarditis. Chest 2005, 128, 764–771. [Google Scholar] [CrossRef]
| S. thermophilus Strains | Acidification | Proteolytic Activity | Lipolytic Activity | |
|---|---|---|---|---|
| ΔpH 6 h | ΔpH 24 h | |||
| GM1 | 0.37 ± 0.05 a | 1.64 ± 0.04 a | − | − |
| GM2 | 0.05 ± 0.01 b | 1.32 ± 0.14 b | − | − |
| GM3 | 0.05 ± 0.01 b | 1.55 ± 0.06 a | + | − |
| GM4 | 0.11 ± 0.01 b | 1.63 ± 0.09 a | + | − |
| Strains | Hemolysis | DNase | Gelatinase | Antibiotic Resistance * | Virulence Genes † |
|---|---|---|---|---|---|
| GM1 | γ | - | - | OXA, VAN, KAN | - |
| GM2 | γ | - | - | OXA, VAN, KAN, GEN | - |
| GM3 | γ | - | - | OXA, TET, STR, ERY | - |
| GM4 | γ | - | - | Sensitive to all | - |
| Auto-aggregation (%) | 46.8 ± 0.9 |
| Co-aggregation (%) | |
| Escherichia coli ATCC 25922 | 36.0 ± 0.2 |
| Listeria monocytogenes ATCC 13932 | 31.4 ± 0.3 |
| Staphylococcus aureus ATCC 25923 | 30.9 ± 0.3 |
| Salmonella enterica serovar Typhimurium ATCC14028 | 26.4 ± 0.2 |
| β-galactosidase activity (103 Miller units) | 56.2 ± 2.5 |
| Cholesterol biding (%) | 26.9 ± 3.3 |
| DPPH scavenging activity (%) | 22.7 ± 1.2 |
| Hydroxyl scavenging activity (%) | 5.7 ± 0.2 |
| EPS yield milk (g/L) | 0.13 ± 0.03 |
| EPS yield milk + Lac (g/L) | 2.58 ± 0.05 |
| Dextranase resistance of EPS (%) | 70.3 ± 1.2 |
| Sugar Composition (%mol) | Total Sugars (mg/g) | |||||||
|---|---|---|---|---|---|---|---|---|
| Rha | Rib | Ara | Man | Gal | Glc | Uronic Acids | ||
| EPS GM4 | tr * | 6 | 2 | 29 | 2 | 53 | 8 | 509 ± 12 |
| Deduced Linkages | mol% |
|---|---|
| t-Glcp | 21.6 |
| 4-Glcp | 57.1 |
| 6-Glcp | 0.9 |
| 4,6-Glcp | 5.4 |
| Total Glcp | 85.0 |
| 2-Manp | 6.6 |
| 6-Manp | 1.1 |
| 2,6-Manp | 7.1 |
| Total Manp | 14.7 |
| Total | Peak 1 | Peak 2 | |
|---|---|---|---|
| Mn (g/mol) | 12,027 | 95,102 | 5614 |
| Mw (g/mol) | 111,500 | 188,959 | 10,110 |
| Mw/Mn (g/mol) | 9.271 | 1.987 | 1.801 |
| EPS (mg/mL) | DPPH Scavenging Activity (%) | Hydroxyl Scavenging Activity (%) |
|---|---|---|
| 1 | 30.10 ± 2.43 | 2.41 ± 0.40 |
| 2 | 41.32 ± 0.74 | 7.26 ± 1.02 |
| 3 | 52.46 ± 0.57 | 12.86 ± 0.67 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2025 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Jurášková, D.; Pires, V.C.; Ribeiro, S.C.; Ferreira, S.S.; Bernardo, F.; Evtyugin, D.; Coimbra, M.A.; Silva, C.C.G. Exopolysaccharide (EPS)-Producing Streptococcus thermophilus: Functional and Probiotic Potential. Foods 2025, 14, 3013. https://doi.org/10.3390/foods14173013
Jurášková D, Pires VC, Ribeiro SC, Ferreira SS, Bernardo F, Evtyugin D, Coimbra MA, Silva CCG. Exopolysaccharide (EPS)-Producing Streptococcus thermophilus: Functional and Probiotic Potential. Foods. 2025; 14(17):3013. https://doi.org/10.3390/foods14173013
Chicago/Turabian StyleJurášková, Dominika, Vanessa C. Pires, Susana C. Ribeiro, Sónia S. Ferreira, Fábio Bernardo, Dmitry Evtyugin, Manuel A. Coimbra, and Celia C. G. Silva. 2025. "Exopolysaccharide (EPS)-Producing Streptococcus thermophilus: Functional and Probiotic Potential" Foods 14, no. 17: 3013. https://doi.org/10.3390/foods14173013
APA StyleJurášková, D., Pires, V. C., Ribeiro, S. C., Ferreira, S. S., Bernardo, F., Evtyugin, D., Coimbra, M. A., & Silva, C. C. G. (2025). Exopolysaccharide (EPS)-Producing Streptococcus thermophilus: Functional and Probiotic Potential. Foods, 14(17), 3013. https://doi.org/10.3390/foods14173013









